Abstract
The high diversity of microbial communities hampers predictions about their responses to global change. Here we investigate the potential for using a phylogenetic, trait-based framework to capture the response of bacteria and fungi to global change manipulations. Replicated grassland plots were subjected to 3+ years of drought and nitrogen fertilization. The responses of leaf litter bacteria and fungi to these simulated changes were significantly phylogenetically conserved. Proportional changes in abundance were highly correlated among related organisms, such that relatives with approximately 5% ribosomal DNA genetic distance showed similar responses to the treatments. A microbe’s change in relative abundance was significantly correlated between the treatments, suggesting a compromise between numerical abundance in undisturbed environments and resistance to change in general, independent of disturbance type. Lineages in which at least 90% of the microbes shared the same response were circumscribed at a modest phylogenetic depth (τD 0.014–0.021), but significantly larger than randomized simulations predict. In several clades, phylogenetic depth of trait consensus was higher. Fungal response to drought was more conserved than was response to nitrogen fertilization, whereas bacteria responded equally to both treatments. Finally, we show that a bacterium’s response to the manipulations is correlated with its potential functional traits (measured here as the number of glycoside hydrolase genes encoding the capacity to degrade different types of carbohydrates). Together, these results suggest that a phylogenetic, trait-based framework may be useful for predicting shifts in microbial composition and functioning in the face of global change.
Similar content being viewed by others
Introduction
Over a decade ago, Lavorel and Garnier (2002) proposed a trait-based framework to predict how global change will affect community composition and ecosystem functioning. A trait is any morphological, biochemical, physiological, structural or behavioral characteristic of an organism (Díaz et al., 2013) and, at least for larger organisms such as plants and animals, traits tend to be relatively consistent within a species. The framework divides traits into two types: response and effect traits. Response traits of a species determine how its abundance changes in the face of a new environment; the collection of a community’s response traits thus underlies changes in community composition (that is, the relative abundance of species). In contrast, effect traits (hereafter, ‘functional’ traits as in Srivastava et al. (2012)) are species characteristics that influence ecosystem properties or processes such as nutrient cycling and trace gas emissions. In theory, knowledge about the response and functional traits of all species in a community could be used to predict how composition will shift in response to environmental change and how, in turn, this shift will alter ecosystem functioning (Garnier et al., 2004; Allison and Martiny, 2008; Suding et al., 2008; Williams et al., 2010; Díaz et al., 2013).
A clear limitation of this framework is that it is not generally feasible to quantify the traits of each and every species in a community, particularly those that are microbial and highly diverse (Bruggeman et al., 2009). To overcome this, some researchers have proposed considering the trait framework in a phylogenetic context (Allison and Martiny, 2008). Plant and animal traits often show a phylogenetic signal, whereby the similarity between two species’ traits is correlated with their genetic relatedness (Harvey and Pagel, 1991; Freckleton et al., 2002; Blomberg et al., 2003; Srivastava et al., 2012; Langille et al., 2013). Thus, a phylogeny may be useful for predicting the traits of related, unstudied organisms. Indeed, recent work suggests that phylogenetic relatedness among plant species is correlated with their similarity in functional traits, leading to a positive relationship between a plant community’s phylogenetic diversity and its productivity (Cadotte et al., 2008, 2009; Flynn et al., 2011).
For microorganisms, the phylogenetic conservation of traits is less clear. Horizontal gene transfer, gene loss and convergent evolution may confound the relationship between traits and phylogeny (Doolittle, 1999; Snel et al., 2002; Fitzpatrick, 2012). Given the central role of microorganisms in biogeochemical cycling, however, the application of a trait-based framework could be immensely valuable as a path to incorporate microbial diversity in ecosystem models (Allison, 2012). Whereas some microbial traits, such as denitrification, appear to have been gained and lost repeatedly throughout the bacterial domain (Lu et al., 2014), recent work finds that many microbial traits are indeed phylogenetically patterned (Lennon et al., 2012; Berlemont and Martiny, 2013; Martiny et al., 2013; Zimmerman et al., 2013). A comparative genomic analysis demonstrated particularly strong phylogenetic conservatism for oxygenic photosynthesis, methane oxidation and sulfate reduction (Martiny et al., 2013), examples of traits that could directly influence ecosystem functioning. Similarly, the ability of leaf-decomposer fungi to metabolize various organic N compounds seems to be genetically correlated (McGuire et al., 2010). Comparatively less is known about the phylogenetic patterning of microbial response traits or whether they are linked with functional traits. However, broad taxonomic ranks of bacteria (for example, classes, orders and phyla) appear to be ‘ecologically coherent,’ as they often covary over space, habitat and season (Philippot et al., 2010). These patterns suggest that certain microbial responses to particular environmental variables may be evolutionarily conserved.
