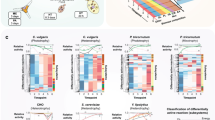
Abstract
The environmental conditions that describe an ecosystem define the amount of energy available to the resident organisms and the amount of energy required to build biomass. Here, we quantify the amount of energy required to make biomass as a function of temperature, pressure, redox state, the sources of C, N and S, cell mass and the time that an organism requires to double or replace its biomass. Specifically, these energetics are calculated from 0 to 125 °C, 0.1 to 500 MPa and −0.38 to +0.86 V using CO2, acetate or CH4 for C, NO3− or NH4+ for N and SO42− or HS− for S. The amounts of energy associated with synthesizing the biomolecules that make up a cell, which varies over 39 kJ (g cell)−1, are then used to compute energy-based yield coefficients for a vast range of environmental conditions. Taken together, environmental variables and the range of cell sizes leads to a ~4 orders of magnitude difference between the number of microbial cells that can be made from a Joule of Gibbs energy under the most (5.06 × 1011 cells J−1) and least (5.21 × 107 cells J−1) ideal conditions. When doubling/replacement time is taken into account, the range of anabolism energies can expand even further.
Similar content being viewed by others
Introduction
In an effort to better understand the origin, evolution and extent of life, significant effort has gone into finding the limits of temperature, pressure, pH, salinity and other compositional and physical variables that define habitability (Rothschild and Mancinelli, 2001; Pikuta et al., 2007; Canganella and Wiegel, 2011; Colwell and D’Hondt, 2013; Picard and Daniel, 2013; Takai et al., 2014). However, one variable in particular—energy—has received much less attention (Amend and Teske, 2005; Shock and Holland, 2007; Hoehler and Jørgensen, 2013; LaRowe and Amend, 2015a, b). Virtually every aspect of microbial behavior requires energy, including growth, nutrient uptake, motility, excretion of biomolecules, synthesis of external structures such as stalks, sheaths, tubes and extracellular polymeric saccharides, and changes in stored nutrients. Although some of these fall within the definition of maintenance functions (van Bodegom, 2007), many, such as growth, require the synthesis of biomolecules. Consequently, the energy required to make biomolecules is a fundamental limit to the habitability of an environment.
Because the Gibbs energy of all chemical reactions is a function of temperature, pressure and composition, determining the energetic limits of anabolism will differ depending on these environmental variables (for example, Amend and Shock, 1998, 2000; McCollom and Amend, 2005; LaRowe and Regnier, 2008; Amend and McCollom, 2009; Amend et al., 2011, 2013; LaRowe and Amend, 2014; Osburn et al., 2014; Teske et al., 2014; Price et al., 2015). In addition, the amount of energy that it takes to produce and sustain a given number of microorganisms depends on the mass of cells and the relative proportion of biomolecule types that make up these cells. As a result, the energy boundary for life is a moving one that changes depending on a large number of factors. The purpose of this study is to quantify the influence of environmental variables on the amount of energy it takes to synthesize microbial biomass.
Our approach considers a great range of growth conditions to account for the fact that life has been found in hydrothermal systems, deep marine and freshwater sediment, hydrocarbon reservoirs, unconsolidated sedimentary rock, shale, sandstone, fractured granite, deep aquifers, caves, mines, paleosols, aquitards, permafrost and more (Amy and Haldeman, 1997; Fredrickson and Fletcher, 2001; Steven et al., 2006; Orcutt et al., 2011; Strapoć et al., 2011; Edwards et al., 2012). Other attempts to account for the energy required to make biomass rely upon laboratory experiments that are designed to promote growth, conditions that are rarely encountered in nature (for a review see, LaRowe and Amend, 2015a). In addition, numerous energy-based models developed to predict biomass yields have limited applicability because of being restricted to standard and/or reference states (for reviews see, Heijnen and van Dijken, 1992; LaRowe and Amend, 2015a). Most of these models fix biomass yield coefficients, which lead to predictions of equal biomass production (for the same synthesis reactions) under very different environmental conditions. The methods adopted in this study explicitly quantify the role that physiochemical variables have on biomass synthesis.
Materials and methods
Computational techniques
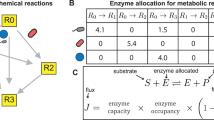
The Gibbs energy required to make a microbial cell, ΔGcell, can be calculated with

where ΔGsynth represents the energy required to synthesize biomolecules and ΔGmain stands for the energy that is used for all other functions in support of viability, the so-called maintenance energy. ΔGsynth accounts for the reaction energetics of monomer biomolecules (for example, amino acids, nucleotides, saccharides) from simple starting materials (for example, CO2, acetate, CH4,  , HS−,
, HS−,  ,
,  ) and the energetics associated with polymerization of these monomers into biomacromolecules (for example, proteins, DNA, polysaccharides). Although the types and amounts of biomolecules found in microorganisms can vary depending on growth conditions (see below), the most detailed analysis of the biochemical composition of a microbial cell, Escherichia coli (Battley, 1991), is used here as a model for microbial composition. Briefly, Battley (1991) reports the proportions of individual amino acids, nucleotides, lipids, saccharides, amines and other compounds, mostly from E. coli B/r in exponential phase growth on glucose minimal media corrected for the presence of glycogen. These biomolecules and the proportions in which they are found in E. coli, as modified by McCollom and Amend (2005), are listed in Table 1. Values of ΔGsynth are calculated using
) and the energetics associated with polymerization of these monomers into biomacromolecules (for example, proteins, DNA, polysaccharides). Although the types and amounts of biomolecules found in microorganisms can vary depending on growth conditions (see below), the most detailed analysis of the biochemical composition of a microbial cell, Escherichia coli (Battley, 1991), is used here as a model for microbial composition. Briefly, Battley (1991) reports the proportions of individual amino acids, nucleotides, lipids, saccharides, amines and other compounds, mostly from E. coli B/r in exponential phase growth on glucose minimal media corrected for the presence of glycogen. These biomolecules and the proportions in which they are found in E. coli, as modified by McCollom and Amend (2005), are listed in Table 1. Values of ΔGsynth are calculated using

where ΔGr,i stands for the Gibbs energy of the reaction describing the synthesis of the ith biomolecule,  represents the number of moles of biomolecule i in a dry gram of bacterial cells (Table 1) and ΔGpoly denotes the Gibbs energy required to polymerize biomolecules into their respective biomacromolecules.
represents the number of moles of biomolecule i in a dry gram of bacterial cells (Table 1) and ΔGpoly denotes the Gibbs energy required to polymerize biomolecules into their respective biomacromolecules.
