Abstract
The assimilation of organic nutrients by autotrophs, a form of mixotrophy, has been demonstrated in the globally abundant marine picocyanobacterial genera Prochlorococcus and Synechococcus. However, the range of compounds used and the distribution of organic compound uptake genes within picocyanobacteria are unknown. Here we analyze genomic and metagenomic data from around the world to determine the extent and distribution of mixotrophy in these phototrophs. Analysis of 49 Prochlorococcus and 18 Synechococcus isolate genomes reveals that all have the transporters necessary to take up amino acids, peptides and sugars. However, the number and type of transporters and associated catabolic genes differ between different phylogenetic groups, with low-light IV Prochlorococcus, and 5.1B, 5.2 and 5.3 Synechococcus strains having the largest number. Metagenomic data from 68 stations from the Tara Oceans expedition indicate that the genetic potential for mixotrophy in picocyanobacteria is globally distributed and differs between clades. Phylogenetic analyses indicate gradual organic nutrient transporter gene loss from the low-light IV to the high-light II Prochlorococcus. The phylogenetic differences in genetic capacity for mixotrophy, combined with the ubiquity of picocyanobacterial organic compound uptake genes suggests that mixotrophy has a more central role in picocyanobacterial ecology than was previously thought.
Similar content being viewed by others
Introduction
The prokaryotic world has conventionally been divided into autotrophs and heterotrophs based on carbon source, and phototrophs and chemotrophs as dictated by energy source. In the past several decades we have learned that a number of bacteria, archaea and even eukarya do not easily fit into these categories and are capable of using a mix of energy and carbon sources, a trophic strategy known as mixotrophy. Examples include photoautoheterotrophy, photoorganoheterotrophy and chemolithoautoheterotrophy (Perez and Matin, 1980; Jones, 2000; Zubkov, 2009). In marine systems, bacterivory by photosynthetic protists is now known to be common in eukaryotes such as dinoflagellates (Jones, 2000). According to Hartmann et al. (2012), the majority of bacterivory in the Atlantic may be carried out by phototrophs. Similarly, the ocean is populated by up to 11% aerobic anoxygenic photoheterotrophic bacteria that combine phototrophy and carbon fixation with organic compound uptake (Kolber et al., 1999; Sieracki et al., 2006; Jiao et al., 2007; Kirchman et al., 2014). Heterotrophic bacteria that utilize light to pump protons via rhodopsin are abundant globally as well (Béjà et al., 2000, 2001; Rusch et al., 2007), making up an estimated 13% of the photic zone bacteria in the Mediterranean and Red Seas (Sabehi et al., 2005), 50% in the Sargasso Sea (Campbell et al., 2008), and 48% in 116 marine and terrestrial samples examined by Finkel et al. (2013). In addition to using proteorhodopsin, some of these bacteria can fix up to 30% of their total carbon (Palovaara et al., 2014), suggesting a significant contribution to global carbon fixation.
Some freshwater cyanobacteria have long been known to employ photoautoheterotrophy (Rippka, 1972; Joset-Espardellier et al., 1978; Chen et al., 1991; Paerl, 1991; Zubkov, 2009) and evidence is growing that marine picocyanobacteria—the most abundant marine phototrophs (Partensky et al., 1999)—also have this capacity. The dominant genera, Prochlorococcus and Synechococcus, are known to take up amino acids (Church et al., 2004; Zubkov et al., 2003, 2004, 2008; Michelou et al., 2007; Mary et al., 2008; Gómez-Pereira et al., 2013; Evans et al., 2015), glucose (Gómez-Baena et al., 2008; Muñoz-Marín et al., 2013) and dimethylsulfoniopropionate (Vila-Costa et al., 2006), and analysis of 12 genomes has shown that certain strains have genes for amino acid, sugar, oligopeptide and phosphonate uptake (Rocap et al., 2003; Martiny et al., 2006; Kettler et al., 2007).
The ability of these picocyanobacteria to take up organic compounds raises the question of how mixotrophic capacity is distributed phylogenetically within this group. Prochlorococcus and marine Synechococcus (hereafter designated marine picocyanobacteria) can be divided into phylogenetic clusters that generally correspond to physiologically distinct ecotypes (West and Scanlan, 1999; West et al., 2001; Ahlgren et al., 2006; Johnson et al., 2006; Zinser et al., 2006, 2007; Malmstrom et al., 2010; Sohm et al., 2015). In Prochlorococcus these ecotypes have different light and temperature optima, which results in a partitioning of the water column with depth, and different relative abundances along latitudinal gradients (Moore et al., 1998; Moore and Chisholm, 1999; Rocap et al., 2003; Johnson et al., 2006; Zinser et al., 2007). Synechococcus ecotypes can be defined by open ocean and coastal phylogenetic clusters (Dufresne et al., 2008; Ahlgren and Rocap, 2012) as well as by temperature- and nutrient concentration-related groups (Sohm et al., 2015). Because of this ecotype partitioning along phylogenetic lines, if picocyanobacterial mixotrophic capacity is partitioned by phylogenetic group it suggests a role for mixotrophy in niche adaptation. In flagellates for example, mixotrophy can provide alternative sources of energy in light-limiting conditions and alternative sources of nitrogen or phosphorous in nutrient-limiting conditions (Rothhaupt, 1996a, b). Furthermore, if mixotrophy is also universally distributed among marine picocyanobacteria, it indicates that it has a more central role in the ecology of these genera than previously thought.
In order to determine the extent of mixotrophic ability among picocyanobacteria in the global oceans, we examined a collection of 67 Prochlorococcus and Synechococcus isolate reference genomes (Supplementary Table S1). Because these strains were isolated from a limited number of locations, we also examined data from the largest marine metagenomic survey to date, the Tara Oceans expedition (Karsenti et al., 2011; Sunagawa et al., 2015). This census of the ocean microbiome allowed us to estimate the global picocyanobacterial genetic capacity for mixotrophy and examine how it is distributed among different oceanographic regimes at a global scale.
Materials and methods
Isolation and sequencing of isolate genomes
Sixty-seven Prochlorococcus and Synechococcus genomes were used as reference genomes in this study (Supplementary Table S1). The genomes of MIT1306, MIT1312, MIT1318, MIT1320, MIT1323, MIT1327, MIT1342, MITS9504, MITS9508 and MITS9509 are described by Cubillos-Ruiz (2015) and Thompson (2015) and are publically available in the National Center for Biotechnology Information (NCBI) Genbank database.
