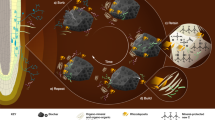
Abstract
Pyrogenic organic matter (PyOM) additions to soils can have large impacts on soil organic carbon (SOC) cycling. As the soil microbial community drives SOC fluxes, understanding how PyOM additions affect soil microbes is essential to understanding how PyOM affects SOC. We studied SOC dynamics and surveyed soil bacterial communities after OM additions in a field experiment. We produced and mixed in either 350 °C corn stover PyOM or an equivalent initial amount of dried corn stover to a Typic Fragiudept soil. Stover increased SOC-derived and total CO2 fluxes (up to 6x), and caused rapid and persistent changes in bacterial community composition over 82 days. In contrast, PyOM only temporarily increased total soil CO2 fluxes (up to 2x) and caused fewer changes in bacterial community composition. Of the operational taxonomic units (OTUs) that increased in response to PyOM additions, 70% also responded to stover additions. These OTUs likely thrive on easily mineralizable carbon (C) that is found both in stover and, to a lesser extent, in PyOM. In contrast, we also identified unique PyOM responders, which may respond to substrates such as polyaromatic C. In particular, members of Gemmatimonadetes tended to increase in relative abundance in response to PyOM but not to fresh organic matter. We identify taxa to target for future investigations of the mechanistic underpinnings of ecological phenomena associated with PyOM additions to soil.
Similar content being viewed by others
Introduction
Large inputs of pyrogenic organic matter (PyOM) in fire-affected ecosystems can constitute up to 80% of total soil organic carbon (SOC) (Lehmann et al., 2008). Whether PyOM is produced naturally in fires (Czimczik and Masiello, 2007), intentionally for carbon (C) management and/or as an agricultural amendment (Lehmann, 2007; Laird, 2008), it is important to understand how it affects the C cycle (Whitman et al., 2010). PyOM additions to soil can significantly affect plant growth and crop yields (Jeffery et al., 2011) and SOC dynamics (Maestrini et al., 2014; Watzinger et al., 2014; Whitman et al., 2015). For example, among other effects, PyOM can change soil pH (Gul et al., 2015), add nutrients to soils (Enders et al., 2012) and alter soil water-holding capacity (Abel et al., 2013). Although PyOM additions to soils can alter inorganic C dynamics through changes to soil pH, a key mechanism for changes in SOC mineralization is altered microbial mineralization of SOC in response to PyOM (Maestrini et al., 2014; Whitman et al., 2015). Hence, an understanding of the effects of PyOM on the soil microbial community composition and function is required to understand the effects of PyOM on SOC stocks.
There are many ways PyOM could affect soil microorganisms (Kuzyakov and Bol, 2004; Lehmann et al., 2011; Ameloot et al., 2013), including, but not limited to, microbial use of PyOM as a source of energy or nutrients, changes in soil physical or chemical characteristics, impacts on plant growth and interference with microbial signaling (Masiello et al., 2013). Recent research has only begun to identify PyOM effects on soil microbial communities, and it is clear that PyOM additions to soil can induce changes in soil microbial community composition. Most current evidence has been gathered using fingerprinting approaches, such as terminal restriction fragment length polymorphism (Bingeman et al., 1953; Jin, 2010; Kolton et al., 2011) or denaturing gradient gel electrophoresis (Kolton et al., 2011; Chen et al., 2013), or by surveying phospholipid fatty acids to assess microbial diversity at low phylogenetic resolution (Dunavin, 1969; Jindo et al., 2012; Gomez et al., 2014; Watzinger et al., 2014; Mitchell et al., 2015). In addition, clone libraries targeting functional genes (Song et al., 2014) and some high-throughput DNA sequencing approaches have been applied to survey microorganisms in PyOM systems. For example, significant differences were found between soil bacterial communities in Amazonian Dark Earth soils (amended with PyOM thousands of years ago), PyOM isolated from the Amazonian Dark Earth soils and adjacent unamended Acrisols, by surveying small subunit ribosomal RNA (SSU rRNA) genes (Taketani et al., 2013). A handful of other studies have used high-throughput sequencing approaches to characterize PyOM effects on soil bacterial community composition (for example, Nielsen et al., 2014; Xu et al., 2014). However, owing to the low number of studies and the wide diversity of PyOM materials, addition rates, soils and environmental conditions, it is difficult to draw generalizable conclusions about the effects of PyOM on soil microbial community composition, particularly at the level of individual taxa.
In this study, we investigated the effects of PyOM additions on SOC mineralization and soil bacterial community composition in a field setting over 12 weeks. We also included a treatment where plots received a mass of fresh corn stover equivalent to the mass required to produce the PyOM we added. This treatment serves as a system-level control that addresses the question, ‘What if a given amount of biomass were not used to produce PyOM, but were applied directly to the soil?’ We predicted that there would be significant differences in C dynamics between the two systems, with fresh biomass decomposing faster and having a greater effect on SOC mineralization. In addition, we predicted that the addition of fresh biomass would induce the greatest changes in the bacterial community composition, with organisms that access easily mineralizable C sources proliferating initially, and the organisms that are able to decompose aromatic or insoluble substrates, such as lignin or cellulose, emerging later. We expected that the majority of the microorganisms that increase in relative abundance in response to PyOM additions will also increase in response to fresh organic matter additions, but they will respond to PyOM to a smaller degree, as a smaller fraction of the PyOM-C is easily mineralizable.
Materials and methods
Experimental design
We conducted a field trial, with soil left unamended, soil amended with 13C-labeled 350 °C corn stover (Zea mays (L.)) PyOM or soil amended with fresh corn stover (Supplementary Tables 1 and 2). The 13C label allowed for the C sources to be partitioned between SOC and PyOM-C or corn stover-derived C (Whitman and Lehmann, 2015). The corn stover addition was designed so that the dried original corn biomass was equivalent to that which would have been required to produce the mass of corn PyOM that was applied to each plot (that is, 4.1 Mg ha−1 corn-derived PyOM were applied, which, with 0.365 mass fraction conserved during PyOM production, translates into 11.2 Mg ha−1 corn stover, which is representative of a productive corn crop in the United States (Shinners and Binversie, 2007)).
The field site is located in Cornell’s research fields in Mt Pleasant, NY, USA, and is a Mardin soil (coarse-loamy, mixed, active, mesic Typic Fragiudept) (Supplementary Table 2). The soil has been historically planted to a potato, rye, clover rotation, for the past >30 years, but was kept in rye-clover rotation for the past 5 years, with one planting of sudangrass 3 years ago. The plot was sprayed with Roundup (glyphosate) herbicide in the fall of 2012 (1.8 l ha−1), plowed on 3 May 2013, and kept weed-free by hand-weeding and water-permeable landscape fabric through the summer until trial initiation.