The energy calculations are normalized per dry gram of biomass, J (g cell)−1, because cell masses and the proportions of biomacromolecules in them vary considerably (see below). The energy required to form all the biomacromolecules that constitute a dry gram of microbial cells, ΔGpoly, was calculated using the energy that it takes to form all of the polypeptides in a dry gram of cells (Amend et al., 2013) and the following assumptions. Since 55% of the dry weight of a bacterial cell (E. coli) is protein and nearly all of the biomolecules listed in Table 1 are thought to be in biopolymers such as RNA, DNA, lipopolysaccharides and peptidoglycan (Battley, 1991), it was assumed that the energy required to produce polypeptides from amino acids is proportional to that for polymerizing the other biomacromolecules. This is based on the fact that biopolymerization reactions are dehydration reactions, the energetics of which should not change much between one set of biomolecules and another. That is, if 191 J is required to polymerize all of the amino acids into protein in a dry gram of cells at 25 °C and 0.1 MPa (Amend et al., 2013), protein comprises 55% of the dry weight of a single cell and 45% of the rest of the dry weight is other polymers, then 156 J (g cells)−1=(45/55) × 191 J (g cells)−1 is required to polymerize all the RNA, DNA, lipopolysaccharides and peptidoglycan in a cell.
Values of ΔGr,i and ΔGpoly were calculated as a function of temperature and pressure using

where  and Qr refer to the standard molal Gibbs energy and the reaction quotient of the indicated reaction, respectively, R represents the gas constant and T denotes temperature in Kelvin. Values of
and Qr refer to the standard molal Gibbs energy and the reaction quotient of the indicated reaction, respectively, R represents the gas constant and T denotes temperature in Kelvin. Values of  are calculated as a function of temperature and pressure using the revised-HKF equations of state (Helgeson et al., 1981; Tanger and Helgeson, 1988; Shock et al., 1992), the SUPCRT92 software package (Johnson et al., 1992) and thermodynamic data taken from Shock and Helgeson (1988); Shock et al. (1989); Shock and Helgeson (1990); Shock (1995); Sverjensky et al. (1997); Amend and Helgeson (1997a, b); Amend and Plyasunov (2001); Schulte et al. (2001); Dick et al. (2006); LaRowe and Helgeson (2006) and LaRowe and Dick (2012). Values of Qr are calculated using
are calculated as a function of temperature and pressure using the revised-HKF equations of state (Helgeson et al., 1981; Tanger and Helgeson, 1988; Shock et al., 1992), the SUPCRT92 software package (Johnson et al., 1992) and thermodynamic data taken from Shock and Helgeson (1988); Shock et al. (1989); Shock and Helgeson (1990); Shock (1995); Sverjensky et al. (1997); Amend and Helgeson (1997a, b); Amend and Plyasunov (2001); Schulte et al. (2001); Dick et al. (2006); LaRowe and Helgeson (2006) and LaRowe and Dick (2012). Values of Qr are calculated using

where ai stands for the activity of the ith species and νi corresponds to the stoichiometric coefficient of the ith species in the reaction of interest. The thermodynamic properties and revised-HKF equation of state parameters for some of the biomolecules considered in this study were estimated using existing algorithms (McCollom and Amend, 2005; Amend and McCollom, 2009). These algorithms are reproduced in the Supplementary Information along with a table of the resulting standard molal thermodynamic properties of these compounds at 25 °C and 0.1 MPa and the revised-HKF equation of state parameters required to calculate these properties as a function of temperature and pressure (Supplementary Table S1).
The maintenance energy in Equation (1) is calculated using

where td/r refers to the amount of time that a cell or population takes to double or replace all of its biomass and Pd stands for the maintenance power of the cell or population in units of Watts (for example, W cell−1 or W cm−3). To compare properly the energetics of maintenance, ΔGmain, and biomass synthesis, ΔGsynth, the values of ΔGsynth must be converted from units of kJ per gram of cell biomass (kJ (g cell)−1) into the same units of ΔGmain, J cell−1. This requires knowing the number of cells in a dry gram of cellular biomass and the mass of an individual dry cell. In the literature, masses of microorganisms are typically given in units of fg C cell−1, but to transform carbon masses into total cell masses, the stoichiometry of the major elements in a cell must be known:

It should be noted that the values of the mass of carbon in microbial cells,  , and the stoichiometry of biomass,
, and the stoichiometry of biomass,  and
and  , required to evaluate Equation (6) differ significantly as a function of several variables such as species type, growth stage, nutrient limitation and habitat (see Supplementary Information). A review of the literature reveals that cell carbon masses in bacteria range from at least 3 to 310 fg C cell−1, while cellular stoichiometries result in average nominal carbon oxidation states, NOSC, from +0.89 to −0.45 (LaRowe and Van Cappellen, 2011). Using the lowest estimated cell carbon mass, 3 fg C cell−1 (Vrede et al., 2002), the stoichiometry of E. coli given by Rittman and McCarty (2001) and Equation (6), the total dry mass of a very small cell would be 5.7 fg. On the other hand, using the largest estimated carbon mass, 310 fg C cell−1 (Bratbak, 1985), the cellular stoichiometry given by Redfield et al. (1963) and Equation (6), the resulting dry mass of a large bacterial cell is 865 fg. Here, we use a dry cell mass of 122 fg cell−1. This corresponds to a stoichiometry C5H7NO2 (Rittman and McCarty, 2001) and a cell carbon mass of 65 fg C cell−1 (Hoehler and Jørgensen, 2013), which reflect a weighted average of those reported in the literature.