Sampling
The Tara Oceans expedition is the first global oceanographic expedition to combine analysis of nutrient concentration, temperature, salinity and particulate data with deep sequencing of environmental DNA for metagenomic reconstruction at multiple depths (4 to ~800 m, Supplementary Table S2, http://doi.pangaea.de/10.1594/PANGAEA.840721). The samples used in this study were taken at 68 sites in the Atlantic, Pacific, Indian and Southern Oceans, as well as in the Mediterranean and Red Seas (Supplementary Table S2, Sunagawa et al., 2015). Sample collection and preparation were described previously (Logares et al., 2014; Pesant et al., 2015; Sunagawa et al., 2015). The current analysis focuses on 139 DNA samples from the two smallest size fractions (0.2–1.6 μm and 0.2-3 μm). These samples were taken from up to three different depths: ~5 m, the deep chlorophyll maximum, and the mesopelagic zone.
Metagenomic DNA extraction, sequencing and assembly
DNA extraction and Illumina sequencing were described previously (Logares et al., 2014; Sunagawa et al., 2015). A total of 7.2 terabases were sequenced and processed using the MOCAT software package (Kultima et al. 2012) to yield metagenomic assemblies and gene predictions as summarized in Sunagawa et al. (2015). In order to estimate gene abundances, high-quality reads were mapped onto a non-redundant reference database, the Ocean Microbial Reference Gene Catalog (OM-RGC) including the genes from the Tara Oceans expedition, the Global Ocean Sampling expedition (Yooseph et al., 2007), the Moore Marine Microbial Sequencing project, the Moore Viral Genomes, the Pacific Ocean Virome study (Hurwitz and Sullivan, 2013) and the NCBI Viral Reference Genomes data set (Sunagawa et al., 2015). Mapped reads had a minimum of 95% nucleotide identity to a reference gene and a minimum length of 45 bp. Gene abundances were estimated from read depths that were then normalized by the reference gene length and the total number of bases per sample in order to take into account sequencing depth.
Extraction of 16S mitags/OTU classification
Reads mapped to 16S ribosomal RNA (rRNA) sequences were extracted from the metagenomic reads as described previously (Logares et al., 2014) and were designated 16S mitags. Sequences with 100 or more high-quality bases were then mapped via UCLUST (v. 1.2.22) (Edgar, 2010) to an abridged SILVA database (v. 108 clustered at 97% nucleotide identity). Reads were required to have 97% or higher identity to the reference sequence, and were assigned to their best hit reference sequence. OTU abundances were estimated based on read counts per 16S rRNA gene. These raw counts were then normalized to the amount of DNA sequencing per sample by dividing them by the total number of 16S mitags for each sample. All data and count tables are available at: http://ocean-microbiome.embl.de/companion.html.
Taxonomic classification of assembled contigs
In order to classify assembled contigs, the non-redundant set of proteins of the OM-RGC was compared to UniProt using Rapsearch2 (Ye et al., 2011). Hits with an E-value <10−3 were kept and the last common ancestor was determined for each gene as described by Hingamp et al. (2013).
Quantification of taxonomic groups
In order to quantify Prochlorococcus ecotype and Synechococcus cluster abundances in each sample, we considered using abundances of single-copy genes that had been used for this purpose previously by Li et al. (2010) and Martiny et al. (2009). Most of the genes from the Li et al. analysis failed to properly separate Synechococcus clusters, whereas abundances based on the eight single-copy genes used by Martiny et al. failed to correlate well with flow cytometry counts. Thus, we selected only the rpoC1 RNA polymerase gamma subunit gene and the psbO photosystem II manganese-stabilizing protein gene from the Li et al. analysis for quantification. These gene abundances correlate well with flow cytometry counts (Pearson correlations from 0.43 to 0.54) and maximum likelihood trees of their nucleotide sequences generally agree with picocyanobacterial internally transcribed spacer trees, allowing for minor topological differences (Supplementary Figure S1). The rpoC1 gene has previously been used to classify ecotypes of both Prochlorococcus and Synechococcus (Palenik, 1994; Ferris et al., 1998; Mühling et al., 2005, 2006), and psbO is a photosystem II protein that is not homologous to other known proteins, making it an ideal candidate for quantification of cyanobacterial genomes (Raymond and Blankenship, 2004). The relative abundances of rpoC1 and psbO in Tara Oceans samples were highly correlated (0.84 Pearson r2). On the basis of the maximum likelihood trees of these genes and those found in the reference genomes, we were able to assign Tara Oceans single-copy genes to specific ecotypes or clusters.
Identification of transporter genes
In order to identify transporter genes, we utilized the Transporter Classification Database (TCDB http://www.tcdb.org/) (Saier et al. 2006). We specifically looked for (oligo)peptide, amino acid and sugar uptake transporters, excluding peptide transporters involved in signaling. A list of the TCDB families used is given in the Supplementary Materials (Supplementary Table S3). A blastp homology search was carried out against the TCDB and hits were retained that had more than 30% amino-acid identity over 70% or more of the reference gene length with an E-value of less than 1 × 10−5. The Pro1404 glucose transporter (Gómez-Baena et al., 2008) was manually added to the database. We recognize that a 30% amino-acid identity threshold may not always be high enough to indicate identical substrate specificity. Thus, the substrate specificity of the transporters in this study should be considered putative. We chose to use the 30% threshold because the TCDB is greatly lacking in transporter genes from cyanobacteria. A higher threshold would likely miss a substantial number of transporter genes in marine picocyanobacteria.
For the reference genomes, genes were clustered into orthologous groups, CyCOGs, as described by Kelly et al. (2012). A list of CyCOG clusters of genes used in these analyses is provided in Supplementary Table S4. For the Tara Oceans OM-RGC, transporter gene abundances were normalized to 16S mitag abundances, assuming one copy of the 16S rRNA gene per genome for Prochlorococcus and two copies for Synechococcus. Samples with low abundance of 16S rRNA genes in Prochlorococcus or Synechococcus were omitted from the analyses if they contained less than 10 × average read coverage per SILVA accession identified in the sample.
Gene context
Genes consistently found in clusters across prokaryotic taxa are frequently functionally related (Dandekar et al., 1998; Huynen et al., 2000; Rogozin et al., 2004; Karimpour-Fard et al., 2008; Yelton et al., 2011). In order to further support the annotation of organic compound transporter genes, we examined their gene context to determine if they were associated with other subunits of the same transporters or metabolic genes that use the transporter substrates. For the two ABC transporters examined (3.A.1.3.18 and 3.A.1.5.-), most subunit genes were found in gene clusters (Supplementary Table S5). Overall three subunit genes were missing from three separate genomes for the 3.A.1.5.- peptide transporter and two subunit genes were missing from two genomes for the 3.A.1.3.18 amino-acid transporter. In addition, both the glucose:H+ symporter, glcP (2.A.1.1.32) and the recently discovered Pro1404 glucose porter (Gómez-Baena et al., 2008) were found in clusters of genes with related functions. glcP is adjacent to a sugar porin (1.B.19.1.4) in most Prochlorococcus genomes and two out of three Synechococcus genomes with this gene (Supplementary Figure S2). This porin is homologous to the glucose inducible sugar porin oprB in Pseudomonas (Hancock and Carey, 1980; Saravolac et al., 1991).
