Abstract
It is well known that bacteria often exist in naturally formed multispecies biofilms. Within these biofilms, interspecies interactions seem to have an important role in ecological processes. Little is known about the effects of interspecies interactions on gene expression in these multispecies biofilms. This study presents a comparative gene expression analysis of the Xanthomonas retroflexus transcriptome when grown in a single-species biofilm and in dual- and four-species consortia with Stenotrophomonas rhizophila, Microbacterium oxydans and Paenibacillus amylolyticus. The results revealed complex interdependent interaction patterns in the multispecies biofilms. Many of the regulated functions are related to interactions with the external environment and suggest a high phenotypic plasticity in response to coexistence with other species. Furthermore, the changed expression of genes involved in aromatic and branched-chain amino acid biosynthesis suggests nutrient cross feeding as a contributing factor for the observed synergistic biofilm production when these four species coexists in a biofilm.
Similar content being viewed by others
Introduction
Natural biofilms accommodate multiple species where interactions are crucial in shaping dynamic communities (Hall-Stoodley et al., 2004; Burmølle et al., 2014; Lee et al., 2014). Previously, we showed cooperative interactions between four soil isolates in a multispecies biofilm resulting in increased cell numbers and biofilm formation relative to single-species biofilms (Ren et al., 2015). A strong biofilm producer, Xanthomonas retroflexus, was combined with Stenotrophomonas rhizophila, Microbacterium oxydans and Paenibacillus amylolyticus and despite their incapability of single-species biofilm formation, the three other strains were indispensable for the strong synergy, which makes this consortium a powerful model for studying interactions in multispecies biofilms.
Here, the gene expression profile of X. retroflexus in single-species biofilm was compared to dual- and four-species biofilms with S. rhizophila, M. oxydans and P. amylolyticus. We utilized a RNA-Seq-based metatranscriptomic approach and found significant effects of interspecies interactions on gene expression.
Results
Transcriptome libraries were created from single-, dual- and four-species biofilms (in three, three and five replicates, respectively) of X. retroflexus combined with S. rhizophila, M. oxydans and P. amylolyticus (Supplementary Materials and Methods). In total, 125 Mb of the transcripts were mapped to coding sequences of the X. retroflexus genome with sample sizes ranging between 3.16 and 17.38 Mb (Supplementary Tables S1 and S2). To ensure that the statistical analyses were based on a sufficient signal, the top thousand most abundant genes, with a mapping of at least 400 transcripts, were tested (⩾8.10-fold coverage). About 84% of the total base pairs of the protein-coding transcriptome of X. retroflexus mapped to these thousand genes.
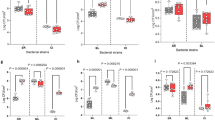
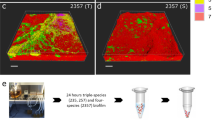
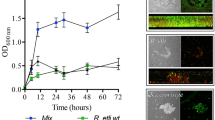
Protein-coding transcripts from S. rhizophila and M. oxydans comprised a low fraction of the total metatranscriptome compared with X. retroflexus (meanS. rhizophila=10.5 ±0.6% and meanM. oxydans=2.1 ±0.9%), whereas P. amylolyticus transcripts dominated in samples from the dual-species biofilm (meanP.amylolyticus=68.0 ±12.4%) (Figure 1a). When all four species were combined, the fraction of P. amylolyticus transcripts decreased (meanP. amylolyticus=52.0 ±4.8%) and the total amount of biofilm increased significantly (Figures 1a and b). Investigating the gene expression of X. retroflexus, the five replicates of the four-species biofilm and the three replicates of the dual-species biofilm with P. amylolyticus created two distinct clusters in a principal coordinate analysis (PCoA) of the Bray–Curtis dissimilarity between the samples (Figure 1c). These emphasize a very consistent and unique expression profile of the multispecies biofilms.
An RNA sequencing experiment was performed on a multispecies biofilm model system isolated from soil. X. retroflexus (Xr) was cultured in three replicates of single- and dual-species biofilms and five replicates of a four-species biofilm with S. rhizophila (Sr), M. oxydans (Mo) and P. amylolyticus (Pa). (a) Mean relative abundance of protein-coding transcripts from the four different species in the five different biofilms. (b) The total biofilm formation of the five combinations after 24 h, relative to the total biofilm measured for the best single-species biofilm former (in all cases X. retroflexus). The plot is based on data published in Ren et al. 2015, where the biofilm was quantified by use of crystal violet assay. (c) PCoA of the Bray–Curtis dissimilarity of the gene expression in X. retroflexus in single- dual- and four-species biofilms. The five dot colors equal the colored bars for the biofilm combinations presented in Figure 1b.
