Abstract
Different microbial cell types typically specialize at performing different metabolic processes. A canonical example is substrate cross-feeding, where one cell type consumes a primary substrate into an intermediate and another cell type consumes the intermediate. While substrate cross-feeding is widely observed, its consequences on ecosystem processes is often unclear. How does substrate cross-feeding affect the rate or extent of substrate consumption? We hypothesized that substrate cross-feeding eliminates competition between different enzymes and reduces the accumulation of growth-inhibiting intermediates, thus accelerating substrate consumption. We tested this hypothesis using isogenic mutants of the bacterium Pseudomonas stutzeri that either completely consume nitrate to dinitrogen gas or cross-feed the intermediate nitrite. We demonstrate that nitrite cross-feeding eliminates inter-enzyme competition and, in turn, reduces nitrite accumulation. We further demonstrate that nitrite cross-feeding accelerates substrate consumption, but only when nitrite has growth-inhibiting effects. Knowledge about inter-enzyme competition and the inhibitory effects of intermediates could therefore be important for deciding how to best segregate different metabolic processes into different microbial cell types to optimize a desired biotransformation.
Similar content being viewed by others
Introduction
Different metabolic processes are often segregated into different microbial cell types (Costa et al., 2006; Johnson et al., 2012; Zelezniak et al., 2015). A canonical example is substrate cross-feeding. Substrate cross-feeding occurs when one cell type partially consumes a primary substrate into a metabolic intermediate and another cell type then further consumes the intermediate. Substrate cross-feeding controls numerous environmentally and economically important microbial processes, including the mineralization of organic carbon (Schink, 1997; McInerney et al., 2009), the degradation of environmental contaminants (de Souza et al., 1998; Møller et al., 1998; Pelz et al., 1999; Drzyzga and Gottschal 2002; Holmes et al., 2006) and the recycling of fixed nitrogen into dinitrogen gas (N2) (Martienssen and Schops, 1999; Van de Pas-Schoonen et al., 2005; Costa et al., 2006). While substrate cross-feeding is frequently observed within natural and engineered microbial communities, its effects on microbial processes are often unclear. How does substrate cross-feeding affect the rate or extent of substrate consumption?
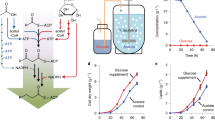
We propose a hypothesis about how substrate cross-feeding could accelerate substrate consumption, and thus affect microbial processes. Our hypothesis is based on the following main assumption: the enzyme that transforms the primary substrate competes with the enzyme that transforms the intermediate for the same finite pool of intracellular resources. Different enzymes may compete for elemental building blocks required for enzyme biosynthesis (e.g., carbon, nitrogen or phosphorous building blocks), for energy transfer molecules or co-factors required for enzyme activity (e.g., ATP, NADH) or for cytosolic or periplasmic space for enzymes to occupy (reviewed in Johnson et al., 2012). If the enzyme that transforms the primary substrate has preferential access to those intracellular resources (e.g., via higher affinity, regulatory mechanisms), then the rate of transformation of the parent substrate would be greater than that for the intermediate (Figure 1a, early time). The consequence is the accumulation of the intermediate (Figure 1b; note that we use batch culture for illustrative purposes, but inter-enzyme competition would also result in the accumulation of the intermediate in chemostat culture). After the parent substrate is sufficiently depleted, intracellular resources are then increasingly available to the enzyme that transforms the intermediate and the rate of transformation of the intermediate would increase (Figure 1a, late time). Indeed, experiments and mathematical simulations support inter-enzyme competition as an explanation for the accumulation of intermediates within some microbial populations (Thomsen et al., 1994; Almeida et al., 1995b).
Hypothesis for how segregating metabolic processes among different cell types could accelerate substrate consumption. Consider a metabolic pathway where a primary substrate (S) is transformed by an enzyme (Es) into an intermediate (I). The intermediate is then further transformed by another enzyme (EI) into an end product (P). (a) If ES and EI compete for the same finite pool of an intracellular resource and ES has preferential access to that resource, then I will accumulate at early times and only be consumed at later times. (b) The consequence is the transient accumulation of I. (c) Conversely, if ES and EI are segregated into different cell types, then competition is eliminated and EI has increased access to the intracellular resource. (d) The consequence is the increased consumption and reduced accumulation of I.
An important expectation of inter-enzyme competition, then, is that competition is eliminated when the enzyme that transforms the parent substrate and the enzyme that transforms the intermediate are segregated into different microbial cell types (Figure 1c). The enzyme that transforms the intermediate would then have increased access to intracellular resources that would otherwise be diverted to the enzyme that transforms the parent substrate. The consequence is that the rate of transformation of the intermediate would increase, thus reducing the accumulation of the intermediate and decreasing its growth-inhibiting effects (Figure 1d).
If inter-enzyme competition occurs then it leads to the following prediction: substrate cross-feeding should accelerate substrate consumption as the growth-inhibiting effects of the intermediate increase. This is because substrate cross-feeding breaks inter-enzyme competition and reduces the accumulation of the intermediate (Figures 1b and d), thus decreasing its growth-inhibiting effects. We note that this prediction does not state that substrate cross-feeding results in faster substrate consumption than complete consumption, as this transition point depends on a variety of environmental and biological factors. It also does not predict when substrate cross-feeding is likely to evolve. Instead, it simply states that substrate cross-feeding increases the rate of substrate consumption relative to complete consumption as the growth-inhibiting effects of the intermediate increase. Empirical evidence to support this prediction, however, remains anecdotal. For example, while substrate cross-feeding is often observed for the consumption of environmental pollutants that produce toxic and growth-inhibiting intermediates (de Souza et al., 1998; Møller et al., 1998; Pelz et al., 1999; Drzyzga and Gottschal 2002; Holmes et al., 2006), there is no empirical evidence that substrate cross-feeding itself accelerates the consumption of those pollutants.
