Abstract
Little research has been conducted on microbial diversity deep under the Earth’s surface. In this study, the microbial communities of three deep terrestrial subsurface aquifers were investigated. Temporal community data over 6 years revealed that the phylogenetic structure and community dynamics were highly dependent on the degree of isolation from the earth surface biomes. The microbial community at the shallow site was the most dynamic and was dominated by the sulfur-oxidizing genera Sulfurovum or Sulfurimonas at all-time points. The microbial community in the meteoric water filled intermediate aquifer (water turnover approximately every 5 years) was less variable and was dominated by candidate phylum OD1. Metagenomic analysis of this water demonstrated the occurrence of key genes for nitrogen and carbon fixation, sulfate reduction, sulfide oxidation and fermentation. The deepest water mass (5000 year old waters) had the lowest taxon richness and surprisingly contained Cyanobacteria. The high relative abundance of phylogenetic groups associated with nitrogen and sulfur cycling, as well as fermentation implied that these processes were important in these systems. We conclude that the microbial community patterns appear to be shaped by the availability of energy and nutrient sources via connectivity to the surface or from deep geological processes.
Similar content being viewed by others
Introduction
Although microbial life in the deep biosphere was discovered in the early 1930s (Lipman, 1931), this environment has only recently become the subject of more extensive biological investigation. This lack of study was mainly due to the serious logistic challenge of obtaining valid biological samples without introducing contaminants via drilling and equipment (Wilkins et al, 2014). To distinguish the ‘shallow’ terrestrial biosphere from the ‘deep’, the classification scheme used is based on the degree of hydrological connectivity to the surface rather than actual depth (Lovley and Chapelle, 1995). Accordingly, organisms at the surface and in aquifers directly connected to the surface are rapidly influenced by fluctuations in nutrient availability and input of photosynthetically derived organic carbon. In contrast, the deep biosphere is oligotrophic and independent of recent photosynthetic primary production (Edwards et al., 2012). As fresh organic carbon from the surface is not available in these deep environments, the resident microorganisms rely on energy sources deposited in the sediment, transported by water through long-term recharge from the surface, or leached from the surrounding rocks. With the exception of local patches of enhanced organic material levels that reside in pocket formations in the rock, it is likely that microorganisms deplete nutrients from the water and convert organic matter from labile to refractory as it travels down from the surface (Lovley and Chapelle, 1995). All of these factors push the cells toward long generation times and slow metabolic turnover rates (Jørgensen, 2012). Nevertheless, recent estimates state that potentially 2–19% of the Earth’s total biomass lies in the deep terrestrial subsurface (McMahon and Parnell, 2014).
Microorganisms can gain energy through oxidation of organic matter and reduced electron donors such as Mn(II), Fe(II), ammonia and sulfide. In the absence of oxygen, organic compounds will be either fermented and/or oxidized coupled to the reduction of nitrate, Mn(IV), Fe(III), sulfate and carbon dioxide (Lovley and Chapelle, 1995). It is hypothesized that when the above-mentioned compounds are not available or have been consumed, hydrogen produced by either fermentation or abiotic radiolysis of water is used as the main energy source (Hallbeck and Pedersen, 2008). Possible key factors in these oligotrophic hydrogen fueled environments include sulfate/sulfur reducers; acetogenic bacteria that couple hydrogen oxidation with carbon dioxide reduction to acetate (homoacetogens); and methanogens that either yield methane from hydrogen and carbon dioxide (autotrophic methanogens) or from acetate produced by the homoacetogens (acetoclastic methanogens).
The field site for this study is the Äspö Hard Rock Laboratory (Äspö HRL), operated by the Swedish Nuclear Fuel and Waste Management Co. (SKB). The Äspö HRL is an underground research facility located in the southeast of Sweden, which consists of a 3600 m long tunnel going down to a depth of 450 m under the Äspö island (Ionescu et al, 2015). The geological formation of this area is referred to as the Fennoscandian Shield, which is dated to be between 1.6 and 3.1 billion years old (Ström et al., 2008). The bedrock mainly consists of granite and quartz-monzodiorite, suffered from different orogenies and late glaciations which caused several fracture reactivations events. Different Quaternary aged fluids like marine, brackish or glacial waters penetrated these fractures thus leading to the formation of various aquifers. The Äspö site has been extensively studied in terms of its geology, hydrology and chemistry (Rönnback, 2005; Söderbäck, 2008; Drake and Tullborg, 2009).
Results from previous microbiological studies suggest that this anaerobic (microaerophilic) oligotrophic environment is mainly populated by nitrate-, ferric-, sulfate- and manganese-reducing microorganisms along with acetogens and methanogens (Kotelnikova and Pedersen, 1997; Motamedi and Pedersen, 1998; Kotelnikova and Pedersen, 1998; Jägevall et al., 2011; Pedersen, 2012). One metagenomics study investigating water types of different ages at the Äspö HRL identified communities of heterotrophic, mixotrophic and autotrophic populations suggested to exhibit a plethora of different growth strategies (Wu et al., 2015). Other studies include investigations on carbon cycling genes (Purkamo et al., 2015) and the taxonomy and function of microbes located in the crystalline crust of Outokumpu, Finland (Nyyssonen et al., 2014), all of which suggest a predominant chemolithoautotrophic lifestyle.
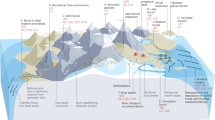
Three water masses at the Äspö HRL that differed in water chemistry and retention time were chosen (Ionescu et al., 2015) (Figure 1). Modeling of mixing components (that is, isotopes d18O; 87Sr/86Sr; 3H; and d34S) and rare earth element fractionation patterns did not indicate a connection or mixing between the respective aquifers (Hengsuwan et al., 2015). The isotopic signature (d18O; 87Sr/86Sr; 3H) of the shallow aquifer (HA1327B) at a depth of 183 m points to a recharge of Baltic Seawater and meteoric water within weeks to years (Ionescu et al., 2015). This water is characterized by low salinity (0.55%), high iron (30 μM) and the highest sulfide (10.5 μM) content of the three aquifers. The intermediate aquifer (KA2162B:1) is located 290 m below the surface and has a hydrological retention time of about 5 years and mainly holds meteoric water with slightly higher salinity (0.59%) along with lower iron (12 μM) and sulfide (0.9 μM) concentrations (Ionescu et al., 2015). The deep water mass (KF0069A01) at 450 m below the surface consists of 4000- to 5000-year-old highly saline glacial melt and meteoric waters (salinity 2.5%) with the lowest iron (<1.5 μM) and sulfide (0.1 μM) contents (Ionescu et al., 2015). The lack of tritium in the deep aquifer shows that there is no connection to modern groundwater sources (Hengsuwan et al., 2015).