Apart from a microbe’s innate tolerance to environmental change, its success in a new context will depend on gene-by-environment interactions and interactions with other organisms. To account for these complexities, the suitability of a phylogenetic response trait framework for global change predictions must be tested in the field. For instance, recent declines or extinctions of plant (Willis et al., 2008) and bird (Davis et al., 2010) species are phylogenetically patterned, and freshwater zooplankton disturbed by human activities were comprised of closer relatives than expected by chance (Helmus et al., 2010). In microbial communities, the short-term response of grassland soil bacteria to abrupt changes in moisture appears to be phylogenetically conserved at approximately the phylum rank (Placella et al., 2012; Barnard et al., 2013). The response of soil bacteria to multiple cycles of drying–rewetting also appears to be conserved, albeit at a finer phylogenetic scale (Evans and Wallenstein, 2014). It remains unknown, however, whether microbial responses to longer-term, press disturbances such as those expected via human-driven global change, are also phylogenetically conserved. It would also be useful to know whether the traits that constrain a microbe’s response are correlated with its functional traits, and thus enable predictable changes in microbial functioning.
To address these unknowns, we investigated the response of bacteria and fungal communities to added nitrogen and reduced precipitation (drought) in a global change field experiment. We focused on leaf litter microbial communities, which play an important role in global carbon cycling (Cornelissen et al., 2007). Microbial decomposition of dead plant material regulates the rate that terrestrial carbon is released as CO2 into the atmosphere versus its accumulation in soils.
We tested four predictions about microbial response traits. First, we predicted that the response of microbes (changes in their relative abundance) to simulated global change would be clustered with respect to phylogeny. Second, we hypothesized that bacterial responses would be less phylogenetically conserved than those of fungi because of presumably higher rates of vertical gene transfer (Doolittle, 1999) and other factors weakening the correlation between phylogeny and response. Third, we expected that responses to nitrogen addition (in both fungi and bacteria) would be less phylogenetically conserved compared with those of drought, as traits conferring resistance to drought (involving specialized cell wall architecture and osmoregulation among others) likely involve more genes than those responding to nitrogen availability. Gene systems consisting of many interacting proteins are less likely to be transferred between organisms resulting in a positive correlation between trait complexity and phylogenetic conservatism (Wellner et al., 2007; Martiny et al., 2013).
Finally, we sought to determine whether shifts in microbial composition in response to global change may lead to predictable functional changes. We predicted that a litter microbe’s response would correlate with its potential functional traits. To assay functional traits, we estimated a taxon’s genomic capacity for litter decomposition based on sequenced genomes. We quantified the type and number of glycoside hydrolases (GHs), genes encoding enzymes that degrade glycosidic bonds between carbohydrates, including those comprising plant cell walls.
Materials and methods
Sampling and molecular methods
Our experiment was conducted in a grassland savannah located near Irvine, CA (33.74N, 117.70W), described in detail elsewhere (Allison et al., 2013). We used a subset of plots from an existing field manipulation of precipitation and nitrogen inputs that began in February 2007. The field manipulation involves two levels of precipitation (ambient or reduced) applied at the plot scale and two levels of nitrogen (ambient or added) applied to subplots within precipitation treatments. This design is replicated in eight experimental blocks. Within each block, we only sampled subplots with (a) ambient precipitation and ambient nitrogen (‘control’ plots), (b) reduced precipitation ambient nitrogen (‘drought’ plots) and (c) ambient precipitation added nitrogen (‘nitrogen fertilization’ plots). Thus, we did not study any effects of interactions between precipitation and nitrogen. Rainfall was reduced by covering the 6.7 × 9.3 m whole plots with clear polyethylene during a subset of the rainstorms each winter to achieve a ~55% reduction in annual precipitation (from 283 to 141 mm per year; Supplementary Table S1). Every year, nitrogen fertilization subplots received 20 kg N/ha as soluble CaNO3 before the growing season and 40 kg N/ha as 100-day release CaNO3 during the growing season.
Three leaf litter subsamples were collected from each replicate plot using a random point intercept strategy every four months between April 2010 and February 2011 (N=8 blocksx3 treatmentsx4 seasons=96 samples). Sampled litter contained a mixture of dead standing and well-weathered organic material. Within-replicate litter was dried, pooled and homogenized in a coffee grinder with a stainless steel insert. Environmental genomic DNA was extracted from 50 mg of homogenized litter using a CTAB extraction/spin column purification method as described in DeAngelis et al. (2010). The communities of fungi and bacteria in each sample were PCR amplified using MID encoded fusion primers containing the primer pairs LROR and LR5_F (fungi; D1 and D2 regions of the 28S) or 515f and 907r (bacteria; v4 of the 16S) using PCR conditions as in Amend et al. (2012) and Beller et al. (2012), respectively. Amplicons were combined by locus into equimolar pools, cleaned using a magnetic bead protocol and the two pools were sequenced on separate regions of a 454 GS FLX+ sequencing plate at the Duke University IGSP facility. SFF files and associated metadata are deposited in NCBI SRA under deposition number SRP041807.
