Abstract
Multiple ectomycorrhizal fungi (EMF) compete to colonise the roots of a host plant, but it is not known whether their success is under plant or fungal control, or a combination of both. We assessed whether plants control EMF colonisation by preferentially allocating more carbon to more beneficial partners in terms of nitrogen supply or if other factors drive competitive success. We combined stable isotope labelling and RNA-sequencing approaches to characterise nutrient exchange between the plant host Eucalyptus grandis and three Pisolithus isolates when growing alone and when competing either indirectly (with a physical barrier) or directly. Overall, we found that nitrogen provision to the plant does not explain the amount of carbon that an isolate receives nor the number of roots that it colonises. Differences in nutrient exchange among isolates were related to differences in expression of key fungal and plant nitrogen and carbon transporter genes. When given a choice of partners, the plant was able to limit colonisation by the least cooperative isolate. This was not explained by a reduction in allocated carbon. Instead, our results suggest that partner choice in EMF could operate through the upregulation of defence-related genes against those fungi providing fewer nutrients.
Similar content being viewed by others
Introduction
Ectomycorrhizal fungi (EMF) have a key role in forest ecosystem functioning, colonising the roots of most tree species (Tedersoo et al., 2010). They receive carbon from their host plant in exchange for providing multiple benefits such as access to nutrients and water, protection against pathogens or tolerance to heavy metals (Smith and Read, 2008). Multiple EMF species and individuals with different functional attributes compete to colonise the plant root system. Several studies have reported negative effects of EMF competition in root colonisation and plant biomass (for example, Wu et al., 1999; Kennedy et al., 2007, 2009; Hortal et al., 2008, 2016). However, existing studies provide an incomplete understanding of factors driving EMF competition (Kennedy, 2010).
Competition between EMF during root colonisation could occur either directly between the EMF, or indirectly, via the host plant. Direct competition can be evaluated by growing fungi in a one-compartment microcosm, while indirect competition can be evaluated using two physically separated compartments that are connected to the same host (Kennedy, 2010). If colonisation is the same in both experiments, then interfungal interaction is likely minimal and the plant controls EMF colonisation. Alternatively, if colonisation changes in direct versus indirect competition, then that would suggest that fungal–fungal interactions drive competitive outcomes independent of the host plant.
Considerable progress has been made in understanding how plants may control the dynamics of another key group of symbiotic fungi: arbuscular mycorrhizal fungi (AMF). It has been shown that market theory of nutrient shuttling prevails in some arbuscular mycorrhizal fungi associations with the plant rewarding the fungus that provides the most nutrients with more carbohydrates and therefore minimising infection by less-cooperative symbionts (Bever et al., 2009; Kiers et al., 2011). This response is not universal, however, and may only apply to certain conditions (Walder and van der Heijden, 2015). Similar preferential allocation operates in the legume–rhizobium symbiosis (Simms et al., 2006; Sachs et al., 2010) and partner choice is considered important to stabilise the mutualism (Bever, 2015).
The evidence for nutrient trading dynamics in regulating EMF colonisation is equivocal. Corrêa et al. (2008) showed that carbon supply to the EMF species Pisolithus tinctorius continued even when nitrogen supply decreased. Carbon supply to the fungus does not actually represent a significant cost for EMF plant partners (Corrêa et al., 2012). Valtanen et al. (2014) showed that fungi supplying the plant with more nitrogen did not receive more carbon than those supplying lower amounts of nitrogen and Albarracín et al. (2013) suggested fungal autonomy with respect to carbon and nitrogen utilisation. In addition, Näsholm et al. (2013) and Hasselquist et al. (2016) found evidence of reduced nitrogen transfer to the plant with increased carbon. This evidence against reciprocal rewards in the EM symbiosis opens the questions of what actually drives nutrient flux between partners and what causes one symbiont to colonise more roots than another.
The existence of strong nonreciprocal behaviour and antagonistic interactions (Walder and Van der Heijden, 2015) also suggests that an alternative explanation to preferential allocation is needed. Active repression of uncooperative symbionts after infection through sanctions has been shown to be important in rhizobial symbioses (West et al., 2002; Kiers et al., 2003) and has been proposed, but not demonstrated, as a means by which host plants might control the functioning of EMF associations (Hoeksema and Kummel, 2003). In particular, the latter suggested that plants could facilitate the persistence of more beneficial fungi by enhancing the mortality of root tips colonised by competitively superior and less beneficial fungi.
In the present study, we used a microcosm-based system where Eucalyptus grandis seedlings were colonised by a range of Pisolithus isolates competing either directly or indirectly. Using this simplified model system, we sought to: (i) further our understanding of whether the host plant, the fungal partner or both control the outcomes of EMF competition, (ii) further define the carbon and nitrogen trading and transport dynamics occurring in EMF associations and (iii) evaluate the importance of alternative mechanisms of plant control over EMF colonisation. We sought to address these aims by integrating a complementary set of physiological measurements, stable isotope tracing and gene expression analyses. Taken together, these data address key knowledge gaps in the areas of nutrient trading, competition outcomes and control over root colonisation, all essential for understanding the relationship dynamics among trees and their ensemble of symbiotic partners.