The trial initiation date was 16 August 2013 (day 0). Square plots (0.7 m × 0.7 m for the soil-only and PyOM, 0.45 m × 0.45 m for the stover) were surrounded by 0.7-m wide weed-free borders, maintained by hand weeding. Treatments were organized using a spatially balanced complete block design with 16 replicates of the unamended and PyOM plots, and 8 replicates of the stover plots (van Es et al., 2007). Soil (6.1 kg or ~1–2 cm) was removed from the surface of each plot, combined with stover or PyOM additions, if needed, and mixed in a V-mixer (Twin shell dry blender, Patterson-Kelley, East Stroudsburg, PA, USA). Amended plots received 4.1 Mg ha−1 of PyOM or 11.2 Mg ha−1 of dried original corn stover. After mixing, soils were returned to their respective plots and evenly spread at the surface. Soil was gently tamped down using a flat piece of plywood. Two soil respiration collars made from 194 mm diameter white polyvinylchloride pipes were installed directly adjacent to each other at the center of each plot with the collar protruding 30 mm and reaching 30 mm into the ground. Plots, including soil collars, were covered with water-permeable landscape fabric except during measurement for the first 2 weeks, after which fabric was removed and plots were kept weed-free by hand-weeding multiple times a week. The plots were not fertilized or watered during the trial, but were exposed to natural rainfall (temperature and precipitation are plotted in Supplementary Figure 1).
Biomass production
Two sets of corn plants (Zea mays (L.)) were grown, one in an enriched 13CO2 atmosphere growth chamber and the other in an ambient 13CO2 greenhouse. The labeled plants were grown in potting mix in a Percival AR-100L3 CO2-controlled growth chamber (Percival, Perry, IA, USA). The plants were exposed to cycles of 18-h light/6-h darkness. During light cycles, the atmosphere was maintained at 400 p.p.m. CO2. During the dark cycle, CO2 was allowed to accumulate through respiration, and was then drawn down by photosynthesis during the next light cycle. This was done in order to reduce net respiratory losses of labeled 13CO2. Plants were pulse labeled with 13 l of 99% 13CO2 at regular intervals over the course of their growth in order to produce an even label. Pulse labels were delivered by opening the 13CO2 cylinder to fill a balloon with ~500 ml 13CO2. The balloon remained attached to the cylinder so that the 13CO2 slowly diffused out of the balloon, delivering the pulse at a rate so that the total atmospheric concentration of CO2 was not affected. Plants in the growth chambers and the greenhouse were harvested just before they reached reproductive maturity and were oven-dried at 70 °C.
PyOM production and amendment mixing
Oven-dried corn plants were ground in a hammer mill (Viking Manufacturing Company, Manhattan, KS, USA) to <2 mm. The milled corn was pyrolyzed in a modified Fisher Scientific Isotemp programmable muffle furnace (Thermo Fisher Scientific, Waltham, MA, USA) (described in detail in Guereña et al. 2015) by ramping at 5 °C min−1 to 350 °C, then holding at 350 °C for 45 min, under Ar (Supplementary Table 1). The 13C-labeled and natural abundance corn PyOM materials were mixed together to produce a δ13C value of +37.5‰. For the corn biomass-only plots, mixtures of 13C-labeled and natural abundance corn with a δ13C value of +1.7‰ were created. Mixing was done in plot-level batches to ensure that each plot received exactly these proportions of labeled and unlabeled materials.
CO2 flux and 13CO2 measurements
Soil CO2 flux was measured using a LI-6400XT infra-red gas analyzer with a 6400-09 soil CO2 flux chamber attachment (LI-COR, Lincoln, NE, USA). Three flux measurements were taken in succession for each plot and averaged. Measurements were taken on days 0, 1, 2, 3, 4, 5, 6, 7, 9, 11, 12, 14, 16, 18, 26, 30, 34, 38, 41, 45, 49, 53, 57, 62, 66, 74 and 81. Additional gas samples were taken for 13CO2 analysis and emissions partitioning on day 12 and on day 66 using modified static Iso-FD chambers (Nickerson et al., 2013; Whitman and Lehmann, 2015) (Supplementary Figure 2).
Soil sampling
Soil samples for bacterial community analyses were taken on days 1, 12 (chosen because it was after the first major rainfall; Supplementary Figure 1) and 82. Samples were sieved to <2 mm, and immediately frozen in Whirl-Paks in liquid N2 and stored at −80 °C. On days 1 and 12, two 25-mm deep soil probe samples were pooled, while on day 82, soil within the entire collar was destructively sampled.
Isotopic partitioning of CO2 samples
We partitioned the total CO2 fluxes between SOC and stover or PyOM-C on days 12 and 66. To determine the relative contributions of SOC and added C (either PyOM or corn stover) to soil CO2 fluxes, a standard isotope partitioning approach was applied (Balesdent and Mariotti, 1996). For example, for the plots with PyOM additions, the isotopic signature of the total emissions will be:

where δ represents the 13C signature of CO2 from total CO2 (δT), soil alone (δS), or PyOM (δPyOM) and fS and fPyOM represent the fraction of total emissions made up by soil and PyOM, respectively (Werth and Kuzyakov, 2010). As δT, δS, and δPyOM were all measured, and we know

we can solve this system of two equations for the two unknown values, fS and fPyOM.
Data processing and statistical analyses for CO2 fluxes
On rare occasions toward the end of the trial, we recorded negative CO2 fluxes, which we interpret as experimental error because of low flux rates, and have excluded these values from analyses. In addition, we excluded two data points where recorded fluxes were 56 and 16 s.d. away from the mean of the remaining plots. All statistical analyses were performed in R (R Core Team, 2015). CO2 fluxes were evaluated using a linear mixed effects model, with amendment, day, interaction between amendment and day, and plot ID (a repeated-measures approach) as factors, using the R package ‘lme4’ (Bates et al., 2015). To make post-hoc comparisons, we performed pairwise comparisons between the different soil amendments for a given day with a Tukey's adjustment of P-values, using the ‘lsmeans’ R package (Lenth, 2014).