The Pro1404 glucose porter is a major facilitator superfamily transporter that has been implicated in glucose uptake in Prochlorococcus (Gómez-Baena et al., 2008; Muñoz-Marín et al., 2013). This transporter was consistently found in all picocyanobacterial reference genomes in a gene cluster with a glycogen debranching enzyme (E.C. 3.2.1.-) (Supplementary Figure S2), which functions in glycogen degradation to glucose monomers in Escherichia coli (Jeanningros et al., 1976; Dauvillée et al., 2005) and has also been shown to affect glycogen branching patterns in Synechococcus elongatus PCC7942 (Suzuki et al., 2007). Thus, we expect that the proteins coded by the Pro1404 gene and the glycogen debranching enzyme gene both function in providing the cell with glucose.
Statistical and phylogenetic analyses
Kolmogorov–Smirnov and Shapiro–Wilk tests indicated that the transporter gene abundance data were not normally distributed. Thus we used nonparametric tests in all statistical analyses unless otherwise noted. The correlations used were Spearman correlations and hypothesis tests between populations were Mann–Whitney–Wilcoxon tests. Phylogenetic trees were made with the RaxML software v. 7.3.0 (Stamatakis, 2006) and are maximum likelihood trees with 100 bootstraps. Evolutionary reconstruction of ancestral traits was carried out with the ace method of the ape (v. 3.3) R statistical package (Team RDC, 2012). The evolutionary model used assumes equal rates of transitions from one state to another and uses maximum likelihood ancestral state estimation (Cunningham et al., 1998).
Results and discussion
Mixotrophic capacity in cultured strains
Genetic capacity for organic compound uptake and degradation
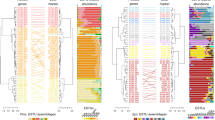
In order to determine if isolate picocyanobacteria have the capacity for mixotrophy we looked for genes for uptake and degradation of organic compounds. All 67 picocyanobacterial reference genomes contained transporters for amino acid, sugar and peptide uptake, indicating a universal capacity for mixotrophy (Figure 1), and suggesting a persistent advantage conferred by organic compound uptake across the marine environments where these picocyanobacteria live. Genes specific to degradation of the sugars and amino acids taken up by these transporters were also identified (Supplementary Table S6). The presence of these genes along with transporter genes suggests that these strains have the ability to break down organic compounds for use in central carbon, and in some cases nitrogen, metabolism. Unsurprisingly, glucose degradation genes were identified in all genomes along with glucose transporter genes. Of the 36 cases where alanine transporter genes were found, 35 also had an alanine degradation gene, alanine dehydrogenase. This enzyme is reversible, but has been implicated in alanine degradation in Synechococcus elongatus (Lahmi et al., 2006). The glutamate porter is found in 56 genomes, but the glutamate degradation dehydrogenase is found in only the low-light IV (LLIV) Prochlorococcus and the CC9605 Synechococcus strains. Only genomes that contain the glutamate and alanine transporters contain the degradation genes with one exception, WH5701, which contains an alanine dehydrogenase. This suggests that these strains are not only capable of taking up amino acids, but can also degrade them to obtain ammonium for use in biosynthesis. The co-occurrence of the degradation genes with the transporters also is consistent with the annotation of these genes as glutamate and alanine transporters.
Mixotrophic capacity by phylogenetic group
Within the Prochlorococcus ecotypes, there is a trend of reduction in gene number of amino-acid transporters from the common ancestor with Synechococcus to the high-light II (HLII) ecotype (Figure 1). This trend suggests gradual gene loss during the genome streamlining that began during the divergence from Synechococcus and involved genes that do not confer strong selective advantages in new niches (Partensky et al., 2010; Sun and Blanchard, 2014). Because of this streamlining, transporter gene numbers are correlated with genome size (amino acid and sugar transporters Pearson correlation 0.63 and 0.53, P-value <0.01). The hypothesis of gradual organic compound transporter gene loss is supported by evolutionary reconstruction of the number of amino acid and sugar transporters from the common ancestor of Prochlorococcus to the extant strains and is consistent with the amino-acid transporter gene trees in these strains (Supplementary Figures S3 and S4).
These results suggest a reduced selection pressure for mixotrophic capacity in cells that dominate surface waters, where the environment is characterized by higher light intensities and lower nutrient concentrations than deeper euphotic zone waters. The increase in amino acid and sugar transporter genes in cells that are most prevalent in low light, open ocean environments is consistent with the use of organic compounds to supplement energy and carbon under light limitation, but also indicates that despite the potential for nitrogen limitation, surface oligotrophic ocean waters do not favor the maintenance of a full suite of organic compound uptake genes. This may be because many of the substrates for these transporters are found at such low concentrations in low nutrient surface ocean waters (Keil and Kirchman, 1999; Kaiser and Benner, 2008) that the energy and nutrients required to maintain these transporters is greater than the advantage they confer.
We next looked for patterns in the distribution of organic compound transporter genes among Synechococcus phylogenetic clusters. All major Synechococcus groups have more genes per genome for uptake of organic compounds than HLI, HLII and LLI Prochlorococcus ecotypes (Figure 1). The 5.1A Synechococcus subcluster has fewer organic uptake genes than 5.2 and 5.3 Synechococcus genomes (Figure 1, Supplementary Table S7). The 5.1A group has previously been shown to dominate the Synechococcus population in oligotrophic environments (Dufresne et al., 2008). However, recent work indicates that adaptation to oligotrophic waters may not be a characteristic of the large 5.1A and 5.1B groups, but rather a clade-level adaptation (Zwirglmaier et al., 2008; Ahlgren and Rocap, 2012; Huang et al., 2012). Thus we looked for the specific clades that dominated oligotrophic samples in the Tara data set-waters with <0.1 mg chlorophyll a per m3 as defined by Behrenfeld and Falkowski (1997). Clades II and III (both 5.1A clades) dominated the Synechococcus community in oligotrophic versus mesotrophic waters (P-values <0.05 and <0.01, respectively). The reference genomes from these oligotrophic clades had fewer organic compound transporter genes versus the other Synechococcus groups, though this difference was not significant (Figure 2a). We made the same comparison between coastal and open ocean samples, as determined by Longhurst biome (Clade II dominated open ocean waters in the Tara data set; P-value <0.05). In that case, coastal reference genomes on average contained more organic compound transporter genes than open ocean genomes (Figures 2b; P-value <0.05).
