Abstract
Symbiotic partnerships between heterotrophic hosts and intracellular microalgae are common in tropical and subtropical oligotrophic waters of benthic and pelagic marine habitats. The iconic example is the photosynthetic dinoflagellate genus Symbiodinium that establishes mutualistic symbioses with a wide diversity of benthic hosts, sustaining highly biodiverse reef ecosystems worldwide. Paradoxically, although various species of photosynthetic dinoflagellates are prevalent eukaryotic symbionts in pelagic waters, Symbiodinium has not yet been reported in symbiosis within oceanic plankton, despite its high propensity for the symbiotic lifestyle. Here we report a new pelagic photosymbiosis between a calcifying ciliate host and the microalga Symbiodinium in surface ocean waters. Confocal and scanning electron microscopy, together with an 18S rDNA-based phylogeny, showed that the host is a new ciliate species closely related to Tiarina fusus (Colepidae). Phylogenetic analyses of the endosymbionts based on the 28S rDNA gene revealed multiple novel closely related Symbiodinium clade A genotypes. A haplotype network using the high-resolution internal transcribed spacer-2 marker showed that these genotypes form eight divergent, biogeographically structured, subclade types that do not seem to associate with any benthic hosts. Ecological analyses using the Tara Oceans metabarcoding data set (V9 region of the 18S rDNA) and contextual oceanographic parameters showed a global distribution of the symbiotic partnership in nutrient-poor surface waters. The discovery of the symbiotic life of Symbiodinium in the open ocean provides new insights into the ecology and evolution of this pivotal microalga and raises new hypotheses about coastal pelagic connectivity.
Similar content being viewed by others
Introduction
Symbiosis is central to the evolution and ecology of ecosystems, and is present in virtually all living organisms (Maynard Smith, 1989; Margulis and Fester, 1991). This intimate and long-term association between organisms of different species can range from parasitism, where one partner benefits at the expense of the other, to mutualism, where both partners benefit (de Bary, 1878). Photosymbiosis, defined as a close symbiotic relationship with a photosynthetic partner (generally the symbiont), has led to the acquisition of transient and even permanent photosynthesis in different eukaryotic lineages (Keeling, 2010). In marine and freshwater ecosystems, this type of symbiosis is common (Stoecker et al., 2009) and is considered mutualistic: the symbiont transfers photosynthetic products to its host that, in turn, provides a nutrient-rich microenvironment and protection from parasites and predators (Muscatine et al., 1984; Yellowlees et al., 2008; Davy et al., 2012). One of the best-known examples of marine photosymbiosis involves the photosynthetic dinoflagellate Symbiodinium that lives within a wide diversity of benthic hosts, including metazoans (for example, corals, jellyfishes, anemones, sponges, mollusks) and protists (for example, ciliates and foraminiferans; LaJeunesse, 2001) in tropical reef ecosystems. Symbioses involving Symbiodinium are ecologically and economically important as they sustain the very productive and biologically diverse reef ecosystems worldwide (Stanley, 2006) that in turn provide close to $375 billion in goods and services each year (Costanza et al., 1997). The genus Symbiodinium is genetically diverse, having evolved throughout the Cenozoic into nine divergent lineages or clades (A to I), each containing several subclade genotypes that can have distinct physiological capacities, spatial distribution and host spectra (Rowan, 2004; Coffroth and Santos, 2005; Pochon and Gates, 2010).
