Abstract
Linking microbial community composition with the corresponding ecosystem functions remains challenging. Because microbial communities can differ in their functional responses, this knowledge gap limits ecosystem assessment, design and management. To develop models that explicitly incorporate microbial populations and guide efforts to characterize their functional differences, we propose a novel approach derived from reliability engineering. This reliability modeling approach is illustrated here using a microbial ecology dataset from denitrifying bioreactors. Reliability modeling is well-suited for analyzing the stability of complex networks composed of many microbial populations. It could also be applied to evaluate the redundancy within a particular biochemical pathway in a microbial community. Reliability modeling allows characterization of the system’s resilience and identification of failure-prone functional groups or biochemical steps, which can then be targeted for monitoring or enhancement. The reliability engineering approach provides a new perspective for unraveling the interactions between microbial community diversity, functional redundancy and ecosystem services, as well as practical tools for the design and management of engineered ecosystems.
Similar content being viewed by others
Introduction
Understanding the relationships between community composition, community structure and ecosystem function is challenging, particularly for microbially-mediated processes. The small size of microorganisms, the unparalleled diversity of microbial communities and the often uncertain linkages between microbial phylogeny and function hinder our ability to predict ecosystem processes based on microbial community structure. Because microorganisms play critical nutrient cycling roles in all ecosystems, models that explicitly incorporate microbial populations and their functional differences could improve ecosystem assessment, design and management.
Despite the common, and likely valid, assumption that microbial communities have high levels of functional redundancy, microbial community composition and structure can sometimes have important effects on ecosystem functions (Schimel, 2001; Reed and Martiny, 2007; Allison and Martiny, 2008; Bell et al., 2009; Strickland et al., 2009). For example, different soils show distinct temperature optima for nitrite oxidation and denitrification, suggesting that their microbial communities have functional differences (Saad and Conrad, 1993). While process-based biogeochemical models play an important role in understanding ecosystem functions, they typically do not capture the adaptive potential of biological systems and therefore often require calibration for each location.
The ability to model microbial ecosystems and their associated functions is hampered by the limited understanding of fundamental relationships in microbial ecology, including those between commonly measured characteristics (such as microbial diversity and community composition) and the resulting ecosystem function. These relationships appear to be complex, based on the varied results obtained to date (reviewed in Griffin et al., 2009). For instance, overall functional and taxonomic richness were positively associated with each other in complex wastewater treatment plant microbial communities (Johnson et al., 2015). Similarly, as diversity was decreased in another study, specialized functions such as nitrification and denitrification also decreased (Griffiths et al., 2000). However, in the same study, general functions such as decomposition had the opposite response; they increased as diversity decreased. These results are consistent with the idea that the reliability of narrowly distributed functions will be most sensitive to changes in microbial community composition (Schimel and Gulledge, 1998). In contrast, reduced diversity did not reduce the rate of the specialized functions of denitrification or ammonia oxidation in soil microcosms, where the inocula had been diluted to vary the diversity, provided the inoculum was sufficient to allow regrowth to the original abundance of the functional group (Wertz et al., 2006). Without an understanding of these fundamental relationships between community structure and function, microbial ecology is lacking a foundation, and researchers may pursue increasingly detailed molecular analyses of microbial communities without a basis for understanding when or if that information is of functional relevance.
In ecosystem models, microbial activity is often approached as a ‘black box’ (for example, a single rate for decomposition), despite continuing evidence supporting the need for more detail (for example, Schimel, 2001; Allison and Martiny, 2008; Treseder et al., 2012). The range of scales (micrometer to global) and the magnitude of microbial diversity present challenges to the development of more detailed models, but the evidence for differences in microorganisms’ responses to disturbances suggest that these challenges must be met. To incorporate microbial diversity into ecosystem models in a way that improves their predictive power and yet remains agile and computationally feasible will require us to identify those functions and situations where microbial community composition is critical to predicting function.
