Abstract
Natural transformation in bacteria facilitates the uptake and genomic integration of exogenous DNA. This allows horizontal exchange of adaptive traits not easily achieved by point mutations, and has a major role in the acquisition of adaptive traits exemplified by antibiotic resistance determinants and vaccination escape. Mechanisms of DNA uptake and genomic integration are well described for several naturally transformable bacterial species; however, the selective forces responsible for its evolution and maintenance are still controversial. In this study we evolved transformation-proficient and -deficient Acinetobacter baylyi for 175 days in serial transfer cultures where stress was included. We found that natural transformation-proficient populations adapted better to active growth and early stationary phase. This advantage was offset by the reduced performance in the late stationary/death phase. We demonstrate fitness trade-offs between adaptation to active growth and survival in stationary/death phase caused by antagonistic pleiotropy. The presented data suggest that the widely held assumption that recombination speeds up adaptation by rapid accumulation of multiple adaptive mutations in the same genetic background is not sufficient to fully account for the maintenance of natural transformation in bacteria.
Similar content being viewed by others
Introduction
Horizontal gene transfer (HGT) through natural transformation allows bacteria to actively take up exogenous DNA from the environment that can be recombined into the genome to generate novel genotypes. Natural transformation, as well as other modes of HGT (transduction and conjugation), ensures that adaptive evolution in bacterial species competent for transformation is not limited to the selection of slowly accumulating point mutations and genetic rearrangements (Ochman et al., 2000; Redfield, 2001; Thomas and Nielsen, 2005). Retrospective genome sequence analyses demonstrate that recombination following HGT has an important role in shaping bacterial genomes (Johnsborg et al., 2007; Didelot and Maiden, 2010). Bacteria competent for natural transformation have a potential advantage during niche expansion (Levin and Bergstrom, 2000; Ochman et al., 2000) and may, from one generation to the next, respond to abrupt environmental shifts that require genomic changes inaccessible by point mutations. This is exemplified by microbial adaptation to the immune system of vertebrates and resistance to the use of antimicrobial agents (Dubnau, 1999). Whereas the potential adaptive effects of natural transformation are clear, and for several species the main mechanistic aspects of DNA uptake and genomic integration have been elucidated, the selective forces responsible for the maintenance of the natural transformation machinery are still controversial (Redfield, 2001; Narra and Ochman, 2006; Baltrus, 2013).
Two of the existing hypotheses for the maintenance (and possibly evolution) of natural transformation suggest that DNA is taken up for nutritional purposes (Stewart and Carlson, 1986; Redfield, 1993a; Macfadyen et al., 2001; Redfield, 2001) or as a template for recombination-mediated repair of DNA double-stranded breaks (Bernstein et al., 1987; Hoelzer and Michod, 1991). According to these hypotheses, the integration of exogenous DNA with adaptive traits following uptake is coincidental. This contrasts the widely accepted view that natural transformation evolved and is maintained to 'explore the fitness landscape' in surrounding populations (Dubnau, 1999). Another hypothesis states that natural transformation is maintained due to competence-induced growth-arrest between rare episodes of beneficial DNA uptake (Johnsen et al., 2009). Perhaps the most intuitive hypothesis is a prokaryotic version of the classical explanation for the evolution of sex whereby DNA uptake increases the adaptation rates in bacterial populations through break-up of linkage disequilibrium and reduction of clonal interference in populations, leading to the accumulation of multiple beneficial mutations in single clones (Fisher, 1930; Muller, 1932; Vos, 2009). Theoretical reports reveal that recombination mediated by natural transformation can be selected for by inducing higher rates of adaptive evolution, but that this selective effect is frequency dependent (Levin and Cornejo, 2009) and depends on strong clonal interference between adaptive mutations (Moradigaravand and Engelstädter, 2013). The experimental evidence in support of this hypothesis is, however, limited to a single study in Helicobacter pylori (Baltrus et al., 2007). Other experimental studies in Acinetobacter baylyi, and Streptococcus pneumoniae populations suggest that this ‘transformation-for-recombination’ hypothesis is not universal in bacteria (Bacher et al., 2006; Engelmoer et al., 2013). It is likely that the different outcomes of the abovementioned reports on evolutionary benefits of natural transformation are due to species-specific differences in the selective forces maintaining natural transformation and/or variation in the experimental setups. Recent experimental evidence suggests conditional benefits of recombination as demonstrated under stressful growth conditions in sexually reproducing yeast (Gray and Goddard, 2012), and recently for S. pneumoniae where periods of stress offset the initial costs of competence (Engelmoer et al., 2013).
Our model organism A. baylyi (Barbe et al., 2004) is convenient for studying the possible benefits of natural transformation for three reasons: (1) competence is expressed upon dilution into new medium, (2) it stays competent during most of the log phase and (3) most of the cells in a population are competent for DNA uptake at the same time (Bacher et al., 2006; Metzgar et al., 2004).
In this report we test the ‘transformation-for-recombination’ hypothesis in A. baylyi cultures experimentally evolving for 175 days in minimal medium including extended periods in stationary and death phase between the serial transfer events. We show for the first time that the evolutionary benefits of natural transformation are growth phase-specific. Natural transformation-proficient populations adapt better to active growth and early stationary phase (where competence is expressed) but this advantage is offset during extended periods of stationary and death phase. Our data also suggest that fitness trade-offs between performance during exponential growth and survival in stationary/death phase are caused by antagonistic pleiotropy.