Our objective was to experimentally test this prediction, and thus test a mechanism that explains how segregating different metabolic processes into different microbial cell types might affect microbial processes. To accomplish this, we genetically engineered a novel substrate cross-feeding system based on the denitrification pathway of the Gram-negative bacterium Pseudomonas stutzeri A1501 (Yan et al., 2008). In the absence of oxygen, P. stutzeri can support its growth by sequentially reducing nitrate (NO3−) to nitrite (NO2−), nitric oxide (NO), nitrous oxide (N2O) and finally to dinitrogen gas (N2), where the nitrogen oxides serve as terminal electron acceptors (Lalucat et al., 2006). We constructed three isogenic mutant strains of P. stutzeri A1501 that differ in their ability to reduce nitrate and nitrite (note that asparagine is supplied in excess as the nitrogen source for biosynthesis; Figure 2 and Supplementary Table 1). One strain completely reduces nitrate to dinitrogen gas (strain A1601; designated as the completely consuming strain), a second strain contains a loss-of-function deletion in the nar gene cluster that encodes nitrate reductase (Nar) and partially reduces nitrite to dinitrogen gas (strain A1602; designated as the nitrite-consuming strain) and a third strain contains a loss-of-function deletion in the nir gene cluster that encodes nitrite reductase (Nir) and partially reduces nitrate to nitrite (strain A1603; designated as the nitrite-producing strain; Figure 2 and Supplementary Table 1). We then grew the nitrite-producing and -consuming strains together in nitrite cross-feeding co-cultures or the completely consuming strain alone and measured how rapidly nitrogen oxides were consumed.
Experimental system established for this study. We constructed three isogenic mutant strains of P. stutzeri A1501 that differ in their ability to consume nitrate (NO3−) and nitrite (NO2−) as terminal electron acceptors. Strain A1601 completely consumes nitrate to dinitrogen gas (N2), strain A1602 has a loss-of-function deletion in the nar gene cluster and cannot consume nitrate and strain A1603 has a loss-of-function deletion in the nir gene cluster and cannot consume nitrite. Definitions: Nar, nitrate reductase encoded by the nar gene cluster; Nir, nitrite reductase encoded by the nir gene cluster; Nor, nitric oxide (NO) reductase encoded by the nor gene cluster; and Nos, nitrous oxide (N2O) reductase encoded by the nos gene cluster. Arrows indicate the metabolic steps that each strain can perform.
There are two critically important features of our experimental system. First, for some denitrifying microorganisms, the nitrate (NO3−) and nitrite (NO2−) reductases are thought to compete for the same finite pool of intracellular electron donors, but the nitrate reductase has preferential access to this intracellular resource (Thomsen et al., 1994; Almeida et al., 1995b). Second, we can manipulate the growth-inhibiting effects of the cross-fed metabolic intermediate—nitrite—and measure the consequences on substrate consumption. Nitrite accumulates within the periplasm during denitrification (Lalucat et al., 2006; Klueglein et al., 2014) and can inhibit growth by at least two different pH-dependent mechanisms, both of which act indirectly. First, as the pH decreases, nitrite increasingly protonates to nitrous acid (HNO2) and uncouples proton translocation (Sijbesma et al., 1996; Zhou et al., 2011). Second, as the pH decreases, nitrite increasingly and spontaneously generates nitric oxide radicals (NO) that interact with enzymes to form metal-nitrosyl complexes (Zumft, 1993). Nitrite inhibition is typically negligible at pH>7.5 but severely reduces growth at pH<6.8 (Almeida et al., 1995b; Sijbesma et al., 1996; Baumann et al., 1997; Zhou et al., 2011). We exploited this pH dependence to experimentally manipulate the magnitude of nitrite inhibition by adjusting the pH of the culture medium. This approach is effective because the pH of the culture medium is approximately equal to the pH of the periplasm for Gram-negative bacteria (Wilks and Slonczewski, 2007). We then measured how the magnitude of nitrite inhibition affects the speed at which completely consuming cells and co-cultures of nitrite cross-feeding cells consume nitrogen oxides, where we expect that nitrite cross-feeding increasingly accelerates the consumption of nitrogen oxides as the growth-inhibiting effects of nitrite increase.
Materials and methods
Bacterial strains, genetic manipulations and growth conditions
We obtained wild-type P. stutzeri A1501 from the Biological Resource Center of Institut Pasteur (Paris, France; www.crbip.pasteur.fr) and used this strain to construct all of the isogenic mutant strains used in this study. P. stutzeri A1501 was originally isolated from a rice patty soil and its genome has been fully sequenced and annotated (Yan et al., 2008). The genetic modifications of all mutant strains are summarized in Supplementary Table 1 and the primers used to create the genetic modifications are provided in Supplementary Table 2. In brief, we deleted the narG gene to prevent nitrate (NO3−) reduction and the nirS gene to prevent nitrite (NO2−) reduction. We additionally deleted the comA gene from all strains to prevent the internalization of extracellular DNA (Meier et al., 2002), thus reducing the probability that the nitrite-consuming (strain A1602) and -producing (strain A1603) strains would recombine with each other via natural transformation when grown together. Detailed descriptions of the methods used for gene deletion and phenotype validation are provided in the Supplementary Information.
We cultivated all P. stutzeri strains under aerobic conditions with a completely defined asparagine–citrate synthetic medium (ACS medium) (Coyle et al., 1985) in 1 ml mixed batch reactors. We cultivated all P. stutzeri strains under anaerobic conditions with dinitrogen gas (N2)-sparged ACS medium in 25 ml serum bottles fitted with gas-tight stoppers. We provided a detailed description of methods to prepare and inoculate anaerobic ACS medium in the Supplementary Information.
pH dependence of nitrite inhibition
We streaked the completely consuming strain (strain A1601) onto a lysogeny broth agar plate, inoculated one colony into a different test tube containing 1 ml of aerobic ACS medium buffered to pH 7.5 or 6.5, and incubated the test tubes for 24 h at 30 °C with continuous shaking (220 r.p.m.). We then transferred the cultures at a dilution of 1:100 (vol:vol) into 16 wells of a 96-well microtiter plate containing fresh ACS medium to achieve a final volume of 200 μl per well. In total, we prepared four wells each containing ACS medium buffered to pH 7.5, ACS medium buffered to pH 6.5, ACS medium buffered to pH 7.5 and amended with 10 mm of sodium nitrite (NaNO2), and ACS medium buffered to pH 6.5 and amended with 10 mm of sodium nitrite. After inoculation, we incubated the microtiter plate at 30 °C with continuous shaking (220 r.p.m.) and measured the OD600 every 10 min for 1440 min with an Eon plate reader (BioTek Instruments, Luzern, Switzerland). We quantified the maximum growth rate for each well by fitting a zero-order growth model to 10 consecutive data points that coincide with the most rapid period of growth. We performed similar experiments under anaerobic conditions (see the Supplementary Information).