The Äspö HRL sampling site showing the three analyzed boreholes. 3D-figure modified after Laaksoharju and Wold (2005).
In contrast to a previous study where bioreactors were connected to the three boreholes to identify biofilm producers within the aquifer (Ionescu et al., 2015), the aim of this study was to observe temporal changes in community structure of the pristine aquifers over a time span of several years. Our results highlight that connectivity to the surface and input from solar-exposed ecosystems are a major driver of microbial community composition and dynamics.
Materials and methods
Sampling and DNA extraction
Sampling and extraction of microbial DNA has been previously described (Ionescu et al, 2015). The three aquifers were sampled at least yearly from 2006 until 2010 at the following Äspö HRL tunnel sites: HA1327B, a shallow aquifer at −183 m (tunnel section A; 1327 m from the entrance); an intermediate aquifer at KA2162B:1 (−290 m; 2156 m from the tunnel entrance); and a deep aquifer at KF0069A01 (−455 m depth; 3600 m from the tunnel entrance). To avoid external contamination, all sampling equipment was sterilized. As the water continuously flowed through the boreholes for several years at a rate of 0.8–1.0 l min−1, keeping the drill hole constantly full, contamination inserted during the drilling was highly unlikely. Samples were stored frozen at −20 °C until further processing (Ionescu et al., 2015). Following filtration of 250–5000 ml of aquifer water onto polycarbonate filters (0.22 μm pore size; 47 mm Ø; Whatman, Maidstone, UK), an iron dissolution solution (0.35 M acetic acid; 0.2 M sodium citrate; 25 mM sodium dithionite; Thamdrup et al., 1993) was added to the filtration tower. After 5 min of incubation, the solution was removed by vacuum and the filter was washed with 100 ml of sterile 1 × phosphate-buffered saline (Ionescu et al., 2015). DNA extraction was performed with a hot-phenol method (Ionescu et al., 2012) and the resulting DNA extracts had concentrations in the range of 0.2–150 ng μl–1 (Supplementary Data S1). At each sampling point, detailed physical and chemical parameters were measured (Hengsuwan et al., 2015), revealing that these chemical parameters remained relatively stable over the study period (Supplementary Datas S2-S3).
16S rRNA gene amplicon sequencing
Microbial 16S ribosomal RNA (rRNA) gene amplicons were sequenced on the MiSeq (Illumina, San Diego, CA, USA) platform according to published procedures (Sinclair et al., 2015). In brief, partial 16S rRNA gene sequences were obtained using a two-step PCR procedure with primers targeting the V3 and V4 regions of the 16S rRNA gene. The applied primers Bakt_341F (5'-CCTACGGGNGGCWGCAG-3') and Bakt_805R (5'-GACTACHVGGGTATCTAATCC-3') were originally designed for pyrosequencing (Herlemann et al., 2011). The first amplification was carried out for 20 cycles using primers without barcodes to minimize primer-induced PCR bias (Sinclair et al., 2015). The obtained PCR products were then diluted and used as template in a second 10-cycle PCR with identical primers, except that both forward and reverse primers included 7-bp DNA barcodes that were unique for each amplified sample. This two-step amplification regime also reduces PCR-based artifacts (Thompson et al., 2002). PCR products carrying unique sample-specific barcodes were then pooled and purified using Qiagen gel purification kit (Qiagen, Hilden, Germany) and submitted to the SciLifeLab SNP/SEQ sequencing facility (Uppsala University) where the routine TruSeq protocol was applied (Illumina I, 2013). A previously described in-house sequence analysis and annotation pipeline was used (Sinclair et al., 2015). The raw amplicon data were deposited at the ENA database (ERP011840).
Shotgun metagenomics
DNA from one sampling campaign in 2008 of the intermediate site (borehole KA2162B:1) and one sample from 2007 of the deep borehole (KF0069A01) was submitted to the Science for Life Laboratory SNP/SEQ facility in Uppsala, Sweden, where metagenome library construction (Lundin et al., 2010; Borgström et al., 2011) and Illumina HiSeq sequencing was performed. First, a non-quantifiable (very low) amount of genomic DNA was sheared using a focused-ultrasonicator (Covaris E220, Woburn, MA, USA) from five samples. Sequencing libraries were prepared with the Thruplex FD Prep kit from Rubicon Genomics (Ann Arbor, MI, USA) according to the manufacturer's protocol (R40048-08, QAM-094-002). Library size selection was made with AMPpure XP beads (Beckman Coulter, Beverly, MA, USA) at a 1:1 ratio. The prepared sample libraries were quantified using KAPA Biosystem next-generation sequencing library qPCR kit and run on a StepOnePlus (Thermo Fisher Scientific, Waltham, MA, USA) real-time PCR instrument. This resulted in two libraries usable for sequencing on the Illumina HiSeq sequencing platform utilizing a TruSeq paired-end cluster kit v3 and Illumina’s cBot instrument to generate a clustered flowcell for sequencing. Sequencing of the flowcell was performed on the Illumina HiSeq2500 sequencer using Illumina TruSeq SBS sequencing kits v3, following a 2 × 100-bp high-output run protocol.
The obtained reads were trimmed with Sickle (version 1.33) (Joshi and Fass, 2011) and subsequent assembly was performed with MEGAHIT (v0.3.2) (Li et al., 2015). Genes were predicted using the Prokka software (Seemann, 2014) and mapped back to protein coding sequences (CDS) by Bowtie 2 (Langmead and Salzberg, 2012) to obtain a read count per CDS. HMMER (version 3.1b1) (Finn et al., 2011) was used with default parameters to match Pfams to the CDS. The resulting data set was screened for Pfams associated with key genes for fermentation, sulfate reduction, sulfide oxidation, methanogenesis, hydrogenase activity, nitrogen and carbon fixation. The raw metagenome data were deposited at the ENA database (ERP011782).