Materials and methods
Fungal and plant material
Two P. microcarpus isolates (R10 and SI9) and one P. albus isolate (SI12, previously attributed to P. microcarpus) were selected based on their ability to colonise E. grandis and their negative response to the presence of a competitor (Hortal et al., 2016). Seeds of E. grandis were surface sterilised and germinated as per Hortal et al. (2016). The main root of each seedling was cut to promote splitting of the root system. After 12 weeks, seedlings with well-developed split root systems were selected.
Microcosm design
Individual E. grandis seedlings were transferred to 9 cm Petri dishes containing ½ strength modified Melin–Norkrans agar with 0.1% glucose (Hortal et al., 2016) in such a way that the leaves remained outside the dish using a hole burned through the side and the roots inside on top of a sterile cellophane membrane (Figure 1). Seedlings were inoculated with either a single Pisolithus isolate (R10, SI12 or SI9; eight replicates per treatment) or two isolates in all pairwise combinations (R10-SI12, R10-SI9 or SI12-SI9; 10 replicates per treatment) under either indirect or direct competition resulting in nine treatments in total. Indirect competition microcosms were established in a two-compartment Petri dish with the root system of the plant equally divided between the two compartments. For the inoculations, one (for single isolate treatments) or two (one per isolate in the competition treatments) circular fungal plugs (0.5 cm diameter) were obtained from the edge of actively growing colonies and placed at either side from the seedling root. In all microcosms we removed the agar of two 3-mm-wide vertical trenches at 2.5 cm of each side of the seedling. This created two side compartments that only fungal mycelium could access. The whole system was contained inside a larger empty Petri dish. Control microcosms with non-inoculated seedlings (eight replicates) as well as microcosms with mycelium only (three replicates per isolate) were also established following the same design. Microcosms were incubated in a growth cabinet at 25 °C under a 16 h light cycle (PAR 100 μmol m−2 s−1) for 6 weeks.
The microcosm set-up showing an indirect competition treatment using a two-compartment Petri dish with a physical barrier in the middle. The seedling root was split between the two compartments; each of them inoculated with one fungal isolate. The agar of a trench (indicated with arrows) was removed to create a side compartment (indicated with an F) only accessible to the fungus where the 15NH4Cl solution was placed. The leaves remained outside and the whole system was included inside a larger Petri dish with holes covered with micropore tape. Microcosms of the direct competition treatment were the same but without the middle physical barrier while those with no competition had one fungal isolate only.
Stable isotope labelling
For 15N labelling, two sterile pieces of filter paper (Whatman no. 1; Whatman, GE Healthcare Life Sciences, Little Chalfont, UK) 1 cm long x 0.5 cm wide were placed on top of each other 5 mm from the edge of the fungal colony growing in the fungal compartment and saturated with 40 μl of a 30 mM 99% atom excess 15NH4Cl solution in an acetate buffer (pH 4.8). In the microcosms with two fungal isolates, only one of the isolates received the 15N-labelled solution. Four or five labelled microcosms of each isolate in each competition combination (that is, none, indirect or direct) were produced.
On the following day, microcosms were transferred to a clear polycarbonate chamber designed for the 13CO2 labelling (for more details see Supplementary Information and Supplementary Figure 1). The internal Petri dishes were completely sealed while leaves were given access to 13CO2. The chamber was placed in a controlled environment growth room (BioChambers Inc., Winnipeg, MB, Canada) at 25 °C and a 14-h light cycle (PAR 500 μmol m−2 s−1). Twelve millilitres of 99% atom excess 13CO2 were injected into the completely sealed chamber using a syringe through a rubber septum. Plants were allowed to photosynthesise under these conditions for 5 h. The same procedure of 13CO2 labelling was repeated on the following day. We detected a decrease in 12CO2 and 13CO2 concentration in air samples collected 5 h after the injection compared with samples taken right after the injection, confirming that there had been an uptake of 13C label by the seedlings (Supplementary Information). Non-inoculated seedlings and microcosms with only mycelium were labelled following the same procedures. Sixteen microcosms from across the different treatments remained unlabelled to determine background levels of 15N and 13C in both mycelium and leaves.
Harvesting and parameters measured
Harvesting took place 6 days after 15N labelling (that is, 4 days after the second 13CO2 injection) following Fellbaum et al. (2012), Grelet et al. (2009), Högberg et al. (2008), Kiers et al. (2011) and Pickles et al. (2016). We counted the number of mycorrhizas formed by each isolate in each microcosm. Differences in mycorrhiza morphology among the three isolates (Hortal et al., 2016) plus their position in the microcosm allowed us to attribute each mycorrhiza to a particular isolate as the colonies did not overlap. Microcosms were scanned and the area of the colonies was measured using the Image J software (Rasband, 1997).
