Abstract
The Black Queen Hypothesis, recently proposed to explain an evolution of dependency based on gene loss, is gaining ground. This paper focuses on how the evolution of dependency transforms interactions and the community. Using agent-based modeling we suggest that species specializing in the consumption of a common good escape competition and therefore favor coexistence. This evolutionary trajectory could open the way for novel long-lasting interactions and a need to revisit the classically accepted assembly rules. Such evolutionary events also reshape the structure and dynamics of communities, depending on the spatial heterogeneity of the common good production. Let Black be the new black!
Similar content being viewed by others
Main
Popular among theories of ecology and evolution, the Red Queen Hypothesis (Van Valen, 1973) has recently been echoed by a new hypothesis: the Black Queen Hypothesis (BQH; Morris et al., 2012; Figure 1), which concerns the evolution of dependency between organisms.
Overview of the BQH. The yellow and green spots represent two types of microorganisms, namely A and B1 in the next figures, both producing a common good. A mutant that has lost the capacity to produce the common good is shown in blue (B2 in the next figures). The shade of the background (dark gray) expresses the concentration of the common good in the environment. (a) Initial state: A and B are present. (b) As the common good is extensively produced in the environment, a mutant strain (B2, blue) no longer able to produce the common good emerges; it is dependent (that is, beneficiary) on other microorganisms producing it (that is, helpers). (c) As the beneficiary’s fitness is improved (no energy invested in the production), it invades the population (d) and a new equilibrium is reached in the community between the helpers (yellow) and beneficiaries (blue), which supplanted their ancestors. Figure inspired from Bjørn Østman, 2012, http://pleiotropy.fieldofscience.com/2012/05/black-queen-hypothesis.html.
Although the Red Queen Hypothesis sets the basis for (mostly antagonistic) co-evolution, the BQH renews and puts into perspective current understanding of the evolution of interactions between free-living organisms within microbial communities. This novel reductive theory describes evolutionary mechanisms potentially leading to the connectedness between organisms in a community. More precisely, it provides theoretical interpretations on the evolution of dependencies through adaptive gene loss in free-living organisms.
In the BQH context, some free-living organisms called beneficiaries ‘avoid’ having a function in order to optimize their adaptation to the environment. This loss of function is made possible because other organisms in their close environment (helpers) publicly and continuously provide for the function, offering a (partially) stable environment. The mechanism underlying this particular loss of function is genome reduction through gene loss. This type of evolutionary process is of major significance in the context of long-lasting interactions, as beneficiary species develop strong dependency on the function provided by helper species. The fact that most bacterial species cannot be grown in monocultures, referred to as the ‘uncultured microbial majority’ (Giovannoni et al., 2014), could result from such dependency-based gene loss, as the species are unable to grow when extracted from their community.
After its introduction, the BQH was echoed in papers describing the evolution of organisms through adaptive functions and gene loss (Ellers et al., 2012; Giovannoni, 2012; D’Souza et al., 2014; Giovannoni et al., 2014; Luo et al., 2014), the evolution of interactions (Estrela et al., 2012; Hussa and Goodrich-Blair, 2013), notably cooperation (Sachs and Hollowell, 2012) and evolution of the community (for example, Sachs and Hollowell, 2012; Mitri and Richard Foster, 2013; Hanson et al., 2014). It also triggered one dedicated evolutionary experiment (Morris et al., 2014) and uncovered new possibilities regarding the outcomes of other evolutionary experiments (D’Souza et al., 2014; Hosoda et al., 2014).
One notable point of the BQH is the association of adaptive genome reduction with free-living organisms (that is, organisms living independently of any host), a phenomenon that had not been apparent before. This evolutionary event of adaptive genome reduction stems from a particular type of interaction: the use of a common good (that is, a freely available element present in the environment) (Figure 1).
Being a recent hypothesis, the BQH has not been thoroughly tested. Nevertheless, we focused on the development of new ideas related to this hypothesis. We detail herein the possible mechanisms driving the observed loss of genes, and then use a modeling approach to fathom the consequences of this adaptive gene loss on population dynamics, at the community scale and in different ecological contexts. Thus, our aim was to explore the BQH beyond its original definition. A corollary of the BQH is also introduced.