Bacterial community analyses
DNA was extracted from 0.25 g moist soil samples using the MoBio PowerLyzer PowerSoil kit (Carlsbad, CA, USA), following the kit’s directions. The DNA was quantified using the Quant-iT PicoGreen dsDNA Assay Kit (Life Technologies, Madison, WI, USA) with a multimode microplate reader (Molecular Devices, Sunnyvale, CA, USA). DNA yields were normalized on the basis of grams of dry soil extracted. SSU rRNA genes were PCR amplified in triplicate for each sample. PCR was conducted with 12.5 μl Q5 Hot Start High Fidelity 2X mastermix (New England Biolabs, Ipswich, MA, USA), 5 μl of DNA template diluted with water at a ratio of 1:50, 5 μl water and 2.5 μl primer mixtures to a total volume of 25 μl. Each PCR consisted of a 98 °C hold for 30 s, followed by 30 cycles of (5 s at 98 °C, 20 s at 20 °C and 10 s at 72 °C), with a final extension for 2 min at 72 °C. Modified 515F and 907R primers (Supplementary Tables 3 and 4) were used to target the V4/V5 regions of the 16S ribosomal RNA gene. Unique barcodes were added to the primers used for each sample so that the SSU rRNA sequences from each sample could be demultiplexed post sequencing (Supplementary Tables 3 and 4). Each PCR product was run on a 0.5% agarose gel, along with the negative control, to determine whether amplification was successful. Replicate PCR products were pooled, and DNA concentrations were normalized across all samples using SequalPrep normalization plates (Applied Biosystems, Foster City, CA, USA). The pooled sample was purified using a Wizard SV Gel and PCR Clean-Up System (Promega, Fitchburg, WI, USA). This sample was submitted with sequencing primers (Supplementary Table 5) for paired ends 2 × 300-bp sequencing on the Illumina MiSeq v3 platform (Illumina, Inc., San Diego, CA, USA) at Cornell’s Biotech Core Facility.
Bacterial community bioinformatics
Paired read merging, demultiplexing and quality control
We merged the forward and reverse reads using the paired end read merger (Zhang et al., 2014) and demultiplexed them by barcode. DNA sequences were managed using screed databases (Nolley and Brown, 2012).
We removed merged reads with more than one expected error using USEARCH (Edgar, 2013). We also removed any sequences that had ambiguous base calls. We further identified erroneous sequences with alignment based quality control (Schloss et al., 2009), by aligning our sequences to the SILVA Reference Alignment as provided by the Mothur developers and removing reads that did not align to the expected region of the SSU rRNA gene. We then removed any sequences that were <370-bp or >376-bp long or had homopolymers (runs of the same base in a row) >8 nucleotides in length. Quality control removed 6 265 604 reads, leaving 10 237 689 high-quality sequences. With 3 treatments, 2 with 16 replicates and 1 with 8 replicates, and 3 time-points, we had a total of 120 samples. Individual samples contained from 8830 to 194 356 total sequences, with a mean of 69 419±44 070.
Operational taxonomic unit (OTU) picking
Reads were clustered into OTUs using the UPARSE methodology (Edgar, 2013) with an OTU sequence identity cutoff of 97%. In the UPARSE workflow, chimeras are detected and discarded when OTU centroids are selected. Of the quality-controlled reads, 81% could be mapped to OTU centroids. We taxonomically annotated OTU centroids using the ‘uclust’ based taxonomic annotation framework in QIIME (v1.8) (Caporaso et al., 2010) with default parameters (Caporaso et al., 2010; Edgar, 2013). OTUs annotated as ‘Archaea’, ‘mitochondria’ or ‘Eukarya’ were removed in downstream analyses. One sample (an unamended plot from day 82) was left with only eight sequences, so it was excluded from further analyses.
Community analysis
OTU centroids were taxonomically annotated within the Greengenes taxonomic nomenclature using the ‘uclust’ taxonomic annotation framework in QIIME with default parameters (Caporaso et al., 2010). Centroids from 97% sequence identity clusters of Greengenes database SSU rRNA gene sequences (version 13_8) and corresponding annotations were used as reference for taxonomic annotation. We used the R package ‘vegan’ (Oksanen et al., 2015) to perform a nonmetric multidimensional scaling analysis on the weighted UniFrac distance between communities across all time points and amendments (Lozupone et al., 2011). Weighted UniFrac accounts for the relative abundance of each OTU, not just considering its presence/absence. We then tested whether there were significant effects on the bacterial community because of amendment types and day of sampling with a nonparametric multivariate analysis of variance (NPMANOVA) on weighted UniFrac distances, using the ‘adonis’ function from the R package ‘vegan’ (Oksanen et al., 2015). As we found day and amendment both had significant effects (P<0.001), we performed a separate weighted UniFrac analysis and NPMANOVA for each day and amendment type (comparing each amendment with either each other or with no additions, for each day), adjusting P-values using a Bonferroni correction for six comparisons (Padj=P × 6).
We used the R package ‘DESeq2’ (Love et al., 2014) to calculate the differential abundance (log2-fold change in relative abundance of each OTU) for each amendment type as compared with the unamended plots for both sampling days (McMurdie and Holmes, 2014). We independently filtered out OTUs that were sparsely represented across samples (that is, those OTUs for which the DESeq2-normalized count across samples (‘baseMean’) was <0.6). Sparse OTUs will not contain sufficient sequence counts to provide statistically significant results and their removal reduces the number of multiple comparisons performed, thereby mitigating problems associated with multiple comparisons with some extent. We adjusted the P-values with the Benjamini and Hochberg correction method and selected a study-wide false discovery rate of 10% to denote statistical significance (Love et al., 2014). We defined ‘responding OTUs’ as OTUs with a differential abundance >1 and an adjusted P-value of <0.1. We performed a BLAST (nucleotide blast, version 2.2.29+, default parameters) search with OTU centroid sequences of responding OTUs, against the Living Tree Project database (version 115) (Yarza et al., 2008). The Living Tree Project database contains 16S rRNA gene sequences for all sequenced archaeal and bacterial type strains.
It is important to note that the detection of bacterial DNA in the soil does not indicate that these microbes are active, let alone necessarily alive (Blazewicz et al., 2013). In addition, although the detection of a significant increase in relative abundance against the diverse and abundant background soil community likely does indicate growth and cell replication, some organisms may increase their activity in response to OM inputs, but not grow or divide, thereby evading detection via this technique.
Results
Soil C dynamics
Total CO2 fluxes from plots that received uncharred stover additions were significantly (mixed model repeated-measures design, Tukey-adjusted post-hoc comparisons, P<0.05) higher than all other plots for the first 26 days and for 19 of the 27 days for which fluxes were determined (Figure 1). Stover additions had the greatest effects on CO2 fluxes (CO2 fluxes six times those of unamended soils) on days 7 and 11, after strong rain events (Supplementary Figure 1). Plots with PyOM additions experienced significantly higher CO2 fluxes than plots with no additions for the first 12 days, after which there were no significant differences (Figure 1). The increases in fluxes with PyOM additions were much less marked than those in the stover-amended plots, and were never >2.1 times the CO2 emissions from unamended soils.