Approximately 10 mg of mycorrhizal root tips from a subset of microcosms (see below) were collected in random order within a 2 h timeframe and immediately placed in liquid nitrogen for RNA extraction. The mass of leaves and mycelium for all microcosms was determined after drying at 40 °C for 72 h. Mycelium density was calculated by dividing the mass of the colony by its area. Dry leaf and mycelium samples were manually ground into a fine powder using a micropestle and 1 mg was weighed into tin capsules. We checked for any 15N diffusion to the central compartment and 13C diffusion into the fungal compartment in the agar of microcosms containing non-inoculated seedlings as well as in the mycelium of competing unlabelled fungi. Samples were analysed for atom % 15N and 13C as well as nitrogen and carbon content (%) using an Automated Nitrogen Carbon Analyser system (Sercon Limited, Crewe, UK) consisting of a Sercon 20-22 mass spectrometer (Sercon Limited) and an elemental analyser (Sercon Limited) at the West Australian Biogeochemistry Centre (University of Western Australia, Crawley, WA, Australia). Atom % excess 15N and 13C were calculated as the difference between labelled and unlabelled samples. Following Pena and Polle (2014), 15N and 13C content was calculated as follows:
Leaf 15N content (ng excess)=(leaf dry biomass (g) x N concentration (ng g−1) x 15N atom % excess)/100.
Mycelium 13C content (ng excess)=(mycelium dry biomass (g) x C concentration (ng g−1) x 13C atom % excess)/100
RNA-sequencing
RNA was extracted from three samples of non-colonised root tips of control E. grandis seedlings (for baseline plant expression levels), three samples of mycelium from each of the three Pisolithus isolates (for baseline fungal expression levels) and three samples of mycorrhizal root tips per isolate alone (3 isolatesx3 replicates) and in competition (3 isolatesx2 competitorsx2 competition treatmentsx3 replicates). In all cases, samples had been labelled with 15NH4Cl solution. RNA was extracted using the Qiagen RNeasy Plant Mini Kit (Qiagen, Hilden, Germany) following the manufacturer’s instructions. Transcriptional analysis of all tissues was tested using RNA-sequencing via conventional Poly-A library preparation for Illumina sequencing. Library construction and 125-bp paired-end reads sequencing was performed on an Illumina HiSeq 2500 flow cell by the Western Sydney University Next Generation Sequencing Facility. Details on data analysis and availability are provided in Supplementary Information. For carbon and nitrogen transport, we extracted genes annotated as having a role in nutrient movement (Supplementary Tables 1–4). Gene expression fold-change values (versus control values) for each sample for all primary transcripts can be found in Supplementary Tables 5 (plant) and 6 (fungus).
Statistical analyses
We first used a subset of data corresponding to microcosms inoculated with one isolate (no competitor) to check for differences among Pisolithus isolates. Differences in measured parameters were analysed with general linear models (GLMs) that included isolate (R10, SI12 or SI9) as a fixed factor. To relate atom excess results with RNA-sequencing data, we assessed the correlations between 15N content or 13C content and the coordinates of each sample in the first two axes of a principal coordinates analysis (PCoA) with Bray–Curtis similarity index using the expression levels of either nitrogen or carbon transporter genes in the fungal and plant transcriptomes.
We then assessed differences induced by the presence of a competitor per each of the isolates independently. Differences were analysed with GLMs including competition (none, indirect, direct) as a fixed factor. Correlations between variables were assessed using Pearson's coefficients. All statistical analyses were performed with R (R Development Core Team, 2016) using the interface implemented in the InfoStat software (Di Rienzo et al., 2012), except for the PCoA that was run in the Past software (Hammer et al., 2001). Additional details on statistical analyses are provided in Supplementary Information, F- and P-values for GLMs are shown in Supplementary Table 7 and R- and P-values for correlations in Supplementary Table 8.
Results
Root colonisation and nutrient exchange with the different EMF isolates in the absence of a competitor
When isolates R10, SI12 and SI9 were given access to labelled nitrogen, they all showed similar uptake of 15NH4+, although there was a trend suggesting that uptake by the mycelium of SI9 could be the highest (Figure 2a, P=0.08). Plants colonised by either SI12 or SI9 had higher leaf 15N content than plants colonised by R10 (Figure 2b), indicating that SI12 and SI9 were better at transferring nitrogen to the plant. Isolate R10 also had the highest percentage of nitrogen in its mycelium when in contact with host roots, supporting that it is able to uptake nitrogen but that it retains this nitrogen rather than transfer it to its host (Figure 2c). Despite differences in nitrogen transfer ability, all isolates received similar amounts of 13C from the plant (Figure 2d). We tested 13C enrichment in a few colonised root tips and they had similar values of enrichment as roots of control non-inoculated seedlings (data not shown), indicating that 13C was not retained in the mycorrhizas. There was no difference in overall % nitrogen in plant leaves or in leaf biomass between plants colonised by the three isolates (Supplementary Figures 2a and b).