Size matters in the BQH
A key element in this hypothesis is the loss of genes, which expresses a selected genome reduction. Although evolution is usually associated with genome complexification (Wolf and Koonin, 2013), in the BQH the source of evolutionary opportunities is simplification through gene loss. Typically, gene loss results from two different forces: genetic drift and positive selection.
Genetic drift or positive selection for gene loss in the BQH?
Genetic drift refers to changes in allele frequencies of a population due to random sampling. The ability of drift to influence allele frequencies is inversely proportional to population size. By definition, natural selection increases fitness, whereas genetic drift operates at random, and only occasionally confers patent fitness benefits.
If a function becomes useless, the selection pressures on genes involved in that function are lifted. In the absence of purifying selection acting on these genes, circum-neutral mutations can accumulate (McCutcheon and Moran, 2011) and be randomly retained by genetic drift in a small population, leading to the decay and eventual loss of these genes. Given the large size of the populations encompassed by the BQH, genetic drift is excluded as the main driver of evolution (Morris et al., 2012). One example of the BQH concerns the most abundant photosynthetic organisms on Earth: Prochlorococcus (Partensky et al., 1999).
General genome reduction and removal of specific gene
Gene loss is known to be selected via two distinct operating modes: it can be favored by (i) general genome reduction and (ii) the removal of specific targeted gene(s). The two systems most likely act together.
Positive selection for genome reduction refers to genome streamlining sensu stricto (Ochman and Moran, 2001; Giovannoni et al., 2005), which is defined as the selection process that ‘[…] acts to reduce genome size because of the metabolic burden of replicating DNA with no adaptive value [...]’ (Giovannoni et al., 2005). Indeed, every function has a constitutive cost, considering both genomic content and metabolism (Giovannoni et al., 2005, Kreft and Bonhoeffer, 2005; Lahti et al., 2009; Driscoll et al., 2011) but this cost is usually offset by the fitness benefits the function provides (Giovannoni et al., 2014). If a function loses its beneficial effects it will eventually be purged to reduce energy costs. Because retaining a bigger genome is costly (maintenance, replication and regulation), positive selection for genome reduction is assumed to be the main driver of genome reduction, at least in oligotrophic environments (Dufresne et al., 2005; Hottes et al., 2013; Giovannoni et al., 2014). A reduction or elimination of redundant and useless genetic material will occur from one generation to the next. Consistently, free-living microorganisms are reported to often experience genome reduction via a loss of paralogs for multicopy genes (Porter and Crandall, 2003).
Thus, in the genome streamlining theory (Ochman and Moran, 2001; Giovannoni et al., 2005), selection favors gene loss, as smaller genomes provide more adaptive advantages than bigger ones.
Recent experimental tests of evolutionary dynamics have shown that it may not be genome reduction itself, but particular and individual gene loss that confers the greatest advantages (Cooper et al., 2001; Lee and Marx, 2012; D’Souza et al., 2014; Pande et al., 2014), especially if the cell’s lack of metabolite production is compensated by the habitat (D’Souza et al., 2014). The fitness gained from a given gene deletion is dependent on its metabolic function (for example, which metabolite it codes for) and on the position of the deletion in the metabolic pathway. For example, the deletion of genes at the end of a biosynthetic pathway could be more advantageous than the deletion of anterior genes (D’Souza et al., 2014). Thus, the energy costs linked with anabolism could be efficiently reduced by the removal of specific targeted genes.
Specialization toward common goods consumption
A special interaction: the circulation of a common good
Every species in a community is linked to one or several other species, thereby forming an intricate web of direct and indirect interactions, metaphorically described as a tapestry in which the weaving (that is, interactions between species) is as important as the species themselves (Estes et al., 2013).
In the BQH (Morris et al., 2012) species evolutionary dynamics are based on indirect interactions through common goods utilization. Common goods are freely available elements present in the environment for both the producing species and other species around. The producers of common goods can still have preferential access to these goods (Estrela et al., 2015), thereby avoiding the emergence of a ‘tragedy of the commons’ situation (Hardin, 1968). Thus, the interaction between ‘helper’ and ‘beneficiary’ species is a case of indirect symbiosis existing through the flux of a common good, and the evolution of interactions between the two species can be considered as a side effect of common good consumption.
Within the community, helper individuals transform their microbial vicinity into a stable and homogeneous place (by continuous production of a common good) allowing beneficiary mutant organisms to follow this adaptive path of specialization.