Sequence processing and phylogenetic tree inference
Sequences were quality control processed, ‘denoised’ and binned into operational taxonomic units (OTUs) at 97% sequence identity using the UCLUST algorithm (Edgar et al., 2011) as implemented in the QIIME analytical pipeline (Caporaso et al., 2010). Although this cutoff is the de facto standard for circumscribing bacteria OTUs at approximately the species or genus level, the fungal large ribosomal subunit gene is less variable, so the same dissimilarity value likely circumscribes fungal OTUs at a higher taxonomic level. Following initial quality filtering, 595 602 and 387 242 sequences were retained and circumscribed into 1211 and 734 OTUs (bacteria and fungi, respectively). Samples contained a mean of 5598±2743 (s.d.; bacteria) or 3850±1234 (fungi). To calculate community diversity statistics, samples were normalized to the same sampling depth by randomly selecting 2613 sequences (bacteria) or 1988 sequences (fungi) over 100 resamples. Mean indexes calculated among resampled replicates were reported.
Representative sequences from each OTU were assigned taxonomy using the RDP classifier with its native LSU database (fungi) or Greengenes (bacteria). Assignments to a given taxonomic rank were retained given >80% probability of correct assignment. Sequences were aligned in MAFFT v 6.8 (Katoh et al., 2009) with outgroup taxa. One hundred maximum likelihood trees were inferred using RAxML 7.2.8 (Stamatakis et al., 2008) using a GTR+CAT model to infer the tree, and a GTR+Γ model to evaluate the resulting topologies. Outgroup taxa were pruned before subsequent analyses. To maintain consistency with previous publications using consenTRAIT metrics, these were calculated as mean values calculated across 100 neighbor-joining trees inferred using PHYLIP (Felsenstein, 2005).
Statistical calculations
Increases and decreases in OTU relative abundance following global change treatments were calculated as log2-fold ratios between communities in manipulated and non-manipulated control plots. As log transformation of zero values are undefined, OTUs that were absent in a given treatment were tallied as present with a relative abundance equivalent to 50% that of a singleton in the most sequence-rich sample. Each OTU was screened for significant increase or decrease among replicates using a negative binomial frequency distribution model in the program DESeq2 (V. 1.2.8; Anders and Huber, 2010) within R, based on non-normalized library counts. Significance values were corrected for multiple tests using the Benjamini–Hochberg procedure with an adjusted alpha of 0.2. Phylogenetically independent contrasts among the response traits were calculated using the ape package (Paradis et al., 2004) in R, and were used in the correlation analysis.
The relationship between sequence abundance and change in proportional abundance because of global change disturbance was compared using a Mantel correlogram as calculated within the Vegan package of R (V. 2.0-7; Oksanen et al., 2013). Autocorrelation among values of log2-fold changes are plotted against a lag of genetic distance and compared with 999 randomizations. Alpha significance values were progressively corrected among sequence dissimilarity classes.
In order to determine which lineages were impacted, and at what phylogenetic depths, we used an index of trait conservatism (consenTRAIT; Martiny et al., 2013). As the analysis considers binary responses, increases and decreases relative to control communities are considered separately. Phylogenetic consensus was noted for each node in the phylogenetic tree when 90% or greater of subtending leaf nodes (tips) displayed the same response to global change treatments (increase or decrease in relative abundance). To determine the mean phylogenetic depth at which microbial response traits were clustered, the phylogenetic ‘depth’ of a consensus lineage (τD) was measured as the average phylogenetic branch length between the lineage’s OTUs and the root that subtends them, averaged over 100 neighbor-joining bootstrap trees. Calculations and comparisons with randomized community node age distribution followed procedures of Martiny et al. (2013). Nodes containing a greater proportion of consensus than expected under random were identified using the Analysis of Traits function in the program Phylocom (V. 4.2; Webb et al., 2008).
We expected communities subjected to experimental manipulations to contain OTUs with a higher degree of phylogenetic relatedness than predicted via random sampling of all OTUs present in our plots. To test for phylogenetic clustering among manipulated communities, we used two metrics as in Webb et al. (2002). The nearest taxon index (NTI) is a standardized measure of the mean phylogenetic distance between all individuals and their closest relatives in a community. The net relatedness index (NRI) is a measure of the mean phylogenetic distance between two individuals drawn randomly from a community. The NTI and NRI were calculated using Picante (V. 1.6-0; Kembel et al., 2010) within R. Analyses were run with and without sequence abundance information. With abundance data included, the calculations measure the relatedness of individuals within communities and how these change with along with shifts in OTU relative abundance. Omitting abundance data, the analyses test the relatedness of OTUs within communities and how presence/absence relates to the environment. To test whether phylogenetic clustering differed from random, we constructed a local microbial pool of potential community members by combining all OTUs found among all replicates. Observed NTI and NRI were compared with a null distribution of 1000 communities drawn at random from the local pool selected by shuffling OTU labels. Standardized effect size scores (Z; the difference between observed and expected values, divided by the s.d. of the expected values) are reported as means of all replicates per factor. The Z values are equivalent to negative NTI or NRI statistics.
To examine the relationship between GH family genes and response, bacteria OTUs in our study were linked to 8840 sequenced genomes available in the SEED database (Disz et al., 2010), by excising their 16S gene regions and compiling them into a database for closed-reference clustering at 97% using the UCLUST algorithm (Edgar 2010). Fifty-three genomes were linked to OTUs from our study using this method (Supplementary Information Data), and were retained for correlation with response values. GH family genes were binned by category using the CAZy database classification scheme, and tallies within each category were treated as fixed effects within a linear model using a Brownian motion phylogenetic error term (as in Ho and Ané, 2014) with response (log2fold ratios of change compared with controls) as the dependent variable.