Time to stationary-phase experiments
We streaked the completely consuming strain (strain A1601), the nitrite (NO2−)-consuming strain (strain A1602) and the nitrite-producing strain (strain A1603) onto lysogeny broth agar plates, inoculated one colony of each strain into a different test tube containing 1 ml of aerobic ACS medium set to pH 7.5 or 6.5, and incubated the test tubes for 24 h at 30 °C with continuous shaking (220 r.p.m.). We then inoculated the aerobic cultures (the completely consuming strain alone or co-cultures of nitrite cross-feeding strains together at a 50:50 vol:vol mixture) at a dilution of 1:25 (vol:vol) into serum bottles containing anaerobic ACS medium to achieve a final volume of 20 ml. The ACS medium contained 10 mm of sodium nitrate (NaNO3) and the pH was set to 7.5 or 6.5. These cultures then served as precultures for the following experiment. In total, four anaerobic precultures were prepared: the completely consuming strain at each pH and co-cultures of nitrite cross-feeding strains together at each pH. Once the anaerobic precultures had reached the stationary phase, we used them to inoculate bottles containing fresh anaerobic ACS medium containing 10 mm of sodium nitrate set to pH to 7.5 or 6.5 (1:100 dilution (vol:vol) in 20 ml) and measured the time for the cultures to reach the stationary phase. In total, we prepared three serum bottles each for the completely consuming strain and the co-cultures of nitrite cross-feeding strains at pH 7.5, and six bottles each at pH 6.5. We increased the replication at pH 6.5 because we knew from preliminary experiments that the cultures would sporadically and prematurely stop growing at this pH. After inoculation, we incubated the serum bottles at 30 °C with continuous shaking (220 r.p.m.) and measured the OD600 by transferring aliquots to a 96-well microtiter plate and analyzing the plate with a Synergy Mx plate reader (BioTek). We continued the experiment until there was no further increase in the OD600. We verified that cells entered the stationary phase as a consequence of consuming all of the available nitrogen oxides. Namely, we found that the stationary-phase cultures resumed growth after we added additional nitrate to the cultures.
Nitrate and nitrite measurements
We streaked the completely consuming strain (strain A1601), the nitrite (NO2−)-consuming strain (strain A1602) and the nitrite-producing strain (strain A1603) onto lysogeny broth agar plates, inoculated one colony of each strain into a different test tube containing 1 ml of aerobic ACS medium set to pH 7.5 and 10 mm of sodium nitrate (NaNO3) and incubated the test tubes for 24 h at 30 °C with continuous shaking (220 r.p.m.). We then inoculated the aerobic cultures (the completely consuming strain alone or co-cultures of nitrite cross-feeding strains together at a 50:50 vol:vol mixture) at a dilution of 1:100 (vol:vol) into three serum bottles each containing anaerobic ACS medium amended with 10 mm of sodium nitrate to achieve a final volume of 20 ml. After 24 h when the cultures had reached the stationary phase, we amended the cultures with 5 mm of additional sodium nitrate and sampled small volumes (~200 μl) from the cultures every 20 min while they were growing at 30 °C with continuous shaking (220 r.p.m.). We then removed the cells by centrifugation and stored the supernatants at −20 °C until chemical analysis. We amended the stationary-phase cultures with 5 mm of sodium nitrate to minimize the effects of cell growth on denitrification kinetics, where on average individual cells replicated less than one time. We provide detailed descriptions of the chemical analysis methods in the Supplementary Information.
Mathematical model
We developed a mathematical model to predict the kinetics of cell growth, nitrate (NO3−) consumption, and nitrite (NO2−) production and consumption. The model includes a term that simulates competition between the nitrate and nitrite reductases for reduced electron carriers (see Equation (7) in the Supplementary Information; Almeida et al., 1995b). The model also explicitly accounts for mass transfer between the periplasm and the culture medium and for the pH-dependent inhibitory effects of nitrite. We provide a complete description of the model and its parameters in the Supplementary Information.
Results
Competition between the nitrate and nitrite reductases
We first tested for inter-enzyme competition between the nitrate (NO3−) and nitrite (NO2−) reductases under our experimental conditions. To test this, we exploited a well-known feature of denitrifying microorganisms growing in batch culture; nitrate and nitrite are consumed sequentially, resulting in the transient accumulation of the intermediate nitrite (Betlach and Tiedje, 1981; Carlson and Ingraham, 1983). Previous experimental and theoretical studies hypothesized that the nitrate reductase has a stronger affinity for those reduced electron carriers and out-competes the nitrite reductase when nitrate is present, resulting in the preferential consumption of nitrate and the transient accumulation of the intermediate nitrite (Figure 1b; Thomsen et al., 1994; Almeida et al., 1995b). We reasoned that if inter-enzyme competition is biologically significant, then segregating the nitrate and nitrite reductases into different nitrite cross-feeding cells should eliminate inter-enzyme competition. The consequence would be the simultaneous consumption of nitrate and nitrite and the reduced accumulation of the intermediate nitrite (Figure 1d).
We experimentally tested this expectation by measuring the accumulation of the intermediate nitrite (NO2−) in batch cultures containing completely consuming cells (strain A1601) or co-cultures of nitrite cross-feeding cells (strains A1602 and A1603). We set the pH of the culture medium to 7.5 because we found that nitrite does not cause growth inhibition under those conditions (see results below). This allowed us to isolate the effects of competition between the nitrate and nitrite reductases from the growth-inhibiting effects of nitrite itself on the dynamics of substrate consumption. We first grew completely consuming cells alone or co-cultures of nitrite cross-feeding cells together with 10 mm of nitrate until they reached the stationary phase. This allowed each nitrite cross-feeding strain to achieve a relative frequency that reflects its realized yield from its particular substrates. This was approximately 50% of each strain, which is consistent with the expected yields from nitrate and nitrite reduction (i.e., each reduction step consumes two electrons). We then amended the stationary-phase cultures with an additional 5 mm of nitrate and measured the concentrations of nitrate and nitrite over time.