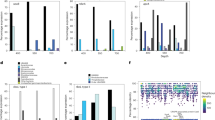
Average number of organic compound transporters. Synechococcus clades were assigned to open ocean versus coastal sites (a) and oligotrophic versus meso- and eutrophic sites (b) based on abundance of their single-copy genes in the Tara data. The average number of transporter genes per genome for reference genomes in the assigned clades are shown above. Clade II was the only open ocean clade. All other clades are coastal clades. The oligotrophic clades are clades II and III. All other clades are meso/eutrophic clades. Error bars are s.d.’s. Individual bar height indicates number of transporters per genome.
Mixotrophic potential in picocyanobacteria in the global oceans
Comparison of reference genome data set with wild populations
The availability of global metagenomic data allowed us to test whether our picocyanobacterial reference isolate genomes were representative of abundances of organic compound transporter genes globally. Specifically we aimed to determine whether global averages of organic compound uptake genes per genome were similar to the average numbers of these genes in our reference data set. To this end, we estimated the average number of transporter genes per Prochlorococcus and Synechococcus genome in the global Tara data, normalized to 16S mitag abundances, and compared these estimates to normalized transporter gene numbers in reference genomes. Transporter numbers in reference genomes were averaged within each ecotype and then normalized by multiplying them by the proportion of their respective ecotype in the Tara data set, based on single-copy gene abundances. The results indicate that the Prochlorococcus reference data set is adequately representative of in situ populations (Figure 3). The overall average number of organic compound uptake genes in the Tara Synechococcus data was similar to those in reference genomes. However, the number of amino acid uptake genes was higher in the in situ data, whereas the number of sugar and peptide transporters was lower. This suggests that the Synechococcus reference genomes do not cover the diversity seen globally. Certain taxa within the Synechococcus genus that are not represented in reference genomes likely have widely varying numbers of organic compound transporters. It is also possible that taxonomic identification failed to identify in situ Synechococcus transporter genes because of the inadequacy of the reference data set, but this scenario is unlikely, due to the low homology threshold used (30% amino-acid identity) in annotating the metagenomic genes.
Average number of transporter genes per genome in the Tara data set versus weighted average by ecotype in the reference genome data set. Tara refers to the overall average number of genes per genome (16S mitag) in the metagenomic data. Reference refers to the estimated average number of genes per genome based on averages of the reference genomes by ecotype weighted by the proportion of each ecotype in the Tara data set. Error bars are s.d.’s. Individual bar height indicates number of transporters per genome.
Geographic distribution of organic compound transporters
The global nature of the Tara Oceans data allowed us to assess the worldwide geographic distribution of picocyanobacterial mixotrophic capacity as we are defining it. We determined that organic compound uptake transporters assigned to Prochlorococcus and Synechococcus were ubiquitous in the data set (Figure 4). This finding provides the strongest evidence to date that the capacity for mixotrophy is the dominant trophic strategy in marine picocyanobacteria. Every sample that contained Prochlorococcus or Synechococcus 16S rRNA genes also contained picocyanobacterial peptide uptake transporter genes. There was only one sample that contained Prochlorococcus 16S genes but no Prochlorococcus amino acid or sugar uptake transporter genes—a sample from one of the three strong oxygen minimum zone samples. Similarly, one Southern Ocean sample lacked Synechococcus organic compound transporter genes. These oxygen minimum zone and Southern Ocean samples consistently had among the lowest number of Prochlorococcus and Synechococcus nutrient transporters (data not shown). Oxygen minimum zones similar to the one in question are dominated by novel LLV and LLVI Prochlorococcus lineages (Lavin et al., 2010) not present in current reference genome data sets. Organic compound uptake transporter genes may not have been detected for this reason or because of some unknown ecological difference between oxygen minimum zones and other open ocean habitats.
Number of picocyanobacterial organic compound transporters per genome in the Tara Oceans data set. Ocean Data View v. 4.6.2 (Schlitzer, 2002) projection using DIVA gridding with 30 by 30 scale length. Stations are in black.
In addition to their ubiquity, picocyanobacterial organic compound transporters also represent a large portion of the total organic compound transporters in samples dominated by Prochlorococcus. In these samples, up to 13.8% of amino-acid transporters, 31.1% of peptide transporters and 6.4% of sugar transporters were assigned to picocyanobacterial taxa.
No clear differences in transporter abundances by ocean were recognizable. However, three ocean ‘hot spots’ were identified where organic uptake transporter gene abundances were very high. These stations were characterized by very low (<3%) HLII single-copy gene abundances as a proportion of all Prochlorococcus single-copy genes. The Prochlorococcus single-copy genes from each station were as follows: 95% LL and HLI strains at Station 68, 98% HLI and unassigned HL strains at Station 94 and 95% unassigned HL strains at Station 128.
Relationship of transporter gene abundances to environmental parameters
To explore the hypothesis that organic compound uptake is more advantageous to picocyanobacteria under low light and relatively higher nutrient conditions, we examined relationships between number of organic compound uptake transporter genes and environmental variables including depth, nitrate and nitrite concentration, proximity to the coast, and the proportions of the different picocyanobacterial ecotypes present. We controlled for high light ecotype abundance with partial Spearman correlations because this ecotype is strongly correlated with depth and nitrate concentration (−0.5 and −0.35 correlation, respectively; P<1e−4) and because high light ecotype single-copy genes are on average 20 times more abundant than low light ones in this data set (at depths of 50 m or more). It is important to note that the proportion of HLI and HLII genes varied between samples even when holding the total abundance of HL single-copy genes constant. We found a significant positive correlation between nitrate concentration and amino-acid transporter gene abundances (Supplementary Table S8). As low light ecotypes are rare in the Tara data, the correlation with nitrate may be due to the differential distribution of HLI and HLII ecotypes at the surface. Because there is no clear mechanistic link between increased nitrate concentrations and organic compound uptake, it is likely that the correlation is actually related to another covarying environmental variable. One possibility would be the concentrations of amino acids themselves, which are likely to be more expendable in higher nitrate environments.
In addition to correlations with nitrate concentrations, significant negative correlations were found between the proportion of single-copy genes from the HLII ecotype, and the number of organic compound transporters per genome. The ecotype correlations were consistent with results from the isolate genomes where HLII genomes have the smallest number of organic compound uptake genes (Figure 1). Finally, the mean number of transporters per Synechococcus genome was higher in coastal waters (based on Longhurst biome) than in the open ocean (Table 1).