Mean (a) mycelium 15N content, (b) leaf 15N mycelium content, (c) mycelium nitrogen percentage, (d) mycelium 13C content, (e) number of mycorrhizas and (f) mycelium mass in microcosms inoculated with isolate R10, SI12 or SI9. Different letters in a graph indicate significant differences (P<0.05) among treatments by GLM and DGC post hoc comparison. Data are mean±1 s.e.
Isolate R10 formed the highest number of mycorrhizas, followed by SI12 and SI9 (Figure 2e). R10 also formed the largest colonies (Supplementary Figure 2c), but its mycelium was much less dense than the other two isolates (that is, mycelium mass per cm2 was the lowest; Supplementary Figure 2d). Thus, overall, colony mass was similar for the three isolates (Figure 2f). Contents of leaf 15N and mycelium 13C were not significantly correlated with either mycelium mass or number of mycorrhizas. Therefore, an isolate forming a larger number of mycorrhizas does not necessarily result in getting more carbon or providing more nitrogen.
Regulation of plant and fungal nutrient transporter genes in the absence of a competitor
RNA was sequenced from the mycelium and mycorrhizal root tips of R10, SI12 and SI9 to determine the expression of plant and fungal nutrient transporters. The mycelium of isolate R10 showed the lowest expression of genes encoding nitrogen importers, while isolates SI9 and SI12 expressed different classes of nitrogen importers (Figure 3a). R10 colonised root tips showed much lower expression of two predicted amino-acid/nitrate efflux fungal genes (Pismi.96126 and Pismi.686226) when compared with SI9 and SI12 (Figure 3b, section i). Roots colonised by SI9 and SI12 showed increased gene expression of different plant nitrogen transporters such as oligopeptide and lysine/histidine importers (arrows for SI9 and regulon EucNC for SI12 in Figure 3b, section ii). These results fit well with the lowest nitrogen transfer to the plant by isolate R10 and are supported by significant correlations between mycelium 15N content and the overall expression of fungal nitrogen importers as well as between leaf 15N content and the expression of fungal N efflux genes (expressed as axes of the corresponding PCoAs).
(a) Average relative expression of nitrogen transport genes in the fungal transcriptome of colonies growing in pure culture. (b) Nitrogen efflux genes of each fungal isolate (i) and nitrogen transport genes of E. grandis (ii) in root tips colonised by individual Pisolithus isolates; (c) carbon transport genes in the fungal (i) and plant (ii) transcriptome of colonised root tips. Results are expressed as log 2 fold change. Genes in regulon EucNA, EucNB and EucNc were predominantly upregulated, repressed or highly expressed in SI12, respectively. Regulons PisoCi and PisoCii show genes induced in SI9 or SI12, respectively, while regulons EucCA, EucCB and EucCC show genes highly expressed in R10, SI9 or SI12, respectively.
Isolates SI12 and SI9 had opposing groups of fungal carbon transporter genes that were induced when colonising E. grandis (regulons PisoCi versus PisoCii in Figure 3c, section i). The individual Pisolithus isolates induced very distinct sets of carbon transporters in the host plant that were almost unique to each isolate (Figure 3c, section ii, regulons EucCA, EucCB and EucCC). We found a significant correlation between mycelium 13C content and the overall expression of plant carbon transporters (second axis of the PCoA).
Changes in nutrient exchange and colonisation induced by the presence of a competitor
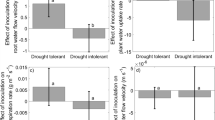
When more than one Pisolithus isolate was introduced to the same root system competition effects were seen. Isolate R10 delivered significantly more nitrogen to the plant when it was in direct competition with another isolate, as opposed to colonising alone or in indirect competition (Figure 4a). Mycelium of SI9 showed higher 13C content when paired with another isolate in either indirect or direct competition than when alone (Figure 4b). Overall, there was no significant correlation between leaf 15N content and mycelium 13C content (Figure 5).
Mean (a) leaf 15N content, (b) mycelium 13C content and (c) number of mycorrhizas in plants inoculated with isolate R10, SI12 or SI9 in the absence of competition (none), in indirect or direct competition. Each vertical panel corresponds to a given isolate as indicated by the top headings. Different letters in a graph indicate significant differences (P<0.05) among competition treatments per each isolate by GLM and DGC post hoc comparison. NS represents no significant differences among competition treatments. Data are mean±1 s.e.
Isolate R10 formed fewer mycorrhizas when another isolate was present in either indirect or direct competition than alone (Figure 4c). No differences in the number of mycorrhizas under indirect versus direct competition were observed for any of the isolates.
Differential regulation of plant and fungal nutrient transporter genes induced by the presence of a competitor
Compared to when alone, we found increased expression of two Pisolithus nitrogen efflux transporters in R10 under direct competition and in SI12 under indirect competition (Supplementary Figure 3a, section i). These were the same nitrogen efflux transporter genes that were highly expressed in SI9 when alone. The host exhibited differential regulation of both nitrogen and carbon transporter genes in response to the presence of two isolates compared with one isolate only (Supplementary Figures 3a and b, sections ii).