The rise of the mutant
To further understand the Black Queen’s dynamics we used an agent-based model (Supplementary Information). We based our simulations on one or two species, then we introduced a mutant with higher fitness than its ancestor in order to observe (i) the temporal and spatial patterns of its invasion in the community (ii) the dynamics of the simulated organisms.
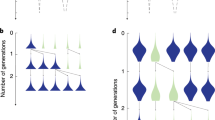
These simulations showed that when a fitter mutant unable to synthesize the common good emerges in a population (that is, a loss-of-function mutant) this new strain will supplant its ancestors. But if the ancestors are the only helpers around, the beneficiary mutant population never excludes its original population due to its vital dependence on the production of the common good (Figure 2a). Thus, the BQH also fits for a single species model. Nevertheless, if another species in the community produces enough of the common good, the mutants (because they are fitter) will replace the ancestors (Figure 2b).
Trajectories for two species populations (A and B1) when a loss-of-function mutant (B2) arises in one population. The trajectories shown represent a single simulation that measures the population of each species over time as defined by the species’ rules given in the Supplementary Information. (a) As the fitter mutants B2 (blue) emerge within their original population B1 (green), they will increase to the detriment of this original population. However, because the mutants still depend on the common good produced by the helpers (B1s), they can only spread if there are enough B1s present to produce the common good, leading to an equilibrium state of helper and mutant populations. (b) When the fitter mutants arise from their original population B1, if another species (A, in yellow) also produces the common good, then the B2 population will not be exclusively dependent on the B1s, and (B2s) will entirely supplant the B1s, if the B1s provide enough of the common good to sustain the B2s.
In our simulations, the helpers’ population is always sustained and their density (Supplementary Information) reaches a steady state (Figure 3a). The dependence of the beneficiaries on the helpers forces both populations to be in equilibrium. This confirms the BQH prediction (Morris et al., 2012), whereby a loss-of-function mutant will be able to expand within its ancestral population if the function loss gives the mutant a growth advantage over its ancestors but the mutant retains a need for the common good.
Steady-state populations of the original population B1 (helper) and of the fitter mutant B2 (beneficiary), with changing parameter values of density and of quantity of common good required. The population of B1s in green (the original population, which became a helper population) and B2s in blue (the beneficial mutants) is shown after they have reached equilibrium. Fifty simulation replicates were performed for each modality, and data were collected after 500 time steps (Supplementary Information). The default values for the fixed parameters were: (a, b) reproduction latency=3, lifespan=3; (a) minimum of common good required=5; and (b) density=3. Both (a) the density value (in arbitrary units, corresponding to the upper limit of individuals living in a small radius) and (b) the quantity of required common good by beneficiaries (arbitrary units) are altered to see how the equilibrium changes. Whiskers show confidence intervals (α=0.5) of the means. (a) The density is indirectly representative of the nutrient resource available: if more resource is available, more organisms can live together on a given surface unit. Because the beneficiaries B2 are fitter, they will supplant the B1s when more nutrients are available (Kruskal–Wallis test, P<2.10−16) while B1 density hardly changes (Kruskal–Wallis test, P= 0.0191). However, because the B2s are dependent on the common good, they can only spread if there is enough common good produced, thus the helpers’ population (B1) cannot be excluded. (b) When the minimum quantity of common good required by beneficiaries B2s is higher (that is, the mutants have a greater need in common good) the B2s will be more dependent on the helpers B1. As the need in common good for the B2s increases, their population density at steady state is lowered (Kruskal–Wallis test, P<2.10−16) and conversely for the B1 (Kruskal–Wallis test, P<2.10−16).
Helper or beneficiary?
What, in a community, determines which species will evolve into a beneficiary or into a helper? The current proposition by Morris et al. (2012) is that beneficiaries are species that evolve the most rapidly. We propose a new idea.
Our simulation reveals that when the minimum value of a common good needed for the mutants to multiply increases, it lowers the density of the mutants at steady state, whereas it increases the density of the helper species (Figure 3b). This is tied to the fact that increasing the need for a common good increases the dependency of the mutants on helper species producing this common good. As dependency is constraining, we suggest that the fittest of the loss-of-function mutants will be the one on which a rise in dependency will have the least effect, that is, for which the loss of function is less enslaving. Nevertheless, if the function is accessory, it could be entirely lost by the community, and the hypotheses would no longer stand, suggesting that the BQH might only be valid for functions vital to the organisms. We suggest that a potential loss-of-function mutant may be a species with a ‘silently advantageous’ trait, which will only fully express its benefits after a concomitant mutation. Such a trait could be a minor need for the common good or a faster consumption rate, for example.