Statistical analysis
Before comparative diversity analyses, operational taxonomic unit (OTU) tables of the different aquifers were rarefied to the smallest sample size. When comparing community composition, rarefaction was performed separately for each individual water mass. Alpha diversity indices such as ChaoI, Simpson’s diversity index (1/D) and Simpson’s evenness (E1/D) index were calculated using the vegan package in R (Dixon, 2003). Nonmetric dimensional scaling on Bray–Curtis dissimilarity matrices was used to visualize community patterns. The functions adonis, betadis and envfit from the vegan package (Dixon, 2003), as well as function aovp from the lmPerm package (Wheeler, 2010) were used to relate patterns in bacterial diversity with environmental properties.
Species co-occurrence networks
Species–species association networks were constructed using CoNet (Faust et al., 2012) as implemented in Cytoscape 3.3.0 (Shannon et al., 2003) software. This program was chosen as it allows the merging of networks calculated via several association methods of which we chose: Kendall, Pearson and Spearman correlations, as well as Steinhaus and variance log ratio similarities. The latter results in values between 0 and 1, whereas the rest range between −1 and 1. Only edges confirmed by at least two methods each with a score either smaller than −0.6 (negative association) or larger than 0.6 (positive association) were retained in the final network. The resulting network was visualized and graphically organized using Cytoscape 3.3.0 (Shannon et al., 2003).
Results
16S rRNA gene amplicon sequencing data
From the amplicon sequencing run, 2.4 million reads were obtained of which 1.5 million remained after sequence cleanup and quality control. Ninety percent of these reads have a base call accuracy of 99.9%. The 472-bp length sequences were assigned to 6561 OTUs at 98% identity level.
Bacterial community clustering
Comparison of bacterial OTU clustering with environmental variables via nonmetric multidimensional scaling analysis showed a strong positive correlation with sulfate content and salinity (Figure 2). In contrast, iron content was not significantly related to community composition. Furthermore, the nonmetric multidimensional scaling analysis grouped the samples into three distinct clusters based on the origin of the sample, with the two shallowest aquifers being most similar to each other (nonmetric dimensional scaling stress=0.11). This distinction between water masses was confirmed by permutational multivariate analysis of variance (R2=0.31, P<0.001). In addition, the analysis revealed that community composition differed significantly between years (R2=0.30, P<0.001). As there was an interaction effect between water mass and year on community structure (R2=0.26, P<0.001), it was tested if the temporal community dynamics differed between the boreholes. Permuted beta-dispersion to test homogeneity of multivariate dispersion among the water masses confirmed that the dynamics of the two shallower aquifers were different from that of the deepest, more variable borehole (Figure 3a). Species richness and evenness also significantly differed between the boreholes. No significant variability between replicates of the same sampling campaign was observed suggesting that these differences were not caused by sampling. The highest and lowest Chao1 richness were observed in the shallow and deep aquifer, respectively (Figure 3b). In contrast, the highest Simpson diversity index (1/D) was found in the intermediate aquifer (Figure 3c). Simpson’s evenness index (E1/D) increased with the depth of the aquifers (Figure 3d).
Microbial community structure of the three boreholes and associated environmental factors. Circles represent the shallow water (HA1327B), triangles for the intermediate water (KA2162B:1) and crosses for the deepest aquifer (KF0069A01). A nonmetric multidimensional scaling analysis of Bray–Curtis distances was overlaid with the result of envfit to relate patterns in bacterial diversity with environmental properties.
Bacterial community composition
The sequence data, particularly from the deep aquifer contained rRNA sequences matching known skin or reagent contaminants: Propionibacterium, Corynebacterium, Staphylococcus, Streptococcus, Bradyrhizobium and Hoeflea. Following a phylogenetic analysis to evaluate whether these sequences are contaminants or rather environmental strains, the first four taxa were removed from further analyses of intra-species association analysis. The Bradyrhizobium and Hoeflea sequences were 99–100% identical to different environmental strains with physiological properties suitable for the studied aquifers and were thus retained. Nevertheless, data regarding these taxa should be viewed with caution.
The shallow aquifer (HA1327B) featured a high proportion of 16S rRNA gene reads most similar to Epsilonproteobacteria, most of which were classified as Sulfurovum sp. and Sulfurimonas sp. (Figure 4). Both of these genera are putatively involved in sulfur and nitrate metabolism, as well as hydrogen oxidation (Campbell et al., 2006). Another abundant group in the shallow aquifer was candidate division OD1 (Parcubacteria; 8–16% of the 16S rRNA reads), which are anaerobic/microaerophilic microbes typically associated with sulfur containing milieu (Peura et al., 2012; Kantor et al., 2013). The highest OTU diversity (~ 100 OTUs) affiliated with this group, some of which showed ongoing dynamics of their own. The occurrence of OD1 was strongly negatively correlated to Sulfurimonas (r=–0.76) and had a moderately positive correlation to Sulfurovum (r=0.54).
The highest proportion of 16S rRNA gene reads in the intermediate water mass (KA2162B:1) were most similar to sequences from candidate phylum OD1 (15–63% relative abundance). However, the intermediate aquifer OD1 population had a reduced OTU richness (~60 OTUs) of which half of the OTUs were also present in the shallow aquifer. The second most abundant class in the intermediate water mass was represented by 16S rRNA gene sequences most similar to the Gram-negative, nitrogen-fixing genus Bradyrhizobium (≤16%). Overall, despite the water being replenished every 5 years, there were no clear shifts in abundant groups during the course of the 6-year experiment.
The deepest sampling site (borehole KF0069A01) had a fluctuating community composition with the highest proportion of unknown 16S rRNA gene sequences (≤12%), whereas sequences assigned to candidate phylum OD1 were found in relatively high abundances (≤14%). Although candidate phylum OD1 had the least diversity (nine OTUs) in borehole KF0069A01, the 16S rRNA gene sequences constituted unique OTUs not found at the other sampling sites. Some of the most abundant OTUs classified as OD1 were only distantly related to sequences in the available databases and possibly represent novel lineages. In addition, 16S rRNA gene sequences were also identified and most closely resembled the purple non-sulfur bacteria Rhodospirillaceae (≤54%) and Roseovarius spp. (≤6%). Unexpectedly, OTUs related to Cyanobacteria, mostly annotated within the Oscillatoriales and Synechococcaceae, were identified in almost every sample from the deepest aquifer. Despite that the water at this site has not seen sunlight for thousands of years, it nevertheless had the highest relative abundance of Cyanobacteria of the three sites. The cyanobacterial 16S rRNA gene reads contributed up to 16% of all reads in the deep aquifer, 1% of the reads in the intermediate aquifer and only 0.4% in the shallow water mass.