Mean CO2 flux rates over time. Error bars ±1 s.e., n=8–16. Dotted line indicates plots that received fresh stover additions. Dashed orange line indicates plots that received PyOM additions. Solid yellow line indicates plots that had no additions. *Indicates significant differences between plots with PyOM additions and stover or no-addition plots, whereas ‡ indicates significant differences between plots that received fresh stover additions and PyOM or no-addition plots (mixed model repeated measures design, Tukey-adjusted post-hoc comparisons, P<0.05). X indicates days where microbial community was sampled. • indicates days where 13CO2 flux was partitioned between SOC and amendments. The color reproduction of this figure is available at the The ISME Journal online.
On day 12, 13C partitioning revealed that corn stover additions significantly increased SOC-derived CO2 fluxes as compared with soils with no additions, but PyOM additions did not (Supplementary Figure 3). On day 66, there were no significant differences in SOC-derived CO2 fluxes between all soils, and overall fluxes were much lower on this date (Figure 1 and Supplementary Figure 3).
Bacterial community analyses
We did not detect significant differences in DNA yield because of the addition of PyOM or stover directly after amendment additions (day 1), but DNA yield was significantly higher in plots with stover additions on days 12 and 82 (Supplementary Table 6; analysis of variance and Tukey’s honest significant difference, P<0.05). There were significant changes in the bacterial community composition over time (NPMANOVA, P<0.006, R2=0.19) and with amendment type (NPMANOVA, P<0.006, R2=0.20) (Figure 2). Bacterial community composition in unamended plots remained relatively consistent over time (NPMANOVAs comparing day 1 with day 12 (P<0.06, R2=0.18) and with day 82 (P<0.29, R2=0.11)), whereas PyOM-amended plots varied slightly and corn stover-amended plots varied greatly over time (Figure 2). The different treatments were not observed to alter bacterial community composition on day 1 (only 24 h after OM additions). By day 12, the stover treatment caused a significant change in community composition (NPMANOVA, P<0.006, R2=0.75), although no effect of PyOM was observed. By day 82, the bacterial communities in the PyOM-amended plots (NPMANOVA, P<0.02, R2=0.19) and corn stover-amended plots (NPMANOVA, P<0.02, R2=0.56) were both distinct from the unamended plots and from each other (NPMANOVA, P<0.01, R2=0.36).
The stover treatment caused multiple phyla to change significantly in relative abundance as compared with control plots on days 12 and 82 (Figure 3). In contrast, PyOM additions only led to a significant change in the relative abundance of the phyla Armatimonadetes (decreased), and Bacteroidetes (increased) and only on day 82 (Figure 3). However, we caution that decreases in relative abundance do not necessarily correspond to decreases in absolute abundance—rather, they could result from an increase in the absolute abundance of other phyla. Hence, we transformed the relative abundance data from stover treatments by scaling by the mass of extracted DNA per soil sample, but found that the only changes were that declines in relative abundance for Acidobacteria and Gemmatimonadetes with stover additions on day 82 were no longer statistically significant (Supplementary Note 1).
Relative abundance (unscaled by total microbial DNA) of top 12 phyla observed in the unamended, stover-amended and PyOM-amended soils (n=8–16). Values beyond 1.5 times the interquartile range are indicated (x), as are values that differ significantly from unamended (*, t-test, P<0.05, Bonferroni-corrected for 72 comparisons).
Many OTUs, from several phyla, responded significantly to stover and/or PyOM additions (Figure 4; Supplementary Tables 7 and 8; Supplementary Figures 6–14 and 20). We use the term ‘responders’ to refer specifically to those OTUs that increase significantly in relative abundance by more than doubling in response to stover and/or PyOM additions as compared with corresponding plots that did not receive amendments. We identified 806 responders to either stover and/or PyOM from among the 7770 total OTUs observed across all soil samples. There were more responders to stover (677 OTUs) than to PyOM (264 OTUs) (Figures 2, 4 and 5). A total of 8% of all OTUs responded specifically to stover (Figure 5, blue region), 2% responded specifically to PyOM (Figure 5, pink region), and 2% responded to both PyOM and stover (Figure 5, purple region and Supplementary Figures 4 and 5). The total number of responders to both stover and PyOM increased over time and nearly all phyla had more OTUs respond at day 82 than day 12. The only notable exception is Firmicutes, which had more stover responders at day 12 (36 OTUs) than day 82 (11 OTUs), and which only had 1 OTU that responded to PyOM (Supplementary Figure 6).
Log2-fold change in relative abundance of OTUs as compared with unamended plots. Each circle represents a single OTU and dashed and dotted lines represent increases or decreases of 2x and 10x, respectively. Colors are scaled from yellow to red in decreasing P-value, with gray points indicating OTUs with Benjamini and Hochberg (BH)-adjusted P-values>0.10. The color reproduction of this figure is available at the The ISME Journal online.
Log2-fold change in relative abundance of OTUs in response to stover or PyOM as compared with unamended plots. Data are the same as those depicted in Figure 4. Note different scales on axes. Each point represents the response for a single OTU across replicates, colored by phylum. Dashed oval overlays indicate response groupings, where blue indicates strong stover responders, pink indicates strong PyOM responders and purple indicates common responders. Gray histograms at the sides of the plot indicate density of OTU points. The color reproduction of this figure is available at the The ISME Journal online.
Many OTUs from Firmicutes, Proteobacteria and Bacteroidetes responded to stover additions (Figures 4 and 5; Supplementary Figures 6). Among Proteobacteria, 125 OTUs responded to stover additions on day 12, increasing to 235 responders on day 82 (primarily OTUs from the orders Rhizobiales, Burkholderiales, Sphingomonadales, Xanthomonadales and Pseudomonadales on both days, and Rhodospirillales and Myxococcales on day 82) (Supplementary Figure 7). Among Bacteroidetes, 60 OTUs responded to stover additions on day 12, increasing to 128 on day 82 (primarily orders Saprospirales, Cytophagales and Sphingobacteriales, on both days and Flavobacteriales on day 12) (Supplementary Figure 8). In Firmicutes, 36 OTUs responded to stover on day 12, declining to 11 OTUs by day 82 (orders Bacillales and Clostridiales at both times) (Supplementary Figure 6).
Only 19 OTUs were observed to respond to both stover and PyOM at day 12, but this increased to 113 OTUs by day 82. The OTUs that responded to both stover and PyOM included Proteobacteria (66 OTUs; Supplementary Figure 7), Bacteroidetes (24 OTUs; Supplementary Figure 8) and Verrucomicrobia (11 OTUs; Supplementary Figure 9). In particular, a number of OTUs from Verrucomicrobia were observed to increase greatly in relative abundance in response to both stover and PyOM as compared with untreated control plots (Figure 6 and Supplementary Figures 9 and 15).
Log2-fold change of the relative abundance of OTUs in PyOM or stover plots vs soil-only plots on days 12 and 82, for Firmicutes, Gemmatimonadetes and Verrucomicrobia phyla. Each point represents the response for a single OTU across replicates, colored by order. Circles represent day 12 and triangles represent day 82. Square brackets indicate candidate order in Greengenes taxonomic nomenclature.