About 158 genes were significantly regulated at a minimum of eightfold expression change when comparing the single-species to the dual- and four-species biofilms (Supplementary Materials and Methods). The fold changes of these genes are displayed in Figure 2a and support the observations from the PCoA. Many of the upregulated genes gradually increased from the P. amylolyticus dual-species biofilms to the four-species biofilms and this pattern seemed to be highly correlating with the first principal coordinate (Figures 1c and 2a). A total of 141 genes were differentially expressed in the four-species biofilms and 52 of these were specific for this biofilm (fold change >8 in the four-species and fold change <2 in the dual-species biofilms). Of these 52, only four were upregulated, whereas 48 were downregulated.
(a) Heatmap of the log2 fold change (LFC) between X. retroflexus gene expression in the single-species biofilm and multispecies biofilms. The dual-species biofilms with S. rhizophila (Sr), M. oxydans (Mo), P. amylolyticus (Pa) and the four-species biofilm (All), are all displayed and included 157 genes with a fold change >8. The inner circle shows differentially upregulated (blue/green) and downregulated (brown) genes in the four-species biofilms. Darker colors represent genes uniquely regulated in the four-species biofilm (fold change >8 in the four-species and fold change <2 in the dual-species biofilms). The size of external circles reflects the contribution of the genes in the separation of the samples in the PCoA (Figure 1c) by displaying the significance of the adjusted P-values (Benjamini–Hochberg) of the Pearson correlation between the genes and PCo1. The dendrogram represents a hierarchical clustering of the LCFs. (b) Likert chart of the eggNOG functional categorization of the genes that are differentially expressed in the four-species biofilm. The color codes are equal to Figure 2a. (c) Differentially regulated genes in the branched-chain and in the aromatic amino acid biosynthesis pathways for the four-species biofilm. Alpha-ketobutyrate and pyruvate are both processed with hydroxyethyl-thiamine pyrophosphate (TPP) through the same enzymatic apparatus to synthesize either isoleucine or valine and leucine. The observed downregulation of ilvA (threonine deaminase) indicates a decrease in alpha-ketobutyrate synthesis, which shifts the system away from the isoleucine pathway. The upregulation of ilvE (branched-chain amino acid aminotransferase) might indicate a resulting increase in valine and leucine production. The biosynthesis of aromatic amino acids is initiated by the production of chorismate. The downregulation of trpE (anthranilate synthase component I) and trpB (tryptophan synthase beta chain) indicates a lower conversion of chorismate to anthranilate and a decrease in the last step of the tryptophan synthesis, resulting in lower tryptophan levels. The transcripts of trpG (anthranilate synthase component II) and trpA (tryptophan synthase alpha chain) were of too low abundance to give a significant signal. LFCs relative to the single-species biofilm are displayed in the single-row heatmaps as in Figure 2a. Genes indicated with a ‘#’ were among the top 1000 most abundant transcripts, which were tested for differential expression.
To get an overview of the changed functions in the four-species biofilm, the 141 responding genes were annotated to eggNOG functional groups and Figure 2b illustrates the distributions within the categories (Supplementary Table S3; Powell et al., 2014). Of the 12 genes annotated to ‘Amino Acid Metabolism and Transport’, two were specifically involved in branched-chain amino acid synthesis and two were involved in tryptophan synthesis. The downregulation of ilvA (threonine deaminase) and upregulation of ilvE (branched-chain amino acid aminotransferase) indicate a decrease in alpha-ketobutyrate production, reducing isoleucine production and shifting the pathway to valine and leucine synthesis when all four species are present (Figure 2c; Levinthal et al., 1973).
Chorismate is a common precursor for aromatic amino acids. The downregulation of trpE (anthranilate synthase component I) and trpB (tryptophan synthase beta chain) indicates lower priority of the tryptophan pathway (Figure 2c) in the multispecies biofilm. A unique downregulation of a gene (kynU kynureninase) involved in the tryptophan catabolism further supports this prioritization. The remaining seven genes within the eggNOG functional category were annotated to either ambiguous functions or proved hard to interpret (Supplementary Table S3).
Ten genes encoding transcription factors, two chemotaxis genes, genes associated with the cell membrane and cell wall, 21 transporters, efflux pumps and signal transduction functions were regulated, indicating an active response to changes in the environment (Phelan et al., 2011).
Discussion
This study compared the transcription profiles of X. retroflexus grown in single-, dual- and four-species biofilms in combination with S. rhizophila, M. oxydans and P. amylolyticus. When engaged in the four-species biofilm, a unique expression pattern for X. retroflexus emerged, which differed from the sum of the changes observed with the three strains individually. This indicates that complex interspecies interactions induce unique and consistent changes in the global gene expression, reflecting the emergent behavior of such multispecies communities.
P. amylolyticus induced the largest expression response in X. retroflexus and comprised a larger fraction of the total protein-coding transcripts, compared with the other dual-species biofilms. Even though the genes induced by P. amylolyticus in dual-species incubations were consistently upregulated in the four-species biofilms, synergistic effects were not observed in terms of higher biofilm production in the dual-species incubations.