Interspecies associations
Only the shallow aquifer had a clear succession with a strong negative correlation between the genera Sulfurimonas and Sulfurovum (r=–0.74). During the first 8 months of the study, Sulfurimonas was found in almost every sample and dominated the system with up to 83% of all of the total 16S rRNA gene reads, whereas Sulfurovum was either undetectable or in low abundance (Figure 5). Over the course of approximately 1 year, this relationship was reversed with 16S rRNA gene reads most similar to Sulfurimonas becoming undetectable, whereas Sulfurovum increased from 0% to 56% of all reads. Another effect of the decrease of Sulfurimonas was a short-lived increase of sequences most related to the genus Thiobacillus (maximum <10% relative abundance). Concomitant with this shift, there was a decrease in sulfate content and to a lesser extent in oxygen concentration (Supplementary Data S2). To identify less evident species–species associations, we used a network analysis approach (Figures 5,6,7). Alongside the succession between Sulfurimonas and Sulfurovum in the shallow aquifer (Figure 5, negative correlation), these genera interacted differently with two groups of sulfate reducers. Both Desulfarculaceae and Desulfovibrionaceae were negatively associated with Sulfurimonas and positively associated with Sulfurovum. OTUs affiliated to Rhodospirillaceae, Phenylobacterium, Hyphomicrobiaceae and candidate divisions OD1 and OP11 were positively associated to Marinicella that was in turn negatively associated with Sulfurovum (Figure 5).
Interspecies association network of the shallow aquifer bacterial community, supported by two or more statistical methods (see legend) with a result of <−0.6 (negative association) or <0.6 (positive association). The temporal change in abundance of associated OTUs is given in the lower graphs. For illustration purposes, each OTU shown in the temporal graphs was normalized to its own maximum abundance. This value is provided in the graph legend.
Interspecies association network of the intermediate aquifer bacterial community, supported by two or more statistical methods (see legend) with a result of <−0.6 (negative association) or <0.6 (positive association). The temporal change in abundance of associated OTUs is given in the lower graphs. For illustration purposes, each OTU shown in the temporal graphs was normalized to its own maximum abundance. This value is provided in the graph legend.
Interspecies association network of the deep aquifer bacterial community, supported by two or more statistical methods (see legend) with a result of <−0.6 (negative association) or <0.6 (positive association). The temporal change in abundance of associated OTUs is given in the lower graphs. For illustration purposes, each OTU shown in the temporal graphs was normalized to its own maximum abundance. This value is provided in the graph legend.
The intermediate aquifer species association network was dominated by positive associations, such as between Sphingomonas, Sediminibacterium and Bradyrhizobium (Figure 6). One exception was Brevundimonas that was negatively associated with several candidate division OD1 and OP11 OTUs and an unresolved Gammaproteobacteria order.
In the deepest aquifer, Hoflea was negatively associated with Bradyrhizobium and positively associated with Roseovarius and an abundant Gammaproteobacteria OTU with unresolved taxonomy. Pseudomonas was negatively associated with Sphingomonas and a Rhodobacteraceae OTU, whereas the latter two were positively associated to each other (Figure 7).
Metabolic potential of the deep aquifers
Only metagenomes from the intermediate and deepest water mass were obtained, producing 81 million and 84 million reads after quality control and filtering, respectively. The metabolic potential of these two systems was inferred from the presence of certain phylogenetic groups found in the amplicon data and specific marker genes associated with their potential energy and biomass production were searched in the metagenomes to confirm these inferences.
The intermediate water mass contained gene sequences encoding pathway components for energy production via the fermentation of glucose (pgi, pfk, fda and fba), maltose (malQ), starch/glycogen (glgA and Rv3032), pyruvate and acetate (porCD), as well as nitrogen assimilation (gltB) and fixation (nifH, nifU) and nitrate reduction (nirk). Key genes for sulfide oxidation (soxFVW), sulfate reduction (dsrAB and aps) and hydrogenase activity (hypCDF) were present, along with contigs that had high similarity with marker genes for methanogenesis from acetate (pta and ackA), methanol (mtaA), as well as CO2 and H2 (fwdB). Furthermore, contigs with high similarity to cdhC, a key gene for the reductive acetyl-CoA pathway for carbon fixation into biomass (Yau et al, 2013) were detected (Supplementary Data S4). The deepest aquifer metagenome also contained the same marker genes for the above-mentioned pathways plus other genes coding for enzymes involved in hydrogenase activity (hypA), pyruvate fermentation to acetate (porC), enzymes involved in carbohydrate metabolism (rbsK, porC, gpmA and pgk/tpi) and methanogenesis from methanol (mtaB; Supplementary Data S5). A complete analysis of the metagenomic data from the intermediate and deep site will be discussed elsewhere.
Discussion
Bacterial diversity patterns
Modeling has suggested that the three aquifers were unconnected (Hengsuwan et al., 2015) and evaluation of the physicochemical parameters measured in this study amended with data from the SKB Sicada database showed that water chemistry only featured minor fluctuations during the sampling period (Supplementary Data S2 and S3). Thus, we concluded that the aquifers are unconnected and therefore, the patterns in diversity and community composition discussed below are independent among sites.
The shallow aquifer was the most fluctuating environment (based on chemical and physical properties) over the 6-year sampling period as it is under the influence of monthly water recharge from cracks in the surrounding rocks, carrying fresh nutrients and biomass from the Baltic Sea. Given the high richness and low evenness in the shallow aquifer, we propose that the bacterial community in this water mass consists of few dominant species and a majority of less abundant populations. As succession events such as the highlighted Sulfurimonas-Sulfurovum example was evident in the shallow aquifer, it is likely that the composition of the low-abundance majority changes with time. The high richness in the shallow aquifer is most likely a result of two factors: (1) an increased number of ecological niches because of availability of more resources and gradients and (2) a carryover from the source waters of organisms that do not form abundant populations in this aquifer.