Materials and methods
Strains and media
The bacterial strains used in this study are listed in Table 1. Primers are listed in Supplementary Table SI 1. If not specified elsewhere, the strains and populations were grown in Luria Bertani (LB; Sambrook et al., 1989) or S2-minimal medium (Juni, 1974) supplied with lactate (0.1%) as carbon source (Nielsen et al., 1997). In addition, two carbon sources that A.baylyi is not able to utilize were added to provide an opportunity for adaptation, methyl-benzoate (2.5 mM) and arabinose (20 mM). S2-medium with the three carbon sources was designated S2-LMBA. Growth assays designed to explore growth on alternative carbon sources are described in Supplementary Information.
Strain construction
The various A. baylyi strains described in Table 1 are all derived from A. baylyi strain ADP1 (NC_005966) and were constructed by standard molecular biology techniques, including the pZR79/80 chromosomal insert system (AY003885.1) developed by (Kok et al., 1999), as well as the splicing-PCR approach as described in (Metzgar et al., 2004; Bacher et al., 2006). The comFEBC operon, involved in DNA binding and transport of the DNA through the periplasm (Averhoff and Graf, 2008), was knocked out to make a transformation-deficient phenotype of the ancestor (Com−; Bacher et al., 2006).
Experimental evolution
Three groups of six populations each were subjected to serial transfers in S2-LMBA. Two groups contained transformation-proficient populations (designated Com+) and the third group comprised transformation-deficient populations (designated Com−). To explore the potential benefits of heterologous DNA in the evolving populations, a DNA mix from Spitzbergen soil, pig feces, Acinetobacter baumannii AZR3410 (Ray et al., 2009) and Acinetobacter johnsonii LUH540 (Young and Ornston, 2001) was added to one of the transformation-proficient groups during the serial transfers (see Supplementary Information for description; group Com+DNA). The other transformation-proficient group (Com+) could only acquire DNA from within the population. To detect cross-contamination half of the populations (odd numbers) were founded from single colonies of the ancestor without kanamycin resistance (ADP1200Com+ or ADP1200Com−) and the other half (even numbers) from the otherwise isogenic ancestor with kanamycin resistance (ADP1200Com+KanR or ADP1200Com−KanR). All populations are listed in Table 1.
Serial transfers
All the 18 populations were propagated in 10 ml culture volumes in separate 100 ml Erlenmeyer flasks. Minimal medium S2-LMBA as described above was always used. Twice a week (every 84–90th hour) the cultures were diluted 1:100 in fresh medium and incubated at 37 °C and 120 rounds per minute. Populations belonging to group Com+DNA were supplemented with 5 μg exogenous DNA as described above every second passage (once a week). Samples (5 ml) were frozen down in glycerol (20%) after 35, 100 and 175 days of serial transfers. The medium supports early stationary phase ancestor density of ~1.0 × 109 cells ml−1, which is reached after about 10 h under the described growth conditions (SI Supplementary figure 4). The 100-fold re-growth between each transfer represents minimum 6.6 generations of binary fissions per transfer (13.2 generations per week). The number of generations is likely an underestimate since the estimates of re-growth did not take into account the population dynamics in the stationary and death phase.
Fitness measurements: head-to-head competitions and growth rates
After preconditioning (see Supplementary Information) the two competitors were mixed in equal volumes (75 μl of each) in a flask containing 9.9 ml of S2-LMBA with DNase (0.1 mg ml−1) to exclude HGT during the competitions (see Supplementary Information for description of control experiment). After mixing, the densities of each competitor (CFU ml−1) were determined by selective plating (LB and LBKan50 or LBSpec20) at T0, T24 and T90. Selection rate constants (r) were estimated for the entire 90-h passage as described in (Travisano and Lenski, 1996) with the modification of using ‘per passage’ as denominator instead of ‘per day’. Standard Darwinian relative fitness (w) measurements (Lenski et al., 1991) were estimated during the first 24 h of competitions as described in Starikova et al. (2012) except for the incubation at 37 °C and shaking at 120 rounds per minute. To quantify the potential differences in stationary and death phase between competing clones and/or populations we measured the difference in survival (su). This fitness parameter was calculated as:

Where NE(90), NE(24), NA(90) and NA(24) are the densities of the evolved and the ancestral strains after 24 and 90 h in competition, respectively. Competitions between evolved populations and ancestors with appropriate markers were done in at least six replicates after 175 days (50 passages) and in at least three replicates from time points of 35 and 100 days (10 and 29 passages, respectively). Evolved populations without kanamycin resistance were competed against ADP1200Com+KanR/ADP1200Com−KanR and evolved populations with kanamycin resistance were competed against control strains with spectinomycin resistance, ADP1200Com+SpecR/ADP1200Com−SpecR. Adjustments for marker effects on fitness were based on control competitions between the ancestral backgrounds with and without the selectable markers (Lind et al., 2010; Supplementary Table SI 2). See Supplementary Information for a description of the growth rate assays and competition experiments executed to test whether the adaptation to stationary phase was independent of the interactions in the growth phase.