Here, we propose the application of reliability theory to microbial ecosystem data, and we assert that doing so will be useful for linking community structure, adaptability and performance, thereby improving our ability to predict and increase stability of ecosystem services provided by natural and engineered ecosystems. Reliability theory is a field of engineering using probability and information on the performance and lifespan or time-to-failure of individual components to model and predict the stability of complex systems (Elsayed, 1996; Crowder et al., 2000). It can be used to identify, for example, which equipment in a factory should be replicated to have the largest reduction in downtime. When applied to microbial ecosystem data, reliability modeling can identify microbial populations or groups of populations associated with good or poor performance, identify the populations least resistant or resilient to disturbance, and assist in predicting performance. These results could then inform ecosystem models, for example, through inputs of specific failure frequency distributions or through modifications of controlling equations based on key parameters affecting microbial communities and their associated functions. The reliability engineering approach to modeling ecosystem function is described here and then briefly illustrated with a case study of denitrifying bioreactors.
Approach
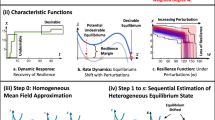
The basic approach to reliability modeling of an ecosystem is described here and illustrated in Figure 1. Whereas in traditional applications of reliability modeling the components might be stages within a manufacturing process, here the components are biological. Depending on the question at hand, the biological components could be general groupings of organisms (for example, trophic levels), more specific groupings of organisms (for example, functional groups, individual populations or groups of populations within the ecosystem), or enzymes (for example, each individual step in a biochemical pathway). For simplicity, the approach is described here at the middle level of resolution, where the components are microbial populations within an ecosystem.
To apply the reliability approach, one needs time-series data of a desired function or performance metric (system performance) and corresponding data on the microbial populations (component abundance). Reliability is then defined by setting a threshold below which the system or component is defined to have failed (Step 3, Figure 1). For an engineered ecosystem, the threshold might correspond to a regulatory limit or a minimum acceptable level of performance. For a natural ecosystem, the objective might be to capture periods of unusually low activity, in which case the threshold could be set at some fraction of the average activity of the population or performance factor of interest. For microbial population data, this approach assumes that a substantial drop in abundance results in a similar decrease in that population’s contribution to ecosystem function.
As compared to industrial applications of reliability modeling, microbial communities are likely to contain many more components. To make the analysis more tractable, microbial populations that show correlated dynamics can be grouped (Step 2, Figure 1). Grouping assumes that there are functional relationships among microbial populations showing correlated dynamics. One way to identify groups of populations is through Local Similarity Analysis (LSA) (Ruan et al., 2006). LSA compares each microbial population curve to the system performance, providing a test statistic (LS score) and P-value. In contrast to the original algorithm, which was designed to identify the period of maximum similarity for a particular population, the LSA code was modified here to allow negative scores, accounting for periods of high dissimilarity in computing similarity over the entire set of observations. Microbial populations with either positive (Group P) or negative (Group N) correlations with good system performance can be identified with this approach.
Mathematical reliability functions describing the system performance and the populations are determined independently (Steps 4–7, Figure 1). Applying the thresholds to the system performance and component abundance data for a particular population or group results in determination of times-to-failure. When a time-to-failure extends outside the monitoring period, this is called censored data. Censored data can either be excluded or methods appropriate for handling censored data may be applied (Crowder et al., 2000). Inclusion of censored data is a conservative approach, as it will result in a shorter time-to-failure and thus a potential underestimation of reliability. For Group P, the times-to-failure were determined from spans of presence (time periods where the abundance was above the threshold). In contrast, for Group N, times-to-failure were determined from spans of absence (time periods where the abundance was below the threshold). The times-to-failure are then used to generate a population or system reliability function, for example, by empirically fitting with the Weibull probability distribution function, which accommodates distributions of varied shapes. A χ2 goodness of fit test is applied in each case to ensure that the Weibull function was indeed suitable to model the observed data.