The decreasing richness with depth of the aquifer alongside an increase in evenness likely reflects the outcome of connectivity-dependent mass effects, reducing the number of available ecological niches and minimizing the presence of populations unable to establish stable communities. The old saline glacial melt and meteoric waters in the deepest aquifer are more or less isolated from such transient bacteria. In this case, 5000 years of isolation and species filtering are the main factors leading to marked species loss.
In contrast to richness, evenness increased with depth. This could be a consequence of increased nutrient limitation and seclusion. The high dissolved organic carbon/dissolved organic nitrogen ratio (<70; Supplementary Data S2) means that the organic matter is rather recalcitrant, suggesting carbon was also limiting. As species immigration becomes extremely rare with depth, interactions between community members would be tighter and more indispensable under such oligotrophic conditions (Hillebrand et al., 2008; Wittebolle et al., 2009). Preceding succession events could have already taken place and eliminated dominant species through competition and/or predation. Other studies have indicated high viral activity in the Äspö aquifers (Kyle et al., 2008; Eydal et al., 2009), which implies that such mechanisms could be a significant factor in shaping microbial communities in these depths.
In a parallel investigation of bioreactors connected to the same three boreholes (Ionescu et al., 2015), a decrease in richness and Shannon index of the biofilm forming biomass was observed with increasing depth. Since the time this study was conducted the bioreactors were functioning for ca 4 years, the decrease in both indices suggests that given sufficient time for the communities to become self-dependent, patterns of richness are eventually reflected by diversity indices.
We are aware that the assertion of a low resource environment leading to low community turnover in the deeper water masses is contradicted by the result of the beta-dispersion analysis (Figure 3a), which shows that the highest community turnover was in fact in the deepest aquifer. However, we have to take into account that as the specimens were taken from borehole outlets, each sample may have consisted of a mixture of detached biofilm patches from within the aquifer and planktonic cells. Although we tried to bypass this methodological issue by collecting replicate samples at each time point, the patchiness and low biomass may have produced this result, rather than reflecting a dynamic community.
Community composition
The shallow water mass was dominated by 16S rRNA gene sequences assigned to genera involved in sulfur cycling and within the monitored time frame a succession was observed in which a dominating Epsilonproteobacterium, Sulfurimonas-like population was replaced by a Sulfurovum-like population. Epsilonproteobacteria are known to dominate sulfidic ecosystems such as deep-sea hydrothermal environments (Nakagawa and Takai, 2008; Yamamoto and Takai, 2011) and can also act as symbionts of marine invertebrates (Grzymski et al., 2008). This group is proposed to be the primary producer in aphotic sulfuric environments that in turn provide organic carbon to the rest of the community through CO2 fixation via the reverse tricarboxylic acid cycle (Campbell et al., 2006; Mattes et al., 2013; Hamilton et al., 2014). The genera Sulfurovum and Sulfurimonas are phylogenetically well separated but largely overlapping in their functionality. Both are metabolically diverse, microaerophilic chemolithoautotrophs that can use sulfur compounds both as electron donor or acceptor and additionally use nitrate or oxygen reduction pathways (Yamamoto and Takai, 2011).
We present here, to the best of our knowledge, the first description of a succession event between these two genera, which are commonly observed together in the environment (Nakagawa and Takai, 2008; Yamamoto and Takai, 2011). This shows that despite their apparent overlapping functionality and niche, they represent distinct ecotypes. The decline of the Sulfurimonas-like taxa follows an increase in oxygen and sulfide levels in the aquifer, thus it is possible that these conditions favored Sulfurovum, leading to the community shift. Otherwise, the measured chemical data did not show major shifts that could explain this pattern. Interestingly, the three main sulfate reducers detected in this aquifer also increased in relative abundance upon the change from a Sulfurimonas to a Sufurovum dominated community (Figure 5).
A second set of organisms that overlap the Sulfurimonas and disappear with the increase in Sulfurovum abundance consisted of Marinicella, Phenylobacterium, and unresolved members of the Rhodospirillaceae and Hyphomicrobiaceae families. We hypothesize that the changes in dissolved organic matter quality is partly responsible for these changes in bacterial community composition. Phenylobacterium, for example, is known to be limited to a narrow range of organic compounds (Eberspächer and Lingens, 2006). Marinicella grows best on rich media supplemented with casein (Romanenko et al., 2010) and Rhodospirillaceae and Hyphomicrobiaceae also depend on an organic-rich environment. Their disappearance accompanied a decrease in dissolved organic matter quantity, as well as quality as indicated from the increasing dissolved organic carbon/dissolved organic nitrogen ratio (Supplementary Data S2). The latter ratio, affected by the consumption of amino groups from proteins (Thurman, 1985), is a measure for the lability of the organic material with higher numbers being characteristic of more recalcitrant compounds (Hopkinson et al., 2002). The suggested change in dissolved organic matter properties could also lead to increased abundance of fermenting sulfate reducers such as the Desulfarculaceae.
Another abundant group, previously associated with anaerobic and sulfur-rich environments (Wrighton et al., 2012), is the candidate phylum OD1, which was highly abundant in all three water masses, although there was minor overlap in OTUs with the deep aquifer, which hosted a unique set of OD1 populations. This agrees with the diverse metabolic potential within the phylum, ranging from degradation of complex carbon molecules (fermentation) to hydrogen production and sulfur reduction (Wrighton et al., 2012). Network analysis also revealed three main sets of associated OTUs (Figure 6). However, it was not possible to explain the clustering and the temporal distribution based on the physiological traits of the few hitherto metabolically characterized taxa or by the measured environmental parameters.