Results
Individual OTU responses
Both drought and nitrogen addition produced clear microbial ‘winners’ and ‘losers’—for example, OTUs whose average relative abundance increased or decreased in response to the experimental treatments across all four seasons. We found that drought significantly impacted the relative abundance of more OTUs (108 OTUsbacteria (8.9%); 77 OTUsfungi (10.4%)) than did nitrogen addition (53 OTUsbacteria (4.3%); 57 OTUsfungi (7.8%)), where OTUs are defined as ⩾97% similarity of ribosomal sequences. Further, an OTU's change in relative abundance in response to nitrogen addition was correlated with its response to drought (Figure 1); a microbe that responded negatively to drought also tended to respond negatively to nitrogen addition. Considering just the subset of OTUs whose change in relative abundance significantly differed under both treatments, the relationship between the drought and nitrogen responses were even stronger (inset Figure 1).
Phylogenetically independent contrasts of log2fold ratios of global change effect size were significantly correlated between drought and nitrogen amended communities (N= 562, r2bacteria= 0.36; N=496, r2fungi=0.47). The correlation was stronger among OTUs whose changes in abundance were determined statistically significant for both treatments (inset N=10, r2bacteria=0.73; N=8, r2fungi=0.90). All correlations were significant at P=0.0001.
Phylogenetic depth of the responses
Given that OTUs were impacted by global change treatments, we investigated whether these impacts were correlated with the microbes’ genetic similarity. We then tested how this correlation depended on the genetic resolution of the taxonomic definition (that is, the percent ribosomal sequence similarity; Figure 2). Results across both bacteria and fungi and for both treatments were qualitatively similar. Microbial responses were most positively correlated with genetic similarity when OTUs were defined at a fine genetic scale and generally decreased as a function of sequence dissimilarity. Fungal response to drought clustered deepest of all; OTUs aligned at ⩽7.6% dissimilarity or less responded more similarly than would be expected from random. Fungal response to nitrogen fertilization was significantly clustered among OTUs ⩽5.0% dissimilar. The bacterial response to both treatments was also correlated among OTUs ⩽5.0% dissimilar, although the strength of the correlation was less than that of fungi. Bacteria responding to nitrogen addition also showed a significant positive correlation among OTUs of intermediate (~12% dissimilar) relatedness, a result of coordinated responses among distinct clusters scattered throughout the bacteria phylogeny. Thus, in the face of both nitrogen and precipitation manipulations, microbial OTUs within clusters of ⩽5.0% sequence dissimilarity tended to respond in a similar manner. This result indicated that microbial responses to these environmental changes were phylogenetically patterned at a broader taxonomic level than the typical operational definition of 3% sequence dissimilarity.
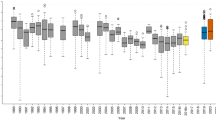
Effect size of the response to global change correlates with genetic similarity. Here the Mantel correlogram of change in relative abundance is plotted as a factor of genetic distance. Genetic distance is calculated as the proportion of sites that differ between aligned sequence pairs. Genetic distance classes in which the autocorrelation coefficient significantly differs from zero are filled. Observed values were compared against 999 bootstrap resamples. Overlapping points were offset slightly for visualization.
Microbial response impacted the phylogenic structure of communities as well. Drought significantly impacted community composition and diversity in all cases, whereas nitrogen only significantly impacted some indices of fungal community structure (Supplementary Table S2). The NTI analysis indicated that all replicates consisted of more closely related OTUs than random draws from a local pool of potential community members would predict (Supplementary Table S3). ConsenTRAIT analysis was consistent with the NTI results. At the whole community level, responses of fungi to both treatments were phylogenetically clustered at a greater depth than what would be expected given a randomized null. Conversely, for bacteria, only lineages with an increase in relative abundance were phylogenetically clustered (Table 1). Depth of the 90% consensus was modest (τD 0.014–0.021; average genetic distance of approximately 0.03–0.04 among OTUs), comparable to the cluster depth of bacterial lineages able to utilize complex carbon substrates, or traits involving roughly five or fewer genes (Martiny et al., 2013). Consensus lineages cluster depths followed a roughly normal distribution, and the deepest lineages were dispersed throughout the bacterial and fungal trees (Figure 3).
Phylogenetic distribution of response traits in global change treated communities. The inner ring is the log2-fold change in proportional abundance compared with the control treatment. Red is a decrease in abundance, blue is an increase in abundance. Colored nodes indicate ‘clustered’ lineages in which 90% or greater of the OTUs show consensus for increase or decrease in abundance. Open circles indicate nodes where values fall outside of the 95% confidence interval for consensus expected under a randomized null distribution. The outer ring indicates the order-level taxonomy of the OTU determined using a Bayesian classification algorithm. In cases where the probability of a correct assignment was <80%, no taxonomy was indicated. For a small minority of OTUs, taxonomic assignment did not match tree topology, however, no manual curation was attempted.