In order to estimate the average number of organic compound uptake genes per phylogenetic group in natural populations, we assigned single copy cyanobacterial genes to groups with maximum likelihood trees (Supplementary Figure S1). We then estimated group averages by taking the mean of the number of transporters per genome in samples containing more than 50% single-copy genes from this phylogenetic group. Because LLII, III and IV ecotypes were generally very low abundance, we were not able to estimate in situ averages for them. The data show a trend of increasing number of amino-acid transporter genes per genome from the HLII to the LLI Prochlorococcus ecotypes and no clear trend for sugar and peptide transporters, consistent with results from cultured strains (Table 2). The organic compound uptake transporter ‘hot spots’ previously identified are also consistent with this trend, as they are characterized by an unusually low proportion of HLII single-copy genes.
Diversity of Tara Oceans picocyanobacterial organic compound transporters
The elevated numbers of organic compound transporters in Prochlorococcus and Synechococcus that live in lower light, higher nutrient environments relative to other strains could be indicative of several different types of evolutionary strategies. Genome streamlining within the Prochlorococcus lineage may have eliminated paralogous transporter genes that served to increase expression in a common ancestor. Alternatively, multiple transporters may serve to transport the same general substrate (for example, amino acids) but different specific substrates (for example, glutamate versus aspartate), allowing the cells to take advantage of whichever resource is currently available. To differentiate between these possibilities, we looked for paralogs. This analysis found only one paralogous transporter gene in the reference genomes: the 2.A.27.2.1 probable Glu/N-acetylglutamate uptake porter (Supplementary Figure S3). In all other cases, higher numbers of transporters per genome were associated with a higher diversity of transporter classes and substrates (Figure 5). The same is true of the metagenomic data: samples with high numbers of picocyanobacterial transporters per genome also have a more diverse set of picocyanobacterial transporter classes and putative substrates (Figure 6; Spearman P-value <0.05). Collectively, these observations suggest that picocyanobacterial strains with more transporter genes have the capacity to take up a wider variety of organic substrates. The association of increased diversity with increased number of transporter genes indicates that the cyanobacteria with more of these genes have a more generalist trophic strategy.
Diversity of organic compound transporter genes in Tara Oceans metagenomic data. Number of transporter genes per Prochlorococcus or Synechococcus genome in each sample versus number of transporter gene systems or families in the sample, as defined by the TCDB. The line shows the mean number of transporters per genome. (a) Prochlorococcus amino-acid transporter gene families. (b) Synechococcus amino acid transporter gene families. (c) Prochlorococcus peptide transporter gene system. (d) Synechococcus peptide transporter system.
Conclusions
Picocyanobacterial mixotrophy genes are more abundant in low light-adapted ecotypes of Prochlorococcus and coastal groups of Synechocococcus. The differences among phylogenetic groups can be attributed to the loss of a diverse subset of transporter genes as picocyanobacteria expanded to new, high light, more oligotrophic habitats. Because of this evolutionary trajectory, we infer that a mixotrophic strategy is most advantageous to marine picocyanobacteria in low light, mesotrophic environments such as the deep euphotic zone above the deep chlorophyll maximum. These environments have higher nutrient concentrations than the surface ocean, but are still likely nutrient limited ted. ndances (Cullen, 2015). Under these conditions, cyanobacteria can supplement their energy stores with organic carbon compounds and take advantage of alternative nitrogen sources that are readily available in amino acids and peptides. Results for Prochlorococcus are compelling, but the lack of a good reference database necessitates further work on Synechococcus.
In addition to demonstrating a clear relationship between phylogeny and capacity for organic compound uptake in the genomes of cultured isolates, we have also determined that mixotrophic capacity is ubiquitous in the Tara Oceans metagenomic data set, covering all major oceans. We conclude from the near-universal distribution of picocyanobacterial genes involved in organic compound transport, that mixotrophy is a widespread strategy among these phototrophs. Given that picocyanobacteria are estimated to contribute 25% of global marine net primary productivity (Flombaum et al., 2013), their potential contribution to the assimilation of organic carbon could be significant. Furthermore, because picocyanobacteria and obligate heterotrophs make up the vast majority of marine prokaryotic communities, we postulate that almost all prokaryotes in the surface oceans are heterotrophs or mixotrophs, a finding that calls for revision of oceanic carbon and energy flux estimates between trophic levels.
References
Ahlgren NA, Rocap G . (2012). Diversity and distribution of marine Synechococcus: Multiple gene phylogenies for consensus classification and development of qPCR assays for sensitive measurement of clades in the ocean. Front Microbiol 3: 1–24.
Ahlgren NA, Rocap G, Chisholm SW . (2006). Measurement of Prochlorococcus ecotypes using real-time polymerase chain reaction reveals different abundances of genotypes with similar light physiologies. Environ Microbiol 8: 441–454.
Behrenfeld MJ, Falkowski PG . (1997). Photosynthetic rates derived from satellite-based chlorophyll concentration. Limnol Oceanogr 42: 1–20.
Béjà O, Aravind L, Koonin EV, Suzuki MT, Hadd A, Nguyen LP et al. (2000). Bacterial rhodopsin: evidence for a new type of phototrophy in the sea. Science (80-) 289: 1902–1906.
Béjà O, Spudich EN, Spudich JL, Leclerc M, DeLong EF . (2001). Proteorhodopsin phototrophy in the ocean. Nature 411: 786–789.
Campbell BJ, Waidner LA, Cottrell MT, Kirchman DL . (2008). Abundant proteorhodopsin genes in the North Atlantic Ocean. Environ Microbiol 10: 99–109.
Chen TH, Chen TL, Hung LM, Huang TC . (1991). Circadian rhythm in amino acid uptake by Synechococcus RF-1. Plant Physiol 97: 55–59.
Church MJ, Ducklow HW, Karl DM . (2004). Light dependence of [ 3 H ] leucine incorporation in the oligotrophic North Pacific Ocean light dependence of [ 3 H ] leucine incorporation in the oligotrophic North Pacific Ocean. Appl Environ Microbiol 70: 4079–4087.
Cubillos-Ruiz AF, Thompson JW, Becker JW, Chisholm SW, Shi Y, Van der Donk WA et al. (2015). Ecology and evolution of lanthipeptides in marine picocyanobacteria. (Doctoral dissertation). Retrieved from DSpace MIT Thesis Database.
Cullen JJ . (2015). Subsurface chlorophyll maximum layers: enduring enigma or mystery solved? Ann Rev Mar Sci 7: 207–239.