Plant defence-related genes differentially regulated during fungal competition
The presence of a competitor induced a reduction in the number of mycorrhizas formed by isolate R10 and we found differential expression in 953 plant genes in roots colonised by R10 in competition compared with root colonised by R10 alone. Some of these were also differentially regulated in roots colonised by either SI9 or SI12 in competition, while a smaller subset of 145 genes were uniquely regulated in R10 colonised roots (Figure 6a and Supplementary Table 9). These genes encode proteins from a variety of functional categories, including 12 disease resistance proteins (including NB-ARC domain containing and TIR-NBS-LRR class) and pathogenesis-related genes (Figure 6b). Thirty of these 145 genes show protein family (PFAM) enrichment in the specific areas of ‘defence response, incompatible interaction’ or ‘regulation of plant-type hypersensitive response’ (false discovery rate <0.001; Supplementary Table 9).
(a) Relative expression (log 2 fold change) of 145 genes differentially regulated in plants colonised by R10 when in indirect or direct competition compared with when colonised by R10 alone and uniquely regulated when colonised by R10 compared with SI9 and SI12 in competition. (b) Functional classification of these 145 genes.
Discussion
No link found between nitrogen and carbon allocation in EM symbiosis under non-limiting nitrogen conditions
We found no overall significant correlation between the amount of nitrogen delivered to the plant and the carbon allocated to any of the Pisolithus isolates tested (Figure 5). ‘Bad’ (R10) and ‘good’ (SI12 and SI9) symbionts, in terms of nitrogen provision, received similar amounts of 13C from the host plant (Figure 2). While it is possible that carbon was actually transferred but respired before harvesting took place (4 days), other studies have used a similar timeframe to allow for carbon transfer and detection in the fungal tissue (Högberg et al., 2008; Grelet et al., 2009; Kiers et al., 2011; Fellbaum et al., 2012; Pickles et al., 2016).
Our results support recent suggestions that nitrogen flux to the plant is not the main control on carbon flux to the fungus in the EM symbiosis (Corrêa et al., 2008, 2012; Albarracín et al., 2013; Valtanen et al., 2014). Because of the environmental dependence of mutualism, these results might have been different under experimental conditions where nitrogen was limiting. Bever (2015) predicted that in a fertile system such as ours, symbionts evolve toward reduced mutualism. However, the absence of correlation between carbon allocated to EMF and nitrogen transferred to the plant has been also described in nitrogen-limited systems (Hasselquist et al., 2016).
Differences in expression of specific carbon/nitrogen shuttling genes explain the outcome of EMF–plant nutrient fluxes
The high import of 15N label by the exploratory hyphae of SI9 was linked to high expression of an ammonium importer, while the other isolates appeared to depend upon other nitrogen transporters (Figure 3). These results reinforce previous work that demonstrated that different EMF isolates exhibit differences in nitrogen use (Anderson et al., 1999; Lilleskov et al., 2002; Pena and Polle, 2014). R10 had the lowest expression of nitrogen importers of the isolates tested. Its reduced capacity to absorb nitrogen from its surroundings may explain why it is the least cooperative of the symbionts tested in terms of nitrogen delivery to the host. Given that nitrogen transfer to the host plant is such a crucial part of the EM symbiosis, it is interesting that only two fungal amino-acid efflux transporters were induced in mycorrhizal root tips (Figure 3). A similar restriction in the repertoire of nitrogen transporters induced during symbiosis has been observed in the interaction between Laccaria bicolor and its host Populus trichocarpa (Plett et al., 2015). Of interest, hosts with the greatest 15N label in the leaves also exhibited the largest contingent of induced amino-acid transporters, underlining the importance of amino acids as a nutrient vector during EM symbiosis. Overall, our findings contribute to better understanding the regulation of transport at symbiotic interfaces (Garcia et al., 2016).
Despite similarities in total 13C recovered in the mycelium of isolates SI12 and R10, we found that the two isolates were characterised by nearly opposing host carbon transporter expression profiles in mycorrhizal root tips (Figure 3). This would suggest that there is redundancy in the roles of host carbon transporters during EM symbiosis. The amount of 13C in mycelium was positively correlated with the overall expression pattern of plant carbon transporter genes, while the amount of 15N in leaf was correlated with fungal nitrogen transporter genes expression. This might suggest that the partner that provides the benefit has greater control over it than the partner that receives it.