, required to evaluate Equation (6) differ significantly as a function of several variables such as species type, growth stage, nutrient limitation and habitat (see Supplementary Information). A review of the literature reveals that cell carbon masses in bacteria range from at least 3 to 310 fg C cell−1, while cellular stoichiometries result in average nominal carbon oxidation states, NOSC, from +0.89 to −0.45 (LaRowe and Van Cappellen, 2011). Using the lowest estimated cell carbon mass, 3 fg C cell−1 (Vrede et al., 2002), the stoichiometry of E. coli given by Rittman and McCarty (2001) and Equation (6), the total dry mass of a very small cell would be 5.7 fg. On the other hand, using the largest estimated carbon mass, 310 fg C cell−1 (Bratbak, 1985), the cellular stoichiometry given by Redfield et al. (1963) and Equation (6), the resulting dry mass of a large bacterial cell is 865 fg. Here, we use a dry cell mass of 122 fg cell−1. This corresponds to a stoichiometry C5H7NO2 (Rittman and McCarty, 2001) and a cell carbon mass of 65 fg C cell−1 (Hoehler and Jørgensen, 2013), which reflect a weighted average of those reported in the literature.
In cases where values of ΔGsynth are negative, an infinite number of biomolecules could be made. To make sense of these negative values of ΔGsynth and the large range of cell masses that seem to exist, energy-based yield coefficients have been developed, which are consistent with a recently proposed bioenergetic model that posits that any power available above and beyond the power required for maintenance has the potential to become new biomass (LaRowe and Amend, 2015a). The yield coefficient, which relates this excess power to biomass production, is based on the energy required to synthesize biomolecules, and calculated using

Range of environmental conditions
In this study, we evaluate the effects of temperature (0–125 °C), pressure (0.1–500 MPa) and composition on the energetics of biomolecular synthesis. We note that composition can influence the energetics of anabolism in a number of different ways, with the identities of the C, N and S sources having the greatest impact (see McCollom and Amend (2005) and below). We consider the full range of oxidation states in sources of C (CO2, acetate, CH4), N ( and
and  ) and S (
) and S ( and HS−). Because phosphorous is typically found only in the +5 oxidation state, we use only phosphate in the reactions describing biomass synthesis. To focus on the effects of differing redox states on the energetics of anabolism, we fixed the activities of the C-, N-, S- and P-source compounds (as well as all other reactants and products, see Table 2).
and HS−). Because phosphorous is typically found only in the +5 oxidation state, we use only phosphate in the reactions describing biomass synthesis. To focus on the effects of differing redox states on the energetics of anabolism, we fixed the activities of the C-, N-, S- and P-source compounds (as well as all other reactants and products, see Table 2).
The prevailing oxidation state of the host environment also dictates anabolism energetics. For instance, in a relatively oxidizing environment, such as surface waters, any organism that fixes CO2,  and
and  into biomass must acquire a sufficient stream of electrons to reduce these compounds to the common oxidation states in biomolecules (−3 for N, −2 for S, between +2 and −2 for C). On the other end of the redox spectrum, organisms that fix CH4,
into biomass must acquire a sufficient stream of electrons to reduce these compounds to the common oxidation states in biomolecules (−3 for N, −2 for S, between +2 and −2 for C). On the other end of the redox spectrum, organisms that fix CH4,  and HS− into biomass would not have to reduce their N and S sources, and would have to partially oxidize their carbon source. Let us examine these two scenarios with the synthesis of the amino-acid cysteine (C3H7NO2S), in Reaction (8) from the most oxidized and in Reaction (9) from the most reduced set of precursor compounds:
and HS− into biomass would not have to reduce their N and S sources, and would have to partially oxidize their carbon source. Let us examine these two scenarios with the synthesis of the amino-acid cysteine (C3H7NO2S), in Reaction (8) from the most oxidized and in Reaction (9) from the most reduced set of precursor compounds:

and

Note that the free electrons in these half-reactions are on opposite sides in Reactions (8) and (9).
The quantitative impact of the redox state of the environment on the energetics of electron transfer reactions (including biomass synthesis) is commonly assessed using the activity of electrons ( ), even though there are no free, hydrated electrons in solution (Truesdell, 1968). Values of
), even though there are no free, hydrated electrons in solution (Truesdell, 1968). Values of  are directly related to the redox potential of an environment, typically reported as Eh (mV), through a version of the Nernst equation,
are directly related to the redox potential of an environment, typically reported as Eh (mV), through a version of the Nernst equation,

where  , F stands for the Faraday constant, and R and T are as defined above. The range of Eh values considered in this study is +0.858 V to −0.384 V, corresponding to the most oxidizing and reducing natural settings in which microorganisms have been found—adjusted to pH 7 because the energetics of the biomolecule synthesis reactions have been calculated for pH 7. More specifically, +0.858 mV corresponds to O2-saturated fresh water at 0.01 °C and pH 7. The most reducing value, −0.384 V, has been converted from an ORP measurement of −0.656 V at 25 °C and pH 11.6 taken at the Cedars alkaline spring in northern California, United States of America (Morrill et al., 2013). It should be noted that for a given concentration of O2(aq), calculated values of Eh are a function of pH and temperature. For example, if the O2(aq) concentration in freshwater is at the saturation limit but in a pH 1 fluid (instead of pH 7), then the value of Eh at 0.01 °C would be +1.18 V (instead of +0.858). Furthermore, it should be noted that other redox state scales, such as those using the fugacities of H2 and O2, can be easily converted to values of pe using Equation (10) and the thermodynamic properties of the appropriate half-reactions (see McCollom and Amend, 2005; Dick, 2009; Dick and Shock, 2011).
, F stands for the Faraday constant, and R and T are as defined above. The range of Eh values considered in this study is +0.858 V to −0.384 V, corresponding to the most oxidizing and reducing natural settings in which microorganisms have been found—adjusted to pH 7 because the energetics of the biomolecule synthesis reactions have been calculated for pH 7. More specifically, +0.858 mV corresponds to O2-saturated fresh water at 0.01 °C and pH 7. The most reducing value, −0.384 V, has been converted from an ORP measurement of −0.656 V at 25 °C and pH 11.6 taken at the Cedars alkaline spring in northern California, United States of America (Morrill et al., 2013). It should be noted that for a given concentration of O2(aq), calculated values of Eh are a function of pH and temperature. For example, if the O2(aq) concentration in freshwater is at the saturation limit but in a pH 1 fluid (instead of pH 7), then the value of Eh at 0.01 °C would be +1.18 V (instead of +0.858). Furthermore, it should be noted that other redox state scales, such as those using the fugacities of H2 and O2, can be easily converted to values of pe using Equation (10) and the thermodynamic properties of the appropriate half-reactions (see McCollom and Amend, 2005; Dick, 2009; Dick and Shock, 2011).