Thus, in species adopting a beneficiary trajectory, the level of dependency on the helper might be low or potentially facilitated.
Effects of the Black Queen trajectory: transformation of interactions and of the community
We propose that the BQH offers new perspectives as to how evolution can modulate community life. Notably, the emergence of a mutation within a population can (i) transform its interactions with other species and (ii) deepen community life and modify the global dynamics of the community.
The Black Queen as a way to elude competition
Niche partitioning (through environmental filtering or through species interactions sorting) and neutral processes (Hubbell, 2001) are classically acknowledged to drive community assemblies and explain diversity patterns. When competition occurs in spatially and temporally homogeneous environments, coexistence is mainly assumed to result from complementarity in resource use (Webb et al., 2002). In addition to these classical explanations of assembly rules we suggest that the transition from competition to dependency relationships may also be a driver of community structure.
In our simplified two-species model, the emergence of a helper-dependent mutant shifts competition toward coexistence. In this situation, helpers and beneficiaries will reach a state of equilibrium (Figure 2b) that bypasses exclusive competition.
The fact that long-term coexistence is reached through the evolution of dependency has already been demonstrated: De Mazancourt and Schwartz (2010) showed that resource trade enhances coexistence even if it decreases the abundance of one of the species (consistent with Figure 3b). In a similar way, Turcotte et al. (2012) showed in a context of waste-product exploitation that dependency increases the coexistence of species. Cross-feeding interactions are also known to evolve in competitive environments (for example, Friesen et al., 2004; Louca and Doebeli, 2015).
To put it simply, in a system where several species are in competition, if a beneficial dependency emerges between two species (through resource trade, waste-product or common good use), their coexistence will be optimized. Here natural selection will favor the establishment of tighter relationships and confer an advantage to coexistence.
One change changes it all
In simulations where two species are able to sustain themselves irrespectively of the other’s presence, an emerging loss-of-function mutant can invade the population and eventually replace the ancestral strain of that population (Figure 2b). We observe that the population density at steady state depends on life history traits such as the quantity of common good required (Figure 3a), lifespan (Figure 4a) and reproduction latency (Figure 4b). Increases and decreases in population sizes are attributed to a density dependence effect (Figures 3a, 4a and b) related to the parameters used in the model. The dynamics of the loss-of-function mutant and extinction of the original population will thus be dependent on such life history traits to reach a steady state.
Steady-state populations of the original population B1 (helper) and of the fitter mutant B2 (beneficiary) with changing values of lifespan and reproduction latency parameters, a and b, respectively. The population of helpers (B1s) and beneficiaries (B2s) is shown after they have reached equilibrium. The lifespan (corresponding to the time available for reproduction, in arbitrary units) and the reproduction latency (the time before individuals can reproduce, in arbitrary units) were altered to see how the equilibrium changed. The population(s?) of helpers (B1) and beneficiaries (B2) were sampled at 50 different time points after introducing the mutant (B2) and when an equilibrium state (500 time steps) was reached for each set of parameters. Each bar shows the mean of the 50 time points and the whiskers represent the 95% confidence interval. The default values of the fixed parameters were as follows: lifespan=3; reproduction latency=3; density=3; and the minimum quantity of common good required by beneficiaries=5. (a) With increasing lifespan (Kruskal–Wallis test, P<2.10−16) a B1 individual is guaranteed to have enough common good in the final time steps of its life. A B2 individual, however, needs to coexist with a B1 on the same patch for there to be enough common good. That is, the chances of a beneficiary individual reproducing decrease because of the greater dependency on helpers. (b) Similar reasoning as for lifespan holds for the reproduction latency.
Metabolic dependencies are potentially a major driver of species co-occurrence (Zelezniak et al., 2015). Until recently, the repercussions of such evolution at the community level were mostly overlooked, whereas it is now becoming evident that they are an integral part of community systems (Hairston et al., 2005; Johnson and Stinchcombe, 2007; Schoener, 2011) implying that evolutionary dynamics should systematically be taken into account when characterizing communities.