The reliability functions for various populations are then combined to develop an overall reliability function for the biological system. Existing biological knowledge can guide the model configuration, suggesting arrangement of the components in parallel and series relationships to simulate ecosystem structures (Step 8, Figure 1). Parallel relationships could reflect functional redundancy, whereas trophic groups would likely be placed in series based on their known relationships. As a specific example, if the objective was to model the reliability of the denitrification pathway, enzymes catalyzing the same reaction would be placed in parallel, while the different steps in the pathway would be arranged in series (Figure 2). A k-out-of-n structure, where a specified number (k) of the components in that group (n) must be functioning (without regard to which ones they are), can also be used to account for redundancy, particularly in instances where total throughput (or processing capacity) may be limited. To incorporate the effects of Groups P and N, one might employ a series structure, where presence of Group P and absence of Group N are both required for system function. If there is no known biological basis for suggesting a configuration, multiple configurations should be evaluated.
Reliability analysis of a biological system of known structure is illustrated here for denitrification. The biological denitrification pathway, with redundant enzymes indicated by the ovals for each step, results in a reliability configuration with redundant enzymes arranged in parallel and the distinct steps arranged in series.
For each configuration, the component reliability functions are mathematically combined to calculate system-level reliability (Step 9, Figure 1). Finally, the resulting ecosystem-level models are compared to the reliability of the desired ecosystem function (Step 10, Figure 1). Goodness of fit can be evaluated by using the sum of squared error tests. A detailed description of these methods and the EcoReliAnT software that implements them is available (Bartolerio 2011), along with their application to a simple test dataset from laboratory reactors. The software is available at http://hdl.handle.net/2142/88445.
Case study
To illustrate the application of reliability modeling to a complex microbial community, a dataset from a denitrifying bioreactor treating subsurface drainage from fields in Decatur, Illinois (FP07) was used (Porter et al., 2015). The data analyzed here were collected between 5 January 2009 and 19 May 2009 and include nitrate removal as a performance metric. Dynamics of the denitrifying bacterial populations were monitored using terminal restriction fragment length polymorphism of the functional gene nosZ (last step in Figure 2) (Rich et al., 2003). For reliability analysis, the relative abundance values for each fragment length (used to define microbial operational taxonomic units–OTUs) were averaged across five sampling ports in the bioreactor for each time point prior to input into the EcoReliAnT software. In this case study, the OTUs were assumed to represent populations.
Both for nitrate removal and for individual microbial OTUs, the thresholds were set to the average value minus two-fifths of the standard deviation. Because of the limited data, censored data were retained and accounted for, and LSA over the entire dataset was used to group microbial OTUs with correlated dynamics. LS score and P-value thresholds of 0.2 and 0.1, respectively, were applied. Despite these actions, the available data were still extremely limited. The system performance reliability function was determined based on three times-to-failure, while the Group P function was based on six, and the Group N function was based on 15. Ideally, a longer dataset with more failures would be used, but for the purposes of illustrating the approach this is sufficient.
In this case study, there was no pre-existing knowledge of how to structure the components, so several common approaches were tested. Two are presented here to illustrate the process. In the first case, the Groups P and N reliability functions were compared directly with the system reliability function (Figure 3a). In the second case, Group P was placed in series with Group N. In combination with the ways these groups were defined, this configuration requires that Group P must be present and Group N must be absent for good function (Figure 3b). For both groups, a k-out-of-n structure was applied, with the n values were fixed at the number of OTUs in each group. Any number of Group P OTUs could actually be performing the desired function (or interfering with it in the case of Group N), but the numbers derived from LSA analysis (using the same thresholds for both groups) provided a reasonable starting place.
Comparison of microbial reliability functions to the denitrifying bioreactor system performance reliability function. (a) Direct comparison of reliability functions from Groups P and N to performance reliability. (b) Sample reliability analysis component configuration: Group P 1-out-of-2 structure in series with a Group N 2-out-of-6 structure. (c) Comparison of reliability function from the configuration shown in (b) to performance reliability.