Results and discussion
Synthesis energy, ΔGsynth
Values of ΔGsynth associated with the synthesis of 1 g of microbial biomass are shown in Figure 1 as a function of temperature under very oxidizing (+0.858 V, red lines) and reducing (−0.384 V, blue lines) conditions. The different panels reflect different sources of C, N and S. It can be seen quantitatively how different combinations of redox state and biomolecular precursor compounds influence the energetics of growth. It should be emphasized that values of ΔGsynth are the sum of the 46 monomer synthesis reactions (Table 1) in the proportions that they exist in a bacterial cell. Values of ΔGr,i are positive (endergonic) for some and negative (exergonic) for others. That is, just because the overall value of ΔGsynth is negative does not mean that no energy is required to make the biomolecules that constitute a cell. Additionally, it cannot be said with certainty whether or not some of the exergonic biosynthetic reactions considered here can be coupled to endergonic ones. To minimize our assumptions, we think of exergonic anabolic reactions simply as decreasing the cost of biosynthesis and freeing up energy for those that are endergonic.
Gibbs energies of the synthesis, ΔGsynth, of all of the biomolecules listed in Table 1 in the proportions that they exist in E. coli from the sources of carbon, nitrogen and sulfur indicated in each panel, including the energetics of polymerization, as a function of temperature (see Equation (2)). For each combination of C, N and S sources, values of ΔGsynth were computed at two redox states, +0.858 V (oxidizing) and −0.384 V (reducing), as labeled in each panel.
Our results show that values of ΔGsynth depend strongly on the combination of the redox state of the precursor compounds and that of the environment, and weakly on temperature. Overall, values of ΔGsynth vary by 39 kJ (g cell)−1 with the most energy-demanding conditions occurring when the environment is oxidizing and CO2,  and
and  are the precursor compounds (+25 (g cell)−1) and the most conducive when CH4,
are the precursor compounds (+25 (g cell)−1) and the most conducive when CH4,  and HS− are coupled with oxidizing conditions (−14 kJ (g cell)−1). When CO2 is the carbon source, the amount of energy required to build a gram of cells is larger under oxidizing than under reducing conditions (Figures 1a and b). However, this difference in ΔGsynth in Figures 1a and b shrinks when
and HS− are coupled with oxidizing conditions (−14 kJ (g cell)−1). When CO2 is the carbon source, the amount of energy required to build a gram of cells is larger under oxidizing than under reducing conditions (Figures 1a and b). However, this difference in ΔGsynth in Figures 1a and b shrinks when  and
and  are replaced by
are replaced by  and HS−, respectively. When acetate provides C, the same trend is observed as when CO2 is the carbon source, but the gap between the oxidized and reduced scenarios is much smaller, particularly when
and HS−, respectively. When acetate provides C, the same trend is observed as when CO2 is the carbon source, but the gap between the oxidized and reduced scenarios is much smaller, particularly when  and HS− supply N and S (Figures 1c and d). Opposite of CO2 and acetate, biomass synthesis under reducing conditions requires more energy in reducing environments than in oxidizing ones when CH4 is the carbon supply (Figures 1e and f).
and HS− supply N and S (Figures 1c and d). Opposite of CO2 and acetate, biomass synthesis under reducing conditions requires more energy in reducing environments than in oxidizing ones when CH4 is the carbon supply (Figures 1e and f).
The impact of pressure on ΔGsynth is negligible. This is illustrated in Figure 2, where acetate,  and HS− are the sources of C, N and S under oxidizing conditions (+0.858 V) at two pressures (Psat for the red line and 0.5 GPa for the dashed black line). Note the very minor difference in ΔGsynth at these two pressures. It should be noted that pressure can influence other variables that in turn have an impact on ΔGsynth. For instance, at higher pressures, gases are generally more soluble, resulting in changes to the reaction quotient (Q) in the computation of Gibbs energies (see Equation (3)).
and HS− are the sources of C, N and S under oxidizing conditions (+0.858 V) at two pressures (Psat for the red line and 0.5 GPa for the dashed black line). Note the very minor difference in ΔGsynth at these two pressures. It should be noted that pressure can influence other variables that in turn have an impact on ΔGsynth. For instance, at higher pressures, gases are generally more soluble, resulting in changes to the reaction quotient (Q) in the computation of Gibbs energies (see Equation (3)).
Comparison of the Gibbs energies of the synthesis, ΔGsynth, of all of the biomolecules listed in Table 1 in the proportions that they exist in E. coli from acetate (CH3COO−),  and HS− at +0.858 V, including the energetics of polymerization, as a function of temperature at two extreme values of pressure, saturation pressure and 5000 bars (0.5 GPa). Saturation pressure corresponds to the minimum pressure required to prevent water from boiling. Values of ΔGsynth are not shown below 25 °C at 5000 bars because the equation of state for water used in these calculations is not valid under these conditions.
and HS− at +0.858 V, including the energetics of polymerization, as a function of temperature at two extreme values of pressure, saturation pressure and 5000 bars (0.5 GPa). Saturation pressure corresponds to the minimum pressure required to prevent water from boiling. Values of ΔGsynth are not shown below 25 °C at 5000 bars because the equation of state for water used in these calculations is not valid under these conditions.
Although values of ΔGpoly are small (<0.4 kJ (g cell)−1), the proportion of ΔGpoly to ΔGr,i varies considerably. For instance, the energy required to synthesize biomolecules from CO2,  and
and  under oxidizing conditions at 0.01 °C (red line, Figure 1a) is about 64 times larger than the energetics of polymerizing these biomolecules at the same temperature. However, when instead,
under oxidizing conditions at 0.01 °C (red line, Figure 1a) is about 64 times larger than the energetics of polymerizing these biomolecules at the same temperature. However, when instead,  and HS− are the sources of N and S and the environmental conditions are reducing (blue line, Figure 1b), values of ΔGpoly are essentially equal to those of ΔGr,i between 25 and 40 °C. Although these examples represent the extremes of the ΔGr,i: ΔGpoly ratio considered in this study, it illustrates how variable the burden of polymerization energetics can be for biomass synthesis.
and HS− are the sources of N and S and the environmental conditions are reducing (blue line, Figure 1b), values of ΔGpoly are essentially equal to those of ΔGr,i between 25 and 40 °C. Although these examples represent the extremes of the ΔGr,i: ΔGpoly ratio considered in this study, it illustrates how variable the burden of polymerization energetics can be for biomass synthesis.