Photosymbioses are also pervasive in planktonic ecosystems, especially in tropical and subtropical oligotrophic waters (Taylor, 1982; Stoecker et al., 2009; Decelle et al., 2015), but remain poorly studied compared with benthic ecosystems. Because of their mixotrophic capacity, photosymbiotic protists play a significant dual role as primary producers and predators in pelagic ecosystems (Swanberg and Caron, 1991; Michaels et al., 1995). Many hosts build mineral skeletons of calcium carbonate, silica or strontium sulfate, hence contributing to the biogeochemical cycles of these elements (Bernstein et al., 1987). For instance, the rhizarian Radiolaria and Foraminifera are known to develop obligate photosymbioses in surface waters with diverse eukaryotic microalgae, such as photosynthetic dinoflagellates (for example, Brandtodinium, Pelagodinium) and haptophytes (for example, Phaeocystis) (Gast and Caron, 2001; Shaked and de Vargas, 2006; Decelle et al., 2012; Probert et al., 2014). Some heterotrophic dinoflagellates have also been found to host prasinophyte (Sweeney, 1976) or pelagophyte (Daugbjerg et al., 2013) microalgae. Paradoxically, although several photosynthetic dinoflagellate taxa have been described to be common symbionts in the oceanic plankton, to date the ubiquitous coastal symbiont Symbiodinium has never been found in symbiosis in the pelagic realm.
Ciliates are also known to acquire phototrophy through photosymbiosis with eukaryotic or prokaryotic microalgal cells (Stoecker et al., 2009; Esteban et al., 2010; Dziallas et al., 2012). In freshwater habitats, many ciliate species (for example, Paramecium bursaria, Coleps hirtus viridis) are found in symbiosis with the green alga Chlorella, whereas in marine ecosystems rare examples of photosymbiosis with eukaryotic microalgae have been reported (Stabell et al., 2002; Stoecker et al., 2009; Johnson, 2011). The benthic ciliates Maristentor dinoferus and Euplotes uncinatus host Symbiodinium endosymbionts in coral reefs (Lobban et al., 2002, 2005) and some Oligotrichida ciliates associate with green algae (prasinophytes) in estuarine environments (Stoecker et al., 1988–1989). These photosymbiotic ciliates almost exclusively dwell in coastal waters or benthic habitats, whereas there are virtually no examples of symbiotic partnerships involving eukaryotic microalgae in the surface open ocean. In this study, we used a combination of microscopy and molecular tools to characterize a novel pelagic photosymbiosis between a calcifying ciliate host and Symbiodinium endosymbionts. Exploration of this symbiosis in the samples and metabarcoding data set of the worldwide Tara Oceans expedition (Karsenti et al., 2011; De Vargas et al., 2015) allowed us to study the global specificity, biogeography and ecology of this novel interaction.
Materials and methods
Morphological identification of the host
Plankton samples and metadata used in this study originated from the Tara Oceans expedition (2009–2012) (Karsenti et al., 2011; De Vargas et al., 2015). From ethanol-preserved plankton samples, 20 ciliate cells were isolated and cleaned under microscopy, and individually placed on a polycarbonate filter without any chemical fixatives for scanning electron microscopy observations (Table 1). Additional cells (n=10) from formaldehyde-preserved samples were also imaged with confocal laser scanning microscope (Leica TCS SP8, Leica Microsystems, Wetzlar, Germany), equipped with an HC PL APO 40 × /1.10 W motCORR CS2 objective. Multiple fluorescent dyes were used sequentially to observe the cellular components of the ciliate (host) and the intracellular microalgae (symbionts), such as the nuclei (blue) and the cellular membranes (green) with Hoechst (Ex405/Em420-470) and DiOC6 (Ex488/Em500-520), respectively. Red autofluorescence of the chlorophyll (Ex638/Em680-700) was also visualized to highlight the chloroplasts of the symbiotic microalgae. Image processing and three-dimensional reconstructions were conducted with Fiji (Schindelin et al., 2012) and IMARIS (Bitplane) software (Bitplane AG, Zurich, Switzerland). Live observations in bright-field microscopy of several ciliate cells (>50) from the Mediterranean Sea (Villefranche-sur-Mer, France, and Naples, Italy) were also conducted and two individual cells (Med 2 and Med 3) were specifically sampled for molecular investigation (Table 1).