Identification of lineages with consistent response traits
The identity and location of the deepest clades differed between the nitrogen and drought treatments (Figure 3). In response to nitrogen fertilization, phylogenetic clustering in fungal communities was deepest among a clade within the Davidiellaceae group (affiliated with Verrucocladiosporium) and a clade containing the Sirobasidiacea (Tremellomycetes), both of which increased in relative abundance. Decreases in relative abundance were coordinated within a deep clade that includes Cryptococcus (also within the Tremellomycetes) and a clade within the Nectriacaeae/Bionectriaceae group of the Hypocreales. Drought treatments, on the other hand, showed the deepest phylogenetic clustering, all increases in relative abundance, among a clade of the Phaeosphaeriaceae (Pleosporales), a clade within the Niessliaceae (Hypocreales), a clade within the Leotiomycetes and a clade within the Russulales (Agaricomycetes).
Within the bacterial communities, a deep cluster within the Acidobacteriales consistently increased in relative abundance with nitrogen addition, as did clusters within the Rhizobiales (Proteobacteria) and Gemmatimonadales (Gemmatimonadetes). Conversely, the Sphingobacteriales (Bacteriodetes) decreased in relative abundance in response to nitrogen addition. The same lineage of Sphingobacteriales increased under drought conditions, as did a clade within the Myxococcales (Proteobacteria). Finally, a distinct lineage of Sphingobacteriales (associated with the Flexibacteraceae) and a clade within the Bacillales (Firmicutes) showed a coordinated decrease in relative abundance.
Correlations between response and functional traits
To examine the relationship between the observed responses and potential functional traits, we identified 53 bacterial OTUs from the litter samples that were closely related (>97% identity of the partial 16S ribosomal RNA gene) to fully sequenced genomes present in the SEED database (Disz et al., 2010). These OTUs tended to be abundant, comprising 18.7% of all bacterial sequences. The sequenced genomes were interrogated for the number of GH family genes, a suite of enzymes responsible for hydrolysis of glycosidic bonds required for decomposition of organic matter.
The response of a bacterial OTU to nitrogen addition was significantly correlated to the number of GH genes involved in cellulose, xylan, starch and sucrose degradation (Figure 4; Supplementary Table S4). These correlations suggested that changes in bacterial community composition in response to nitrogen addition should impact the abundance of functional genes involved in litter degradation. Specifically, OTUs that increased in relative abundance with nitrogen addition carried more cellulose and sucrose degrading genes and fewer xylan and starch degrading genes.
Coefficient plot of linear model explaining relationship between response traits and GH gene family abundance. Gene families are binned by putative substrate affinity (Y axis). Circles indicate coefficient values, inner and outer confidence intervals are one and two standard errors of the coefficient, respectively. Filled circles indicate coefficient values that differ significantly from zero. A positive coefficient indicates that gene abundance was higher among OTUs whose relative abundance increased in response to treatment. The model included a phylogenetic error term to account for evolutionary non-independence.
In contrast, a bacterial OTU’s response to drought was not as related to the GH genes present in the genomes. We detected only one correlation between the drought response and GH gene number; bacteria that increased in relative abundance with drought appear to carry fewer genes related to the breakdown of other plant carbohydrates (Figure 4; Supplementary Table S4).
Discussion
Our first prediction, that changes in microbial relative abundances in response to nitrogen fertilization and drought would be patterned by phylogeny, was supported by trait consensus analyses. At the whole community level, microbes were more closely related than random predictions, and trait depth exceeded random predictions in all categories except for bacteria that decreased in relative abundance. These results show that the response in relative abundance of bacteria and fungi to two global change perturbations—drought and nitrogen addition—is phylogenetically conserved at an intermediate (5–12% ribosomal RNA gene sequence dissimilarity) scale. This depth is greater than conserved traits conferring the production of extracellular enzymes (for example, chitinase or b-N-acetyl-glucosaminidase (Zimmerman et al., 2013)), but less conserved than complex metabolic activities such as oxygenic photosynthesis or methane oxidation (Martiny et al., 2013).
Contrary to our second prediction, the depth and distribution of the response traits, while significantly different from random expectations, were similar between bacteria and fungi. Although direct comparison of bacteria and fungi was difficult given differences in ribosomal loci and rates of evolution, we did not find substantial differences between bacteria and fungi by any of our analyses.
Support was mixed for our third prediction that response to drought and nitrogen addition would differ in their level of phylogenetic conservation. As anticipated, the response of fungal communities to drought appeared to be more conserved than the response to nitrogen fertilization. Indeed, fungal biogeography (Treseder et al., 2014) appears to be driven in part by traits conserved at the phylum level, which also confer drought tolerance. In contrast, the phylogenetic depths of the responses among bacteria were similar for the two treatments.
At the OTU level, we found a significant relationship between a microbe’s responses to both disturbance types. For both bacteria and fungi, an OTU's change in relative abundance in response to nitrogen fertilization was correlated with its response to drought. This correlation may be due to species interactions as opposed to innate resistance to environmental change. Fast growing opportunistic species, for example, may benefit from a decrease in microbial density, resource competition and shifts in nutrient availability.