Only 12 OTUs increased in relative abundance specifically in response to PyOM at day 12, but this increased to 138 OTUs by day 82. The OTUs that responded specifically to PyOM included Proteobacteria (47 OTUs; Supplementary Figure 7), Bacteroidetes (30 OTUs; Supplementary Figure 8), Planctomycetes (13 OTUs; Supplementary Figure 10), Gemmatimonadetes (12 OTUs; Supplementary Figure 11) and Verrucomicrobia (6 OTUs; Supplementary Figure 9). The strongest specific response to PyOM and not stover, by far, was observed for OTUs from Gemmatimonadetes (Figure 5). These included OTUs from the orders Gemmatimonadales and ‘Ellin5290’ (Figure 6 and Supplementary Figure 16). Two of these Gemmatimonadetes OTUs were also among the top 10 most abundant PyOM responders. The most abundant PyOM responders also included two OTUs from Bacteroidetes (from the Oxalobacteraceae), and two OTUs from Proteobacteria (Rhizobiales and Erythrobacteraceae).
Discussion
Soil bacterial community dynamics
The soils that received corn stover amendments showed marked increases in CO2 emissions almost immediately (Figure 1), and bacterial community composition in those plots had changed significantly by day 12 (Figures 2 and 3). Stover additions resulted in significant increases in the relative abundance of OTUs from the phyla Proteobacteria and Bacteroidetes, and the orders Actinomycetales, Bacillales and Clostridiales (Figures 3 and 4), which is consistent with previous studies (Pascault et al., 2013). For example, wheat residue additions to a calcareous silty clay farm soil stimulated Firmicutes OTUs, whereas alfalfa additions stimulated Proteobacteria, Firmicutes and Bacteroidetes (Pascault et al., 2013). It is possible that these microorganisms are adapted to grow rapidly in response to inputs of easily mineralizable organic matter (for example, aliphatic C structures). These early-responding OTUs are likely responsible for the strong increase in total soil CO2 emissions during the first weeks after stover was applied (Figure 1). As corn stover is more easily mineralizable than pre-existing SOC, its addition may be predicted to stimulate a broad spectrum of microorganisms. However, corn stover additions stimulated only a narrow subset of microorganisms (~10% of OTUs).
Although the greatest effect of amendments on CO2 emissions took place within the first 2 weeks (Figure 1), the bacterial community response to organic amendments grew stronger over 82 days. The number of PyOM responders increased 11.5-fold from days 12 to 82, whereas the number of stover responders only increased 1.6-fold during this timeframe. A total of 70% of the PyOM responders that were observed by day 12 also responded to stover (Figure 5 and Supplementary Figure 4). These early PyOM responders likely represent those microorganisms responsible for short-term mineralization of PyOM-C and for the CO2 emissions we observed within the first 2 weeks (although we note that this abundance-based approach would not detect possible changes to the contributions to CO2 efflux from microbes that did not actively grow/divide during the study (Blazewicz et al., 2013)). The fact that 70% of these rapidly responding PyOM responders also responded to stover suggests that they are likely metabolizing easily mineralizable, possibly aliphatic components of PyOM, which are also present within fresh corn stover (Cheng et al., 2008). In contrast, the OTUs that responded to both PyOM and stover on day 82 may represent microbes that were accessing the polyaromatic bulk of the PyOM-C and the remaining, less easily mineralizable stover-C compounds (Whitman et al., 2013). In addition, late-responding OTUs could include microbes that are responding to other effects of the amendments, such as changes in soil physical or chemical properties, or the ecology of the system (for example, changes to the soil food web, competition or mutualisms).
PyOM effects on the soil bacterial community
The OTUs that responded uniquely to PyOM include representatives from 14 phyla (Figure 5, pink region and Supplementary Figures 4). Although the short-term effects of PyOM on SOC dynamics may be driven to a large extent by a relatively small, but easily mineralizable fraction of PyOM-C (Whitman et al., 2014), it is essential to also understand the long-term effects driven by chemical or physical changes to the soil environment, such as pH, soil moisture, nutrient status or the mineralization of more chemically complex PyOM-C sources. For this, the Gemmatimonadetes, particularly those of classes Gemm-5, Gemm-3 and Gemmatimonadales, are a prime target (Figure 6 and Supplementary Figures 11 and 16). The first isolates of the Gemmatimonadetes phylum were described in 2003 (Zhang et al., 2003), and although little is yet known about them ecologically and physiologically, our findings seem consistent with previous observations. For example, Gemmatimonadetes have been found to decrease in relative abundance with the addition of wheat residues (Bernard et al., 2007), were more active in soil microcosms that did not receive leaf litter (Pfeiffer et al., 2013), and were more likely to be decomposing existing SOM than fresh OM (Pascault et al., 2013). Of particular relevance, Gemmatimonadetes increased with the addition of rice straw PyOM made at 500 °C to a farmed Acrisol in a pot trial (Xu et al., 2014). This increase was driven by increases from the class Gemmatimonadetes, with some Gemm-1 and Gemm-3 increasing in relative abundance as well. These studies and our own findings suggest they may be adapted to a lifestyle associated with OM sources that are challenging to mineralize. Increased pH with PyOM additions may also have had a positive effect on Gemmatimonadetes, which have been reported to be more abundant in neutral pH soils (Lauber et al., 2009; Vishnivetskaya et al., 2011), although this effect was not significant for a much larger pH range than that observed in this study (2.6 vs 0.75) (DeBruyn et al., 2011). Gemmatimonadetes may also be adapted for low soil moisture (DeBruyn et al., 2011), but this is not a likely explanation in our study, as we did not measure significant differences in soil moisture in the PyOM-amended soils on any sampling day (data not shown). The top five most abundant OTUs that responded uniquely to PyOM, including members of the Oxalobacteraceae, Rhizobiales and Erythrobacteraceae, could also be good targets for future investigations into microbial interactions with PyOM.