Phylogenetic identification of the symbiotic partners
Sampling and PCR amplifications
Ciliate cells were isolated from ethanol-preserved surface samples collected by a plankton net (20 μm mesh-size) at five Tara Oceans stations in the Red Sea and Indian, Pacific and Atlantic Oceans (Table 2, and see Pesant et al., 2015 and De Vargas et al., 2015 for more details). In addition, two individual ciliates (Med 2 and Med 3) were also collected from live samples in 2011 in the bay of Villefranche-sur-Mer (Mediterranean Sea, France). Cells observed with scanning electron and confocal laser scanning microscopy were not subjected to molecular studies. Ciliate cells were individually isolated using a glass micropipette, carefully washed in three successive baths of 0.22 μm filtered and sterile sea water, transferred into a sterile microtube and preserved in lysis buffer (Tissue and Cell Lysis Solution from MasterPure DNA and RNA Purification Kit, Epicenter, Madison, WI, USA) at −20 °C. DNA extraction was then performed following the protocol of the MasterPure DNA and RNA purification kit (Epicenter). In order to obtain different phylogenetic ribosomal markers, single-cell PCR amplifications were conducted with the Phusion High-Fidelity DNA Polymerase (Finnzymes, Thermo Fisher Scientific, Waltham, MA, USA). The PCR mixture (25 μl final volume) contained 0.5 ng (1 μl) of DNA, 0.35 μM (final concentration) of each primer, 3% of dimethyl sulfoxide and 2 × of GC buffer Phusion MasterMix (Finnzymes, Thermo Fisher Scientific). Amplifications were conducted in a PCR thermocycler (Applied Biosystems, Thermo Fisher Scientific) with the following PCR program: initial denaturation step at 98 °C for 30 s, followed by 36–38 cycles of 10 s at 98 °C, 30 s at the annealing temperature of the primer sets (Supplementary Table S1), 30 s at 72 °C and final elongation step at 72 °C for 10 min. For the nested PCR approach, the amplicons obtained in the first PCR (with 25 cycles) were reamplified with 40 cycles using internal primers. PCR amplicons were visualized on agarose gels, purified with the ExoStar purification kit (Illustra Exostar 1_Step, GE Healthcare Bio-Sciences Corp., Piscataway, NJ, USA) and Sanger sequenced using the ABI-PRISM Big Dye Terminator Sequencing kit (Applied Biosystems, Thermo Fisher Scientific). Amplicon sequences were visualized and assembled using Chromas Pro (version 1.7.5, Technelysium Pty Ltd, Tewantin, QLD, Australia). In addition to washing and rinsing ciliate cells to remove any contaminants, different negative controls were used during the single-cell DNA extraction (a sample with no cell) and the PCR amplifications (a sample without DNA template) to confirm that the PCR products were only from the DNA of the ciliate and the intracellular microalgae. DNA from different phytoplankton cultures, including Symbiodinium, was used to control the primer specificity.
For the ciliate (host), a 640-bp-long fragment of the 18S rRNA gene was PCR amplified using the ciliate-specific primers Cil_384F and Cil_1147R (Dopheide et al., 2008). A second 2000-bp-long fragment, which includes the hypervariable V9 region of the 18S rRNA gene, was obtained with a nested PCR using newly designed specific primers: TiaV4-F1 and Cil28S-R1 (PCR 1) and the internal primers TiaV4-F2 and Cil28S-R2 (PCR 2) (see Supplementary Table S1 and Supplementary Figure S1 for more information about the primers used in this study). Contigs of the different amplicons were obtained and a matrix of 18S rDNA sequences was built with these new sequences and public sequences from GenBank (release 203.0, Benson et al., 2014; Supplementary Table S2).