Mutation and transformation frequencies
Mutation frequency experiments were performed in LB or S2-LMBA at 37 °C and 225 rounds per minute for 24 h selecting for spectinomycin resistance (10 μg ml−1) and calculated as described in (Björkholm et al., 2001). Transformation frequencies were estimated by diluting preconditioned cultures 1:5 in S2-LMBA in 96-well plates (total volume 1 ml) and incubating for 4 h (37 °C, 120 rounds per minute). The cultures were then supplemented with 4 μg genomic DNA (saturating concentration) from ADP1200Com− (trimethoprim resistant), incubated for 3 h and plated out on LB and LBTri250. DNA concentrations in A. baylyi grown in minimal medium can reach several μg ml−1 and is available as a substrate for natural transformation (Lorenz, Gerjets and Wackernagel, 1991). To verify that natural transformation occured under the experimental conditions we added 0.75 μg ml−1 DNA isolated from ADP1200Com+KanR to a freshly diluted ADP1200Com+ culture and incubated as described for the serial transfers above. Transformants were scored after 12 h on LBKan15. Controls for spontaneous kanamycin resistance mutations were included. The transformation frequency was calculated as CFU transformants/CFU recipients.
DNA isolation and sequencing
Chromosomal DNA was prepared according to the Genomic DNA Purification protocol (Qiagen, Hilden, Germany). The ancestral ADP1200Com+ and single isolates from the evolved Com− and Com+ groups were sequenced using Illumina (San Diego, CA, USA) sequencing. This was performed at the Danish National High-Throughput DNA Sequencing Centre as recently described (Overballe-Petersen, Harms et al., 2013; see Supplementary Information). Presence/absence of single-nucleotide polymorphisms (SNPs) was confirmed by Sanger sequencing using the BigDye 3.1 chemistry according to the manufacturer’s instructions (Thermo Fisher Scientific, Carlsbad, CA, USA). Sequencing primers are listed in Supplementary Table SI 1.
Statistical analysis
Initial data inspection revealed heteroscedasticity and dependency in data at the population level. Informed by these initial analyses we chose to analyze the data using a linear quantile mixed model adjusting for temporal pseudoreplication and repeated measures on the same population (for a more detailed description of the model, see Supplementary Information). All reported P-values and parameter estimates are based on the median values unless stated otherwise. All estimates, confidence intervals and their corresponding P-values for both the median and 75th percentile are listed in Supplementary Table SI 3. We tested whether there was an association between survival (su) and relative fitness (w) with a linear mixed model adjusting for repeated measures within populations. The relative fitness value is inherently a ratio and was linearized with a log transformation to make it comparable with the survival measure. Growth rate estimates were analyzed with a one way analysis of variance with a post hoc Tukey HSD for pairwise comparisons using the mean value for each measured population. We used R version 3.0.2 for statistical analysis (RDevelopementCoreTeam, 2013). The lqmm package version 1.04 (Geraci and Bottai, 2013) was used for the linear quantile mixed model and the test for an association between the relative fitness (log10 (w)) and survival (su) was performed in the nlme package version 3.1–111 (Pinheiro et al., 2013).
Results
Growth phase-dependent benefits of natural transformation in A. baylyi
Estimates of selection rate constants (r) between T0 and T90 (one complete passage; Travisano and Lenski, 1996) revealed that most of the populations had begun adapting to the experimental conditions after 35 days of serial passages (Figure 1a). After 100 days the Com− group had adapted significantly better to the environmental conditions than the Com+DNA group (r=6.5 and 3.7, respectively), but not when compared with the Com+ group (r=4.5; P<0.02 and P=0.17, respectively). However, after 175 days of serial passages no significant differences in the selection rate between the three groups were observed (all P-values >0.2), suggesting that there was no net benefit of natural transformation for A. baylyi under these experimental conditions (Figure 1a). All the fitness values on group and single population levels are listed in Supplementary Table SI 5, respectively. In a control experiment ADP1200Com+ generated on average 2655 transformants per milliliter yielding a frequency of 2.7 × 10-5 transformants per recipient (that is, natural transformation occurred below saturated DNA concentrations (Overballe-Petersen, Harms et al., 2013).
Comparison of groups Com−, Com+ and Com+DNA at different time points during serial transfers based on mixed competition experiments with their respective ancestors. (a) The selection rate constant (r) calculated between 0 and 90 h of competition. (b) Relative fitness (w) calculated between 0 and 24 h of competition. (c) Relative survival (su) calculated between 24 and 90 h of competition. Each circle represents one competition and each group is made up of six replicate populations. Note that all median values indicated by horizontal lines in the box plots are descriptive and not adjusted for uneven sample distribution and repeated measurements. * Indicates a significant difference (P <0.05) compared with the Com− group at each time point derived from the linear quantile mixed model. The relationships between individual competitions and replicate populations from each group is presented in Supplementary Table SI 5.
Relative fitness (w) was estimated for the first 24 h of the 90 h mixed culture competition experiments and included lag, log and early stationary phase, as described in (Lenski et al., 1991; see Supplementary Figure SI 4 for a representative mixed competition growth curve). We found that both Com+ and Com+DNA (w=1.81 and 1.65, respectively) performed significantly better when compared with the Com− group (w=1.28) after 175 days (P=0.003 and P=0.01, respectively; Figure 1b). The adaptation to the growth conditions observed in the Com− group can mainly be explained between 0 and 35 days with no additional increase in the relative fitness between 35 days (w=1.37) and 175 days (w=1.28; P>0.39 for both 100 and 175 days compared with 35 days). In both of the Com+ groups we observed a gradual increase in the relative fitness over time.
These results were further supported by monoculture growth rate experiments in which both Com+ groups displayed higher growth rates than the Com− group (Figure 2 and SI Supplementary Table 4). This difference was significant in the Com+ group (P=0.011) but not significant in the Com+DNA group (P=0.059). As expected, both Com+ and Com+DNA populations had higher growth rate than the ancestor (P=0.002 and P=0.011, respectively; Figure 2a). However, the mean growth rate of the Com− group had not significantly improved compared with the ancestor (P=0.86; Figure 2a) and the increased relative fitness for the Com− group could putatively be due to the earlier exit from lag phase, as indicated in Figure 2b.