PyOM produced from different materials under different conditions can result in a wide range of pH values (Enders et al., 2012), and in this study, PyOM additions significantly increased soil pH (Supplementary Figure 17), albeit by less than a full pH unit. Soil pH is strongly correlated with community composition (Lauber et al., 2009; Rousk et al., 2010). In particular, the aptly named phylum Acidobacteria has been shown to be especially sensitive to pH shifts, although its subgroups show variable responses to acidity: subgroups 1, 2 and 3 have been shown to increase at lower pHs, whereas subgroups 4, 5, 6, 7 and 17 have been shown to increase at higher pHs (Rousk et al., 2010; Bartram et al., 2013). Both Bartram et al. (2013) and Rousk et al. (2010) characterized soils from long-term (50+ and 100+ years, respectively) liming trials, so it is not possible to predict from those studies the expected timescale of a soil bacterial community response to pH changes. However, we may ask whether the PyOM-specific response in this study is driven by pH shifts. We found that OTUs from subgroup 4 did generally increase with PyOM additions (Supplementary Figure 18). However, we also found that members of subgroup 6 decreased in relative abundance, whereas subgroup 3 increased with PyOM additions (Supplementary Figure 18), which is counter to what trends in previous studies would predict if these changes were being driven purely by the pH increase with PyOM additions. This does not necessarily contrast with the previous studies—rather, it may show that factors other than pH are likely important for driving the observed changes in subgroups 3 and 6 in this system. Explanations besides pH are particularly likely, as the pH shift by +0.75 units was small in comparison with the potential magnitude of natural pH gradients common in soils (for example, owing to biological activity, rhizosphere effects or wet–dry cycles (Husson, 2012)). Acidobacteria have sometimes been characterized as being poorly equipped to compete in high nutrient conditions (Fierer et al., 2007). If some Acidobacteria are able to mineralize challenging C forms, such as fused aromatic C ring structures, but are poorly adapted for metabolizing easily mineralizable C, for example, this could explain the positive response of OTUs in Acidobacteria subgroup 3 to PyOM additions on day 82, despite the accompanying small pH increase.
Although our study focused on soil bacterial communities, soil fungi have an essential role in SOC cycling (Schimel and Schaeffer, 2012). In addition, soil fungi and their oxidative enzymes have an important role in the degradation of organic matter compounds with condensed aromatic ring structures, such as lignin (Talbot et al., 2011; Burns et al., 2013). This makes fungi an excellent target for future investigations of the effects of PyOM on soil microbial communities.
Soil C dynamics
Stover amendments had a larger effect than PyOM on both soil bacterial community composition (Figures 2, 3, 4, 5, 6) and soil CO2 emissions (Figure 1), and also resulted in significantly greater SOC-derived CO2 emissions on day 12 (Supplementary Figure 3). However, this increase in mineralization of existing SOC disappeared by day 66 (Supplementary Figure 3). These dynamics likely reflect the relative microbial accessibility or solubility of the two amendments (Lehmann and Kleber, 2015).
As the PyOM was produced at a relatively low temperature (350 °C), there was likely a substantial fraction of relatively easily mineralizable C (Zimmerman, 2010; Whitman et al., 2013) that contributed to the increases in total CO2 during the first week after application (Figure 1 and Supplementary Figure 19). This fraction could include aliphatic C compounds, carboxylic acids, cellulose and hemicellulose (Whitman et al., 2013). (It is also possible that a small portion of the PyOM-C losses were due to the dissolution of carbonates (Supplementary Figure 17)). However, unlike with stover, we did not find evidence that increased microbial activity because of PyOM additions affected SOC-derived CO2 emissions (Supplementary Figure 3). This is likely because the PyOM-C was more challenging for microorganisms to mineralize than the stover-C, resulting in slower mineralization of PyOM as compared with the stover. Although there was 1.8 times as much total stover-C added as PyOM-C, CO2 emissions increased with stover additions much more than 1.8 times the amount they increased with PyOM additions (Figure 1 and Supplementary Figure 19). Only the stover provided a large easily mineralizable C subsidy to the microbial community, which stimulated the microbial community, increased microbial biomass and may have increased general enzymatic activity. This, in turn, increased mineralization of existing SOC (Blagodatskaya and Kuzyakov, 2008). The lack of any detectable net effect on SOC mineralization from PyOM additions (often termed a ‘priming effect’ (Bingeman et al., 1953; Woolf and Lehmann, 2012; Whitman et al., 2014)) is not unexpected. Specific combinations of PyOM materials and soils have been found to produce a wide range of effects on SOC mineralization (Maestrini et al., 2014; Whitman et al., 2015). The finding that PyOM application may result in less SOC loss than the application of an equivalent amount of fresh stover has implications for residue and C management at the farm scale. In systems where organic matter sources will rapidly mineralize under baseline conditions, PyOM production from this material and its return to the soil may result in greater net C gains than adding the fresh organic matter directly to the soil, because of the strong impact of fresh organic matter on the soil bacterial community and their C mineralization activity.
Accession codes
References
Abel S, Peters A, Trinks S, Schonsky H, Facklam M . (2013). Impact of biochar and hydrochar addition on water retention and water repellency of sandy soil. Geoderma 202-203: 183–191.
Ameloot N, Graber ER, Verheijen FGA, De Neve S . (2013). Interactions between biochar stability and soil organisms: review and research needs. Eur J Soil Sci 64: 379–390.
Balesdent J, Mariotti A . (1996). Measurement of soil organic matter turnover using 13C natural abundance. Mass Spectrometry of Soils. In: Boutton TW, Yamasaki SI (eds), Marcel Dekker: New York, NY, USA.
Bartram AK, Jiang X, Lynch MD, Masella AP, Nicol GW, Dushoff J et al. (2013). Exploring links between pH and bacterial community composition in soils from the Craibstone Experimental Farm. FEMS Microbiol Ecol 87: 403–415.
Bates D, Maechler M, Bolker B, Walker S . (2015). Fitting Linear Mixed-Effects Models Using lme4. J Stat Softw 67: 1–48.
Bernard L, Mougel C, Maron P-A, Nowak V, Lévêque J, Henault C et al. (2007). Dynamics and identification of soil microbial populations actively assimilating carbon from 13C-labelled wheat residue as estimated by DNA- and RNA-SIP techniques. Environ Microbiol 9: 752–764.
Bingeman CW, Varner JE, Martin WP . (1953). The effect of the addition of organic materials on the decomposition of an organic soil. Soil Sci Soc Am J 17: 34–38.
Blagodatskaya E, Kuzyakov Y . (2008). Mechanisms of real and apparent priming effects and their dependence on soil microbial biomass and community structure: critical review. Biol Fertil Soils 45: 115–131.
Blazewicz SJ, Barnard RL, Daly RA, Firestone MK . (2013). Evaluating rRNA as an indicator of microbial activity in environmental communities: limitations and uses. ISME J 7: 2061–2068.
Burns RG, DeForest JL, Marxsen J, Sinsabaugh RL, Stromberger ME, Wallenstein MD et al. (2013). Soil enzymes in a changing environment: current knowledge and future directions. Soil Biol Biochem 58: 216–234.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7: 335–336.
Chen J, Liu X, Zheng J, Zhang B, Lu H, Chi Z et al. (2013). Biochar soil amendment increased bacterial but decreased fungal gene abundance with shifts in community structure in a slightly acid rice paddy from Southwest China. Appl Soil Ecol 71: 33–44.