The ability to form mycorrhizas is not related to the effectiveness of EMF–plant nutrient exchange
The isolate that was able to form the largest number of mycorrhizas (R10) was neither the isolate that provided the most nitrogen to the host nor the one that received the greatest quantity of carbon from the plant (Figure 2). Bever (2015) proposes that because host plants have to invest in all potential symbionts before identifying which symbiont provides the best reward, one might expect fungal cheaters to specialise in rapid colonisation of new roots to best capture plant investment in symbiosis initiation. This strategy best describes isolate R10, which invests less energy in forming dense mycelium (Supplementary Figure 2) and could therefore be allocating more resources to form mycorrhizas than the other two isolates. It also has the highest nitrogen content, suggesting that it might retain the nitrogen for itself (Figure 2). It would be of interest to further investigate whether these traits in EMF correlate with a propensity to be a mycorrhizal cheater.
EMF competition can improve mutualistic nutrient exchange with the host plant
Plants received more nitrogen from the ‘cheater’ isolate R10 when this isolate was directly competing with another isolate (Figure 4). This increase in nitrogen provision to the plant corresponded with the upregulation of genes responsible for amino-acid efflux (Supplementary Figure 3). Argüello et al. (2016) also found that plants received more phosphorus from less-cooperative arbuscular mycorrhizal fungi when another fungus was present and the model by Franklin et al. (2014) suggests that plant discrimination induces fungal competition for carbon, thus enhancing nitrogen uptake. Our data suggest a plastic response whereby a typically uncooperative EMF isolate would alter behaviour to become more cooperative when under competition with a more beneficial isolate. The fact that this only happened when the two isolates were competing directly might suggest that this response is triggered by the direct interaction with another fungus. The other two fungal isolates tested, SI9 and SI12, did not significantly increase the amount of nitrogen delivered to the plant when placed in direct competition with another isolate. This may be because they were already delivering a fair quantity of nitrogen to the plant, or the results seen with R10 may not be universal. Additional tests with more isolates would be required to see if the results with R10 are more generally applicable. Changes in carbon allocation due to the presence of multiple EMF (Figure 4) were also observed, but again further investigation is needed.
Host plants specifically target defensive signalling to root segments colonised by less-cooperative symbionts
Our findings suggest that fungal success in colonising root tips is controlled at least in part by the plant, because both indirect and direct fungal–fungal interactions had a similar competitive outcome (Figure 4). Otherwise, we would have expected a reduction when competing directly but not when competing indirectly (Kennedy, 2010). Isolate R10 (the least cooperative isolate tested) exhibited significantly reduced colonisation levels when another isolate was present. Similar results have been reported for rapidly growing cheater rhizobial symbionts (Sachs et al., 2010). However, our patterns of carbon allocation did not explain this reduction.
Instead, our results suggest that plants may avoid EMF cheaters by upregulating defence-related genes (Figure 6). We found that there was a significant over-representation of upregulated defence-related genes in roots colonised by R10, but not in other parts of the same root system simultaneously colonised by either SI9 or SI12. These same defence genes were not upregulated in R10 colonised roots in the absence of a competitor, indicating that the plant may only actively reject unhelpful symbionts if another more favourable option is available. Interestingly, the plant did not completely repudiate all colonisation by R10, despite its comparatively poor ability to support plant nutrition. This would support previous work showing that plants are not fully immune to colonisation by less beneficial fungi (Douglas, 2008; Kiers and Denison, 2008; Grman, 2012). Alternatively, this isolate could be providing other benefits to the host plant not accounted here. In any case, the plant defensive response and corresponding reduction in the number of mycorrhizas did not significantly reduce the amount of carbon transferred to R10 (Figure 4), supporting the idea that carbon might not be a cost for the plant (Corrêa et al., 2012). We did not detect upregulation of fungal genes related to production of anti-fungal compounds when in competition (data not shown), which provides additional support for plant control over colonisation rather than fungal control.
Conclusions
We found that, when given a choice of partners, the plant had the ability to limit root tip colonisation by the least cooperative symbiont and therefore influence the outcome of EMF competition. Interestingly, this reduction in colonisation did not result in a reduction in carbon allocation to the fungus providing the lowest amount of nitrogen. Our results also suggest that EMF competition might encourage mutualistic behaviour in some circumstances. These nutrient exchange dynamics were closely related to the expression of key plant and fungal nutrient transporter genes that, overall, resulted in no link between the amount of nitrogen delivered to the host plant and the carbon allocated to the EM fungus. Instead, our results suggest that partner choice in EMF could operate through the upregulation of defence compounds that would prevent infection and discourage cheaters, a mechanism that deserves further investigation.
References
Albarracín MV, Six J, Houlton BZ, Bledsoe CS . (2013). A nitrogen fertilization field study of carbon-13 and nitrogen-15 transfers in ectomycorrhizas of Pinus sabiniana. Oecologia 173: 1439–1450.
Anderson IC, Chambers SM, Cairney JWG . (1999). Intra-and interspecific variation in patterns of organic and inorganic nitrogen utilization by three Australian Pisolithus species. Mycol Res 103: 1579–1587.
Argüello A, O’Brien MJ, van der Heijden MGA, Wiemken A, Schmid B, Niklaus PA . (2016). Options of partners improve carbon for phosphorus trade in the arbuscular mycorrhizal mutualism. Ecol Lett 19: 648–656.