Cunningham CW, Omland KE, Oakley TH . (1998). Reconstructing ancestral character states: a critical reappraisal. Trends Ecol Evol 13: 361–366.
Dandekar T, Snel B, Huynen M, Bork P . (1998). Conservation of gene order: a fingerprint of proteins that physically interact. Trends Biochem Sci 23: 324–328.
Dauvillée D, Kinderf IS, Li Z, Kosar-Hashemi B, Samuel MS, Rampling L et al. (2005). Role of the Escherichia coli glgX gene in glycogen metabolism. J Bacteriol 187: 1465–1473.
Dufresne A, Ostrowski M, Scanlan DJ, Garczarek L, Mazard S, Palenik B et al. (2008). Unraveling the genomic mosaic of a ubiquitous genus of marine cyanobacteria. Genome Biol 9: R90.
Edgar RC . (2010). Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26: 2460–2461.
Evans C, Gómez-Pereira PR, Martin AP, Scanlan DJ, Zubkov MV . (2015). Photoheterotrophy of bacterioplankton is ubiquitous in the surface oligotrophic ocean. Prog Oceanogr 135: 139–145.
Ferris MJ, Palenik B . (1998). Niche adaptation in ocean cyanobacteria. Nature 396: 226–228.
Finkel OM, Béjà O, Belkin S . (2013). Global abundance of microbial rhodopsins. ISME J 7: 448–451.
Flombaum P, Gallegos JL, Gordillo RA, Rincón J, Zabala LL, Jiao N et al. (2013). Present and future global distributions of the marine Cyanobacteria Prochlorococcus and Synechococcus. Proc Natl Acad Sci USA 110: 9824–9829.
Gómez-Baena G, López-Lozano A, Gil-Martínez J, Lucena JM, Diez J, Candau P et al. (2008). Glucose uptake and its effect on gene expression in Prochlorococcus. PLoS One 3: e3416.
Gómez-Pereira PR, Hartmann M, Grob C, Tarran GA, Martin AP, Fuchs BM et al. (2013). Comparable light stimulation of organic nutrient uptake by SAR11 and Prochlorococcus in the North Atlantic subtropical gyre. ISME J 7: 603–614.
Hancock REW, Carey AM . (1980). Protein D1: a glucose-inducible, pore-forming protein from the outer membrane of Pseudomonas aeruginosa. FEMS Microbiol Lett 8: 105–109.
Hartmann M, Grob C, Tarran GA, Martin AP, Burkill PH, Scanlan DJ et al. (2012). Mixotrophic basis of Atlantic oligotrophic ecosystems. Proc Natl Acad Sci USA 109: 5756–5760.
Hingamp P, Grimsley N, Acinas SG, Clerissi C, Subirana L, Poulain J et al. (2013). Exploring nucleo-cytoplasmic large DNA viruses in Tara Oceans microbial metagenomes. ISME J 7: 1678–1695.
Huang S, Wilhelm SW, Harvey HR, Taylor K, Jiao N, Chen F . (2012). Novel lineages of Prochlorococcus and Synechococcus in the global oceans. ISME J 6: 285–297.
Hurwitz BL, Sullivan MB . (2013). The Pacific Ocean Virome (POV): a marine viral metagenomic dataset and associated protein clusters for quantitative viral ecology. PLoS One 8: e57355.
Huynen M, Snel B, Lathe W, Bork P . (2000). Predicting protein function by genomic context: quantitative evaluation and qualitative inferences. Genome Res 10: 1204–1210.
Jeanningros R, Creuzet-Sigal N, Frixon C, Cattaneo J . (1976). Purification and properties of a debranching enzyme from Escherichia coli. Biochim Biophys Acta 438: 186–199.
Jiao N, Zhang Y, Zeng Y, Hong N, Liu R, Chen F et al. (2007). Distinct distribution pattern of abundance and diversity of aerobic anoxygenic phototrophic bacteria in the global ocean. Environ Microbiol 9: 3091–3099.
Johnson ZI, Zinser ER, Coe A, McNulty NP, Woodward EMS, Chisholm SW . (2006). Niche partitioning among Prochlorococcus ecotypes along ocean-scale environmental gradients. Science (80-) 311: 1737–1740.
Jones RI . (2000). Mixotrophy in planktonic protists: an overview. Freshw Biol 45: 219–226.
Joset-Espardellier F, Astier C, Evans EH, Carr NG . (1978). Cyanobacteria grown under photoautotrophic, photoheterotrophic, and heterotrophic regimes: sugar metabolism and carbon dioxide fixation. FEMS Microbiol Lett 4: 261–264.
Kaiser K, Benner R . (2008). Major bacterial contribution to the ocean reservoir of detrital organic carbon and nitrogen. Limnol Oceanogr 53: 99–112.
Karimpour-Fard A, Leach SM, Gill RT, Hunter LE . (2008). Predicting protein linkages in bacteria: which method is best depends on task. BMC Bioinformatics 9: 397.
Karsenti E, Acinas SG, Bork P, Bowler C, De Vargas C, Raes J et al. (2011). A holistic approach to marine eco-systems biology. PLoS Biol 9: e1001177.
Keil R, Kirchman D . (1999). Utilization of dissolved protein and amino acids in the northern Sargasso Sea. Aquat Microb Ecol 18: 293–300.
Kelly L, Huang KH, Ding H, Chisholm SW . (2012). ProPortal: a resource for integrated systems biology of Prochlorococcus and its phage. Nucleic Acids Res 40: 632–640.
Kettler GC, Martiny AC, Huang K, Zucker J, Coleman ML, Rodrigue S et al. (2007). Patterns and implications of gene gain and loss in the evolution of Prochlorococcus. PLoS Genet 3: e231.
Kirchman D, Stegman M, Nikrad M, Cottrell M . (2014). Abundance, size, and activity of aerobic anoxygenic phototrophic bacteria in coastal waters of the West Antarctic Peninsula. Aquat Microb Ecol 73: 41–49.
Kolber ZS, Plumley FG, Lang AS, Beatty JT, Blankenship RE, Vandover CL et al. (1999). Contribution of aerobic photoheterotrophic bacteria to the carbon cycle in the ocean. Science 292: 2492–2495.
Kultima JR, Sunagawa S, Li JH, Chen WN, Chen H, Mende DR et al. (2012). MOCAT: a metagenomics assembly and gene prediction toolkit. PLoS One 7: e47656.
Lahmi R, Sendersky E, Perelman A, Hagemann M, Forchhammer K, Schwarz R . (2006). Alanine dehydrogenase activity is required for adequate progression of phycobilisome degradation during nitrogen starvation in Synechococcus elongatus PCC 7942. J Bacteriol 188: 5258–5265.