Maintenance energy, ΔGmain
Maintenance energy is generally defined as the energy consumed by organisms that does not result in new biomass. Values of maintenance power (Pd) vary by over five orders of magnitude depending on the metabolism, species and experimental procedure (LaRowe and Amend, 2015a)—values determined while the microorganisms of interest were growing. In natural settings, maintenance powers are likely much smaller than those measured in the laboratory, in part because laboratory growth experiments must be conducted at relatively high energy levels to be observable (Hoehler and Jørgensen, 2013). To better describe maintenance energy in natural, low-energy settings, Hoehler and Jørgensen (2013) coined the term ‘basal maintenance power’ for the power used by microorganisms to remain viable. Although basal maintenance power is difficult to measure directly, recent calculations suggest that it could be two orders of magnitude lower compared with the lowest reported values of Pd in the literature (LaRowe and Amend, 2015a, b). Hence, we must first define whether organisms are growing in typical high-energy laboratory-like conditions or simply subsisting in natural low-power settings to correctly calculate values of ΔGmain.
The contribution of turnover/doubling time to laboratory-like and natural, low-energy maintenance powers is illustrated in Figure 3 by displaying values of ΔGmain, for two cases. The first case, indicated by the solid line, refers to values of ΔGmain calculated using Equation (8) and a value of Pd for aerobic heterotrophy equal to 4.9 × 10−14 J s−1, the median value for this type of metabolism collected by LaRowe and Amend (2015a). As the doubling time for this aerobic heterotroph approaches 1 month, the amount of energy required for maintenance reaches 1.3 × 10−7 J cell−1, an amount of energy that exceeds the amount of energy required to synthesize all of its biomolecules from CO2 or acetate serving as the carbon source for microorganism the size of E. coli (see below). For an organism that is not growing, or is doing so in a very low-energy environment, and thus subsisting at its basal Pd, far less energy goes into ΔGmain. This scenario is shown with the dashed line in Figure 1. A basal maintenance power of 1.9 × 10−19 J s−1 is used in conjunction with Equation (8) to generate this line, a value of Pd that is two order of magnitude lower compared with the lowest reported maintenance energy in the literature (Marschall et al., 2010). An organism existing in this energy state would only use 6 × 10−9 J cell−1 over the course of 1000 years, more than two orders of magnitude less energy than the aerobic heterotroph uses for ΔGmain in 1 month.
Maintenance energy, ΔGmain, as a function of time calculated using Equation (5) and a relatively high maintenance power, Pd (the median value for aerobic heterotrophy as collected by LaRowe and Amend, 2015a), of 4.9 × 10−14 J s−1 (solid line) and a basal maintenance power that is two orders of magnitude lower compared with the lowest Pd reported in the literature (Marschall et al., 2010), 1.9 × 10−19 J s−1.
Biomass yield
The number of cells that can be made per Joule of Gibbs energy, Y, are listed in Table 3 in 25 °C intervals. Values of Y were calculated using Equation (7), a cell mass of 122 fg cell−1 and the positive contributions to ΔGsynth. Only positive values of ΔGr,i and ΔGpoly are used because at least some of the biomolecule synthesis reactions are exergonic, and should not be added to the positive ones since doing so effectively assumes that the exergonic synthesis reactions are coupled to the endergonic ones; there is no evidence that this is the case. The yield coefficients listed in Table 3, mirror the large-scale pattern seen in values of ΔGsynth in Figure 1 (see Supplementary Figure S1 for plots analogous to Figure 1). For combinations of redox states and molecular sources of C, N and S that favor biomolecule synthesis (ΔGsynth<0), yield coefficients are large (for example, 2.36 × 1010 cells J−1 for an oxidizing environment and CH4,  and HS− are the sources of C, N and S), while those with positive values of ΔGsynth are considerably smaller (for example, 3.69 × 108 cells J−1 in an oxidizing system and CO2,
and HS− are the sources of C, N and S), while those with positive values of ΔGsynth are considerably smaller (for example, 3.69 × 108 cells J−1 in an oxidizing system and CO2,  and
and  as the sources of C, N and S). Values of Y under oxidizing conditions show little variation as a function of temperature, whereas those calculated at the lower redox state vary more. This variation is most pronounced for the two cases in which CO2 is the source of carbon. The decreasing yield coefficients with temperature are due to the fact that many of the biomolecule synthesis reaction involving CO2 switch from being exergonic at low temperatures to being endergonic at higher temperatures.
as the sources of C, N and S). Values of Y under oxidizing conditions show little variation as a function of temperature, whereas those calculated at the lower redox state vary more. This variation is most pronounced for the two cases in which CO2 is the source of carbon. The decreasing yield coefficients with temperature are due to the fact that many of the biomolecule synthesis reaction involving CO2 switch from being exergonic at low temperatures to being endergonic at higher temperatures.
It should be kept in mind that the analysis presented above is based on a fixed cell size of 122 fg. In situations in which this mass does not represent the cell mass of interest, Equation (6) can be used to generate mass-appropriate yield coefficients. For instance, if the full range of cell masses computed above, 5.7 to 865 fg cell−1, are combined with the largest (22 200 J (g cell−1)) and smallest (347 J (g cell−1)) values of ΔGsynth at 25 °C, and Equation (7), values of Y would differ by a factor of nearly 9700, 5.21 × 107−5.06 × 1011 cell J−1.