In order to identify the lineage of the endosymbiotic microalgae, a cloning approach was first adopted on the 18S and 28S rRNA genes using the TOPO TA Cloning kit (Invitrogen, Thermo Fisher Scientific) and universal eukaryotic primers 63F and 1818R (Lepère et al., 2011) and D1R and D2C (Scholin et al., 1994) for 18S and 28S rDNA, respectively (Supplementary Figure S1). Specific primers and direct sequencing were subsequently employed for the rest of the isolated ciliates to obtain the 28S rRNA gene (D1–D3 domains; with primers ITS-Dino and LO from Pochon et al., 2001) and the rDNA internal transcribed spacer (ITS; with primers Din464F and ITS-DinoRev from Gómez et al., 2011 and this study, respectively) of the microalgal symbionts (Supplementary Figure S1). Matrices of 28S rDNA and ITS2 sequences were then constructed with reference sequences from previous studies (Supplementary Tables S3 and S4,LaJeunesse et al., 2009; Pochon et al., 2014). Sequences obtained in this study have been deposited in GenBank (accession numbers are given in Table 2 and in Supplementary Table S2 (18S rDNA), Supplementary Table S3 (28S rDNA), Supplementary Table S4 (ITS2) and Supplementary Table S5 (V9 rDNA).
Phylogenetic analyses based on ribosomal gene markers
The three matrices of 18S rDNA (host ciliate) and 28S rDNA and ITS2 (intracellular microalgae) were aligned with MUSCLE implemented in the Seaview software (Edgar, 2004; Gouy et al., 2010), and phylogenetic analyses were conducted with the TOPALI software (Topali V2, Milne et al., 2009). According to Modeltest v0.1.1 (Posada, 2008), the general time-reversible and the Tamura–Nei models of nucleotide substitution were selected for the symbiont 28S rDNA (56 taxa; 795 nucleotide positions) and the ciliate 18S rDNA (34 taxa; 1419 nucleotide positions) alignments, respectively. Phylogenetic inference by maximum likelihood was then performed with PhyML v3.0 (Guindon and Gascuel, 2003), and robustness of inferred topologies was assessed by performing 100 nonparametric bootstraps. Final trees were rooted with outgroups and visualized with FigTree (v. 1.4; http://tree.bio.ed.ac.uk/software/figtree/). For the ITS2 sequences of the symbiont (239-bp-long fragment), a statistical parsimony network was constructed with the TCS software (95% connection limit, 10 or less connection steps, gaps considered as fifth state) (Clement et al., 2000), and visualized and edited with Cytoscape software (Shannon et al., 2003).
Ecological significance of the symbiosis
For both the host ciliate and endosymbiotic microalgae, the hypervariable V9 region of the 18S rRNA gene was specifically extracted from multiple sequence alignments using the primers 1389F and 1510R (Amaral-Zettler et al., 2009) in order to interrogate the V9 metabarcoding data set obtained from 47 stations of the Tara Oceans expedition (Karsenti et al., 2011; De Vargas et al., 2015). This publicly available metabarcoding data set includes millions of V9 rDNA sequences of eukaryotic planktonic organisms obtained by PCR with universal eukaryotic primers and sequenced with Illumina Technology (San Diego, CA, USA). In this study, samples corresponding to the microplankton size fraction (20–180 μm) collected at subsurface mixed-layer waters, where the ciliates were found and isolated, were specifically analyzed. Only V9 reads that were 100% identical to the V9 sequence of the ciliates and the symbiont were retrieved in the Tara Oceans barcode table (doi: 10.1594/PANGAEA.843018). Contextual oceanographic parameters, such as temperature, salinity, nitrite and nitrate concentrations, silica and chlorophyll a concentrations, oxygen minimum and deep chlorophyll maximum depths and photosynthetically active radiation, were also used for statistical analyses (Supplementary Table S6). In order to assess co-occurrence patterns between host and symbiont, and the effect of abiotic parameters, multiple pairwise comparisons were performed using Spearman’s rank correlation tests based on environmental physicochemical parameters, and the distribution and abundance of V9 reads of both partners. P-values were adjusted using the false discovery rate approach (Benjamini and Hochberg, 1995). Correlations between pairs of variables were considered significant when P-values were <0.05. Statistical analyses were performed in the R environment (R Core Team, 2008), and correlations between variables were visualized as a network using the Cytoscape software (Shannon et al., 2003).