Lavin P, González B, Santibáñez JF, Scanlan DJ, Ulloa O . (2010). Novel lineages of Prochlorococcus thrive within the oxygen minimum zone of the eastern tropical South Pacific. Environ Microbiol Rep 2: 728–738.
Li B, Sher D, Kelly L, Shi Y, Huang K, Knerr PJ . (2010). Catalytic promiscuity in the biosynthesis of cyclic peptide secondary metabolites in planktonic marine cyanobacteria. Proc Natl Acad Sci USA 107: 10430–10435.
Logares R, Sunagawa S, Salazar G, Cornejo-Castillo FM, Ferrera I, Sarmento H et al. (2014). Metagenomic 16S rDNA Illumina tags are a powerful alternative to amplicon sequencing to explore diversity and structure of microbial communities. Environ Microbiol 16: 2659–2671.
Malmstrom RR, Coe A, Kettler GC, Martiny AC, Frias-Lopez J, Zinser ER et al. (2010). Temporal dynamics of Prochlorococcus ecotypes in the Atlantic and Pacific oceans. ISME J 4: 1252–1264.
Martiny AC, Coleman ML, Chisholm SW . (2006). Phosphate acquisition genes in Prochlorococcus ecotypes: evidence for genome-wide adaptation. Proc Natl Acad Sci USA 103: 12552–12557.
Martiny AC, Tai APK, Veneziano D, Primeau F, Chisholm SW . (2009). Taxonomic resolution, ecotypes and the biogeography of Prochlorococcus. Environ Microbiol 11: 823–832.
Mary I, Garczarek L, Tarran GA, Kolowrat C, Terry MJ, Scanlan DJ et al. (2008). Diel rhythmicity in amino acid uptake by Prochlorococcus. Environ Microbiol 10: 2124–2131.
Michelou VK, Cottrell MT, Kirchman DL, David L . (2007). Light-stimulated bacterial production and amino acid assimilation by cyanobacteria and other microbes in the North Atlantic ocean. Appl Environ Microbiol 73: 5539–5546.
Moore LR, Chisholm SW . (1999). Photophysiology of the marine Ecotypic cyanobacterium Prochlorococcus: differences among cultured isolates. Limnol Oceanogr 44: 628–638.
Moore LR, Rocap G, Chisholm SW . (1998). Physiology and molecular phylogeny of coexisting Prochlorococcus ecotypes. Nature 393: 464–467.
Mühling M, Fuller NJ, Millard A, Somerfield PJ, Marie D, Wilson WH et al. (2005). Genetic diversity of marine Synechococcus and co-occurring cyanophage communities: evidence for viral control of phytoplankton. Environ Microbiol 7: 499–508.
Mühling M, Fuller NJ, Somerfield PJ, Post AF, Wilson WH, Scanlan DJ et al. (2006). High resolution genetic diversity studies of marine Synechococcus isolates using rpoC1-based restriction fragment length polymorphism. Aquat Microb Ecol 45: 263–275.
Muñoz-Marín MDC, Luque I, Zubkov MV, Hill PG, Diez J, García-Fernández JM . (2013). Prochlorococcus can use the Pro1404 transporter to take up glucose at nanomolar concentrations in the Atlantic Ocean. Proc Natl Acad Sci USA 110: 8597–8602.
Paerl HW . (1991). Ecophysiological and trophic implications of light-stimulated amino acid utilization in marine picoplankton. Appl Environ Microbiol 57: 473–479.
Palenik B . (1994). Cyanobacterial community structure as seen from RNA polymerase gene sequence analysis. Appl Environ Microbiol 60: 3212–3219.
Palovaara J, Akram N, Baltar F, Bunse C, Forsberg J, Pedrós-Alió C et al. (2014). Stimulation of growth by proteorhodopsin phototrophy involves regulation of central metabolic pathways in marine planktonic bacteria. Proc Natl Acad Sci USA 111: 1–9.
Partensky F, Garczarek L, Hess WR, Vaulot D . (1999). Prochlorococcus, a marine photosynthetic prokaryote of global significance. Microbiol Mol Biol Rev 63: 106–127.
Partensky F, Hess WR, Vaulot D, Garczarek L . (2010). Prochlorococcus: advantages and limits of minimalism. Ann Rev Mar Sci 2: 305–331.
Perez RC, Matin A . (1980). Growth of Thiobacillus novellus on mixed substrates (Mixotrophic growth). J Bacteriol 142: 633–638.
Pesant S, Not F, Picheral M, Kandels-Lewis S, Le Bescot N, Gorsky G et al. (2015). Open science resources for the discovery and analysis of Tara Oceans data. Open Sci Resour Discov Anal Tara Ocean data Sci Data 2: 150023.
Raymond J, Blankenship RE . (2004). The evolutionary development of the protein complement of Photosystem 2. Biochim Biophys Acta - Bioenerg 1655: 133–139.
Rippka R . (1972). Photoheterotrophy and chemoheterotrophy among unicellular blue-green algae. Arch Mikrobiol 87: 93–98.
Rocap G, Larimer FW, Lamerdin J, Malfatti S, Chain P, Ahlgren NA et al. (2003). Genome divergence in two Prochlorococcus ecotypes reflects oceanic niche differentiation. Nature 424: 1042–1047.
Rogozin IB, Makarova KS, Wolf YI, Koonin EV . (2004). Computational approaches for the analysis of gene neighbourhoods in prokaryotic genomes. Brief Bioinform 5: 131–149.
Rothhaupt KO . (1996a). Laboratory experiments with a mixotrophic chrysophyte and obligately phago- trophic and phototrophic competitors. Ecology 77: 716–724.
Rothhaupt KO . (1996b). Utilization of substitutable carbon and phosphorus sources by the mixotrophic chrysophyte Ochromonas SP. Ecology 77: 706–715.
Rusch DB, Halpern AL, Sutton G, Heidelberg KB, Williamson S, Yooseph S et al. (2007). The Sorcerer II global ocean sampling expedition: northwest Atlantic through eastern tropical Pacific. PLoS Biol 5: e77.
Sabehi G, Loy A, Jung KH, Partha R, Spudich JL, Isaacson T et al. (2005). New insights into metabolic properties of marine bacteria encoding proteorhodopsins. PLoS Biol 3: e273.
Saier MH Jr, Tran CV, Barabote RD . (2006). TCDB: the Transporter Classification Database for membrane transport protein analyses and information. Nucleic Acids Res 34: D181–D186.