Dependence and consequences on the quantity of common goods produced
We propose that the quantity of common goods produced by helpers could impact the spatial dynamic of the entire community. In a scenario where the common good is abundant, the distribution of helpers and beneficiaries is expected to be homogeneous at steady state. Conversely in our simulation where the common good is produced at a limiting concentration, the dynamic invasion of space is uneven (Supplementary Movie S1). The spatial distribution of beneficiaries, because of their dependency, is expected to adhere closely to the distribution of the helpers, inducing a local depletion of nutrients availability. Thus, the heterogeneous spatial ‘aggregation’ of (micro)organisms and resulting heterogeneity of nutrient availability will lead to temporal changes and patches displacement (Supplementary Movie S1). Spatial and temporal heterogeneity may thus be a consequence of helper/beneficiary interaction.
Going further
The BQH, smoothing the way for long-lasting interactions?
Studies of co-evolution, taken in the broad sense where one organism evolves in relation to another, have tended to focus on the acquisition of traits and functions, with little attention given to the loss of functions in free-living organisms. Consequently, the mechanisms behind the emergence of a compensated loss of function remain unknown. It has been assumed that trait loss ‘[...] is only expected to evolve in long-term, stable co-evolutionary physiological relationships [...]’ (Visser et al., 2010). However, we consider that the BQH offers a new approach: function loss, instead of resulting from long-lasting relationships, could actually be the cause of such long-lasting co-evolutionary relationships. Indeed, in the BQH, a ‘passive’ interaction (production and consumption of a common good) is at the basis of the emergence of tighter dependencies between two species.
We perceived that initially a mere coexistence of species within a community happens to be fortuitously beneficial for one of the species (via the redundancy in production of the common good). Then if the beneficial conditions remain sufficiently stable over time, the (future) beneficiary species will tend to lose the compensated function. More than the trait loss itself (Ellers et al., 2012), we believe that the ‘point of no return’ in the shift from a facultative interaction to an obligatory dependency is the loss of gene(s) underlying the loss of function. In this case, genome reduction sets up the first steps for stable long-lasting metabolic interactions (Pande et al., 2014).
The loss of essential genes is widespread in free-living bacteria and most certainly at the root of inter-organism networks (D’Souza et al., 2014). In line with this, some vitamin exchanges are known to cement species connectedness within the community (Giovannoni, 2012), and it has also been inferred from the BQH model that cooperative interactions could rise ‘automatically’ in this context (Sachs and Hollowell, 2012).
Corollary of the BQH
A species, by being dependent on another one, puts itself in a weakened position. If no helper species is around, the species cannot survive. What is more, such species can be considered as ‘accessory’ for the community as they do not provide for the essential function. Admittedly, accessory species could be wiped out more easily than essential species as their loss will not have an immediate effect on the community. On the other hand, being able to handle essential functions, even if more costly, guarantees some degree of ‘security’. First, it allows for resistance against environmental changes or perturbation. Second, in a dependency context, helpers are indispensable, which will partly prevent them from being replaced by competing species (Nadell et al., 2009). It is therefore of particular interest to keep, or even to acquire, the status of helper.
In their paper, Morris et al. (2012) suggest that helpers are keystone species in the community. We propose a possible corollary of the BQH: species could benefit from ensuring the status of helpers by having mutations that enhance the production of the common good, or more extremely, by acquiring genes producing the common good. The occurrence of mutation, which enhances the common good production, could also explain why some species turn into helpers while others become beneficiaries.
Horizontal gene transfer is a common phenomenon in (micro)organisms (McGinty et al., 2011), often leading to the acquisition of functions, and suggested to be an underlying mechanism of microbial cooperation (Smith, 2001). If the enhanced or acquired function leads to the development of a dependency interaction, then (micro)organisms possessing it may be better off, thanks to the key status acquired within the community. This substantiates the suggested BQH corollary, that is, genome expansion for helpers who tend to embrace a generalist ecological status. Thus, the BQH and its corollary invite a new interpretation of the networks of interactions and ecological status (that is, generalist/specialist) of co-occurring organisms and their evolution.
References
Cooper VS, Schneider D, Blot M, Lenski RE . (2001). Mechanisms causing rapid and parallel losses of ribose catabolism in evolving populations of Escherichia coli B. J Bacteriol 183: 2834–2841.
De Mazancourt C, Schwartz MW . (2010). A resource ratio theory of cooperation. Ecol Lett 13: 349–359.