The fourth prediction that an OTU’s response to global change is correlated with its functional potential was supported in the case of nitrogen addition. As a limiting nutrient, nitrogen availability is, in most cases, correlated with hydrolytic enzyme production because enzymes are relatively N-rich biomolecules (Sterner and Elser, 2002). Thus, nitrogen limitation should strongly impact microbes with high hydrolytic potential (Saiya-Cork et al., 2002) and lead to a positive correlation between an OTU’s response and its number of GH genes. As expected, added nitrogen selected for OTUs that carried more GH genes involved in cellulose and sucrose degradation. However, we also observed two negative correlations. The nitrogen treatment selected against OTUs with higher starch and xylan degradation potential. Generally, GH genes that target xylan are abundant within Bacteriodetes (Berlemont and Martiny, 2014). In this experiment, Bacteriodetes were negatively impacted by nitrogen fertilization and thus, a likely driver of this change in functional potential (Berlemont et al., 2014). Importantly, functional potential may not translate directly to gene or enzyme expression.
Although we observed phylogenetic clustering of response traits in our system, there are several scenarios under which phylogeny will not be an acceptable proxy for traits and thus, limit the application of this information for use in other systems. Both response and functional traits are often combinations of many physiological traits (Wainwright, 2007), encoded by suites of genes that are not tightly linked, and thus, less prone to phylogenetically patterned selection. Further, for global change predictions in particular, environmental disturbances will affect many ecosystem components such as plant community composition and litter quality (Allison et al., 2013), factors that alter microbial composition, but are not captured in this phylogenetic framework. Finally, the responses measured here are also relative to other community members; thus an identical change in absolute abundance may appear very different in another environmental context and within a different community. Differences in ribosomal gene copy number, as well as various artifacts introduced by library preparation steps may further distort the correlations between microbial abundance and that of sequence reads. Although we did not simultaneously measure absolute and relative abundances, nitrogen fertilization does not appear to alter microbial biomass in our system (Allison et al., 2013) likely diminishing differences in responses measured in terms of relative and absolute abundance. However, drought does reduce bacteria cell counts and increase fungal hyphal lengths (Allison et al., 2013). As a result, bacterial ‘winners’ who modestly increased in abundance relative to the overall community, may have actually realized a decrease in overall biomass.
Our experiment demonstrates two ways that a phylogenetic, trait-based framework may be a useful step toward predicting microbial responses to global change. First, the responses to changes in precipitation and nitrogen availability appear to be phylogenetically conserved, a pattern that holds true for both bacterial and fungal diversity. In lieu of studying the individual responses of thousands of species, we may instead assign these species to circumscribed groups whose response to environmental change is already or can be documented. In particular, there is the potential to both identify particularly sensitive lineages and predict the response of closely related species in other systems and locations (for example, Langille et al., 2013). Second, our study illustrates that response traits may be correlated with specific functional traits relevant to carbon and nutrient cycling. That these correlations were more prevalent with changes in nitrogen than precipitation suggests that microbial-driven functioning may be more predictable under some types of global changes as has been suggested in litter microcosms (Matulich and Martiny 2014; Amend et al., 2015). A trait-based framework may enable researchers to reconcile the vast complexity of community composition with biogeochemical cycles and other ecosystem services (Violle et al., 2014), and phylogeny may be a useful tool for parameterizing the breadth of trait diversity. Like for plants and animals, then, determining the extent to which microbial response traits correspond with functional traits is a critical next step toward a more complete understanding of how global change, and shifts in microbial community, will affect future ecosystems.
References
Allison SD, Lu Y, Weihe C, Goulden ML, Martiny AC, Treseder KK et al.(2013). Microbial abundance and composition influence litter decomposition response to environmental change. Ecology 94: 714–725.
Allison SD, Martiny JBH . (2008). Colloquium paper: resistance, resilience, and redundancy in microbial communities. Proc Natl Acad Sci USA 105: 11512–11519.
Allison SD . (2012). A trait-based approach for modelling microbial litter decomposition. Ecol Lett 15: 1058–1070.
Amend AS, Barshis DJ, Oliver TA . (2012). Coral-associated marine fungi form novel lineages and heterogeneous assemblages. ISME J 6: 1291–1301.
Amend AS, Matulich KL, Martiny JB . (2015). Nitrogen addition, not initial phylogenetic diversity, increases litter decomposition by fungal communities. Front Microbiol 6: 109.
Anders S, Huber W . (2010). Differential expression analysis for sequence count data. Genome Biol 11: R106.
Barnard RL, Osborne CA, Firestone MK . (2013). Responses of soil bacterial and fungal communities to extreme desiccation and rewetting. ISME J 7: 2229–2241.
Beller HR, Han R, Karaoz U, Lim H, Brodie EL . (2012). Genomic and physiological characterization of the chromate-reducing, aquifer-derived Firmicute Pelosinus sp. strain HCF1. Appl Environ Microbiol 79: 63–73.
Berlemont R, Allison SD, Weihe C, Lu Y, Brodie EL, Martiny JB et al.(2014). Cellulolytic potential under environmental changes in microbial communities from grassland litter. Front Microbiol 5: 639.
Berlemont R, Martiny AC . (2013). Phylogenetic distribution of potential cellulases in bacteria. Appl Environ Microbiol 79: 1545–1554.