Results and discussion
Morphological identity of the host ciliate
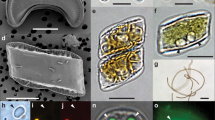
Live observations in light microscopy showed that the ciliates were actively swimming with the help of numerous cilia around the cell (Supplementary Figure S2). From 20 cells, various morphological characters were observed with scanning electron microscopy, notably an armored skeleton composed of calcium carbonate plates (Figures 1a and b), indicating that the ciliate belongs to the family Colepidae Ehrenberg, 1838 (class Prostomatea, order Prorodontida; Lynn, 2008; Chen et al., 2012). A morphological comparison with six previously described ciliate species from the family Colepidae (Coleps hirtus, Nolandia nolandi, Apocoleps magnus, Levicoleps biwae, Tiarina meunieri and Tiarina fusus) indicated that T. fusus is the species that most closely resembles the ciliate isolated in this study (Supplementary Table S7). Both taxa live in marine waters and display a fusiform and elongated body with large longitudinal rows of calcium carbonate plates arranged in six different regions (called ‘tiers’), as well as a wing-like structure that is absent in other Colepidae species (Figures 1a and b; Chen et al., 2012). These morphological commonalities, which are typically used for genus recognition within the Colepidae (Foissner et al., 2008), lead us to argue that the newly isolated ciliate belongs to the genus Tiarina, Berg 1881. In addition to T. fusus, the genus Tiarina is composed of several species including T. meunieri, T. borealis, T. antarctica and T. levigata, but these have very old, incomplete morphological descriptions without reference microscopy images or molecular data. Only T. fusus (Chen et al., 2012) and T. meunieri (previously named Stapperisa fusus, observed in the Arctic and the North Sea; Meunier, 1910; Kahl, 1930) are taxonomically recognized species. Compared with T. fusus, the newly isolated ciliate has a larger length-to-width ratio (1.5 times more), lacks armor spines and possesses long somatic cilia arranged in 18 or more ciliary rows (T. fusus has 15-17 longitudinal rows; Supplementary Table S7). As shown by bright-field (on more than 50 cells) and confocal (on 10 cells) microscopy images (Figures 1d and e and Supplementary Figure S2), another distinct morphological character of the isolated ciliate is the presence of 10 to 25 intracellular microalgal cells that has never been reported in T. fusus (Foissner et al., 2008; Lynn, 2008; Chen et al., 2012). Kahl (1930) observed ‘ingested dinoflagellates’ (‘Nahrung kleine Peredineen’) in the arctic species T. meunieri but this old and succinct description has not been confirmed in the literature. In the newly isolated ciliate, the ovoid microalgae of ∼10 μm in diameter tend to be located next to the macronucleus and the micronucleus of the host (both highlighted in cyan), and seem to be intact as shown by the presence of nondegraded nuclei and plastids, in blue and red respectively (Figures 1d and e). Based on these confocal images, it is not possible to distinguish whether there is a single reticulated or several chloroplasts in each microalgal cell. A very small fraction of cells seemed to be degraded, possibly as a consequence of preservation biases and/or host digestion. Several observations indicate cell division for some of the microalgae (Figure 1e and Supplementary Figure S2). The cellular features observed in confocal laser scanning microscopy on several host individuals (n=10), collected in the Indian and Atlantic Oceans (Tara Oceans stations 41, 64, 72), suggest that the relationship between Tiarina sp. and its symbionts of eukaryotic origin is a putative mutualistic endosymbiosis, rather than predation or kleptoplastidy.
Microscopy images of the photosymbiosis between the ciliate Tiarina sp. (the host) and its intracellular symbiotic microalgae collected in surface oceanic waters. (a, b) Scanning electron microscopy images reveal the calcified skeleton of the ciliate composed of six different tiers: circumoral (COT), anterior secondary (AST), anterior main (AMT), posterior main (PMT), posterior secondary (PST) and caudal (CT). (b) The arrow indicates the wing-like structure that is a morphological feature of the genus Tiarina within the Colepidae family. (c–e) The three-dimensional (3D) reconstructions of symbiotic specimens imaged with confocal laser scanning microscopy (CLSM). (c) The ciliate exhibits numerous cilia, highlighted in green (DiOC6). (d, e) The nuclei of the ciliate (cyan) and the symbiotic microalgae (blue) were reconstructed from the Hoechst fluorescence signal, and the chloroplasts of the microalgae are highlighted by the red chlorophyll autofluorescence. (e) The arrows indicate putative cell division events. Scale bar=20 μm.