Our results are consistent with inter-enzyme competition between the nitrate (NO3−) and nitrite (NO2−) reductases for the same finite pool of intracellular resources. The intermediate nitrite accumulated to relatively high concentrations for the completely consuming cells (Figure 3a, solid blue line) but accumulated to nearly undetectable concentrations for the co-cultures of nitrite cross-feeding cells (Figure 3a, solid red line). Overall, the maximum observed nitrite concentration over the time course of the experiment was 8-fold lower for the co-cultures of nitrite cross-feeding cells than for the completely consuming cells (two-sample Welch test, two-sided P<0.05, n1=n2=3) (Figure 3a). Thus, when the nitrate and nitrite reductases were present together within completely consuming cells, nitrate and nitrite were consumed largely sequentially, which would be expected if the nitrate reductase had preferential access to reduced electron carriers (Thomsen et al., 1994; Almeida et al., 1995b). However, when the nitrate and nitrite reductases were segregated into different nitrite cross-feeding cells and grown together, nitrate and nitrite were consumed simultaneously, which would be expected if competition between the two reductases were eliminated. Our results are therefore consistent with inter-enzyme competition between the nitrate and nitrite reductases and preferential access by the nitrate reductase for intracellular resources, thus providing experimental support for the main assumption of our hypothesis.
Nitrate (NO3−) and nitrite (NO2−) consumption dynamics for completely consuming cells (strain A1601) or co-cultures of nitrite cross-feeding cells (strains A1602 and A1603). (a) We experimentally measured nitrate and nitrite concentrations over time after adding 5 mm of nitrate to stationary-phase cultures. Error bars are one standard deviation (n=3). We additionally simulated the dynamics of nitrate and nitrite concentrations when (b) imposing competition between the nitrate and nitrite reductases, or (c) when setting the maximum rate of nitrate consumption to twice that of nitrite consumption. Definitions: CC, completely consuming cells (strain A1601); CF, co-cultures of nitrite cross-feeding cells (strains A1602 and A1603).
We complemented our experimental results by simulating the dynamics of nitrite (NO2−) accumulation for batch culture. The mathematical model includes a term that simulates inter-enzyme competition between the nitrate and nitrite reductases when they are present together within completely consuming cells, where the rate of nitrate (NO3−) consumption directly represses the rate of nitrite consumption (see Equation (7) in the Supplementary Information; Almeida et al., 1995b). The mathematical model revealed an important insight: when the nitrate and nitrite reductases are present together within completely consuming cells, the competition effect is large immediately after nitrate is added to the batch cultures. Nitrate consumption consequently represses nitrite consumption and the intermediate nitrite accumulates to high concentrations (Figure 3b, solid blue line). However, when the nitrate and nitrite reductases are segregated into different nitrite cross-feeding cells and grown together, the competition effect is necessarily zero. Nitrate consumption does not reduce nitrite consumption and the intermediate nitrite accumulates to low concentrations (Figure 3b, solid red line). These simulations are in remarkable qualitative agreement with our experimental results, where co-cultures of nitrite cross-feeding cells accumulated substantially less nitrite than completely consuming cells (Figure 3a). Thus, the inclusion of competition between the nitrate and nitrite reductases was sufficient to simulate the main qualitative feature of our experimental observations. We note here that we did not fit our data to the model but instead used literature reported kinetic parameters for our simulations (Bryan et al., 1985; Almeida et al., 1995a,b), thus demonstrating the generality of the outcome. Indeed, the qualitative outcome (i.e., that co-cultures of nitrite cross-feeding cells accumulate less nitrite than completely consuming cells) is valid across a biologically relevant parameter space.
Alternative explanations for the accumulation of nitrite
There are alternative explanations other than inter-enzyme competition for the transient accumulation of the intermediate nitrite (NO2−) by completely consuming cells. One often-invoked explanation is that the rate of nitrate (NO3−) consumption is simply greater than the rate of nitrite consumption, which would also result in the transient accumulation of nitrite without requiring inter-enzyme competition (Betlach and Tiedje, 1981). To test the potential validity of this explanation, we again simulated the dynamics of nitrite accumulation in batch culture, but we removed the competition effect between the nitrate and nitrite reductases and set the maximum rate of nitrate consumption to twice that of nitrite consumption. The model generated a qualitatively different outcome. When the nitrate and nitrite reductases are present together within completely consuming cells, nitrate is consumed more rapidly than nitrite and the intermediate nitrite accumulates to moderately high concentrations (Figure 3c, solid blue line). However, when the nitrate and nitrite reductases are segregated into different nitrite cross-feeding cells and grown together, the intermediate nitrite accumulates to even higher concentrations (Figure 3c, solid red line), which is opposite of our experimental observations (Figure 3a). The qualitatively opposite outcome is a consequence of the slower growth of the nitrite-consuming cells in the model simulations. Because of the lower rate of nitrite consumption, the nitrite-consuming cells are present at relatively low cell numbers after all the nitrate is consumed, thus increasing nitrite accumulation. In contrast, the completely consuming cells are present at higher cell numbers after all the nitrate is consumed, thus decreasing nitrite accumulation. Thus, the inability of this alternative explanation to simulate the main qualitative feature of our experimentally observed dynamics provides further support that the nitrate and nitrite reductases do indeed compete with each other for the same finite pool of intracellular resources.