Time and energy
Although the calculations summarized above indicate that the number of microbial cells that can be made from per Joule could vary considerably, this is only the static contribution to ΔGcell (Equation 1). For cells that have large maintenance powers and long doubling/replacement times, the resulting values of ΔGmain could dwarf those of ΔGsynth, which is what the yield coefficients are based on. The importance of doubling/replacement time on the energy budget of a microorganism can be illustrated by comparing values of 1/Y (J cell−1) with ΔGmain. As an example, 1/Y for the most favorable biomolecule synthesis conditions considered in this study (CO2,  ,
,  , reducing at 25 °C) is 4.2 × 10−11 J cell−1 (assuming 122 fg cell−1) and the lowest ΔGmain reported in the literate, 1.9 × 10−17 J s−1. If a cell characterized by these growth parameters turned over its biomass over the course of 26 days, the cumulative maintenance energy would surpass the energy required for making all of its biomolecules. For the largest maintenance power, Pd, collected in the review carried out by LaRowe and Amend (2015a), 4.7 × 10−12 J s−1, and the same value of 1/Y, it would only take 9 s for the cumulative ΔGmain to surpass the energy required for making all of its biomolecules, far less time than the shortest doubling time reported in the literature (9.8 min; Eagon, 1962). If values of Pd in natural, low-energy setting are two orders of magnitude lower compared with the lowest reported in the literature, as suggested by Hoehler and Jørgensen (2013), then even a doubling/replacement time of 7.1 years would result in a value of ΔGmain equal to that of ΔGmain under the most favorable set of synthesis conditions considered in this study.
, reducing at 25 °C) is 4.2 × 10−11 J cell−1 (assuming 122 fg cell−1) and the lowest ΔGmain reported in the literate, 1.9 × 10−17 J s−1. If a cell characterized by these growth parameters turned over its biomass over the course of 26 days, the cumulative maintenance energy would surpass the energy required for making all of its biomolecules. For the largest maintenance power, Pd, collected in the review carried out by LaRowe and Amend (2015a), 4.7 × 10−12 J s−1, and the same value of 1/Y, it would only take 9 s for the cumulative ΔGmain to surpass the energy required for making all of its biomolecules, far less time than the shortest doubling time reported in the literature (9.8 min; Eagon, 1962). If values of Pd in natural, low-energy setting are two orders of magnitude lower compared with the lowest reported in the literature, as suggested by Hoehler and Jørgensen (2013), then even a doubling/replacement time of 7.1 years would result in a value of ΔGmain equal to that of ΔGmain under the most favorable set of synthesis conditions considered in this study.
The cumulative amount of energy that would be needed to maintain a viable cell over increasingly long time spans makes the overall cost of doubling or completely replacing cellular components a greater proportion of ΔGcell. However, this assumes that the values of Pd are constant, which is unlikely. In low-energy environments, the basal maintenance power is likely quite low for long periods of time, and then during growth, it likely increases. An additional point should be made about Equation (1). Although it splits microbial energy usage into a simple dichotomy, it is not possible to completely separate the energies of biomolecule synthesis from maintenance. Not only are numerous so-called maintenance functions required to make biomolecules, such as the conversion of catabolic energy into electron donors such as NADP, and the active transport of metabolites, but some maintenance energy is expended on biomolecule synthesis and/or degradation via changes in stored polymeric carbon, extracellular secretions and the synthesis and turnover of macromolecules such as proteins and RNA (van Bodegom, 2007).
Concluding remarks
The results presented in this communication are intended to illustrate that a quantitative link can be made connecting the physiochemical properties of a natural setting (for example, temperature and composition) to the amount of biomass that exists in it. As noted above, the amount of energy available for microbial purposes is a critical component of this connection, and, as a result, the energy supply in a wide variety of settings has been determined (for example, Amend and Shock, 1998, 2000; McCollom and Amend, 2005; LaRowe and Regnier, 2008; Amend and McCollom, 2009; Amend et al., 2011, 2013; LaRowe and Amend, 2014; Osburn et al., 2014; Teske et al., 2014; Price et al., 2015). In this contribution, the microbial energy demand associated with building biomass under a broad range of conditions has been calculated. Because of the central role of oxidation–reduction processes in living organisms, this quantification is largely based on the energy associated with changing the redox state of C, N and S source compounds into biomolecules as a function of the oxidation–reduction state of the environment. In addition, cell yield coefficients have been reported that take the same environmental conditions into account. Furthermore, the impact of doubling/replacement time on the energetics of making a cell are included in the analysis to show that this may be the largest component of the microbial energy budget in settings that are not characterized by vigorous growth. Taken together, the calculations summarized above can be used to quantify the energy limits for life, and shed light on the conditions that supported the origin of life.
Finally, the results reported in this study support the notion that, from a thermodynamic perspective, the abiotic origin of biomolecules, within the context of terrestrial planetary bodies, does not require high temperatures, but oxidation states that create favorable conditions to synthesize biomolecules from reduced and oxidized precursors depending on the redox state of precursor molecules. This notion, which is illustrated in most of the panels in Figure 1, has been a central tenet of some theories regarding the origin of life (for example, Russell and Hall, 1997; Martin and Russell 2003; Martin et al., 2008; Lane et al., 2010). The panels in this figure show that the sum of the Gibbs energies associated with synthesizing the core biomolecules that make up microorganisms can be negative, even at low temperature, for every combination of C, N and S sources except for those shown in Figure 1d. Although hydrothermal systems have been implicated in the origin of life, it is mostly the rapid mixing of fluids characterized by two very different redox states that promote the energetics of biomolecule synthesis rather than temperature, per se. For instance, Amend and Shock (1998) show that abiotic amino-acid synthesis is more thermodynamically favorable at 100 °C than at 2 °C, but it is the much more reducing conditions at the higher temperature that are the major driving force for biomass synthesis in this example. This is substantiated by Amend et al. (2011), who show that the optimal conditions for synthesizing biomolecules because of mixing of cold seawater and 12 hydrothermal fluids is between 11.2 and 32 °C. Similarly, LaRowe and Regnier (2008) showed that abiotic nucleotide synthesis is thermodynamically possible at low temperatures. Even the polymerization of amino acids into polypeptides is more favored at lower temperature (Amend et al. 2013 and Figure 1) compared with higher ones.
References
Amend JP, Helgeson HC . (1997a). Group additivity equations of state for calculating the standard molal thermodynamic properties of aqueous organic species at elevated temperatures and pressures. Geochim Cosmochim Acta 61: 11–46.
Amend JP, Helgeson HC . (1997b). Solubilities of the common l-α-amino acids as a function of temperature and solution pH. Pure Appl Chem 9:: 35 -–942.
Amend JP, McCollom TM . (2009) Energetics of biomolecule synthesis on early Earth. In: Zaikowski L, Friedrich JM, Seidel SR (eds), Chemical Evolution II: From the Origins of Life to Modern Society. American Chemical Society: Washington DC, USA.