Reliability functions represent a probability distribution, with increasing likelihood of performance failure (or decreasing reliability) over time. Comparing the two groups directly to system performance, Group P has a very similar reliability function, while the shape of the reliability function for Group N is much different than that of system performance (Figure 3a). The better match between Group P and system performance is also reflected in the lower sum of squared errors for Group P versus Group N when comparing the microbial group reliability functions and the system reliability functions (1.28 and 24.13, respectively). When a Group P k-out-of-2 structure was arranged in series with a Group N k-out-of-6 structure (Figure 3c), the best fit k values were determined to be 1 and 2, respectively, and the sum of squared errors was 0.70. Therefore, the reliability of this model best matches the reliability of the system performance when one of the OTUs from Group P is present and two of the OTUs from Group N are absent. This system configuration then defines failure as either both OTUs from Group P no longer being present or five of the six OTUs from Group N being present.
The dataset used in this case study was too small to provide a clear determination of an appropriate model configuration. To further develop a reliability model for these denitrifying bioreactors, it would be necessary to apply these configurations to more data, whether from a longer timescale or from additional bioreactors.
Discussion
Ecosystem reliability models have the potential to provide a critical connection between microbial community surveys and microbially-mediated functions, providing direction for microbial ecology studies that have functional aims and improving our ability to construct design models for microbially-mediated ecosystem functions. The case study presented here illustrates the process of developing ecosystem reliability models, which can be used to identify populations suitable for predicting performance and potentially contributing strongly to the activity of interest. In addition to the form presented here, reliability functions can be used to calculate a mean residual reliability, allowing predictions of how much longer a component of a specified age is likely to continue to function. The beneficial and nuisance populations that have the greatest effect on system reliability can also be identified through importance analysis of reliability models (Elsayed, 1996; Crowder et al., 2000); these populations are logical targets for research aimed at providing more reliable and resilient ecosystem services. Furthermore, there is the potential to integrate reliability models with models of environmental parameters, such as hydrology or temperature, that have rapid effects on ecosystem services, to provide models that can predict both transient fluctuations in performance and catastrophic ecosystem failures.
Ecosystem reliability modeling requires datasets that include both microbial community data and functional measurements at a timescale that captures their fluctuations. Although the tools for collecting both types of data are widely available, such datasets are surprisingly rare. In the microbial literature, the relative lack of functional measurements or activity assays in microbial ecology monitoring projects is striking and emblematic of the current disconnect between microbial community composition and microbially-mediated functionality. Conversely, in engineered ecosystems, where detailed performance data is typically collected, the coverage of microbial community dynamics is generally coarse. Regardless of the type of modeling applied, more consistent collection of both types of data would provide a better foundation for linking structure and function.
Reliability models are empirical in nature and therefore not necessarily representative of underlying mechanisms. For example, in the case study presented here, they cannot distinguish whether the Group N populations are actively affecting performance or simply tend to increase in abundance in the absence of those microorganisms associated with good performance. However, the fact that their inclusion improved the fit with system reliability supports further investigation. If these nuisance populations were shown to directly affect performance, understanding the mechanism(s) involved is also likely to be important.
Conclusions and implications
The dearth of models connecting microbial community composition and function presents a critical problem, because microbial activity is the basis for the nutrient cycles that support life on our planet. There are also opportunities for more sustainable processes for pollutant remediation and water treatment that rely on microbial activity and ecosystem engineering; a better understanding of microbial ecology would improve the design of these systems. Applying reliability engineering to ecosystem analysis provides a way to connect the dynamics of a particular function with the community structure. This approach is well-suited to investigating temporal stability and adaptive response to disturbances or failures, which are critical considerations as we face increasing environmental variation and rates of change.
References
Allison SD, Martiny JBH . (2008). Resistance, resilience, and redundancy in microbial communities. Proc Natl Acad Sci USA 105: 11512–11519.
Bartolerio N . (2011). Development and application of a microbial reliability model to analyze denitrifying biofilter stability. MS thesis, University of Illinois Urbana Champaign, Urbana, IL..
Bell T, Gessner MO, Griffiths RI, McLaren J, Morin PJ, van der Heijden M et al. (2009). Microbial biodiversity and ecosystem functioning under controlled conditions and in the wild. In: Naeem S, Bunker DE, Hector A, Loreau M, Perrings C (eds). Biodiversity, Ecosystem Functioning, and Human Wellbeing: an ecological and economic perspective. Oxford University Press: New York.