An unexpected finding in this investigation was the occurrence of cyanobacteria-like OTUs in the deepest ancient waters, mainly clustering to the order Oscillatoriales. Studies of ancient microbial activity in fracture minerals at 450 m depth dated water injections from the surface to the late Pleistocene, as a result of glacial rebound (Heim et al., 2012). We believe that these cyanobacteria-related OTUs are viable, as several members of the Oscillatoriales order, with high sequence similarity to our data, formed thick mats in bioreactors attached to the same borehole (Ionescu et al., 2014). The same study showed that the cyanobacteria in the reactors are different from those growing on the tunnel walls as a result of cross-tunnel transport, thus strengthening the claim that the bioreactor cyanobacteria originate from the aquifer water. Moreover, in a comparable study, Vishnivetskaya (2009) was able to cultivate ancient Oscillatoriales from the arctic subsurface permafrost in Siberia. Cyanobacteria are ubiquitous in all aquatic environments and it is possible that a cyanobacterial bloom at the surface was trapped in the aquifer thousands of years ago. Whether these phototrophs survived because of their ability to switch to a fermentative lifestyle (Anderson and McIntosh, 1991) or they were less prone to degradation cannot be determined with the data at hand. Alternatively, as Nostocaceae formed the major part of the Cyanobacteria sequences, the formation of akinetes may have allowed for this long-term survival. The relatively high abundance of sequences related to Cyanobacteria may be due to generation times being in the order of hundreds to thousands of years (Hoehler and Jorgensen, 2013), resulting in only a few generations potentially having occurred as the water was trapped and thus, reflecting cyanobacterial blooms in the ancient source water.
Network analysis of the deep aquifer bacterial community suggested an association between the metabolically versatile Sphingomonas (Balkwill et al., 2006) and a Rhodobacteraceae OTU (Figure 7). The closest relatives of the latter in the NCBI database (99% similarity) were Rhodobacter species isolated from hypersaline oil contaminated environments. This supports our assumption that the organic matter was refractory as already suggested by high dissolved organic carbon/dissolved organic nitrogen ratio (Supplementary Data S2). In contrast, the negatively associated Pseudomonas OTU matches sequences from low dissolved organic matter environments and deep subsea sediments.
Metabolic potential of the microbial communities
The metabolic pathways emerging from the metagenomic data go hand in hand with the microbial community discussed. For example, sulfide oxidizers and sulfate reducers were abundant in both upper aquifers. The finding of genes involved in methanogenesis is supported by previous studies at Äspö HRL in which methane production was detected (Kotelnikova and Pedersen, 1998). The use of specific substrates by the microbial community in these aquifers was not part of this study. Nevertheless, it is most likely that at least some of the compounds suggested by the metagenomic analysis are available in the aquifers. Acetate was directly measured in previous studies with values ranging between 1 and 147 mg l−1 (Sicada database, SKB). Schäfer et al. (2015) revealed the presence of sugars such as fucose in biofilm material filling the cracks within the Äspö aquifers. Leefman et al. (2015) studied the formation of biofilms in the Äspö HRL and detected attachment (within 10 min) of amino acids, carbohydrates and carboxylic acids. This suggests the abundant presence of these compounds in the water. Using the predictive results obtained from this metagenomic investigation, specific pathways, their underlying substrates and their products can now be explicitly studied at the Äspö HRL.
Microbial community dynamics in the deep Fennoscandian shield
Energy sources and nutrients are likely only available in pulses at the depths investigated in this study (Jørgensen, 2012), causing extremely long average generation times (Jørgensen, 2011). In contrast to most open systems where species entry is less limited and the environment selects, microbial community dynamics may work differently in deep subsurface aquifers. Here the community was first selected in the original water coming from the surface and then subsequently streamlined on its way down to the subsurface. Cell size would also be an issue as these environments may partially select for a smaller surface-to-volume ratio (Lopez-Garcia et al., 2001; Jørgensen, 2012) because of oligotrophy and presumably also space limitation, as bigger cells will likely get stuck in smaller pore fractions. These factors may have caused the loss of some populations and consequently the ability to carry out key metabolic functions. Once that happens, the community would have been further restructured by predation, competition and resource limitation while having a minimal influence from invasion by new species. Thus, the productivity and efficiency of these communities is not merely a product of energy and nutrient availability, but relies to some degree on stochastic effects, by which organisms were isolated in the first place.
References
Anderson SL, McIntosh L . (1991). Light-activated heterotrophic growth of the cyanobacterium Synechocystis sp. strain PCC 6803: a blue-light-requiring process. J Bacteriol 173: 2761–2767.
Balkwill DL, Fredrickson JK, Romine MF . (2006). Sphingomonas and related genera. In: The Prokaryotes. Springer New York: New York, NY, USA, pp 605–629.
Borgström E, Lundin S, Lundeberg J . (2011). Large scale library generation for high throughput sequencing. PLoS One 6: e19119.
Campbell BJ, Engel AS, Porter ML, Takai K . (2006). The versatile epsilon-proteobacteria: key players in sulphidic habitats. Nat Rev Microbiol 4: 458–468.
Dixon P . (2003). VEGAN, a package of R functions for community ecology. J Veg Sci 14: 927–930.
Drake H, Tullborg E-L . (2009). Paleohydrogeological events recorded by stable isotopes, fluid inclusions and trace elements in fracture minerals in crystalline rock, Simpevarp area, SE Sweden. Appl Geochem 24: 715–732.
Eberspächer J, Lingens F . (2006). The genus phenylobacterium. In: The Prokaryotes. Springer New York: New York, NY, USA, pp 250–256.
Edwards KJ, Becker K, Colwell F . (2012). The deep, dark energy biosphere: intraterrestrial life on earth. Annu Rev Earth Planet Sci 40: 551–568.
Eydal HSC, Jägevall S, Hermansson M, Pedersen K . (2009). Bacteriophage lytic to Desulfovibrio aespoeensis isolated from deep groundwater. ISME J 3: 1139–1147.
Faust K, Sathirapongsasuti JF, Izard J, Segata N, Gevers D, Raes J et al. (2012). Microbial co-occurrence relationships in the human microbiome. PLoS Comput Biol 8: e1002606.
Finn RD, Clements J, Eddy SR . (2011). HMMER web server: interactive sequence similarity searching. Nucleic Acids Res 39: W29–W37 gkr367.
Grzymski JJ, Murray AE, Campbell BJ, Kaplarevic M, Gao GR, Lee C et al. (2008). Metagenome analysis of an extreme microbial symbiosis reveals eurythermal adaptation and metabolic flexibility. Proc Natl Acad Sci USA 105: 17516–17521.
Hallbeck L, Pedersen K . (2008). Characterization of microbial processes in deep aquifers of the Fennoscandian Shield. Appl Geochem 23: 1796–1819.