Cheng C-H, Lehmann J, Engelhard MH . (2008). Natural oxidation of black carbon in soils: Changes in molecular form and surface charge along a climosequence. Geochim Cosmochim Acta 72: 1598–1610.
Czimczik CI, Masiello CA . (2007). Controls on black carbon storage in soils. Global Biogeochem Cycles 21: GB3005.
DeBruyn JM, Nixon LT, Fawaz MN, Johnson AM, Radosevich M . (2011). Global biogeography and quantitative seasonal dynamics of Gemmatimonadetes in soil. Appl Environ Microbiol 77: 6295–6300.
Dunavin LS . (1969). A comparison of Gahi-1 millet and Grazer A sorghum x sudangrass at several pH levels. Proc Soil Crop Sci Soc Florida 29: 163–168.
Edgar RC . (2013). UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10: 996–998.
Enders A, Hanley K, Whitman T, Joseph S, Lehmann J . (2012). Characterization of biochars to evaluate recalcitrance and agronomic performance. Bioresour Technol 114: 644–653.
Fierer N, Bradford MA, Jackson RB . (2007). Toward an ecological classification of soil bacteria. Ecology 88: 1354–1364.
Gomez JD, Denef K, Stewart CE, Zheng J, Cotrufo MF . (2014). Biochar addition rate influences soil microbial abundance and activity in temperate soils. Eur J Soil Sci 65: 28–39.
Gul S, Whalen JK, Thomas BW, Sachdeva V, Deng H . (2015). Physico-chemical properties and microbial responses in biochar-amended soils: Mechanisms and future directions. Agric Ecosyst Environ 206: 46–59.
Guereña DT, Lehmann J, Thies JE, Enders A, Karanja N, Neufeldt H . (2015). Partitioning the contributions of biochar properties to enhanced biological nitrogen fixation in common bean (Phaseolus vulgaris. Biol Fertil Soils 51: 479–491.
Husson O . (2012). Redox potential (Eh) and pH as drivers of soil/plant/microorganism systems: a transdisciplinary overview pointing to integrative opportunities for agronomy. Plant Soil 362: 389–417.
Jeffery S, Verheijen FGA, van der Velde M, Bastos AC . (2011). A quantitative review of the effects of biochar application to soils on crop productivity using meta-analysis. Agric Ecosyst Environ 144: 175–187.
Jin H . (2010). Characterization of microbial life colonizing biochar and biochar-amended soils. PhD dissertation. Cornell University: Ithaca, NY, USA.
Jindo K, Sanchez-Monedero MA, Hernández T, García C, Furukawa T, Matsumoto K et al. (2012). Biochar influences the microbial community structure during manure composting with agricultural wastes. Sci Total Environ 416: 476–481.
Kolton M, Meller Harel Y, Pasternak Z, Graber ER, Elad Y, Cytryn E . (2011). Impact of biochar application to soil on the root-associated bacterial community structure of fully developed greenhouse pepper plants. Appl Environ Microbiol 77: 4924–4930.
Kuzyakov Y, Bol R . (2004). Using natural 13C abundances to differentiate between three CO2 sources during incubation of a grassland soil amended with slurry and sugar. J Plant Nutr Soil Sci 167: 669–677.
Laird DA . (2008). The charcoal vision: a win-win-win scenario for simultaneously producing bioenergy, permanently sequestering carbon, while improving soil and water quality. Agronomy J 100: 178–181.
Lauber CL, Hamady M, Knight R, Fierer N . (2009). Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75: 5111–5120.
Lehmann J . (2007). A handful of carbon. Nature 447: 143–144.
Lehmann J, Kleber M . (2015). The contentious nature of soil organic matter. Nature 528: 60–68.
Lehmann J, Rillig MC, Thies J, Masiello CA, Hockaday WC, Crowley D . (2011). Biochar effects on soil biota—a review. Soil Biol Biochem 43: 1812–1836.
Lehmann J, Skjemstad J, Sohi S, Carter J, Barson M, Falloon P et al. (2008). Australian climate–carbon cycle feedback reduced by soil black carbon. Nat Geosci 1: 832–835.
Lenth RV . (2016). Least-Squares Means: The R Package lsmeans J Stat Softw 69: 1–33.
Love MI, Huber W, Anders S . (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15: 550.
Lozupone C, Lladser ME, Knights D, Stombaugh J, Knight R . (2011). UniFrac: an effective distance metric for microbial community comparison. ISME J 5: 169–172.
Maestrini B, Nannipieri P, Abiven S . (2014). A meta-analysis on pyrogenic organic matter induced priming effect. GCB Bioenergy 7: 577–590.
Masiello CA, Chen Y, Gao X, Liu S, Cheng H-Y, Bennett MR et al. (2013). Biochar and microbial signaling: production conditions determine effects on microbial communication. Environ Sci Technol 47: 11496–11503.
McMurdie PJ, Holmes S . (2014). Waste not, want not: Why rarefying microbiome data is inadmissible. PLoS Comput Biol 10: e1003531.
Mitchell PJ, Simpson AJ, Soong R, Simpson MJ . (2015). Shifts in microbial community and water-extractable organic matter composition with biochar amendment in a temperate forest soil. Soil Biol Biochem 81: 244–254.
Nickerson N, Egan J, Risk D . (2013). Iso-FD: A novel method for measuring the isotopic signature of surface flux. Soil Biol Biochem 62: 99–106.
Nielsen S, Minchin T, Kimber S, van Zwieten L, Gilbert J, Munroe P et al. (2014). Comparative analysis of the microbial communities in agricultural soil amended with enhanced biochars or traditional fertilisers. Agric Ecosyst Environ 191: 73–82.
Nolley A, Brown CT . (2012). Screed - short read sequence utils. Michigan State University. Available from http://screed.readthedocs.org.
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, OHara RB et al. (2015) vegan: Community Ecology Package R package version 2.3-0 Available from http://CRAN.R-project.org/package=vegan.
Pascault N, Ranjard L, Kaisermann A, Bachar D, Christen R, Terrat S et al. (2013). Stimulation of different functional groups of bacteria by various plant residues as a driver of soil priming effect. Ecosystems 16: 810–822.
Pfeiffer B, Fender A-C, Lasota S, Hertel D, Jungkunst HF, Daniel R . (2013). Leaf litter is the main driver for changes in bacterial community structures in the rhizosphere of ash and beech. Appl Soil Ecol 72: 150–160.
R Core Team. (2015) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria. Available from http://www.R-project.org/.
Rousk J, Bååth E, Brookes PC, Lauber CL, Lozupone C, Caporaso JG et al. (2010). Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J 4: 1340–1351.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB et al. (2009). Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75: 7537–7541.