Amend JP, Plyasunov AV . (2001). Carbohydrates in thermophile metabolism: calculation of the standard molal thermodynamic properties of aqueous pentoses and hexoses at elevated temperatures and pressures. Geochim Cosmochim Acta 65: 3901–3917.
Amend JP, Shock EL . (1998). Energetics of amino acid synthesis in hydrothermal ecosystems. Science 281: 1659–1662.
Amend JP, Shock EL . (2000). Thermodynamics of amino acid synthesis in hydrothermal ecosystems on the early Earth. In: Goodfriend G (ed), Perspectives in Amino Acid and Protein Geochemistry. Plenum Press: New York, NY, USA.
Amend JP, Teske A . (2005). Expanding frontiers in deep subsurface microbiology. Palaeogeogr Palaeoclimatol Palaeoecol 219: 131–155.
Amend JP, LaRowe DE, McCollom TM, Shock EL . (2013). The energetics of organic synthesis inside and outside the cell. Philos Trans R Soc Ser B 368: 1–15.
Amend JP, McCollom TM, Hentscher M, Bach W . (2011). Catabolic and anabolic energy for chemolithoautotrophs in deep-sea hydrothermal systems hosted in different rock types. Geochim Cosmochim Acta 75: 5736–5748.
Amy PS, Haldeman DL (eds) (1997). The Microbiology of the Terrestrial Deep Subsurface. CRC Lewis Publishers: Boca Raton, FL, USA.
Battley EH . (1991). An alternative method of calculating the heat of growth of Escherichia coli K-12 on succinic acid. Biotechnol Bioeng 38: 480–492.
Bratbak G . (1985). Bacterial biovolume and biomass estimations. Appl Environ Microbiol 49: 1488–1493.
Canganella F, Wiegel J . (2011). Extremophiles: from abyssal to terrestrial ecosystems and possible beyond. Naturwissenschaften 98: 253–279.
Colwell FS, D’Hondt S . (2013). Nature and extent of the deep biosphere. Rev Miner Geochem 75: 547–574.
Dick JM . (2009). Calculation of the relative metastabilities of proteins in subcellular compartments of Saccharomyces cerevisiae. BMC Syst Biol 3: Article 75.
Dick JM, Shock EL . (2011). Calcualtion of the relative chemical stabilities of proteins as a function of temperature and redox chemistry in a hot spring. PLoS One 6: e22782.
Dick JM, LaRowe DE, Helgeson HC . (2006). Temperature, pressure and electrochemical constraints on protein speciation: group additivity calculation of the standard molal thermodynamic properties of ionized unfolded proteins. Biogeosciences 3: 311–336.
Eagon RG . (1962). Pseudomonas natriegens, a marine bacterium with a generation time of less than 10 minutes. J Bacteriol 83: 736–737.
Edwards KJ, Becker K, Colwell F . (2012). The deep, dark energy biosphere: Intraterrestrial life on Earth. Annu Rev Earth Planet Sci 40: 551–568.
Fredrickson JK, Fletcher M (eds) (2001). Subsurface Microbiology and Biogeochemistry. Wiley: New York, NY, USA.
Heijnen JJ, van Dijken JP . (1992). In search of a thermodynamic description of biomass yields for the chemotrophic growth of microorganisms. Biotechnol Bioeng 39: 833–858.
Helgeson HC, Kirkham DH, Flowers GC . (1981). Theoretical prediction of thermodynamic behavior of aqueous electrolytes at high pressures and temperatures: 4. Calculation of activity coefficients, osmotic coefficients, and apparent molal and standard and relative partial molal properties to 600°C and 5 kb. Am J Sci 281: 1249–1516.
Hoehler TM, Jørgensen BB . (2013). Micorbial life under extreme energy limitation. Nat Rev Microbiol 11: 83–94.
Johnson JW, Oelkers EH, Helgeson HC . (1992). SUPCRT92—a software package for calculating the standard molal thermodynamic properties of minerals, gases, aqueous species, and reactions from 1 bar to 5000 bar and 0°C to 1000°C. Comput Geosci 18: 899–947.
Lane N, Allen JF, Martin W . (2010). How did LUCA make a living? Chemiosmosis in the origin of life. BioEssays 32: 271–280.
LaRowe DE, Amend JP . (2014). Energetic constraints on life in marine deep sediments. In: Kallmeyer J, Wagner K (eds). Life in Extreme Environments: Microbial Life in the Deep Biosphere. de Gruyter: Berlin, Germany.
LaRowe DE, Amend JP . (2015a). Catabolic rates, population sizes and doubling/replacement times of microorganisms in the natural settings. Am J Sci 315: 167–203.
LaRowe DE, Amend JP . (2015b). Power limits for microbial life. Front Extr Microbiol 6: Article 718; doi:10.3389/fmicb.2015.00718.
LaRowe DE, Dick JM . (2012). Calculation of the standard molal thermodynamic properties of crystalline proteins. Geochim Cosmochim Acta 80: 70–91.
LaRowe DE, Helgeson HC . (2006). Biomolecules in hydrothermal systems: calculation of the standard molal thermodynamic properties of nucleic-acid bases, nucleosides, and nucleotides at elevated temperatures and pressures. Geochim Cosmochim Acta 70: 4680–4724.
LaRowe DE, Regnier P . (2008). Thermodynamic potential for the abiotic synthesis of adenine, cytosine, guanine, thymine, uracil, ribose and deoxyribose in hydrothermal systems. Orig Life Evol Bios 38: 383–397.
LaRowe DE, Van Cappellen P . (2011). Degradation of natural organic matter: a thermodynamic analysis. Geochim Cosmochim Acta 75: 2030–2042.
Marschall E, Jogler M, Henssge U, Overmann J . (2010). Large-scale distribution and activity patters of an extremely low-light-adapted population of green sulfur bacteria in the Black Sea. Environ Microbiol 12: 1348–1362.
Martin W, Russell MJ . (2003). On the origins of cells: a hypothesis for the evolutionary transitions from abiotic geochemistry to chemoautotrophic prokaryotes, and from prokaryotes to nucleated cells. Philos Trans R Soc Ser B 358: 59–85.
Martin W, Baross J, Kelley D, Russell MJ . (2008). Hydrothermal vents and the origin of life. Nat Rev Microbiol 6: 805–814.