Hamilton T, Jones DS, Schaperdoth I, Macalady J . (2014). Metagenomic insights into S(0) precipitation in a terrestrial subsurface lithoautotrophic ecosystem. Front Microbiol 5: 756.
Heim C, Lausmaa J, Sjövall P, Toporski J, Dieing T, Simon K et al. (2012). Ancient microbial activity recorded in fracture fillings from granitic rocks (Äspö Hard Rock Laboratory, Sweden). Geobiology 10: 280–297.
Hengsuwan M, Heim C, Simon K, Hansen BT . (2015). Long-term investigation of Sr-isotope and rare earth element fractionation processes within three major aquifers in the Äspö Hard Rock Laboratory (Sweden). Geomicrobiol J 32: 243–254.
Herlemann DP, Labrenz M, Jürgens K, Bertilsson S, Waniek JJ, Andersson AF . (2011). Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J 5: 1571–1579.
Hillebrand H, Bennett DM, Cadotte MW . (2008). Consequences of dominance: a review of evenness effects on local and regional ecosystem processes. Ecology 89: 1510–1520.
Hoehler TM, Jorgensen BB . (2013). Microbial life under extreme energy limitation. Nat Rev Micro 11: 83–94.
Hopkinson CS, Vallino JJ, Nolin A . (2002). Decomposition of dissolved organic matter from the continental margin. Deep Sea Res Part II Top Stud Oceanogr 49: 4461–4478.
Illumina I . (2013). TruSeq DNA Sample Preparation Guide. Http://support.illumina.com/sequencing/sequencing_kits/truseq_rna_sample_prep_kit_v2/documentation.html U. No Title.
Ionescu D, Buchmann B, Heim C, Häusler S, de Beer D, Polerecky L . (2014). Oxygenic photosynthesis as a protection mechanism for cyanobacteria against iron-encrustation in environments with high Fe2+ concentrations. Front Microbiol 5: 1–10.
Ionescu D, Heim C, Polerecky L, Ramette A, Haeusler S, Bizic-Ionescu M et al. (2015). Diversity of iron oxidizing and reducing bacteria in flow reactors in the Äspö Hard Rock Laboratory. Geomicrobiol J 32: 3–4, 207–220.
Ionescu D, Siebert C, Polerecky L, Munwes YY, Lott C, Häusler S et al. (2012). Microbial and chemical characterization of underwater fresh water springs in the Dead Sea. PLoS One 7: e38319.
Jägevall S, Rabe L, Pedersen K . (2011). Abundance and diversity of biofilms in natural and artificial aquifers of the Äspö Hard Rock Laboratory, Sweden. Microb Ecol 61: 410–422.
Jørgensen BB . (2011). Deep subseafloor microbial cells on physiological standby. Proc Natl Acad Sci USA 108: 18193–18194.
Jørgensen BB . (2012). Shrinking majority of the deep biosphere. Proc Natl Acad Sci USA 109: 15976–15977.
Joshi NA, Fass JN . (2011), Sickle: a sliding-window, adaptive, quality-based trimming tool for FastQ files (Version 1.33)[Software].
Kantor RS, Wrighton KC, Handley KM, Sharon I, Hug LA, Castelle CJ et al. (2013). Small genomes and sparse metabolisms of sediment-associated bacteria from four candidate phyla. MBio 4: e00708–e00713.
Kotelnikova S, Pedersen K . (1998). Distribution and activity of methanogens and homoacetogens in Äspö spo deep granitic aquifers at Äspö Hard Rock Laboratory, Sweden. Analysis 26: 121–134.
Kotelnikova S, Pedersen K . (1997). Evidence for methanogenic Archaea and homoacetogenic Bacteria in deep granitic rock aquifers. FEMS Microbiol Rev 20: 339–349.
Kyle JE, Eydal HSC, Ferris FG, Pedersen K . (2008). Viruses in granitic groundwater from 69 to 450m depth of the Äspö hard rock laboratory, Sweden. ISME J 2: 571–574.
Laaksoharju M, Wold S . (2005). The colloid investigation conducted at the Äspö Hard Rock Laboratory during 2000-2004. SKB Technical Report TR-05-20.
Langmead B, Salzberg SL . (2012). Fast gapped-read alignment with Bowtie 2. Nat Methods 9: 357–359.
Leefmann T, Heim C, Lausmaa J, Sjövall P, Ionescu D, Reitner J et al. (2015). An imaging mass spectrometry study on the formation of conditioning films and biofilms in the subsurface (Äspö Hard Rock Laboratory, SE Sweden). Geomicrobiol J 32: 197–206.
Li D, Liu C-M, Luo R, Sadakane K, Lam T-W . (2015). MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31: 1674–1676.
Lipman CB . (1931). Living microorganisms in ancient rocks. J Bacteriol 22: 183–198.
Lopez-Garcia P, Rodriguez-Valera F, Pedros-Alio C, Moreira D . (2001). Unexpected diversity of small eukaryotes in deep-sea Antarctic plankton. Nature 409: 603–607.
Lovley DR, Chapelle FH . (1995). Deep subsurface microbial processes. Rev Geophys 33: 365–381.
Lundin S, Stranneheim H, Pettersson E, Klevebring D, Lundeberg J . (2010). Increased throughput by parallelization of library preparation for massive sequencing. PLoS One 5: e10029.
Mattes TE, Nunn BL, Marshall KT, Proskurowski G, Kelley DS, Kawka OE et al. (2013). Sulfur oxidizers dominate carbon fixation at a biogeochemical hot spot in the dark ocean. ISME J 7: 2349–2360.
McMahon S, Parnell J . (2014). Weighing the deep continental biosphere. FEMS Microbiol Ecol 87: 113–120.
Motamedi M, Pedersen K . (1998). Desulfovibrio aespoeensis sp. nov., a mesophilic sulfate-reducing bacterium from deep groundwater at äspö hard rock laboratory, Sweden. Int J Syst Bacteriol 48: 311–315.
Nakagawa S, Takai K . (2008). Deep-sea vent chemoautotrophs: diversity, biochemistry and ecological significance. FEMS Microbiol Ecol 65: 1–14.
Nyyssonen M, Hultman J, Ahonen L, Kukkonen I, Paulin L, Laine P et al. (2014). Taxonomically and functionally diverse microbial communities in deep crystalline rocks of the Fennoscandian shield. ISME J 8: 126–138.