Calculated growth rates (a) and growth curves including 95% confidence intervals (b) of groups Com−, Com+ and Com+DNA after 175 days of serial transfers and the ancestor. In (a) each circle represents one of six populations per group. The growth rate of Com+ is significantly different from Com− (P= 0.011), whereas Com+DNA is not different from that of Com− (P=0.059).
Survival (su) was calculated as the relative difference in log ratios between 24 and 90 h for each competitor in stationary/death phase and is expressed per hour. Com− populations displayed better survival in stationary phase (su=0.034), compared with the Com+ and Com+DNA treatments after 175 days (su=0.023 and 0.015, respectively; P=0.045 and P=0.008) (Figure 1c). All the evolved population groups displayed better survival when compared with their ancestors (Figure 1c). The Com+DNA group displayed a bi-modal distribution and was only significantly lower than the Com− group treatment at the 50th percentile, but not at the 75th percentile (P=0.008 and P=0.09, respectively; Figure 1c). To verify that the differences observed between Com+ and Com− populations relative to the respective ancestors reflected higher fitness, competitions between fitness representative (Starikova et al., 2012) evolved populations were performed. As expected, evolved Com+ populations outcompeted evolved Com− populations during the first 24 h of the competitions and this advantage was offset favoring the Com− populations between 24 and 90 h (Figure 3). We excluded Com+DNA populations from further analyses because no effect of exogenous DNA on the overall adaptation to the growth conditions was observed (Figure 1), and we thus focused on the directly comparable Com+ and Com− populations for the rest of the study.
Direct competitions between the evolved populations of Com+ (open symbols and dotted line) and Com− (filled symbols and solid line). Selected populations are fitness representatives from their respective groups. Com+ populations (open triangles/circles) were in competition with Com− populations (filled triangles/circles). Both competitions were done in triplicates and each symbol is the result of one competition. The lines represent the central tendency of Com− (solid) and Com+ (dotted) in the competitions. Estimated mean fitness parameters with 95% confidence intervals in these competitions: wCom+=1.45±0.34), suCom+=−0.013 (±0.006), rCom+=−0.22 /passage (±1.2).
Antagonistic pleiotropic effects
A plot of relative fitness (log10 (w)) as a function of survival (su) from all competitions in the Com+ and Com− groups revealed that Com+ populations typically displayed higher w and lower su than the Com− populations after 175 days (Figure 4). We found a significant negative association between log10 (w) and su (β1=−0.056, t=−2.78, P=0.007) supporting antagonistic pleiotropy (genetic trade-off) between ‘early’ and ‘late’ performance. This clear association was not found within the Com+ populations (β1 =−0.03, t=−1.34, P=0.18), most likely due to low statistical power (that is, too few independent measurements). However, the trend supports a negative association between two of the estimated fitness components, relative fitness (w) and survival in stationary/death phase (su). In a control experiment we moved the Com− mutation into an evolved Com+ strain and both strains were competed against the ancestral clones. This did not cause a significant change in either the relative fitness or survival (wmedian=1.34 and 1.34, W=21, P=0.7 (n=12) and sumedian=−0.0033 and 0.0026, W=7, P=0.18 (n=11; Wilcoxon's rank-sum test). Competitions between the Com+ and Com− ancestors also revealed no difference in the relative fitness or survival due to the com− mutation (wmedian= 0.96, W=14, P=0.13 (n=6) and sumedian= 0.0005, W=2, P=0.09 (n=5; Wilcoxon's rank-sum test).
Negative association between relative fitness (log10 (w)) and survival (su) in competitions from the Com− group (circles) and Com+ group (crosses). Competitions are between all the populations from groups Com− and Com+ serially passaged for 175 days and competed with the appropriate ancestor. (β1=−0.056, t=−2.78, P=0.007).
Adaptation to stationary and death phase is density and log phase independent
We let evolved populations and ancestor strains grow separately for the first 24 h of the competitions before they were mixed in equal volumes and incubated for the remaining competition without any new nutrients. When compared with the standard mixed competitions no difference in performance was observed (Supplementary Figure SI 1). Mixed culture competitions where evolved populations were inoculated in lower ratios (1:10 and 1:100) revealed invasion when rare, favoring density-independent adaptation (Supplementary Figure SI 2).
Ability to utilize added carbon sources and adaptation to stationary phase
Growth experiments revealed that the evolved populations had not acquired the ability to use arabinose or methyl-benzoate (data not shown). Most evolved populations displayed high performance in stationary phase resulting in drastic reductions in ancestor densities in mixed culture competitions (Figure 1c and Supplementary Figure SI 4). However, neither liquid- or agar-based experiments with cell-free supernatant from the stationary phase evolved populations showed any evidence of direct inhibition/killing of the ancestor strain ADP1200Com+ (data not shown).
Transformation frequencies in evolved populations
After 175 days of serial passages, transformation frequencies in three of the nine tested populations had decreased 100-fold compared with the ancestor when transformed with DNA from the ancestor strain ADP1200Com− (Supplementary Figure SI 3). For two of these populations this occurred gradually over time (data not shown). The 100-fold decrease in transformation frequencies was also verified using DNA from ADP1200Com+KanR (for two out of three populations) to rule out marker-specific effects (data not shown). Six populations displayed up to 10-fold decreased transformation frequencies compared with the ancestor, consistent with the decreased transformation frequencies reported in (Bacher et al., 2006). The reported reduced transformation frequencies may be due to individual lineages within the populations that lost the ability to take up exogenous DNA or a general loss of the ability throughout the whole population.