Crowder MJ, Kimber AC, Smith RL, Sweeting TJ . (2000) Statistical Analysis of Reliability Data. Chapman and Hall/CRC: Boca Raton, FL.
Elsayed EA . (1996) Reliability Engineering. Addison Wesley Longman Inc: Reading, MA.
Griffin JN, O'Gorman EJ, Emmerson MC, Jenkins SR, Klein A-M, Loreau M et al. (2009). Biodiversity and the stability of ecosystem functioning. In: Naeem S, Bunker DE, Hector A, Loreau M, Perrings C (eds). Biodiversity, Ecosystem Functioning, and Human Wellbeing: an ecological and economic perspective. Oxford University Press: New York, pp 78–93.
Griffiths BS, Ritz K, Bardgett RD, Cook R, Christensen S, Ekelund F et al. (2000). Ecosystem response of pasture soil communities to fumigation-induced microbial diversity reductions: an examination of the biodiversity–ecosystem function relationship. OIKOS 90: 279–294.
Johnson DR, Lee TK, Park J, Fenner K, Helbling DE . (2015). The functional and taxonomic richness of wastewater treatment plant microbial communities are associated with each other and with ambient nitrogen and carbon availability. Environ Microbiol 17: 4851–4860.
Porter MD, Rodriguez LF, Bartolerio N, Andrus JM, Zhang Y, Zilles JL et al. (2015). Seasonal patterns in microbial community composition in denitrifying bioreactors treating subsurface agricultural drainage. Microb Ecol 70: 710–723.
Reed HE, Martiny JBH . (2007). Testing the functional significance of microbial composition in natural communities. FEMS Microbiol Ecol 62: 161–170.
Rich JJ, Heichen RS, Bottomley PJ, Cromack K, Myrold DD . (2003). Community composition and functioning of denitrifying bacteria from adjacent meadow and forest soils. Appl Environ Microbiol 69: 5974–5982.
Ruan Q, Dutta D, Schwalbach MS, Steele JA, Fuhrman JA, Sun F . (2006). Local similarity analysis reveals unique associations among marine bacterioplankton species and environmental factors. Bioinformatics 22: 2532–2538.
Saad OALO, Conrad R . (1993). Temperature dependence of nitrification, denitrification, and turnover of nitric oxide in different soils. Biol Fert Soils 15: 21–27.
Schimel J . (2001). Biogeochemical models: implicit versus expllct microbiology. In: Schulze E-D, Heimann M, Harrison S, Holland E, Lloyd J, Prentice IC et al (eds). Global Biogeochemical Cycles in the Climate System. Academic Press: San Diego, CA.
Schimel JP, Gulledge J . (1998). Microbial community structure and global trace gases. Global Change Biol 4: 745–758.
Strickland MS, Lauber C, Fierer N, Bradford MA . (2009). Testing the functional significance of microbial community composition. Ecol 90: 441–451.
Treseder KK, Balser TC, Bradford MA, Brodie EL, Dubinsky EA, Eviner VT et al. (2012). Integrating microbial ecology into ecosystem models: challenges and priorities. Biogeochem 109: 7–18.
Wertz S, Degrange V, Prosser JI, Poly F, Commeaux C, Freitag T et al. (2006). Maintenance of soil functioning following erosion of microbial diversity. Environ Microbiol 8: 2162–2169.
Acknowledgements
This work was supported by the United States National Science Foundation (CBET-0853820). We thank Matthew Porter and J. Malia Andrus for the denitrifying bioreactor dataset, Jeremy Guest and Chinmay Soman for helpful comments and three anonymous reviewers for their insightful and constructive comments. JLZ, LFR, and ADK developed the approach. LFR and NAB implemented the approach and NAB conducted the analysis. JLZ led the writing and all authors contributed to editing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Zilles, J., Rodríguez, L., Bartolerio, N. et al. Microbial community modeling using reliability theory. ISME J 10, 1809–1814 (2016). https://doi.org/10.1038/ismej.2016.1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2016.1