Pedersen K . (2012). Subterranean microbial populations metabolize hydrogen and acetate under in situ conditions in granitic groundwater at 450m depth in the Äspö Hard Rock Laboratory, Sweden. FEMS Microbiol Ecol 81: 217–229.
Peura S, Eiler A, Bertilsson S, Nykänen H, Tiirola M, Jones RI . (2012). Distinct and diverse anaerobic bacterial communities in boreal lakes dominated by candidate division OD1. ISME J 6: 1640–1652.
Purkamo L, Bomberg M, Nyyssönen M, Kukkonen I, Ahonen L, Itävaara M . (2015). Heterotrophic communities supplied by ancient organic carbon predominate in deep fennoscandian bedrock fluids. Microb Ecol 69: 319–332.
Romanenko LA, Tanaka N, Frolova GM, Mikhailov VV . (2010). Marinicella litoralis gen. nov., sp. nov., a gammaproteobacterium isolated from coastal seawater. Int J Syst Evol Microbiol 60: 1613–1619.
Rönnback P . (2005). Äspö Hard Rock Laboratory: an overview of the geology, hydrology and hydrochemistry. ESS Bull 3: 32–47.
Schäfer N, Schmidt BC, Quéric N-V, Röring B, Reitner J . (2015). Organic compounds and conditioning films within deep rock fractures of the Äspö Hard Rock Laboratory, Sweden. Geomicrobiol J 32: 231–242.
Seemann T . (2014). Prokka: rapid prokaryotic genome annotation. Bioinforma 30: 2068–2069.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al. (2003). Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13: 2498–2504.
Sinclair L, Osman OA, Bertilsson S, Eiler A . (2015). Microbial community composition and diversity via 16S rRNA gene amplicons : evaluating the Illumina platform. PLoS One 10: e0116955.
Söderbäck B . (2008). Geological evolution, palaeoclimate and historical development of the Forsmark and Laxemar-Simpevarp areas Site descriptive modelling SDM-Site. R-08-19. Available at http://www.iaea.org/inis/collection/NCLCollectionStore/_Public/40/015/40015709.pdf?r=1.
Ström A, Andersson J, Skagius K, Winberg A . (2008). Site descriptive modelling during characterization for a geological repository for nuclear waste in Sweden. Appl Geochem 23: 1747–1760.
Thamdrup BO, Finster KM, Hansen JWW, Thamdrup BO, Finster KM, Hansen JWW . (1993). Bacterial disproportionation of elemental sulfur coupled to chemical reduction of iron or manganese. Appl Environ Microbiol 59: 101–108.
Thompson JR, Marcelino LA, Polz MF . (2002). Heteroduplexes in mixed-template amplifications: formation, consequence and elimination by ‘reconditioning PCR’. Nucleic Acids Res 30: 2083–2088.
Thurman EM . (1985) Organic Geochemistry of Natural Waters. Springer Netherlands: Dordrecht, The Netherlands.
Vishnivetskaya TA . (2009) Viable Cyanobacteria and Green Algae from the Permafrost Darkness. Springer: Berlin Heidelberg, Germany.
Wheeler RE . (2010), Package ‘lmPerm’. multResp lmPerm R Proj Stat Comput http://www.r-project.org/.
Wilkins MJ, Daly R, Mouser PJ, Trexler R, Sharma S, Cole DR et al. (2014). Trends and future challenges in sampling the deep terrestrial biosphere. Front Microbiol 5: 481.
Wittebolle L, Marzorati M, Clement L, Balloi A, Daffonchio D, Heylen K et al. (2009). Initial community evenness favours functionality under selective stress. Nature 458: 623–626.
Wrighton KC, Thomas BC, Sharon I, Miller CS, Castelle CJ, VerBerkmoes NC et al. (2012). Fermentation, hydrogen, and sulfur metabolism in multiple uncultivated bacterial phyla. Science 337: 1661–1665.
Wu X, Holmfeldt K, Hubalek V, Lundin D, Åström M, Bertilsson S et al. (2015). Microbial metagenomes from three aquifers in the Fennoscandian shield terrestrial deep biosphere reveal metabolic partitioning among populations. ISME J. Available from http://dx.doi.org/10.1038/ismej.2015.185 (accessed 26 October 2015).
Yamamoto M, Takai K . (2011). Sulfur metabolisms in epsilon- and gamma-proteobacteria in deep-sea hydrothermal fields. Front Microbiol 2: 192.
Yau S, Lauro FM, Williams TJ, DeMaere MZ, Brown MV, Rich J et al. (2013). Metagenomic insights into strategies of carbon conservation and unusual sulfur biogeochemistry in a hypersaline Antarctic lake. ISME J 7: 1944–1961.
Acknowledgements
All authors thank SKB for access to the Äspö HRL tunnel and accompanying facilities; MD and SB acknowledge the Swedish Research Council (contracts 621-2014-4398 and 2012-3892); MD thanks The Crafoord Foundation (contract 20130557), the Nova Center for University Studies, Research and Development, and Familjen Hellmans stiftelse for financial support. Sequencing was carried out at the National Genomics Infrastructure hosted by Science for Life Laboratory. Bioinformatics was carried out utilizing the Uppsala Multidisciplinary Center for Advanced Computational Science resource at Uppsala University (project b2013127).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Hubalek, V., Wu, X., Eiler, A. et al. Connectivity to the surface determines diversity patterns in subsurface aquifers of the Fennoscandian shield. ISME J 10, 2447–2458 (2016). https://doi.org/10.1038/ismej.2016.36
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2016.36
This article is cited by
-
May microbial ecological baseline exist in continental groundwater?
Microbiome (2023)
-
Out of the blue: the independent activity of sulfur-oxidizers and diatoms mediate the sudden color shift of a tropical river
Environmental Microbiome (2023)
-
Unveiling microbial diversity in deep geothermal fluids, from current knowledge and analogous environments
Geothermal Energy (2023)
-
Thiobacillus as a key player for biofilm formation in oligotrophic groundwaters of the Fennoscandian Shield
npj Biofilms and Microbiomes (2023)
-
Microbial communities in the deep-sea sediments of the South São Paulo Plateau, Southwestern Atlantic Ocean
International Microbiology (2023)