Another alternative explanation for the accumulation of nitrite (NO2−) by completely consuming cells is that nitrate (NO3−) represses the transcriptional or translational expression of nitrite reductase. A functional nitrite reductase might therefore not be immediately present when cultures are amended with nitrate, thus creating a lag before nitrite consumption begins. To address this possibility, we performed an additional experiment where we allowed completely consuming cells to consume 2 mm of nitrate and enter the stationary phase. We then amended the cultures with 200 μm of nitrate or nitrite. The stationary-phase cultures immediately consumed both nitrate and nitrite with no observable lag (Supplementary Figure 1). Thus, a functional nitrite reductase was indeed present, and the transcriptional or translational repression of nitrite reductase cannot explain the transient accumulation of nitrite by completely consuming cells.
pH dependence of nitrite inhibition
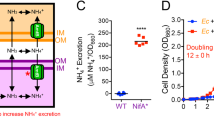
We next tested whether the accumulation of the intermediate nitrite (NO2−) imposes growth-inhibiting effects under our experimental conditions, which is required to test our main prediction. To accomplish this, we exploited the pH-dependent inhibitory effects of nitrite. We set the pH of the culture medium to 7.5 or 6.5 and grew completely consuming cells (strain A1601) alone in the presence or absence of 10 mm of exogenous nitrite. We incubated the cells under fully aerobic conditions because oxygen represses nitrite reductase activity (Vollack et al., 1999), thus preventing nitrite reduction—and consequently detoxification—during the experiment. We measured cell density over time (measured as OD600; Figures 4a and b) and quantified the maximum specific growth rate (umax (min−1)) for each culture (Figure 4c).
pH dependence of nitrite (NO2−) inhibition. We grew completely consuming cells (strain A1601) (a) at pH 7.5 with or without 10 mm of nitrite, or (b) at pH 6.5 with or without 10 mm of nitrite under aerobic conditions and measured the OD600 values over time. We only plotted every fourth data point to facilitate visualization. (c) We measured the zero-order maximum specific growth rate (umax) from 10 consecutive data points that coincide with the most rapid period of growth for each individual culture. The data are presented as Tukey box plots. The P-values are for pair-wise comparisons using the two-sample Welch test (n=4).
We observed three important outcomes. First, decreasing the pH of the culture medium from 7.5 to 6.5 in the absence of nitrite (NO2−) had no significant effect on the maximum specific growth rate of completely consuming cells (two-sample Welch test, two-sided P>0.05, n1=n2=4; Figure 4c). We independently repeated this experiment with a larger sample size and observed the same outcome; reducing the pH from 7.5 to 6.5 in the absence of nitrite did not cause observable growth inhibition (two-sample Welch test, one-sided P>0.8, n1=n2=8; Supplementary Figure 2). Second, the presence of nitrite at pH 7.5 also had no observable effect on the maximum specific growth rate of completely consuming cells (two-sample Welch test, two-sided P>0.05, n1=n2=4; Figures 4a and c). Nitrite therefore did not cause observable growth inhibition at pH 7.5. Third, the presence of nitrite at pH 6.5 completely eliminated the growth of completely consuming cells (two-sample Welch test, two-sided P<0.000001, n1=n2=4; Figures 4b and c). Nitrite therefore imposed severe growth inhibition at pH 6.5. We performed similar experiments in the absence of oxygen and observed qualitatively identical results (see the Supplementary Information). Together, our results demonstrate that (i) nitrite does indeed impose pH-dependent growth inhibition under our experimental conditions and (ii) that the pH of the culture medium can be used to manipulate the magnitude of nitrite inhibition without generating substantial confounding factors caused by differences in the pH itself.
The magnitude of nitrite inhibition determines whether nitrite cross-feeding accelerates substrate consumption
We next experimentally tested the main prediction of our hypothesis: nitrite (NO2−) cross-feeding increasingly accelerates substrate consumption as the growth-inhibiting effects of nitrite increase. To accomplish this, we set the pH of the culture medium to 7.5 or 6.5 and grew completely consuming cells alone (strain A1601) or co-cultures of nitrite cross-feeding cells together (strains A1602 and A1603) with 10 mm of nitrate (NO3−). We measured cell density over time (OD600) (Supplementary Figure 3) and quantified the time required for completely consuming cells (tstat,CC) and co-cultures of nitrite cross-feeding cells (tstat,CF) to reach the stationary phase. We then calculated the ratio tstat,CF/tstat,CC, where a value >1 indicates that completely consuming cells consumed all of the nitrogen oxides more rapidly and a value <1 indicates that co-cultures of nitrite cross-feeding cells consumed all of the nitrogen oxides more rapidly.
We observed two important outcomes. First, at pH 7.5 when nitrite (NO2−) does not inhibit growth (Figure 4c), the value of tstat,CF/tstat,CC was statistically indistinguishable from a value of 1 (one-sample Welch test, two-sided P>0.4, n=3; Figure 5). Thus, when nitrite had no observable negative effects on growth, completely consuming cells and co-cultures of nitrite cross-feeding cells consumed all of the nitrogen oxides equally rapidly. Breaking competition between the nitrate and nitrite reductases via substrate cross-feeding was therefore insufficient alone to affect substrate consumption. Second, at pH 6.5 when nitrite severely inhibits growth (Figure 4c), the value of tstat,CF/tstat,CC was smaller than 1 (one-sample Welch test, two-sided P<0.002, n=3; Figure 5) and smaller than that at pH 7.5 (two-sample Welch test, two-sided P<0.001, n1=n2=3; Figure 5). Thus, when nitrite had severe growth-inhibiting effects, co-cultures of nitrite cross-feeding cells consumed nitrogen oxides more rapidly than completely consuming cells. Our data therefore provide evidence that nitrite cross-feeding does indeed accelerate substrate consumption, but only when nitrite has growth-inhibiting effects.
Effect of pH on substrate consumption for completely consuming cells (strain A1601) or co-cultures of nitrite (NO2−) cross-feeding cells (strains A1602 and 1603). We measured the time for completely consuming cells (tstat,CC) or co-cultures of nitrite cross-feeding cells (tstat,CF) to reach the stationary phase. We reported the ratio tstat,CF/tstat,CC, where a value >1 indicates that the completely consuming cells (strain A1601) consumed all of the nitrogen oxides more rapidly and a value <1 indicates that the co-cultures of nitrite cross-feeding cells (strains A1602 and A1603) consumed all of the nitrogen oxides more rapidly. The data are presented as Tukey box plots. The P-value is for a pair-wise comparison using the two-sample two-sided Welch test (n=3).