Bever JD . (2015). Preferential allocation, physio-evolutionary feedbacks, and the stability and environmental patterns of mutualism between plants and their root symbionts. New Phytol 205: 1503–1514.
Bever JD, Richardson SC, Lawrence BM, Holmes J, Watson M . (2009). Preferential allocation to beneficial symbiont with spatial structure maintains mycorrhizal mutualism. Ecol Lett 12: 13–21.
Corrêa A, Gurevitch J, Martins-Loução MA, Cruz C . (2012). C allocation to the fungus is not a cost to the plant in ectomycorrhizae. Oikos 121: 449–463.
Corrêa A, Strasser RJ, Martins-Loução MA . (2008). Response of plants to ectomycorrhizae in N-limited conditions: which factors determine its variation? Mycorrhiza 18: 413–427.
Di Rienzo JA, Casanoves F, Balzarini MG, Gonzalez L, Tablada M, Robledo CW . (2012), InfoStat version 2012. InfoStat Group, College of Agricultural Sciences, Nacional University of Córdoba, Argentina. Available at: http://www.infostat.com.ar.
Douglas AE . (2008). Conflict, cheats and the persistence of symbioses. New Phytol 177: 849–858.
Fellbaum CR, Gachomo EW, Beesetty Y, Choudhari S, Strahan GD, Pfeffer PE et al. (2012). Carbon availability triggers fungal nitrogen uptake and transport in arbuscular mycorrhizal symbiosis. Proc Natl Acad Sci USA 109: 2666–2671.
Franklin O, Näsholm T, Högberg P, Högberg MN . (2014). Forests trapped in nitrogen limitation—an ecological market perspective on ectomycorrhizal symbiosis. New Phytol 203: 657–666.
Garcia K, Doidy J, Zimmermann SD, Wipf D, Courty P-E . (2016). Take a trip through the plant and fungal transportome of mycorrhiza. Trends Plant Sci 21: 937–950.
Grelet G-A, Johnson D, Paterson E, Anderson IC, Alexander IJ . (2009). Reciprocal carbon and nitrogen transfer between an ericaceous dwarf shrub and fungi isolated from Piceirhiza bicolorata ectomycorrhizas. New Phytol 182: 359–366.
Grman E . (2012). Plant species differ in their ability to reduce allocation to non-beneficial arbuscular mycorrhizal fungi. Ecology 93: 711–718.
Hammer Ø, Harper DAT, Ryan PD . (2001). PAST: Paleontological statistics software package for education and data analysis. Palaeontol Electronica 4: 9.
Hasselquist NJ, Metcalfe DB, Inselsbacher E, Stangl Z, Oren R, Näsholm T et al. (2016). Greater carbon allocation to mycorrhizal fungi reduces tree nitrogen uptake in a boreal forest. Ecology 97: 1012–1022.
Hoeksema JD, Kummel M . (2003). Ecological persistence of the plant–mycorrhizal mutualism: a hypothesis from Species Coexistence Theory. Am Nat 162: S40–S50.
Högberg P, Högberg MN, Göttlicher SG, Betson NR, Keel SG, Metcalfe DB et al. (2008). High temporal resolution tracing of photosynthate carbon from the tree canopy to forest soil microorganisms. New Phytol 177: 220–228.
Hortal S, Pera J, Parladé J . (2008). Tracking mycorrhizas and extraradical mycelium of the edible fungus Lactarius deliciosus under field competition with Rhizopogon spp. Mycorrhiza 18: 69–77.
Hortal S, Powell JR, Plett JM, Simonin A, Anderson IC . (2016). Intraspecific competition between ectomycorrhizal Pisolithus microcarpus isolates impacts plant and fungal performance under elevated CO2 and temperature. FEMS Microbiol Ecol 92: fiw113.
Kennedy P . (2010). Ectomycorrhizal fungi and interspecific competition: species interactions, community structure, coexistence mechanisms, and future research directions. New Phytol 187: 895–910.
Kennedy PG, Hortal S, Bergemann SE, Bruns TD . (2007). Competitive interactions among three ectomycorrhizal fungi and their relation to host plant performance. J Ecol 95: 1338–1345.
Kennedy PG, Peay KG, Bruns TD . (2009). Root tip competition among ectomycorrhizal fungi: are priority effects a rule or an exception? Ecology 90: 2098–2107.
Kiers ET, Denison RF . (2008). Sanctions, cooperation, and the stability of plant–rhizosphere mutualisms. Annu Rev Ecol Evol Syst 39: 215–236.
Kiers ET, Duhamel M, Beesetty Y, Mensah JA, Franken O, Verbruggen E et al. (2011). Reciprocal rewards stabilize cooperation in the mycorrhizal symbiosis. Science 333: 880–882.
Kiers ET, Rousseau RA, West SA, Denison RF . (2003). Host sanctions and the legume–rhizobium mutualism. Nature 425: 78.