McCollom TM, Amend JP . (2005). A thermodynamic assessment of energy requirements for biomass synthesis by chemolithoautotrophic micro-organisms in oxic and anoxic environments. Geobiology 3: 135–144.
Morrill PL, Kuenen JG, Johnson OJ, Suzuki S, Rietze A, Sessions AL et al. (2013). Geochemistry and geobiology of a present-day serpentinization site in California: the Cedars. Geochim Cosmochim Acta 109: 222–240.
Orcutt BN, Sylvan JB, Knab NJ, Edwards KJ . (2011). Microbial ecology of the dark ocean above, at, and below the seafloor. Microbiol Mol Biol Rev 75: 361–422.
Osburn MR, LaRowe DE, Momper L, Amend JP . (2014). Chemolithotrophy in the continental deep subsurface: Sanford Underground Research Facility (SURF), USA. Front Extr Microbiol 5: Article 610.
Picard A, Daniel I . (2013). Pressure as an environmental parameter for microbial life—a review. Biophys Chem 183: 30–41.
Pikuta EV, Hoover RB, Tang J . (2007). Microbial extremophiles at the limits of life. Crit Rev Microbiol 33: 183–209.
Price RE, LaRowe DE, Italiano F, Savov I, Pichler T, Amend JP . (2015). Subsurface hydrothermal processes and the bioenergetics of chemolithoautotrophy at the shallow-sea vents off Panarea Island (Italy). Chem Geol 407–408: 21–45.
Redfield AC, Ketchum BH, Richards FA . (1963). The influence of organisms on the composition of sea water. In: Hill MN (ed), The Sea, vol. 2. Wiley Interscience: New York, NY, USA.
Rittman BE, McCarty PL . (2001) Environmental Biotechnology: Principles and Applications. McGraw-Hill: New York, NY, USA.
Rothschild LJ, Mancinelli RL . (2001). Life in extreme environments. Nature 409: 1092–1101.
Russell MJ, Hall AJ . (1997). The emergence of life from iron monosulphide bubbles at a submarine hydrothermal redox and pH front. J Geol Soc Lond 154: 377–402.
Schulte MD, Shock EL, Wood R . (2001). The temperature dependence of the standard-state thermodynamic properties of aqueous nonelectrolytes. Geochim Cosmochim Acta 65: 3919–3930.
Shock EL . (1995). Organic acids in hydrothermal solutions—standard molal thermodynamic properties of carboxylic acids and estimates of dissociation constants at high temperatures and pressures. Am J Sci 295: 496–580.
Shock EL, Helgeson HC . (1988). Calculation of the thermodynamic and transport properties of aqueous species at high pressures and temperatures—correlation algorithms for ionic species and equation of state predictions to 5 kb and 1000°C. Geochim Cosmochim Acta 52: 2009–2036.
Shock EL, Helgeson HC . (1990). Calculation of the thermodynamic and transport properties of aqueous species at high pressures and temperatures—standard partial molal properties of organic species. Geochim Cosmochim Acta 54: 915–945.
Shock EL, Helgeson HC, Sverjensky D . (1989). Calculation of the thermodynamic and transport properties of aqueous species at high pressures and temperatures—standard partial molal properties of inorganic neutral species. Geochim Cosmochim Acta 53: 2157–2183.
Shock EL, Holland ME . (2007). Quantitative habitability. Astrobiology 7: 839–851.
Shock EL, Oelkers E, Johnson J, Sverjensky D, Helgeson HC . (1992). Calculation of the thermodynamic properties of aqueous species at high pressures and temperatures—effective electrostatic radii, dissociation constants and standard partial molal properties to 1000°C and 5 kbar. J Chem Soc Faraday Trans 88: 803–826.
Steven B, Leveille R, Pollard WH, Whyte LG . (2006). Microbial ecology and biodiversity in permafrost. Extremophiles 10: 259–267.
Strapoć D, Mastalerz M, Dawson K, Macalady J, Callaghan AV, Wawrik B et al. (2011). Biogeography of micorbial coal-bed methane. Ann Rev Earth Planet Sci 39: 617–656.
Sverjensky D, Shock EL, Helgeson HC . (1997). Prediction of the thermodynamic properties of aqueous metal complexes to 1000°C and 5 kb. Geochim Cosmochim Acta 61: 1359–1412.
Takai K, Nakamura K, LaRowe DE, Amend JP . (2014). Life under subseafloor extremes. In: Stein R, Blackman D, Inagaki F, Larsen H-C (eds), Earth and Life Processes Discovered from Subseafloor Environment—A Decade of Science Achieved by the Integrated Ocean Drilling Program (IODP). Elsevier: Amsterdam.
Tanger JC, Helgeson HC . (1988). Calculation of the thermodynamic and transport properties of aqueous species at high pressures and temperatures—revised equations of state for the standard partial molal properties of ions and electrolytes. Am J Sci 288: 19–98.
Teske A, Callaghan AV, LaRowe DE . (2014). Biosphere frontiers: deep life in the sedimented hydrothermal system of Guaymas Basin. Front Extr Microbiol 5: Article 362.
Truesdell AH . (1968). The advantage of using pE rather than Eh in redox equilibrium calculations. J Geolog Educ 16: 17–20.
van Bodegom P . (2007). Micorbial maintenance: a critical review of its quantification. Microbial Ecol 5: 513–523.
Vrede K, Heldal M, Norland S, Bratbak G . (2002). Elemental composition (C, N, P) and cell volume of exponentially growing and nutrient-limited bacterioplankton. Appl Environ Microbiol 68: 2965–2971.
Acknowledgements
Financial assistance was provided by the Center for Dark Energy Biosphere Investigations (C-DEBI, OCE0939564) and the NASA Astrobiology Institute—Life Underground (NAI-LU, NNA13AA92A). This is C-DEBI contribution 284 and NAI-LU contribution 066. Both organizations are based at the University of Southern California.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
LaRowe, D., Amend, J. The energetics of anabolism in natural settings. ISME J 10, 1285–1295 (2016). https://doi.org/10.1038/ismej.2015.227
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.227
This article is cited by
-
Maintenance power requirements of anammox bacteria “Candidatus Brocadia sinica” and “Candidatus Scalindua sp.”
The ISME Journal (2021)
-
Capturing the genetic makeup of the active microbiome in situ
The ISME Journal (2017)