Abstract
Different lifestyles, ranging from a saprophyte to a pathogen, have been reported in bacteria of one species. Here, we performed genome-wide survey of the ecological adaptation in four Burkholderia seminalis strains, distinguished by their origin as part of the saprophytic microbial community of soil or water but also including human and plant pathogens. The results indicated that each strain is separated from the others by increased fitness in medium simulating its original niche corresponding to the difference between strains in metabolic capacities. Furthermore, strain-specific metabolism and niche survival was generally linked with genomic variants and niche-dependent differential expression of the corresponding genes. In particular, the importance of iron, trehalose and d-arabitol utilization was highlighted by the involvement of DNA-methylation and horizontal gene transfer in niche-adapted regulation of the corresponding operons based on the integrated analysis of our multi-omics data. Overall, our results provided insights of niche-specific adaptation in bacteria.
Similar content being viewed by others
Main
Natural habitats, in particular soil and water, have been widely recognized as reservoirs for a variety of plant and human pathogenic bacteria (Rutala and Weber, 1997; Berg et al., 2005). A shift from a natural habitant to a host-associated pathogen lifestyle has been reported in a variety of bacteria (Morris et al., 2007; Mahenthiralingam et al., 2008; Vial et al., 2011; Lee et al., 2013). However, in most cases these natural habitants showed no virulence in plant and human hosts.
This ecological phenomenon was understood in this study by examing the mechanism behind the niche adaptation of four B. seminalis strains DSM 23518 (= LMG 24067T), 0901, S9 and R456, distinguished by their ecological origins as part of the saprophytic microbial community of soil or water but also including human and plant pathogens. This study clearly revealed the preference of each strain for specific niche environment based on both phenotypic characterization and growth in various niches. Indeed, the host plant apricot was specifically infected by strain 0901, while the cell number of strain S9 in water medium, strain R456 in soil extracted medium and strain DSM 23518 in CF medium is significantly higher (P<0.01, ANOVO test) than that of the other strains under the same conditions (Figure 1). Furthermore, adaptation of each strain to specific niche was found to be highly associated with its unique metabolic capacity, as shown by the Biolog test results (Table 1 and Supplementary Table S1). For example, infection of human by strain DSM 23518 was justified by its growth on 1.0% NaCl, similar to physiological saline. Utilization of d-trehalose and l-fucose by strain 0901 was justified by their existence in plant surfaces (Andersson et al., 2001), while utilization of d-mannitol by strain S9 may be due to its high abundance in algae that normally grow in water (Iwamoto and Shiraiwa, 2005). The growth of strain R456 in pH 5.0 coordinates with the result of Stopnisek et al. (2014), who found that acid tolerance accounts for the biogeographical distribution of soil Burkholderia populations.
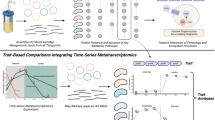
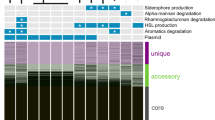
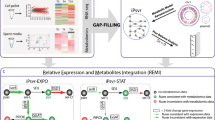
Multi-omics analysis of the four niches-adapted strains 0901, DSM 23518, R456 and S9 of B. seminalis originally isolated from diseased apricot, CF patient’s sputum, rice rhizospheric soil and Westlake water, respectively. (a) Phenotypic characterization of the four niches-adapted strains. E0: SEM observation of cell morphology; E1: growth in water medium. The arrow represents the preference of water strain S9 for growth in water medium; E2: antagonistic to soil-borne pathogen Rhizoctonia solani. The arrow represents growth inhibition by soil strain R456; E3: virulence in an alfalfa infection model. The arrow represents disease symptom caused by human strain DSM 23518; E4: pathogenicity to apricot. The arrow represents infection of apricot by plant strain 0901. (b) Growth of the four niches-adapted strains in CF, SE and water medium. Inoculation of apricot with the four strains is not included since only strain 0901 has pathogenicity to apricot. Growth is presented as the OD600 for three biological replicates vs time. (c) Genomic variation among the four niches-adapted strains determined by sequencing and comparing the complete genomes with sizes of 7.70-8.39 Mb. (d) DNA methylation involved in expression regulation. The vertical bar represents the Log2 RPKM value generated from the RNA-Seq data with a high degree of reliability as shown in Supplementary Figures S6-S7 and Supplementary Table S8. Statistical analysis of the genes with RPKM values >1 reveals the differential expression (marked as blank triangle) of virulence-associated operon bkdA1-bkdA2-bkdB-lpdV under CF condition of strain DSM 23518. The black vertical lines among the gene cluster reveal 7 and 9 DNA methylation sites on bkdA2 and bkdB of strain DSM 23518, respectively, while no methylated sites were found for the two genes in the other three strains.