Berlemont R, Martiny AC . (2014). Genomic potential for polysaccharide deconstruction in bacteria. Appl Environ Microbiol 81: 1513–1519.
Blomberg SP, Garland T, Ives AR . (2003). Testing for phylogenetic signal in comparative data: behavioral traits are more labile. Evolution 57: 717–745.
Bruggeman J, Heringa J, Brandt BW . (2009). PhyloPars: estimation of missing parameter values using phylogeny. Nucleic Acids Res. 37: 179–184.
Cadotte MW, Cardinale BJ, Oakley TH . (2008). Evolutionary history and the effect of biodiversity on plant productivity. Proc Natl Acad Sci USA 105: 17012–17017.
Cadotte MW, Cavender-Bares J, Tilman D, Oakley TH . (2009). Using phylogenetic, functional and trait diversity to understand patterns of plant community productivity. PLoS One 4: e5695.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al.(2010). QIIME allows analysis of high-throughput community sequencing data. Nat Meth 7: 335–336.
Cornelissen JHC, van Bodegom PM, Aerts R, Callaghan TV, van Logtestijn RSP, Alatalo J et al.(2007). Global negative vegetation feedback to climate warming responses of leaf litter decomposition rates in cold biomes. Ecol Lett 10: 619–627.
Davis CC, Willis CG, Primack RB, Miller-Rushing AJ . (2010). The importance of phylogeny to the study of phenological response to global climate change. Phil Trans R Soc B Biol Sci 365: 3201–3213.
DeAngelis KM, Silver WL, Thompson AW, Firestone MK . (2010). Microbial communities acclimate to recurring changes in soil redox potential status. Environ Microbiol 12: 3137–3149.
Disz T, Akhter S, Cuevas D, Olson R, Overbeek R, Vonstein V et al.(2010). Accessing the SEED genome databases via Web services API: tools for programmers. BMC Bioinform 11: 319.
Doolittle WF . (1999). Phylogenetic classification and the universal tree. Science 284: 2124–2128.
Díaz S, Purvis A, Cornelissen JHC, Mace GM, Donoghue MJ, Ewers RM et al.(2013). Functional traits, the phylogeny of function, and ecosystem service vulnerability. Ecol Evol 3: 2958–2975.
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R . (2011). UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27: 2194–2200.
Edgar RC . (2010). Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26: 2460–2461.
Evans SE, Wallenstein MD . (2014). Climate change alters ecological strategies of soil bacteria. Ecol Lett 17: 155–164.
Felsenstein J . (2005). PHYLIP—Phylogeny Inference Package version 3.6.
Fitzpatrick DA . (2012). Horizontal gene transfer in fungi. FEMS Microbiol Lett 329: 1–8.
Flynn DFB, Mirotchnick N, Jain M, Palmer MI, Naeem S . (2011). Functional and phylogenetic diversity as predictors of biodiversity—ecosystem-function relationships. Ecology 92: 1573–1581.
Freckleton RP, Harvey PH, Pagel M . (2002). Phylogenetic analysis and comparative data: a test and review of evidence. Am Nat 160: 712–726.
Garnier E, Cortez J, Billès G, Navas M-L, Roumet C, Debussche M et al.(2004). Plant functional markers capture ecosystem properties during secondary succession. Ecology 85: 2630–2637.
Harvey PH, Pagel MD . (1991) The Comparative Method in Evolutionary Biology. Oxford University Press: : Oxford.
Helmus MR, Keller WB, Paterson MJ, Yan ND, Cannon CH, Rusak JA . (2010). Communities contain closely related species during ecosystem disturbance. Ecol Lett 13: 162–174.
Ho LST, Ané C . (2014). A linear-time algorithm for Gaussian and non-Gaussian trait evolution models. Syst Biol 63: 397–408.
Katoh K, Asimenos G, Toh H . (2009). Multiple alignment of DNA sequences with MAFFT. Methods Mol Biol 537: 39–64.
Kembel SW, Cowan PD, Helmus MR, Cornwell WK, Morlon H, Ackerly DD et al.(2010). Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26: 1463–1464.
Langille MGI, Zaneveld J, Caporaso JG, Mcdonald D, Knights D, Reyes JA et al.(2013). Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31: 814–821.
Lavorel S, Garnier E . (2002). Predicting changes in community composition and ecosystem functioning from plant traits: revisiting the Holy Grail. Funct Ecology 16: 545–556.
Lennon JT, Aanderud ZT, Lehmkuhl BK, Schoolmaster DR . (2012). Mapping the niche space of soil microorganisms using taxonomy and traits. Ecology 93: 1867–1879.
Lu H, Chandran K, Stensel D . (2014). Microbial ecology of denitrification in biological wastewater treatment. Water Res 64: 237–254.
Martiny AC, Treseder K, Pusch G . (2013). Phylogenetic conservatism of functional traits in microorganisms. ISME J 7: 830–838.
Matulich KL, Martiny JBH . (2014). Microbial composition alters the response of litter decomposition to environmental change. Ecology 8: 154–163.
McGuire KL, Bent E, Borneman J, Majumder A, Allison SD, Treseder KK . (2010). Functional diversity in resource use by fungi. Ecology 91: 2324–2332.