Saravolac EG, Taylor NF, Benz R, Hancock REW . (1991). Purification of glucose-inducible outer membrane protein OprB of Pseudomonas putida and reconstitution of glucose-specific pores. J Bacteriol 173: 4970–4976.
Schlitzer R . (2002). Interactive analysis and visualization of geoscience data with Ocean Data View. Comput Geosci 28: 1211–1218.
Sieracki ME, Gilg IC, Thier EC, Poulton NJ, Goericke R . (2006). Distribution of planktonic aerobic anoxygenic photoheterotrophic bacteria in the northwest Atlantic. Limnol Oceanogr 51: 38–46.
Sohm JA, Ahlgren NA, Thomson ZJ, Williams C, Moffett JW, Saito MA et al. (2015). Co-occurring Synechococcus ecotypes occupy four major oceanic regimes defined by temperature, macronutrients and iron. ISME J 10: 1–13.
Stamatakis A . (2006). RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22: 2688–2690.
Sun Z, Blanchard JL . (2014). Strong genome-wide selection early in the evolution of Prochlorococcus Resulted in a reduced genome through the loss of a large number of small effect genes. PLoS One 9: e88837.
Sunagawa S, Coelho LP, Chaffron S, Kultima JR, Labadie K, Salazar G et al. (2015). Structure and function of the global ocean microbiome. Science (80-) 348: 1–10.
Suzuki E, Umeda K, Nihei S, Moriya K, Ohkawa H, Fujiwara S et al. (2007). Role of the GlgX protein in glycogen metabolism of the cyanobacterium, Synechococcus elongatus PCC 7942. Biochim Biophys Acta 1770: 763–773.
Team RDC. (2012) R: A Language and Environment for Statistical Computing.
Thompson JW . (2015) Prochlorococcus: life in light. (Doctoral dissertation). Retrieved from DSpace MIT Thesis Database.
Vila-Costa M, Simó R, Harada H, Gasol JM, Slezak D, Kiene RP . (2006). Dimethylsulfoniopropionate uptake by marine phytoplankton. Science 314: 652–654.
West NJ, Scanlan DJ . (1999). Niche-partitioning of Prochlorococcus populations in a stratified water column in the eastern North Atlantic Ocean. Appl Environ Microbiol 65: 2585–2591.
West NJ, Schonhuber WA, Fuller NJ, Amann RI, Rippka R, Post AF et al. (2001). Closely related Prochlorococcus genotypes show remarkably different depth distributions in two oceanic regions as revealed by in situ hybridization using 16S rRNA-targeted oligonucleotides. Microbiology 147: 1731–1744.
Ye Y, Choi J-H, Tang H . (2011). RAPSearch: a fast protein similarity search tool for short reads. BMC Bioinformatics 12: 159.
Yelton AP, Thomas BC, Simmons SL, Wilmes P, Zemla A, Thelen MP et al. (2011). A semi-quantitative, synteny-based method to improve functional predictions for hypothetical and poorly annotated bacterial and archaeal genes. PLoS Comput Biol 7: e1002230.
Yooseph S, Sutton G, Rusch DB, Halpern AL, Williamson SJ, Remington K et al. (2007). The Sorcerer II global ocean sampling expedition: expanding the universe of protein families. PLoS Biol 5: 0432–0466.
Zinser ER, Coe A, Johnson ZI, Martiny AC, Fuller NJ, Scanlan DJ et al. (2006). Prochlorococcus ecotype abundances in the North Atlantic Ocean as revealed by an improved quantitative PCR method. Appl Environ Microbiol 72: 723–732.
Zinser ER, Johnson ZI, Coe A, Karaca E, Veneziano D, Chisholm SW . (2007). Influence of light and temperature on Prochlorococcus ecotype distributions in the Atlantic Ocean. Limnol Oceanogr 52: 2205–2220.
Zubkov MV, Fuchs BM, Tarran GA, Burkill PH, Amann R . (2003). High rate of uptake of organic nitrogen compounds by Prochlorococcus cyanobacteria as a key to their dominance in oligotrophic oceanic waters. Appl Environ Microbiol 69: 1299–1304.
Zubkov MV, Tarran GA, Fuchs BM . (2004). Depth related amino acid uptake by Prochlorococcus cyanobacteria in the Southern Atlantic tropical gyre. FEMS Microbiol Ecol 50: 153–161.
Zubkov MV . (2009). Photoheterotrophy in marine prokaryotes. J Plankton Res 31: 933–938.
Zubkov MV, Tarran GA, Mary I, Fuchs BM . (2008). Differential microbial uptake of dissolved amino acids and amino sugars in surface waters of the Atlantic Ocean. J Plankton Res 30: 211–220.
Zwirglmaier K, Jardillier L, Ostrowski M, Mazard S, Garczarek L, Vaulot D et al. (2008). Global phylogeography of marine Synechococcus and Prochlorococcus reveals a distinct partitioning of lineages among oceanic biomes. Environ Microbiol 10: 147–161.
Acknowledgements
We thank the US National Science Foundation OCE postdoctoral research fellowship program and the Fulbright Commission, Spain for supporting APY. The work was also supported in part by the European Molecular Biology Laboratory, grants to SWC from the Gordon and Betty Moore Foundation (grant GBMF495) the National Science Foundation (grants OCE-1356460 and DBI-0424599), grants from the Simons Foundation (grant 337262), the Spanish Ministry of Science and Innovation grant to SGA, CGL2011-26848/BOS MicroOcean PANGENOMICS and U FP7-OCEAN.2011-2. Micro3B Marine Microbial Biodiversity, the Bioinformatics and Biotechnology Large Collaborative grant 287589 and is a contribution of the Simons Collaboration on Ocean Processes and Ecology (SCOPE). We are indebted to the scientists and crew of the Tara Oceans expedition.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Yelton, A., Acinas, S., Sunagawa, S. et al. Global genetic capacity for mixotrophy in marine picocyanobacteria. ISME J 10, 2946–2957 (2016). https://doi.org/10.1038/ismej.2016.64
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2016.64
This article is cited by
-
Phototroph-heterotroph interactions during growth and long-term starvation across Prochlorococcus and Alteromonas diversity
The ISME Journal (2023)
-
Elucidating the picocyanobacteria salinity divide through ecogenomics of new freshwater isolates
BMC Biology (2022)
-
Mixotrophy in depth
Nature Microbiology (2022)
-
Single-cell measurements and modelling reveal substantial organic carbon acquisition by Prochlorococcus
Nature Microbiology (2022)
-
Novel functional insights into a modified sugar-binding protein from Synechococcus MITS9220
Scientific Reports (2022)