The integrated analysis of multi-omics data (Supplementary Figures S1-S4, Supplementary Table S2-S4) demonstrates the potential association of the ability to adapt to specific niche with genomic variants, evidenced by the identification of genomic islands (GIs) with a huge number of diversity, and niche-dependent differential gene expression of the four niches-adapted strains. The possibility of species misidentification was ruled out by using both MLSA-based phylogeny (Supplementary Figure S5a) and ANI-based method (Supplementary Table S5), although strain S9 clustered more closely with B. cenocepacia rather than with the other three strains based on phylogenetic analysis of the complete genome sequences (Supplementary Figure S5b). Furthermore, functional enrichment analysis of the differentially expressed genes (DEGs) revealed the importance of the enriched COGs in the adaptation of each strain to specific niche, which was illuminated by the association of the 6 enriched COGs of strain DSM 23518 with bacterial growth, metabolism and virulence (Supplementary Table S6). Among these DEGs involved in strain-specific virulence, niche survival and metabolism, the contribution of horizontal gene transfer to niche-specific adaptation was validated by the result that gene expression pattern clustered by bacterial strains other than niche conditions, while the contribution of DNA methylation was validated by the result that 37, 24, 2 and 5 enriched methylated genes were differentially expressed under corresponding niche conditions of strains S9, DSM 23518, R456 and 0901, respectively. In order to rule out the possibility of random events, more attentions were paid on the differentially expressed operons. Representative examples of each strain were shown as follows (Supplementary Table S7). The growth of strain DSM 23518 in CF medium containing iron-enriched egg yolk and its infection to human was validated by differential expression of Fec-operon and virulence-associated bkdA1-bkdA2-bkdB-lpdV operon (Finelli et al., 2003; Sriramulu et al., 2005; Waite et al., 2006; Drevinek et al., 2008; Patell et al., 2010), while expression of transcriptional regulator encoding gene on Fec-operon and bkdA2 and bkdB on bkdA1-bkdA2-bkdB-lpdV operon was epigenetically regulated by DNA methylation (Figure 1). Furthermore, the infection of strain 0901 to apricot was validated by differential expression of many virulence-associated operons that located in GIs, such as spc ribosomal protein operon (Shiba et al., 1984), while the utilization of trehalose by strain 0901 was justified by the involvement of DNA methylation in the regulation for differential expression of gene cluster associated with trehalose uptake and utilization (d'Enfert and Fontaine, 1997; Foster et al., 2003). The growth of strain R456 in pH 5 was justified by differential expression of operon associated with sulfur oxidation, characterized by a decrease of the pH, while the expression of BsemR456423 encoding d-arabinitol 4-dehydrogenase involved in d-arabitol utilization (Lin, 1961) regulated by DNA methylation justified the result that d-arabitol was unable to be utilized by strain R456, but utilized by the other three strains. In agreement with the result of strain DSM 23518, differential expression was also found in iron uptake operon that located in GIs from strain R456. In addition, the essentiality of iron uptake for the growth of strain S9 was validated by differential expression of orbSHGCDFB operon, which has been found to be important for uptake of iron in the Bcc bacteria (Sokol et al., 2000; Agnoli et al., 2006). The importance of iron transport and uptake for the niche adaptation of different strains in this study was justified by the result of Mathew et al. (2014), who found that the Bcc bacteria have evolved a wide range of strategies to overcome iron shortage and to ensure sufficient uptake since iron is a limiting nutrient in various inhabits, while mutants of Bcc bacteria, impairing in iron uptake, failed to grow under iron-limited conditions.
In conclusion, this result provides new insights into the ecological adaptation process in bacteria, highlighting the importance of strain-specific metabolism and niche survival in adaptation of each strain to specific niche. In particular, the importance of iron, trehalose and d-arabitol utilization during adaptation to specific niche was emphasized by the involvement of DNA methylation and GIs in niche-adapted regulation of the corresponding operons.
Accession numbers
The project accession i.d.’s for raw genomic sequences of strains S9, R456, 0901 and DSM 23518 are PRJNA76133, PRJNA76131, PRJNA63525 and PRJNA76129, respectively. All the transcriptome data were deposited under the project accession id PRJNA235910.
References
Agnoli K, Lowe CA, Farmer KL, Husnain SI, Thomas MS . (2006). The ornibactin biosynthesis and transport genes of Burkholderia cenocepacia are regulated by an extracytoplasmic function σ factor which is a part of the Fur regulon. J Bacteriol 188: 3631–3644.
Andersson U, Levander F, Rådström P . (2001). Trehalose-6-phosphate phosphorylase is part of a novel metabolic pathway for trehalose utilization in Lactococcus lactis. J Biol Chem 276: 42707–42713.
Berg G, Eberl L, Hartmann A . (2005). The rhizosphere as a reservoir for opportunistic human pathogenic bacteria. Environ Microbiol 7: 1673–1685.
d'Enfert C, Fontaine T . (1997). Molecular characterization of the Aspergillus nidulans treA gene encoding an acid trehalase required for growth on trehalose. Mol Microbiol 24: 203–216.
Drevinek P, Holden MTG, Ge Z, Jones AM, Ketchell I, Gill RT et al. (2008). Gene expression changes linked to antimicrobial resistance, oxidative stress, iron depletion and retained motility are observed when Burkholderia cenocepacia grows in cystic fibrosis sputum. BMC Infect Dis 8: 121.