We finally asked whether our mathematical modeling approach could simulate the main qualitative feature of our experiment without fitting the model to our data. More specifically, we asked whether a magnitude of nitrite (NO2−) inhibition exists when nitrite cross-feeding accelerates substrate consumption relative to complete consumption. The model revealed a second important insight. If mass transfer of nitrite between the periplasm and the culture medium is much faster than the maximum reaction rate of the nitrate and nitrite reductases, then the concentrations of nitrite within the periplasm and the culture medium are approximately identical for all cells, irrespective of whether a cell produces or consumes nitrite. Under these conditions, we found that completely consuming cells always consume all of the nitrogen oxides more rapidly than co-cultures of nitrite cross-feeding cells, regardless of the magnitude of nitrite inhibition (i.e., tstat,CF/tstat,CC is always greater than 1 and increases monotonically as nitrite inhibition increases; Figure 6, solid line). However, if mass transfer of nitrite between the periplasm and the culture medium is slower than the maximum reaction rate of the nitrate and nitrite reductases, which is supported by experimental evidence (Klueglein et al., 2014), then the concentrations of nitrite within the periplasm of the nitrite-consuming cells are necessarily less than in the culture medium (i.e., nitrite is consumed faster than it enters the cell). This reduces nitrite inhibition of nitrite-consuming cells and consequently accelerates nitrite consumption, thus reducing nitrite accumulation. Under these conditions, we found a threshold of nitrite inhibition where co-cultures of nitrite cross-feeding cells consume all of the nitrogen oxides more rapidly than completely consuming cells (i.e., a region where tstat,CF/tstat,CC is less than 1; Figure 6). More generally, we found that the value of tstat,CF/tstat,CC decreases monotonically as nitrite inhibition increases (i.e., nitrite cross-feeding increasingly accelerates the consumption of nitrogen oxides as nitrite inhibition increases; Figure 6), which is consistent with our main prediction (Figure 1) and our experimental observations (Figure 5). Thus, the lower concentration of nitrite within the periplasm of nitrite-consuming cells is essential to simulate the main qualitative feature of our results.
Simulations of the effect of nitrite (NO2−) inhibition on substrate consumption for completely consuming cells or co-cultures of nitrite cross-feeding cells. We calculated the time for completely consuming cells (tstat,CC) or co-cultures of nitrite cross-feeding cells (tstat,CF) to reach the stationary phase. We reported the ratio tstat,CF/tstat,CC, where a value >1 indicates that the completely consuming cells consumed all of the nitrogen oxides more rapidly and a value <1 indicates that the co-cultures of nitrite cross-feeding cells consumed all of the nitrogen oxides more rapidly. The solid line is for simulations where mass transfer across the periplasm was set to infinity. The dashed line is for simulations where mass transfer across the periplasm was set to one-half of the maximum rate of nitrite consumption. KI−1 is the inverse of the Andrews inhibition coefficient for nitrite.
Discussion
Our experiments provide evidence for a potentially general principle: The growth-inhibiting effects of metabolic intermediates determine whether segregating different metabolic processes into different cell types promotes more rapid substrate consumption. For our experimental system, segregating metabolic processes eliminates inter-enzyme competition and consequently reduces the accumulation of intermediates, thus accelerating the consumption of substrates that produce growth-inhibiting intermediates. This principle has previously only been supported by anecdotal evidence (de Souza et al., 1998; Møller et al., 1998; Pelz et al., 1999; Drzyzga and Gottschal 2002; Holmes et al., 2006) while direct experimental evidence has so far been lacking, thus reflecting an important gap in our knowledge about the consequences of metabolic specialization on microbial processes.
Our results could be of relevance for a broad range of engineered microbial processes. Consider that the objective of nearly every engineered microbial process is to convert a substrate via one or more metabolic intermediates into a desired end product. For example, one might want to convert a low-cost organic compound into a pharmaceutical or convert a pollutant into an innocuous end product. One strategy to achieve a particular engineering objective would be to engineer a single cell type that completely consumes the substrate into the desired end product. An alternative strategy would be to engineer and assemble together different cell types, where each cell type specializes at performing a different step of the pathway (i.e., a microbial (dis)assembly line). We currently lack basic engineering design principles that predict when such a microbial (dis)assembly line is likely to be advantageous, or how to best segregate different metabolic processes into different cell types to achieve a particular objective (Lindemann et al., 2016). Based on our results, knowledge about inter-enzyme competition and the inhibitory effects of metabolic intermediates could contribute towards enabling such predictions.
Our results also illustrate a mechanism that could help address a long-standing enigma in microbial ecology: How is biodiversity promoted and maintained within microbial communities? Consider that many natural and engineered environments contain many thousands of different microbial strains (Curtis et al., 2002). What prevents a few strains from increasing in frequency and displacing the others? Our results suggest that the production of inhibitory metabolic intermediates is one factor that could promote and maintain biodiversity. Under certain conditions, competition between different metabolic processes and the production of growth-inhibiting intermediates could cause different metabolic processes to segregate into different strains over evolutionary time (Doebeli, 2002; Pfeiffer and Bonhoeffer, 2004; Costa et al., 2006; Johnson et al., 2012). The consequence is the evolution of a collection of metabolically specialized strains, each of which consumes only a subset of the available resources within a particular environment and co-exists with the others (Doebeli, 2002; Pfeiffer and Bonhoeffer, 2004; Costa et al., 2006; Johnson et al., 2012). Given the enormous numbers of substrates and intermediates that are likely to exist in many natural environments, we argue that this mechanism could be an important factor that contributes towards the promotion and maintenance of microbial diversity.