Mutation-fixation experiments
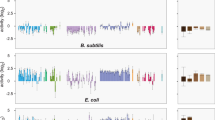
Mutation frequency estimates of the evolved clones revealed similar mutation frequencies (2 × 10−9–7 × 10−10) after 175 days. We sequenced two fitness representative clones, one from the Com+ group and one from the Com− group, as well as the ancestor ADP1200Com+ for comparison. We focused our analyses on SNPs. A total of nine SNPs were found, five in Com+ and four in Com− fitness representative clones (Table 2). After 175 days six of the nine SNPs were fixed or partially fixed in the population that the respective clone came from; two in the Com− isolate and four in the Com+ isolate (Table 2). To determine at what time during the 175 days the mutations appeared, and if/when they were fixed in the populations, we sequenced 10 randomly picked isolates at the preceding time points (35 and 100 days). The SNP fixation-dynamics is shown in Figure 5.
Mutation-fixation pattern of six mutations that were fixed (or partially fixed) after 175 days of serial transfers. Mutations are from Com+ (dotted) or Com− (solid) populations and listed in Table 2. Mutations: znuA-atp1 (filled squares), ACIAD3547 (open circles), ppc (open diamonds), ACIAD3006 (open triangles), ACIAD3007 (stars).
Parallel evolution
Of the nine SNPs (numbers of SNPs in line with a recent experimental evolution report in A. baylyi (Jezequel et al., 2013)) described in Table 2, one SNP in the znuA-atp1 intergenetic region occurred in both sequenced strains. The znuA and atp1 encode a putative transporter and ATPase, respectively. In addition, both strains had non-synonymous/frameshift mutations in the ACIAD3006-3007 loci. We further sequenced both loci in five randomly picked isolates from each of the remaining 10 populations of the Com+ and Com− groups. The znuA-atp1 mutation was present in 6 of the 12 populations in these two groups (Table 3). Extensive parallel evolution was also seen in the ACIAD3006-3007 loci demonstrated by the presence of various SNPs in 11 of the 12 evolved populations in both the Com+ and Com− groups after 175 days (Table 3). ACIAD3006 and ACIAD3007 coding sequences are putative LrgA- and LrgB-like proteins found in several bacteria including Acinetobacter spp. (Bayles, 2007). LrgAB are membrane proteins in a regulatory network that seem to be involved in regulated cell death and lysis (Bayles, 2007) and mutations in these genes could be stationary/phase-specific adaptations.
Discussion
In this report we investigated the effect of natural transformation on adaptation of A. baylyi to a novel stressful environment. Stress included growth above the optimum temperature (Fulsundar et al., 2014), growth in minimal medium and extended periods in stationary and death phase between serial dilutions (90 h passages). We demonstrate for the first time growth phase-dependent costs and benefits of natural transformation in bacteria. Three lines of evidence support that natural selection favored both the Com+ and Com+DNA populations in the first 24 h of the passages including lag, log and early stationary phase. First, estimates of relative fitness in mixed culture competitions between evolved populations and the respective ancestors for the first 24 h of the passage revealed that both transformation-proficient groups adapted significantly better to the growth phase in a novel environment than the Com− populations. Secondly, Com+ and Com+DNA populations displayed higher growth rates in monocultures when compared with the Com− populations. Finally, evolved Com+ populations outcompeted evolved Com− populations in the first 24 h in head-to-head competition experiments.
These results contrast the conclusions in a previous report where Com+ and Com− A. baylyi populations were subjected to a serial transfer regime similar to the first 24 h described in this report (Bacher et al., 2006). It is however important to emphasize that we applied both stressful conditions as well as less severe bottlenecks when compared to (Bacher et al., 2006). This approach reduced the probability for stochastic loss of rare beneficial mutations. Two other studies investigated the evolutionary benefits of natural transformation during active growth (Baltrus et al., 2007; Engelmoer et al., 2013). Our results are consistent with Baltrus et al, demonstrating higher adaptation rates for Com+ populations in a novel environment where arguably stress was introduced in the form of freeze–thaw cycles between daily serial passages in H. pylori (Baltrus et al., 2007). In a recent report competent S. pneumoniae populations performed worse than the competence-deficient populations after 1000 generations of growth in chemostats under benign conditions, but this cost of competence was offset by the introduction of mild stress in the system (Engelmoer et al., 2013). Our results introduce additional complexity to this context-dependent benefit of competence (Engelmoer et al., 2013).
We show that the benefit of competence during log and early stationary phase was offset during subsequent stationary/death phase in mixed culture competitions for 90 h (one complete passage). Thus, no net effects of natural transformation on adaptation rates were demonstrated. Apparently, our choice of 90 h passages balanced the ‘early’ positive and ‘late’ negative effects on the overall adaptation rates in Com+ and Com− populations. Arguably, shorter passages could lead to an overall benefit of natural transformation, whereas longer passages could lead to a net negative effect on the overall adaptation to the experimental conditions relative to the Com− populations. It could be hypothesized that heterogeneous transformation abilities would emerge in the Com+ populations if natural selection favored natural transformation ‘early’ but not ‘late’ in the passages. This is supported by the observed reduced transformation ability in most populations (Supplementary Figure SI 3), suggesting that Com+ populations adjusted the transformation ability to balance the cost of transformation in death phase with the benefit experienced in log, and early stationary phase. This interpretation is consistent with a recent report demonstrating variable transformation frequencies in S. pneumoniae isolates (Evans and Rozen, 2013).