Finelli A, Gallant CV, Jarvi K, Burrows LL . (2003). Use of in-biofilm expression technology to identify genes involved in Pseudomonas aeruginosa biofilm development. J Bacteriol 185: 2700–2710.
Foster AJ, Jenkinson JM, Talbot NJ . (2003). Trehalose synthesis and metabolism are required at different stages of plant infection by Magnaporthe grisea. EMBO J 22: 225–235.
Iwamoto K, Shiraiwa Y . (2005). Salt-regulated mannitol metabolism in algae. Mar Biotechnol 7: 407–415.
Lee Y-J, van Nostrand JD, Tu Q, Lu Z, Cheng L, Yuan T et al. (2013). The PathoChip, a functional gene array for assessing pathogenic properties of diverse microbial communities. ISME J 7: 1974–1984.
Lin ECC . (1961). An inducible D-arabitol dehydrogenase from Aerobacter aerogenes. J Biol Chem 236: 31–36.
Mahenthiralingam E, Baldwin A, Dowson CG . (2008). Burkholderia cepacia complex bacteria: opportunistic pathogens with important natural biology. J Appl Microbiol 104: 1539–1551.
Mathew A, Eberl L, Carlier AL . (2014). A novel siderophore‐independent strategy of iron uptake in the genus Burkholderia. Mol Microbiol 91: 805–820.
Morris CE, Kinkel LL, Xiao K, Prior P, Sands DC . (2007). Surprising niche for the plant pathogen Pseudomonas syringae. Infect Genet Evol 7: 84–92.
Patell S, Gu M, Davenport P, Givskov M, Waite RD, Welch M . (2010). Comparative microarray analysis reveals that the core biofilm‐associated transcriptome of Pseudomonas aeruginosa comprises relatively few genes. Environ Microbiol Rep 2: 440–448.
Rutala WA, Weber DJ . (1997). Water as a reservoir of nosocomial pathogens. Infect Control Hosp Epidemiol 18: 609–616.
Shiba K, Ito K, Yura T, Cerretti DP . (1984). A defined mutation in the protein export gene within the spc ribosomal protein operon of Escherichia coli: isolation and characterization of a new temperature-sensitive secY mutant. EMBO J 3: 631–635.
Sokol PA, Darling P, Lewenza S, Corbett CR, Kooi CD . (2000). Identification of a siderophore receptor required for ferric ornibactin uptake in Burkholderia cepacia. Infect Immun 68: 6554–6560.
Sriramulu DD, Nimtz M, Romling U . (2005). Proteome analysis reveals adaptation of Pseudomonas aeruginosa to the cystic fibrosis lung environment. Proteomics 5: 3712–3721.
Stopnisek N, Bodenhausen N, Frey B, Fierer N, Eberl L, Weisskopf L . (2014). Genus‐wide acid tolerance accounts for the biogeographical distribution of soil Burkholderia populations. Environ Microbiol 16: 1503–1512.
Vial L, Chapalain A, Groleau MC, Déziel E . (2011). The various lifestyles of the Burkholderia cepacia complex species: a tribute to adaptation. Environ Microbiol 13: 1–12.
Waite RD, Paccanaro A, Papakonstantinopoulou A, Hurst JM, Saqi M, Littler E et al. (2006). Clustering of Pseudomonas aeruginosa transcriptomes from planktonic cultures, developing and mature biofilms reveals distinct expression profiles. BMC Genomics 7: 162.
Acknowledgements
This work was supported by the Zhejiang Provincial Natural Science Foundation of China (R13C140001), the National Natural Science Foundation of China (31571971, 31371904 and 31200003), the Zhejiang Provincial Project (2014C32010), the Fundamental Research Funds for the Central Universities, the Agricultural Ministry of China (nyhyzx 201303015, 201003029 and 201003066) and the Key Subject Construction Program of Zhejiang for Modern Agricultural Biotechnology and Crop Disease Control (2010DS700124-KF1101, -KF1203, -KF1309 and -KF1410). We acknowledge the Hangzhou Woosen Biotechnology Co., Ltd, Zhejiang, China for the genome and transcriptome sequencing.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Zhu, B., Ibrahim, M., Cui, Z. et al. Multi-omics analysis of niche specificity provides new insights into ecological adaptation in bacteria. ISME J 10, 2072–2075 (2016). https://doi.org/10.1038/ismej.2015.251
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.251
This article is cited by
-
Microbiome-based enrichment pattern mining has enabled a deeper understanding of the biome–species–function relationship
Communications Biology (2023)
-
Molecular characterization and in-depth genome analysis of Enterobacter sp. S-16
Functional & Integrative Genomics (2023)
-
In-depth genome analysis of Bacillus sp. BH32, a salt stress-tolerant endophyte obtained from a halophyte in a semiarid region
Applied Microbiology and Biotechnology (2022)
-
Genome mining of secondary metabolites from a marine-derived Aspergillus terreus B12
Archives of Microbiology (2021)
-
Pan-genome diversification and recombination in Cronobacter sakazakii, an opportunistic pathogen in neonates, and insights to its xerotolerant lifestyle
BMC Microbiology (2019)