Our study has several important advantages and perceived limitations. In our view, the main advantage and novel aspect is that we could manipulate the inhibitory effects of a single metabolic intermediate and measure the consequences on substrate consumption. A second important advantage is that all three of the investigated strains are isogenic mutants. We were therefore able to exclude confounding factors that might be caused by genomic differences at loci other than the nar and nir gene clusters. One perceived limitation is that all of our experiments and simulations were conducted in batch culture. The main qualitative prediction of our hypothesis, however, is the same for both batch and chemostat culture. Inter-enzyme competition and preferential access by the enzyme that transforms the primary substrate will increase the accumulation of intermediates, regardless of reactor configuration. Moreover, eliminating inter-enzyme competition via substrate cross-feeding will reduce the accumulation of intermediates, again regardless of reactor configuration. Nevertheless, chemostat operation would result in lower concentrations of intermediates because a fraction of the intermediates would be removed via dilution. We therefore expect a smaller parameter space where substrate cross-feeding results in faster substrate consumption than complete consumption. However, the general prediction that substrate cross-feeding increasingly accelerates substrate consumption as the negative effects of intermediates increase, should remain valid.
We emphasize that our results do not address the evolution of substrate cross-feeding itself, which has been the focus of previous studies (Turner et al., 1996; Treves et al., 1998; Doebeli, 2002; Pfeiffer and Bonhoeffer, 2004; Costa et al., 2006). While we identified conditions where substrate cross-feeding accelerates substrate consumption, this is insufficient to predict whether substrate cross-feeding itself would evolve via natural selection from complete consumption under the same conditions. Other evolutionary outcomes are plausible (Pfeiffer and Bonhoeffer, 2004). For our specific experimental system, it is plausible that substrate cross-feeding cells may never displace completely consuming cells, or that intermediate-consuming cells may co-exist with completely consuming cells. The evolutionary outcome ultimately depends on the magnitude of inter-enzyme competition, the growth-inhibiting effects of intermediates and the availability of regulatory solutions that could also minimize the accumulation of intermediates. In batch culture where the diffusion of intermediates away from intermediate-producing cells is rapid, the negative effects of accumulating intermediates might be small. The evolution of substrate cross-feeding might therefore require long periods of time or, as occurred in this study, require relatively high concentrations of growth-inhibiting intermediates (we note here, however, that the nitrogen oxide concentrations observed in our study are not atypical for some ecosystems (Fernández-Nava et al., 2008)). In contrast, in a spatially structured environment where the transport of intermediates away from cells is slowed, the growth-inhibiting effects of intermediates might be much larger. This is because the intermediates would be retained within close proximity to the intermediate-producing cells, thus increasing their inhibitory effects and promoting the rapid evolution of substrate cross-feeding.
Finally, our results have potentially immediate impacts on our understanding of an ecosystem service that is critical for maintaining the quality of water supplies: the consumption of nitrogen oxides. Both natural and engineered microbial communities are known to sometimes accumulate the intermediate nitrite (NO2−), with potentially deleterious effects on environmental quality. The underlying causes of nitrite accumulation, however, remain debated (Betlach and Tiedje, 1981; Almeida et al., 1995b). Our data indicate that, for P. stutzeri, competition between the nitrate (NO3−) and nitrite reductases is one factor that causes nitrite accumulation, and that segregating the nitrate and nitrite reductases into different cell types prevents nitrite accumulation. A further analysis of the cellular compartmentalization of the nitrate and nitrite reductases could therefore help elucidate the underlying causes of nitrite accumulation in natural and engineered ecosystems.
References
Almeida JS, Júlio SM, Reis MA, Carrondo MJ . (1995a). Nitrite inhibition of denitrification by Pseudomonas fluorescens. Biotechnol Bioeng 46: 194–201.
Almeida JS, Reis MA, Carrondo MJ . (1995b). Competition between nitrate and nitrite reduction in denitrification by Pseudomonas fluorescens. Biotechnol Bioeng 46: 476–484.
Baumann B, van der Meer JR, Snozzi M, Zehnder AJ . (1997). Inhibition of denitrification activity but not of mRNA induction in Paracoccus denitrificans by nitrite at a suboptimal pH. Antonie Van Leeuwenhoek 72: 183–189.
Betlach MR, Tiedje JM . (1981). Kinetic explanation for accumulation of nitrite, nitric oxide, and nitrous oxide during bacterial denitrification. Appl Environ Microbiol 42: 1074–1084.
Bryan BA, Jeter RM, Carlson CA . (1985). Inability of Pseudomonas stutzeri denitrification mutants with the phenotype of Pseudomonas aeruginosa to grow in nitrous oxide. Appl Environ Microbiol 50: 1301–1303.
Carlson CA, Ingraham JL . (1983). Comparison of denitrification by Pseudomonas stutzeri Pseudomonas aeruginosa, and Paracoccus denitrificans. Appl Environ Microbiol 45: 1247–1253.
Costa E, Pérez J, Kreft JU . (2006). Why is metabolic labour divided in nitrification? Trends Microbiol 14: 213–219.
Coyle CL, Zumft WG, Kroneck PM, Körner H, Jakob W . (1985). Nitrous oxide reductase from denitrifying Pseudomonas perfectomarina. Purification and properties of a novel multicopper enzyme. Eur J Biochem 153: 459–467.
Curtis TP, Sloan WT, Scannell JW . (2002). Estimating prokaryotic diversity and its limits. Proc Natl Acad Sci USA 99: 10494–10499.
de Souza ML, Newcombe D, Alvey S, Crowley DE, Hay A, Sadowsky MJ et al. (1998). Molecular basis of a bacterial consortium: interspecies catabolism of atrazine. Appl Environ Microbiol 64: 178–184.
Doebeli M . (2002). A model for the evolutionary dynamics of cross‐feeding polymorphisms in microorganisms. Population Ecol 44: 59–70.
Drzyzga O, Gottschal JC . (2002). Tetrachloroethene dehalorespiration and growth of Desulfitobacterium frappieri TCE1 in strict dependence on the activity of Desulfovibrio fructosivorans. Appl Environ Microbiol 68: 642–649.
Fernández-Nava Y, Marañón E, Soons J, Castrillón L . (2008). Denitrification of wastewater containing high nitrate and calcium concentrations. Bioresour Technol 99: 7976–7981.
Holmes VF, He J, Lee PK, Alvarez-Cohen L . (2006). Discrimination of multiple Dehalococcoides strains in a trichloroethene enrichment by quantification of their reductive dehalogenase genes. Appl Environ Microbiol 72: 5877–5883.