Driscoll WW, Pepper JW, Pierson LS, Pierson EA . (2011). Spontaneous gac mutants of Pseudomonas biological control strains: cheaters or mutualists? Appl Environ Microbiol 77: 7227–7235.
D’Souza G, Waschina S, Pande S, Bohl K, Kaleta C, Kost C . (2014). Less is more: selective advantages can explain the prevalent loss of biosynthetic genes in bacteria. Evolution 68: 2559–2570.
Dufresne A, Garczarek L, Partensky F . (2005). Accelerated evolution associated with genome reduction in a free-living prokaryote. Genome Biol 6: R14.
Ellers J, Kiers TE, Currie CR, Mcdonald BR, Visser B . (2012). Ecological interactions drive evolutionary loss of traits. Ecol Lett 15: 1071–1082.
Estes JA, Brashares JS, Power ME . (2013). Predicting and detecting reciprocity between Indirect Ecological Interactions and Evolution. Am Nat 181: S76–S99.
Estrela S, Trisos CH, Brown SP . (2012). From metabolism to ecology: cross-feeding interactions shape the balance between polymicrobial conflict and mutualism. Am Nat 180: 566–576.
Estrela S, Morris JJ, Kerr B . (2015). Private benefits and metabolic conflicts shape the emergence of microbial interdependencies: the origins of microbial interdependencies. Environ Microbiol 10: 1462–2920.
Friesen ML, Saxer G, Travisano M, Doebeli M, Elena S . (2004). Experimental evidence for sympatric ecological diversification due to frequency-dependent competition in Escherichia coli. Evolution 58: 245–260.
Giovannoni SJ, Tripp HJ, Givan S, Podar M, Vergin KL, Baptista D et al. (2005). Genome streamlining in a cosmopolitan oceanic bacterium. Science 309: 1242–1245.
Giovannoni SJ . (2012). Vitamins in the sea. Proc Natl Acad Sci USA 109: 13888–13889.
Giovannoni SJ, Thrash JC, Temperton B . (2014). Implications of streamlining theory for microbial ecology. ISME J 8: 1–13.
Hairston NG, Ellner SP, Geber MA, Yoshida T, Fox JA . (2005). Rapid evolution and the convergence of ecological and evolutionary time. Ecol Lett 8: 1114–1127.
Hanson NW, Konwar KM, Hawley AK, Altman T, Karp PD, Hallam SJ . (2014). Metabolic pathways for the whole community. BMC Genomics 15: 619.
Hardin G . (1968). The tragedy of the commons. Science 162: 1243–1248.
Hosoda K, Habuchi KM, Suzuki S, Miyazaki M, Takikawa G, Sakurai T, Kashiwagi A et al. (2014). Adaptation of a cyanobacterium to a biochemically rich environment in experimental evolution as an initial step toward a chloroplast-like state. PLoS One 9: e98337.
Hottes AK, Freddolino PL, Khare A, ZN1 Donnell, Liu JC, Tavazoie S . (2013). Bacterial adaptation through loss of function. PLoS Genet 9: e1003617.
Hubbell SP . (2001) The Unified Neutral Theory of Biodiversity and Biogeography (MPB-32). Princeton University Press: Princeton, NJ, USA.
Hussa E, Goodrich-Blair H . (2013). It takes a village: ecological and fitness impacts of multipartite mutualism. Annu Rev Microbiol 67: 161–178.
Johnson MT, Stinchcombe JR . (2007). An emerging synthesis between community ecology and evolutionary biology. Trends Ecol Evol 22: 250–257.
Kreft JU, Bonhoeffer S . (2005). The evolution of groups of cooperating bacteria and the growth rate versus yield trade-off. Microbiology 151: 637–641.
Lahti DC, Johnson NA, Ajie BC, Otto SP, Hendry AP, Blumstein DT et al. (2009). Relaxed selection in the wild. Trends Ecol Evol 24: 487–496.
Lee MC, Marx CJ . (2012). Repeated, selection-driven genome reduction of accessory genes in experimental populations. PLoS Genet 8: 2–9.
Louca S, Doebeli M . (2015). Calibration and analysis of genome-based models for microbial ecology. eLife 4: e08208.
Luo H, Swan BK, Stepanauskas R, Hughes AL, Moran MA . (2014). Evolutionary analysis of a streamlined lineage of surface ocean Roseobacters. ISME J 8: 1428–1439.