A total of 141 genes were regulated in the four-species biofilm and many of the functions can be related to the interaction with the external milieu and indicate a high phenotypic plasticity in response to the presence of other species (Frias-Lopez and Duran-Pinedo 2012; Sztajer et al., 2014). At least 52 genes were regulated only in the four-species biofilm and represent a diverse spectrum of functions. The majority (48) was uniquely downregulated, which could imply that the lifestyle in the multispecies biofilm allows X. retroflexus to deprioritize functions vital for the existence in monoculture. This hypothesis is supported by the observed gene regulation in the biosynthesis of costly amino acids and suggests that the cohabitants in multispecies biofilms may share products of energy-consuming pathways. Such division of labor has been shown to result in an overall higher fitness (Schink, 2002; Poltak and Cooper, 2011; Pande et al., 2014, 2015).
Overall, the results show a highly complex gene expression response to interspecies interactions in multispecies biofilms. Further analysis of the metabolome and gene knockout experiments, especially targeting the amino acid biosynthesis pathways, will lead to further understanding of the molecular mechanisms behind the observed synergistic biofilm production (Ren et al., 2015).
References
Burmølle M, Ren D, Bjarnsholt T, Sørensen SJ . (2014). Interactions in multispecies biofilms: do they actually matter? Trends Microbiol 22: 84–91.
Frias-Lopez J, Duran-Pinedo A . (2012). Effect of periodontal pathogens on the metatranscriptome of a healthy multispecies biofilm model. J Bacteriol 194: 2082–2095.
Hall-Stoodley L, Costerton JW, Stoodley P . (2004). Bacterial biofilms: from the natural environment to infectious diseases. Nat Rev Microbiol 2: 95–108.
Lee KWK, Periasamy S, Mukherjee M, Xie C, Kjelleberg S, Rice SA . (2014). Biofilm development and enhanced stress resistance of a model, mixed-species community biofilm. ISME J 8: 894–907.
Levinthal M, Williams LS, Levinthal M, Umbarger HE . (1973). Role of threonine deaminase in the regulation of isoleucine and valine biosynthesis. Nat New Biol 246: 65–68.
Pande S, Merker H, Bohl K, Reichelt M, Schuster S, de Figueiredo LF et al. (2014). Fitness and stability of obligate cross-feeding interactions that emerge upon gene loss in bacteria. ISME J 8: 953–962.
Pande S, Shitut S, Freund L, Westermann M, Bertels F, Colesie C et al. (2015). Metabolic cross-feeding via intercellular nanotubes among bacteria. Nat Commun 6: 6238.
Phelan VV, Liu WT, Pogliano K, Dorrestein PC . (2011). Microbial metabolic exchange—the chemotype-to-phenotype link. Nat Chem Biol 8: 26–35.
Poltak SR, Cooper VS . (2011). Ecological succession in long-term experimentally evolved biofilms produces synergistic communities. ISME J 5: 369–378.
Powell S, Forslund K, Szklarczyk D, Trachana K, Roth A, Huerta-Cepas J et al. (2014). eggNOG v4.0: nested orthology inference across 3686 organisms. Nucleic Acids Res 42: D231–D239.
Ren D, Madsen JS, Sørensen SJ, Burmølle M . (2015). High prevalence of biofilm synergy among bacterial soil isolates in cocultures indicates bacterial interspecific cooperation. ISME J 9: 81–89.
Schink B . (2002). Synergistic interactions in the microbial world. Antonie Van Leeuwenhoek 81: 257–261.
Sztajer H, Szafranski SP, Tomasch J, Reck M, Nimtz M, Rohde M et al. (2014). Cross-feeding and interkingdom communication on dual-species biofilms of Streptococcus mutans and Candida albicans. ISME J 8: 2256–2271.
Acknowledgements
This research was funded by the Danish Council for Strategic Research (Novenia and Center for Gut, Grain and Greens), The Danish Council for Independent Research and Villum Fonden: Young Investigator Programme. We would like to thank Karin P. Vestberg and Annette H. Løth for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Hansen, L., Ren, D., Burmølle, M. et al. Distinct gene expression profile of Xanthomonas retroflexus engaged in synergistic multispecies biofilm formation. ISME J 11, 300–303 (2017). https://doi.org/10.1038/ismej.2016.107
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2016.107
This article is cited by
-
Transcriptional Changes in Bifidobacterium bifidum Involved in Synergistic Multispecies Biofilms
Microbial Ecology (2022)
-
The initial inoculation ratio regulates bacterial coculture interactions and metabolic capacity
The ISME Journal (2021)
-
The Marine Bacterium Shewanella woodyi Produces C8-HSL to Regulate Bioluminescence
Microbial Ecology (2020)
-
Deciphering links between bacterial interactions and spatial organization in multispecies biofilms
The ISME Journal (2019)
-
High-resolution novel method for tracking bacteria in a multi-species biofilm
Archives of Microbiology (2019)