Schimel J, Schaeffer SM . (2012). Microbial control over carbon cycling in soil. Front Microbiol 3: 348.
Shinners KJ, Binversie BN . (2007). Fractional yield and moisture of corn stover biomass produced in the Northern US Corn Belt. Biomass Bioenergy 31: 576–584.
Song Y, Zhang X, Bin MA, Chang SX, Gong J . (2014). Biochar addition affected the dynamics of ammonia oxidizers and nitrification in microcosms of a coastal alkaline soil. Biol Fertil Soils 50: 321–332.
Taketani RG, Lima AB, Conceição Jesus E, Teixeira WG, Tiedje JM, Tsai SM . (2013). Bacterial community composition of anthropogenic biochar and Amazonian anthrosols assessed by 16S rRNA gene 454 pyrosequencing. Antonie van Leeuwenhoek 104: 233–242.
Talbot JM, Yelle DJ, Nowick J, Treseder KK . (2011). Litter decay rates are determined by lignin chemistry. Biogeochemistry 108: 279–295.
van Es HM, Gomes CP, Sellmann M, van Es CL . (2007). Spatially-balanced complete block designs for field experiments. Geoderma 140: 346–352.
Vishnivetskaya TA, Mosher JJ, Palumbo AV, Yang ZK, Podar M, Brown SD et al. (2011). Mercury and other heavy metals influence bacterial community structure in contaminated Tennessee streams. Appl Environ Microbiol 77: 302–311.
Watzinger A, Feichtmair S, Kitzler B, Zehetner F, Kloss S, Wimmer B et al. (2014). Soil microbial communities responded to biochar application in temperate soils and slowly metabolized 13C-labelled biochar as revealed by 13C PLFA analyses: results from a short-term incubation and pot experiment. Eur J Soil Sci 65: 40–51.
Werth M, Kuzyakov Y . (2010). 13C fractionation at the root-microorganisms-soil interface: a review and outlook for partitioning studies. Soil Biol Biochem 42: 1372–1384.
Whitman T, Enders A, Lehmann J . (2014). Pyrogenic carbon additions to soil counteract positive priming of soil carbon mineralization by plants. Soil Biol Biochem 73: 33–41.
Whitman T, Hanley K, Enders A, Lehmann J . (2013). Predicting pyrogenic organic matter mineralization from its initial properties and implications for carbon management. Organic Geochem 64: 76–83.
Whitman T, Lehmann J . (2015). A dual-isotope approach to allow conclusive partitioning between three sources. Nat Communs 6: no.8708.
Whitman T, Scholz SM, Lehmann J . (2010). Biochar projects for mitigating climate change: an investigation of critical methodology issues for carbon accounting. Carbon Manag 1: 89–107.
Whitman T, Singh BP, Zimmerman AR . (2015). Priming effects in biochar-amended soils: Implications of biochar-soil organic matter interactions for carbon storage. Biochar for Environmental Management. In: Lehmann J, Joseph S (eds), Routledge: New York, NY, USA.
Whitman T, Zhu Z, Lehmann J . (2014). Carbon mineralizability determines interactive effects on mineralization of pyrogenic organic matter and soil organic carbon. Environ Sci Technol 48: 13727–13734.
Woolf D, Lehmann J . (2012). Modelling the long-term response to positive and negative priming of soil organic carbon by black carbon. Biogeochemistry 111: 83–95.
Xu H-J, Wang X-H, Li H, Yao H-Y, Su J-Q, Zhu Y-G . (2014). Biochar impacts soil microbial community composition and nitrogen cycling in an acidic soil planted with rape. Environ Sci Technol 48: 9391–9399.
Yarza P, Richter M, Peplies J, Euzeby J, Amann R, Schleifer K-H et al. (2008). The all-species Living Tree project: a 16S rRNA-based phylogenetic tree of all sequenced type strains. Syst Appl Microbiol 31: 241–250.
Zhang J, Kobert K, Flouri T, Stamatakis A . (2014). PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30: 614–620.
Zhang H, Sekiguchi Y, Hanada S, Hugenholtz P, Kim H, Kamagata Y et al. (2003). Gemmatimonas aurantiaca gen. nov., sp nov., a gram-negative, aerobic, polyphosphate-accumulating micro-organism, the first cultured representative of the new bacterial phylum Gemmatimonadetes phyl. nov. Int J Syst Evol Microbiol 53: 1155–1163.
Zimmerman AR . (2010). Abiotic and microbial oxidation of laboratory-produced black carbon (biochar). Environ Sci Technol 44: 1295–1301.
Acknowledgements
We are grateful for the financial support by awards from the Towards Sustainability Foundation, Cornell Sigma Xi, NSERC PGS-D, NSF-BREAD (IOS-0965336), Cornell Biogeochemistry program, Cornell Crop and Soil Science Department, USDA-NIFA Carbon Cycle (2014-67003-22069) and the Cornell Atkinson Center for a Sustainable Future. We also thank Nick Nickerson and Forerunner Research/EOSENSE for advice regarding chamber design; Kelly Hanley, Joseph Amsili, Angela Possinger, Romy Zyngier and Andy Cavin for help with fieldwork; Brent Whitman and Ellen Whitman for helping with lab analyses; and Dr Christine Goodale for helpful discussion and suggestions.
Author contributions
TW, DHB and JL designed the experiment; TW and AE grew the biomass and produced the PyOM; AE designed and built the CO2 chambers; TW conducted the field trial and performed the biogeochemical lab work; AC, CK and TW performed the molecular lab work; AC, CK and CP-R developed the bioinformatics analysis pipeline; TW and CP-R performed the bioinformatics analyses; TW, CP-R, DHB and JL interpreted the data; and TW wrote the manuscript, and all authors commented on the paper.
Accession codes
Sequence data are deposited in the NCBI Sequence Read Archive under accession SRP072444.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Whitman, T., Pepe-Ranney, C., Enders, A. et al. Dynamics of microbial community composition and soil organic carbon mineralization in soil following addition of pyrogenic and fresh organic matter. ISME J 10, 2918–2930 (2016). https://doi.org/10.1038/ismej.2016.68
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2016.68
This article is cited by
-
Rice residue management alternatives in rice–wheat cropping system: impact on wheat productivity, soil organic carbon, water and microbial dynamics
Scientific Reports (2024)
-
Biogeographic patterns and drivers of soil viromes
Nature Ecology & Evolution (2024)
-
Pyrogenic organic matter decreases while fresh organic matter increases soil heterotrophic respiration through modifying microbial activity in a subtropical forest
Biology and Fertility of Soils (2024)
-
Experimentally determined traits shape bacterial community composition one and five years following wildfire
Nature Ecology & Evolution (2023)
-
Biochar application significantly increases soil organic carbon under conservation tillage: an 11-year field experiment
Biochar (2023)