McCutcheon JP, Moran NA . (2011). Extreme genome reduction in symbiotic bacteria. Nat Rev Microbiol 10: 13–26.
McGinty SE, Rankin DJ, Brown SP . (2011). Horizontal gene transfer and the evolution of bacterial cooperation. Evolution 65: 21–32.
Mitri S, Richard Foster K . (2013). The Genotypic View of Social Interactions in Microbial Communities. Annl Rev Genet 47: 247–273.
Morris JJ, Lenski RE, Zinser ER . (2012). The black queen hypothesis: evolution of dependencies through adaptive gene loss. mBio 3: e00036–12.
Morris JJ, Papoulis SE, Lenski RE . (2014). Coexistence of evolving bacteria stabilized by a shared black queen function: experimental evolution of a black queen community. Evolution 68: 2960–2971.
Nadell CD, Xavier JB, Foster KR . (2009). The sociobiology of biofilms. FEMS Microbiol Rev 33: 206–224.
Ochman H, Moran N . (2001). Genes lost and genes found: evolution of bacterial pathogenesis and symbiosis. Science 292: 1096–1099.
Pande S, Merker H, Bohl K, Reichelt M, Schuster S, de Figueiredo L et al. (2014). Fitness and stability of obligate cross-feeding interactions that emerge upon gene loss in bacteria. ISME J 8: 953–962.
Partensky F, Hess WR, Vaulot D . (1999). Prochlorococcus, a marine photosynthetic prokaryote of global significance. Microbiol Mol Biol Rev 63: 106–127.
Porter ML, Crandall KA . (2003). Lost along the way: the significance of evolution in reverse. Trends Ecol Evol 18: 541–547.
Sachs JL, Hollowell C . (2012). The origins of cooperative bacterial communities. MBio 3: e00099–12.
Schoener TW . (2011). The newest synthesis: understanding the interplay of evolutionary and ecological dynamics. Science 331: 426–429.
Smith J . (2001). The social evolution of bacterial pathogenesis. Proc R Soc Lond B 268: 61–69.
Turcotte MM, Corrin MSC, Johnson MTJ . (2012). Adaptive evolution in ecological communities. PLoS Biol 10: e1001332.
Van Valen L . (1973). A new evolutionary law. Evol Theory 1: 1–30.
Visser B, Le Lann C, den Blanken FJ, Harvey JF, van Alphen JJM, Ellers J . (2010). Loss of lipid synthesis as an evolutionary consequence of a parasitic lifestyle. Proc Natl Acad Sci USA 107: 8677–8682.
Webb CO, Ackerly DD, McPeek MA, Donoghue MJ . (2002). Phylogenies and community ecology. Annu Rev Ecol Syst 33: 475–505.
Wolf YI, Koonin EV . (2013). Genome reduction as the dominant mode of evolution: prospects & overviews. BioEssays 35: 829–837.
Zelezniak A, Andrejev S, Ponomarova O, Mende DR, Bork P, Patil KR . (2015). Metabolic dependencies drive species co-occurrence in diverse microbial communities. Proc Natl Acad Sci USA 112: 6449–6454.
Acknowledgements
The work is supported by a grant from ‘l'Agence Nationale de la Recherche’ (ANR-10-STRA-0002) and from the CNRS EC2CO-Microbien funding program (‘Hmmm’ project). We thank D Warwick for comments and suggested modifications to a previous version of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Mas, A., Jamshidi, S., Lagadeuc, Y. et al. Beyond the Black Queen Hypothesis. ISME J 10, 2085–2091 (2016). https://doi.org/10.1038/ismej.2016.22
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2016.22
This article is cited by
-
Radiation impacts gene redundancy and biofilm regulation of cryoconite microbiomes in Northern Hemisphere glaciers
Microbiome (2023)
-
Multi-genome metabolic modeling predicts functional inter-dependencies in the Arabidopsis root microbiome
Microbiome (2022)
-
Dynamic character displacement among a pair of bacterial phyllosphere commensals in situ
Nature Communications (2022)
-
Metabolic adaptation to vitamin auxotrophy by leaf-associated bacteria
The ISME Journal (2022)
-
Genomic evidence of functional diversity in DPANN archaea, from oxic species to anoxic vampiristic consortia
ISME Communications (2022)