We further hypothesized that fitness trade-offs existed between performing well ‘early’ (lag, log and early stationary phase) and worse ‘late’ (stationary and death phase). We demonstrate a significant negative association between relative fitness (w) during growth and survival (su) in the Com+ and Com− groups, strongly suggesting antagonistic pleiotropy between the traits responsible for ‘early’ and ‘late’ performance in the 90 h passage. Previous reports on antagonistic pleiotropy in bacteria demonstrate a role in ecological specialization (Cooper and Lenski, 2000; Cooper et al., 2001; MacLean et al., 2004; Ostrowski et al., 2005; Maharjan et al., 2012). Our data suggest a previously unrecognized interplay between antagonistic pleiotropy and recombination mediated by natural transformation in bacteria. It is tempting to speculate that the concerted action between recombination (speeds up adaptation to active growth) and subsequent fitness trade-offs (reduced performance in stationary/death phase) caused by antagonistic pleiotropy may fuel ecological specialization.
A possible explanation for why the demonstrated fitness trade-off evolved in Com+ populations could be that the DNA available for uptake in log phase (where A. baylyi is competent for natural transformation) primarily originated from cells dying in death phase in the previous passage. Accumulation of exogenous DNA in batch cultures, a feature commonly observed in bacteria (Lorenz and Wackernagel, 1994), has been shown for A. baylyi grown in minimal medium with peak concentrations reached in stationary/death phase (approximately 5 μg ml−1; Lorenz et al., 1991). We assume that DNA from cells with a competitive disadvantage in late stationary/death phase (Redfield, 1988) constitutes the majority of DNA available for uptake. Such an uneven distribution of growth phase-specific adaptive mutations could in theory affect the mutation patterns brought together by natural transformation and had a role in the evolution of fitness trade-offs. In this context it is possible that Com+ and Com− populations were forced to undertake different evolutionary paths. Different evolutionary trajectories between Com+ and Com− populations were also indicated by different SNP-fixation patterns where four and two SNPs fixed in the Com+ and Com− populations, respectively. Two of these mutations and/or mutation loci were identified extensively in several evolving Com+ and Com− populations and such parallel evolution strongly suggest the identification of adaptive mutations (Woods et al., 2006) and support adaptive convergence (Christin et al., 2010; Tenaillon et al., 2012). Although we cannot exclude the accumulation due to successive mutations, the two fixed mutations uniquely found combined in the Com+ lineages could indicate that natural transformation may, under certain circumstances, constrain adaptive convergence. We acknowledge that the small number of mutations identified in the strains subjected to whole-genome sequencing could suggest that the observed differences between Com+ and Com− populations were not directly related to the ability to take up exogenous DNA by natural transformation. However, we emphasize that the adaptive mutations originated from single isolates in mixed populations evolved for 6 months in serial transfer cultures. Moreover, we moved the com− mutation into an evolved Com+ strain and demonstrated no effect on any fitness component measured in this report, strongly suggesting that the genetic manipulation did not affect the evolutionary trajectories taken by the different populations.
Our data did not provide support for a role of interspecies HGT events for rapid adaptation by natural transformation to specific environmental stress conditions. The addition of heterogeneous DNA to the Com+DNA populations did not increase adaptation rates to this novel environment. A potential effect on the increased standing genetic diversity in these populations can, however, not be ruled out as indicated by the bi-modal distribution of survival (su) in stationary/death phase. This would be consistent with the early work on the evolution and maintenance of recombination in Escherichia coli where it was demonstrated that frequent introduction of DNA ‘from without’ did not increase adaptation rates but yielded increased genetic diversity (Souza et al., 1997).
Taken together, the maintenance and evolution of natural transformation depends on a complex interplay between ecological and physiological factors, as recently shown theoretically (Moradigaravand and Engelstädter, 2013; Engelstädter and Moradigaravand, 2014). In conclusion, our data shed light on conditions where the Fisher–Muller model (Fisher, 1930; Muller, 1932; Vos, 2009) is insufficient to account for the maintenance of natural transformation. We report that natural transformation in A. baylyi during adaptation to a stressful laboratory environment for ~6 months is beneficial when resources are abundant/available. However, naturally transformable populations experienced fitness trade-off(s) between high performances during active growth and reduced competitiveness in stationary/death phase. This in turn led to different preferred evolutionary trajectories on adaptation to the novel environment between recombining and non-recombining genotypes.
References
Averhoff B, Graf I . (2008). The Natural Transformation System of Acinetobacter baylyi ADP1: A Unique DNA Transport Machinery. In: Gerischer U (ed). Acinetobacter—Molecular Biology. Caister Academic Press: Norfolk, UK, pp 119–139.
Bacher JM, Metzgar D, de Crécy-Lagard V . (2006). Rapid evolution of diminished transformability in Acinetobacter baylyi. J Bacteriol 188: 8534–8542.
Baltrus D . (2013). Exploring the costs of horizontal gene transfer. Trends Ecol Evol 28: 489–495.
Baltrus D, Guillemin K, Phillips P . (2007). Natural transformation increases the rate of adaptation in the human pathogen Helicobacter pylori. Evolution 62: 10.
Barbe V, Vallenet D, Fonknechten N, Kreimeyer A, Oztas S, Labarre L et al. (2004). Unique features revealed by the genome sequence of Acinetobacter sp. ADP1, a versatile and naturally transformation competent bacterium. Nucleic Acids Res 32: 5766–5779.