Lilleskov EA, Hobbie EA, Fahey TJ . (2002). Ectomycorrhizal fungal taxa differing in response to nitrogen deposition also differ in pure culture organic nitrogen use and natural abundance of nitrogen isotopes. New Phytol 154: 219–231.
Näsholm T, Högberg P, Franklin O, Metcalfe D, Keel SG, Campbell C et al. (2013). Are ectomycorrhizal fungi alleviating or aggravating nitrogen limitation of tree growth in boreal forests? New Phytol 198: 214–221.
Pena R, Polle A . (2014). Attributing functions to ectomycorrhizal fungal identities in assemblages for nitrogen acquisition under stress. ISME J 8: 321–330.
Pickles BJ, Wilhelm R, Asay AK, Hahn AS, Simard SW, Mohn WW . (2016). Transfer of 13C between paired Douglas-fir seedlings reveals plant kinship effects and uptake of exudates by ectomycorrhizas. New Phytol 214: 400–411.
Plett JM, Tisserant E, Brun A, Morin E, Grigoriev IV, Kuo A et al. (2015). The mutualist Laccaria bicolor expresses a core gene regulon during the colonization of diverse host plants and a variable regulon to counteract host-specific defenses. Mol Plant Microbe Interact 28: 261–273.
R Development Core Team R. (2016). A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. ISBN 3-900051-07-0. Available at: http://www.R-project.org.
Rasband WS . (1997) ImageJ. US National Institutes of Health: Bethesda, MD, USA. Available at: http://imagej.nih.gov/ij/.
Sachs JL, Russell JE, Lii YE, Black KC, Lopez G, Patil AS . (2010). Host control over infection and proliferation of a cheater symbiont. J Evol Biol 23: 1919–1927.
Simms EL, Taylor DL, Povich J, Shefferson RP, Sachs JL, Urbina M et al. (2006). An empirical test of partner choice mechanisms in a wild legume–rhizobium interaction. Proc R Soc Lond Ser B 273: 77–81.
Smith SE, Read DJ . (2008) Mycorrhizal symbiosis, 3rd edn. Repr. Elsevier/Academic Press: Amsterdam, Netherlands.
Tedersoo L, May TW, Smith ME . (2010). Ectomycorrhizal lifestyle in fungi: global diversity, distribution, and evolution of phylogenetic lineages. Mycorrhiza 20: 217–263.
Valtanen K, Eissfeller V, Beyer F, Hertel D, Scheu S, Polle A . (2014). Carbon and nitrogen fluxes between beech and their ectomycorrhizal assemblage. Mycorrhiza 24: 645–650.
Walder F, van der Heijden MGA . (2015). Regulation of resource exchange in the arbuscular mycorrhizal symbiosis. Nat Plants 1: 15159.
West SA, Kiers ET, Simms EL, Denison RF . (2002). Sanctions and mutualism stability: why do rhizobia fix nitrogen? Proc R Soc Lond Ser B 269: 685–694.
Wu B, Nara K, Hogetsu T . (1999). Competition between ectomycorrhizal fungi colonizing Pinus densiflora. Mycorrhiza 9: 151–159.
Acknowledgements
Work conducted with the support of an Australian Institute of Nuclear Science and Engineering (AINSE) Research Grant (ALNGRA14531) plus additional funding from the Hawkesbury Institute for the Environment (Western Sydney University) and an Australian Research Council Discovery project (DP160102684). We greatly thank J Berenguel for the design and construction of the chamber used for isotope labelling and J King and C Janitz at the Western Sydney University Next Generation Sequencing Facility for RNA-sequencing services and support. We thank G Skrzypek, D Ford, J Harrison, B Gallagher, G Grelet, B Pickles, C Mitchell, A Bass, M Queralt, R Wright, C Aguirre and G McKenzie for invaluable technical advice and assistance. We also thank F Martin, D Johnson and J Walker for fruitful discussions and suggestions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Hortal, S., Plett, K., Plett, J. et al. Role of plant–fungal nutrient trading and host control in determining the competitive success of ectomycorrhizal fungi. ISME J 11, 2666–2676 (2017). https://doi.org/10.1038/ismej.2017.116
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.116
This article is cited by
-
Community composition of phytopathogenic fungi significantly influences ectomycorrhizal fungal communities during subtropical forest succession
Applied Microbiology and Biotechnology (2024)
-
Tropical tree ectomycorrhiza are distributed independently of soil nutrients
Nature Ecology & Evolution (2024)
-
Mycorrhizal C/N ratio determines plant-derived carbon and nitrogen allocation to symbiosis
Communications Biology (2023)
-
Host population effects on ectomycorrhizal fungi vary between low and high phosphorus soils of temperate rainforests
Mycorrhiza (2023)
-
Combining ectomycorrhizal fungi and plant growth-promoting rhizobacteria to enhance salt tolerance of Metasequoia glyptostroboides
Journal of Forestry Research (2023)