Johnson DR, Goldschmidt F, Lilja EE, Ackermann M . (2012). Metabolic specialization and the assembly of microbial communities. ISME J 6: 1985–1991.
Klueglein N, Zeitvogel F, Stierhof YD, Floetenmeyer M, Konhauser KO, Kappler A et al. (2014). Potential role of nitrite for abiotic Fe(II) oxidation and cell encrustation during nitrate reduction by denitrifying bacteria. Appl Environ Microbiol 80: 1051–1061.
Lalucat J, Bennasar A, Bosch R, García-Valdés E, Palleroni NJ . (2006). Biology of Pseudomonas stutzeri. Microbiol Mol Biol Rev 70: 510–547.
Lindemann SR, Bernstein HC, Song HS, Frederickson JK, Fields MW, Shou W et al. (2016). Engineering microbial consortia for controllable outputs. ISME J (in press).
Martienssen M, Schops R . (1999). Population dynamics of denitrifying bacteria in a model biocommunity. Water Res 33: 639–646.
McInerney MJ, Sieber JR, Gunsalus RP . (2009). Syntrophy in anaerobic global carbon cycles. Curr Opin Biotechnol 20: 623–632.
Meier P, Berndt C, Weger N, Wackernagel W . (2002). Natural transformation of Pseudomonas stutzeri by single-stranded DNA requires type IV pili, competence state and comA. FEMS Microbiol Lett 207: 75–80.
Møller S, Sternberg C, Andersen JB, Christensen BB, Ramos JL, Givskov M et al. (1998). In situ gene expression in mixed-culture biofilms: evidence of metabolic interactions between community members. Appl Environ Microbiol 64: 721–732.
Pelz O, Tesar M, Wittich RM, Moore ER, Timmis KN, Abraham WR . (1999). Towards elucidation of microbial community metabolic pathways: unravelling the network of carbon sharing in a pollutant-degrading bacterial consortium by immunocapture and isotopic ratio mass spectrometry. Environ Microbiol 1: 167–174.
Pfeiffer T, Bonhoeffer S . (2004). Evolution of cross-feeding in microbial populations. Am Nat 163: E126–E135.
Schink B . (1997). Energetics of syntrophic cooperation in methanogenic degradation. Microbiol Mol Biol Rev 61: 262–280.
Sijbesma WFH, Almeida JS, Reis MAM, Santos H . (1996). Uncoupling effect of nitrite during denitrification by Pseudomonas fluorescens: an in vivo31P-NMR study. Biotechnol Bioeng 52: 176–182.
Thomsen JK, Geest T, Cox RP . (1994). Mass spectrometric studies of the effect of pH on the accumulation of intermediates in denitrification by Paracoccus denitrificans. Appl Environ Microbiol 60: 536–541.
Treves DS, Manning S, Adams J . (1998). Repeated evolution of an acetate-crossfeeding polymorphism in long-term populations of Escherichia coli. Mol Biol Evol 15: 789–797.
Turner PE, Souza V, Lenski RE . (1996). Tests of ecological mechanisms promoting the stable coexistence of two bacterial genotypes. Ecol 77: 2119–2129.
Van de Pas-Schoonen KT, Schalk-Otte S, Haaijer S, Schmid M, Op den Camp H, Strous M et al. (2005). Complete conversion of nitrate into dinitrogen gas in co-cultures of denitrifying bacteria. Biochem Soc Trans 33: 205–209.
Vollack KU, Härtig E, Körner H, Zumft WG . (1999). Multiple transcription factors of the FNR family in denitrifying Pseudomonas stutzeri: characterization of four fnr-like genes, regulatory responses and cognate metabolic processes. Mol Microbiol 31: 1681–1694.
Wilks JC, Slonczewski JL . (2007). pH of the cytoplasm and periplasm of Escherichia coli: rapid measurement by green fluorescent protein fluorimetry. J Bacteriol 189: 5601–5607.
Yan Y, Yang J, Dou Y, Chen M, Ping S, Peng J et al. (2008). Nitrogen fixation island and rhizosphere competence traits in the genome of root-associated Pseudomonas stutzeri A1501. Proc Natl Acad Sci USA 105: 7564–7569.
Zelezniak A, Andrejev S, Ponomarova O, Mende DR, Bork P, Patil K . (2015). Metabolic dependencies drive species co-occurrence in diverse microbial communities. Proc Natl Acad Sci USA 112: 6449–6454.
Zhou Y, Oehmen A, Lim M, Vadivelu V, Ng WJ . (2011). The role of nitrite and free nitrous acid (FNA) in wastewater treatment plants. Water Res 45: 4672–4682.
Zumft WG . (1993). The biological role of nitric oxide in bacteria. Arch Microbiol 160: 253–264.
Acknowledgements
We thank two anonymous reviewers for significantly improving the quality of this manuscript; Martin Ackermann, Sebastian Bonhoeffer, Jan Dolinsek, Felix Goldschmidt, Frank Schreiber and Simon van Vliet for useful discussions; William Metcalf for generously providing plasmids and strains used for this study; Anja Bernet and Selina Derksen-Müller for assistance with the genetic manipulations; and Thomas Fleischmann for assistance with the chemical analyses. This work was supported by a grant from the Swiss National Science Foundation (grant number 31003A_149304).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Lilja, E., Johnson, D. Segregating metabolic processes into different microbial cells accelerates the consumption of inhibitory substrates. ISME J 10, 1568–1578 (2016). https://doi.org/10.1038/ismej.2015.243
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.243
This article is cited by
-
Timing of antibiotic administration determines the spread of plasmid-encoded antibiotic resistance during microbial range expansion
Nature Communications (2023)
-
Changes in interactions over ecological time scales influence single-cell growth dynamics in a metabolically coupled marine microbial community
The ISME Journal (2023)
-
Stress-induced metabolic exchanges between complementary bacterial types underly a dynamic mechanism of inter-species stress resistance
Nature Communications (2023)
-
Gut Microbial Communities in Mealworms and Indianmeal Moth Larvae Respond Differently to Plastic Degradation
Journal of Polymers and the Environment (2023)
-
In-depth characterization of denitrifier communities across different soil ecosystems in the tundra
Environmental Microbiome (2022)