Bayles KW . (2007). The biological role of death and lysis in biofilm development. Nat Rev Microbiol 5: 721–726.
Bernstein H, Hopf F, Michod R . (1987). The molecular basis of the evolution of sex. Adv Genet 24: 323–370.
Björkholm B, Sjölund M, Falk PG, Berg OG, Engstrand L, Andersson DI . (2001). Mutation frequency and biological cost of antibiotic resistance in Helicobacter pylori. Proc Natl Acad Sci 98: 14607–14612.
Christin P-A, Weinreich DM, Besnard G . (2010). Causes and evolutionary significance of genetic convergence. Trends Genet 26: 400–405.
Cooper VS, Bennett AF, Lenski RE . (2001). Evolution of thermal dependence of growth rate of Escherichia coli populations during 20 000 generations in a constant environment. Evolution 55: 889–896.
Cooper VS, Lenski RE . (2000). The population genetics of ecological specialization in evolving Escherichia coli populations. Nature 407: 736–739.
Didelot X, Maiden MC . (2010). Impact of recombination on bacterial evolution. Trends Microbiol 18: 315–322.
Dubnau D . (1999). DNA uptake in bacteria. Annu Rev Microbiol 53: 217–244.
Engelmoer DJ, Donaldson I, Rozen DE . (2013). Conservative sex and the benefits of transformation in Streptococcus pneumoniae. PLoS Pathog 9: e1003758.
Engelstädter J, Moradigaravand D . (2014). Adaptation through genetic time travel? Fluctuating selection can drive the evolution of bacterial transformation. Proc R Soc B 281: 20132609.
Evans BA, Rozen DE . (2013). Significant variation in transformation frequency in Streptococcus pneumoniae. ISME J 7: 791–799.
Fisher R . (1930) The genetical theory of natural selection. Clarendon Press: Oxford, UK.
Fulsundar S, Harms K, Flaten GE, Johnsen PJ, Chopade BA, Nielsen KM . (2014). Gene transfer potential of outer membrane vesicles of Acinetobacter baylyi and effects of stress on vesiculation. Appl Environ Microbiol 80: 3469–3483.
Geraci M, Bottai M . (2013). Linear quantile mixed models. Stat Comput 8: 140–154.
Gray JC, Goddard MR . (2012). Sex enhances adaptation by unlinking beneficial from detrimental mutations in experimental yeast populations. BMC Evol Biol 12: 43.
Hoelzer MA, Michod R . (1991). DNA repair and the evolution of transformation in Bacillus subtilis. III. Sex with damaged DNA. Genetics 128: 215–223.
Jezequel N, Lagomarsino MC, Heslot F, Thomen P . (2013). Long-term diversity and genome adaptation of Acinetobacter baylyi in a minimal-medium chemostat. Genome Biol Evol 5: 87–97.
Johnsborg O, Eldholm V, Håvarstein LS . (2007). Natural genetic transformation: prevalence, mechanisms and function. Res Microbiol 158: 767–778.
Johnsen PJ, Dubnau D, Levin BR . (2009). Episodic selection and the maintenance of competence and natural transformation in Bacillus subtilis. Genetics 181: 1521–1533.
Juni E . (1974). Simple genetic transformation assay for rapid diagnosis of Moraxella osloensis. Appl Microbiol 27: 16–24.
Kok RG, Young DM, Ornston LN . (1999). Phenotypic expression of PCR-generated random mutations in a Pseudomonas putida gene after its introduction into an Acinetobacter chromosome by natural transformation. Appl Environ Microbiol 65: t1675–t1680.
Lenski RE, Rose MR, Simpson SC, Tadler SC . (1991). Long-term experimental evolution in Escherichia coli. I. Adaptation and divergence during 2,000 generations. Am Nat 138: 1315–1341.
Levin BR, Bergstrom CT . (2000). Bacteria are different: observations, interpretations, speculations, and opinions about the mechanisms of adaptive evolution in prokaryotes. Proc Natl Acad Sci USA 97: 6981–6985.
Levin BR, Cornejo OE . (2009). The population and evolutionary dynamics of homologous gene recombination in bacteria. PLoS Genet 5: e1000601.
Lind PA, Tobin C, Berg OG, Kurland CG, Andersson DI . (2010). Compensatory gene amplification restores fitness after inter-species gene replacements. Mol Microbiol 75: 1078–1089.
Lorenz MG, Gerjets D, Wackernagel W . (1991). Release of transforming plasmid and chromosomal DNA from two cultured soil bacteria. Arch Microbiol 156: 319–326.
Lorenz MG, Wackernagel W . (1994). Bacterial gene transfer by natural genetic transformation in the environment. Microbiol Rev 58: 563.
Macfadyen LP, Chen D, Vo HC, Liao D, Sinotte R, Redfield RJ . (2001). Competence development by Haemophilus influenzae is regulated by the availability of nucleic acid precursors. Mol Microbiol 40: 700–707.
MacLean RC, Bell G, Rainey PB . (2004). The evolution of a pleiotropic fitness tradeoff in Pseudomonas fluorescens. Proc Natl Acad Sci USA 101: 8072–8077.
Maharjan RP, Ferenci T, Reeves PR, Li Y, Liu B, Wang L . (2012). The multiplicity of divergence mechanisms in a single evolving population. Genome Biol 13: R41.
Metzgar D, Bacher JM, Pezo V, Reader J, Döring V, Schimmel P et al. (2004). Acinetobacter sp. ADP1: an ideal model organism for genetic analysis and genome engineering. Nucleic Acids Res 32: 5780–5790.