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O'Hara RB et al.(2013), Vegan: Community Ecology Package. R package version 2.0-7. Community ecology package version, 2, 0-7.
Paradis E, Claude J, Strimmer K . (2004). APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20: 289–290.
Philippot L, Andersson SGE, Battin TJ, Prosser JI, Schimel JP, Whitman WB et al.(2010). The ecological coherence of high bacterial taxonomic ranks. Nat Rev Microbiol 8: 523–529.
Placella SA, Brodie EL, Firestone MK . (2012). Rainfall-induced carbon dioxide pulses result from sequential resuscitation of phylogenetically clustered microbial groups. Proc Natl Acad Sci USA 109: 10931–10936.
Saiya-Cork KR, Sinsabaugh RL, Zak DR . (2002). The effects of long term nitrogen deposition on extracellular enzyme activity in an Acer saccharum forest soil. Soil Biol Biochem 34: 1309–1315.
Snel B, Bork P, Huynen MA . (2002). Genomes in flux: the evolution of archaeal and proteobacterial gene content. Genome Res 12: 17–25.
Srivastava DS, Cadotte MW, MacDonald AAM, Marushia RG, Mirotchnick N . (2012). Phylogenetic diversity and the functioning of ecosystems. Ecol Lett 15: 637–648.
Stamatakis A, Hoover P, Rougemont J . (2008). A rapid bootstrap algorithm for the RAxML Web servers. Syst Biol 57: 758–771.
Sterner RW, Elser JJ . (2002) Ecological Stoichiometry: The Biology of Elements from Molecules to the Biosphere. Princeton University Press: Princeton, NJ, USA.
Suding KN, Lavorel S, CHAPIN FS, Cornelissen JHC, Díaz S, Garnier E et al.(2008). Scaling environmental change through the community-level: a trait-based response-and-effect framework for plants. Glob Chang Biol 14: 1125–1140.
Treseder KK, Maltz MR, Hawkins BA, Fierer N, Stajich JE, McGuire KL . (2014). Evolutionary histories of soil fungi are reflected in their large-scale biogeography. Ecol Lett 17: 1086–1093.
Violle C, Reich PB, Pacala SW, Enquist BJ, Kattge J . (2014). The emergence and promise of functional biogeography. Proc Natl Acad Sci USA 111: 13690–13696.
Wainwright PC . (2007). Functional versus morphological diversity in macroevolution. Annu Rev Ecol Evol Syst 38: 381–401.
Webb CO, Ackerly DD, Kembel SW . (2008). Phylocom: software for the analysis of phylogenetic community structure and trait evolution. Bioinformatics 24: 2098–2100.
Webb CO, Ackerly DD, McPeek MA, Donoghue MJ . (2002). Phylogenies and community ecology. Annu Rev Ecol Syst 33: 475–505.
Wellner A, Lurie MN, Gophna U . (2007). Complexity, connectivity, and duplicability as barriers to lateral gene transfer. Genome Biol 8: R156.
Williams NM, Crone EE, Roulston TH, Minckley RL, Packer L, Potts SG . (2010). Biological conservation. Biol Conserv 143: 2280–2291.
Willis CG, Ruhfel B, Primack RB, Miller-Rushing AJ, Davis CC . (2008). Phylogenetic patterns of species loss in Thoreau's woods are driven by climate change. Proc Natl Acad Sci USA 105: 17029–17033.
Zimmerman AE, Martiny AC, Allison SD . (2013). Microdiversity of extracellular enzyme genes among sequenced prokaryotic genomes. ISME J 7: 1187–1199.
Acknowledgements
We thank Kristin Matulich for technical and statistical assistance, the Orange County Parks and the Irvine Ranch Conservancy for site access, and Eoin Brodie and several anonymous reviewers for critical feedback. This study was support by a NOAA VSP climate change fellowship (ASA) and by the US Department of Energy, Office of Science, Office of Biological and Environmental Research (BER), under Award Number DE-PS02-09ER09-25 (ACM, SDA, MLG, KKT and JBHM).
Author Contributions
ASA, JBHM and ACM designed research; ASA, YL and CW performed research; ASA, RB and ACM analyzed data; and ASA, ACM, SDA, MLG, KKT and JBHM wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Amend, A., Martiny, A., Allison, S. et al. Microbial response to simulated global change is phylogenetically conserved and linked with functional potential. ISME J 10, 109–118 (2016). https://doi.org/10.1038/ismej.2015.96
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.96
This article is cited by
-
Land management shapes drought responses of dominant soil microbial taxa across grasslands
Nature Communications (2024)
-
Bacterial and archaeal community distributions and cosmopolitanism across physicochemically diverse hot springs
ISME Communications (2023)
-
Plant property regulates soil bacterial community structure under altered precipitation regimes in a semi-arid desert grassland, China
Journal of Arid Land (2023)
-
Nitrate has a stronger rhizobacterial-based effect on rice growth and nitrogen use than ammonium in acidic paddy soil
Plant and Soil (2023)
-
Unbalanced diets enhance the complexity of gut microbial network but destabilize its stability and resistance
Stress Biology (2023)