Moradigaravand D, Engelstädter J . (2013). The evolution of natural competence: disentangling costs and benefits of sex in bacteria. Am Nat 182: E112–E126.
Muller H . (1932). Some genetic aspects of sex. Am Nat 66: 20.
Narra HP, Ochman H . (2006). Of what use is sex to bacteria? Curr Biol 16: R705–R710.
Nielsen KM, Van Weerelt M, Berg TN, Bones AM, Hagler AN, Van Elsas J . (1997). Natural transformation and availability of transforming DNA to Acinetobacter calcoaceticus in soil microcosms. Appl Environ Microbiol 63: 1945–1952.
Ochman H, Lawrence JG, Groisman EA . (2000). Lateral gene transfer and the nature of bacterial innovation. Nature 405: 299–304.
Ostrowski EA, Rozen DE, Lenski RE . (2005). Pleiotropic effects of beneficial mutations in Escherichia coli. Evolution 59: 2343–2352.
Overballe-Petersen S, Harms K, Orlando LA, Mayar JVM, Rasmussen S, Dahl TW et al. (2013). Bacterial natural transformation by highly fragmented and damaged DNA. Proc Natl Acad Sci USA 110: 19860–19865.
Pinheiro J, Bates D, DebRoy S, Sarkar D, Team RDC . (2013), nlme: Linear and Nonlinear Mixed Effects Models. R package version 3.1-111.
Ray JL, Harms K, Wikmark OG, Starikova I, Johnsen PJ, Nielsen KM . (2009). Sexual Isolation in Acinetobacter baylyi Is locus-specific and varies 10 000-fold over the genome. Genetics 182: 1165–1181.
RDevelopementCoreTeam. (2013) R: A language and environment for statistical computing. R Foundation for Statistical Computing: Vienna, Austria, http://www.R-project.com.
Redfield RJ . (1988). Evolution of bacterial transformation: is sex with dead cells ever better than no sex at all? Genetics 119: 213–221.
Redfield RJ . (1993a). Genes for Breakfast: The Have-Your-Cake and-Eat-lt-Too of Bacterial Transformation. J Hered 84: 400–404.
Redfield RJ . (2001). Do bacteria have sex? Nat Rev Genet 2: 634–639.
Sambrook J, Fritsch E, Maniatis T . (1989) Molecular Cloning: a Laboratory Manual2 edn.Cold Spring Harbor Laboratory Press: Cold Spring Harbor NY, USA.
Souza V, Turner P, Lenski R . (1997). Long-term experimental evolution in Escherichia coli. V. Effects of recombination with immigrant genotypes on the rate of bacterial evolution. J Evol Biol 10: 743–769.
Starikova I, Harms K, Haugen P, Lunde TT, Primicerio R, Samuelsen Ø et al. (2012). A trade-off between the fitness cost of functional integrases and long-term stability of integrons. PLoS Pathog 8: e1003043.
Stewart GJ, Carlson CA . (1986). The biology of natural transformation. Annu Rev Microbiol 40: 211–231.
Tenaillon O, Rodríguez-Verdugo A, Gaut RL, McDonald P, Bennett AF, Long AD et al. (2012). The molecular diversity of adaptive convergence. Science 335: 457–461.
Thomas CM, Nielsen KM . (2005). Mechanisms of, and barriers to, horizontal gene transfer between bacteria. Nat Rev Microbiol 3: 711–721.
Travisano M, Lenski RE . (1996). Long-term experimental evolution in Escherichia coli. IV. Targets of selection and the specificity of adaptation. Genetics 143: 15–26.
Vos M . (2009). Why do bacteria engange in homologous recombination? Trends Microbiol 17: 7.
Woods R, Schneider D, Winkworth CL, Riley MA, Lenski RE . (2006). Tests of parallel molecular evolution in a long-term experiment with Escherichia coli. Proc Natl Acad Sci USA 103: 9107–9112.
Young DM, Ornston LN . (2001). Functions of the mismatch repair gene mutS from Acinetobacter sp strain ADP1. J Bacteriol 183: 6822–6831.
Acknowledgements
We thank Robert Hanes and Anne-Hilde Conradi for technical assistance as well as Klaus Harms, Pål Haugen and Bruce R. Levin for many useful discussions. This work was supported by the Tromsø Research Foundation (grant number A5870 to P.J.J.), UiT The Arctic University of Norway (awarded P.J.J. and K.M.N.) and the Research Council of Norway (grant number 204263/F20 to P.J.J.). We thank the editors and five anonymous referees for insightful and valuable comments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Utnes, A., Sørum, V., Hülter, N. et al. Growth phase-specific evolutionary benefits of natural transformation in Acinetobacter baylyi. ISME J 9, 2221–2231 (2015). https://doi.org/10.1038/ismej.2015.35
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.35
This article is cited by
-
Distribution of fitness effects of cross-species transformation reveals potential for fast adaptive evolution
The ISME Journal (2023)
-
Antidepressants promote the spread of extracellular antibiotic resistance genes via transformation
ISME Communications (2022)
-
Artificial sweeteners stimulate horizontal transfer of extracellular antibiotic resistance genes through natural transformation
The ISME Journal (2022)
-
Non-antibiotic pharmaceuticals enhance the transmission of exogenous antibiotic resistance genes through bacterial transformation
The ISME Journal (2020)
-
Costs and benefits of natural transformation in Acinetobacter baylyi
BMC Microbiology (2017)