Phylogenetic identification of the host ciliate
Partial 18S rDNA sequences (that is, a 1419-bp-long fragment from the V3 to the V9 regions) were obtained from 15 individual ciliates collected in six oceanic regions: the Mediterranean Sea (n=2), and the Indian (n=6), Atlantic (n=5) and Pacific (n=2) Oceans (Table 2). The majority of the 18S rDNA sequences were identical irrespective of their geographic location (which we subsequently call genotype 1), except for four sequences (Med_2, Med_3, TI_381 and TI_382) that have one nucleotide difference in the V9 region (which we call genotype 2). Although Tiarina sp. genotype 1 was found in most plankton samples, genotype 2 was specifically observed in the Mediterranean Sea and the South Atlantic Ocean. In the latter region (for example, station 72), both genotypes were found. Based on BLAST analyses (Altschul et al., 1990), we found no environmental Sanger sequences identical to the newly isolated Tiarina sp. (both genotypes) in GenBank (release 207.0; April 2015); the closest sequence corresponded to the ciliate T. fusus (FJ858217).
In the 18S rDNA phylogenetic tree (Figure 2), sequences of genotypes 1 and 2 grouped together and were placed within a highly supported monophyletic clade (bootstrap values=100%) that corresponds to the family Colepidae, confirming the morphological identification. The monophyly of the Colepidae was also recovered in a previous study (Yi et al., 2010). Within this family, phylogenetic analyses showed that sequences of our new ciliate grouped with T. fusus and an environmental clone (EF526888) in a highly supported monophyletic subclade (bootstrap values=100%) that likely corresponds to the genus Tiarina. However, the isolated Tiarina sp. is genetically clearly different from T. fusus, as it is distinguished by 10 and 11 nucleotide differences in the 18S rDNA for genotypes 1 and 2, respectively. These molecular results corroborate the morphological dissimilarities highlighted above and provide further evidence that the new ciliate is a novel Tiarina species that requires formal description.
Maximum likelihood phylogeny inferred from partial 18S rDNA sequences of the ciliate order Prorodontida (1419 aligned nucleotide positions) and the Tamura–Nei (TrN) model of nucleotide substitution. Sequences of individual cells of Tiarina sp. obtained in this study are colored according to their geographic origin. Bootstrap percentages based on 100 pseudoreplicates are indicated at each node when support values are >50%.
Phylogenetic identification of the microalgal symbiont
Ribosomal gene sequencing was also performed to reveal the identity and diversity of the intracellular eukaryotic microalgae observed in the ciliate Tiarina sp. Amplification of the 18S and 28S rRNA genes with universal eukaryotic primers followed by a cloning procedure showed that the microalgae of different isolated host cells systematically belonged to the dinoflagellate genus Symbiodinium. Twenty partial sequences of the 28S rRNA gene (a 795-bp-long fragment covering the D1–D3 regions) were subsequently obtained with Symbiodinium-specific primers (Figure 3). The phylogenetic tree, which has the same topology as in previous studies (Pochon et al., 2012, 2014), demonstrated that Symbiodinium isolated from all ciliates (genotypes 1 and 2) collected in different oceanic regions belong to clade A (maximum support: bootstrap values=100%), the earliest branching Symbiodinium lineage. Within this clade, the new sequences of Symbiodinium are different from any known reference sequences, and separate analyses with shorter 28S rDNA sequences showed that they also do not belong to the ‘Temperate A’ subclade (Savage et al., 2002; Casado-Amezúa et al., 2014; Supplementary Figure S3). These new 28S rDNA sequences of Symbiodinium display genetic dissimilarities (five to nine polymorphic sites) according to their sampling location. This means that the host ciliate Tiarina sp. is systematically found in symbiosis with Symbiodinium clade A worldwide, but can associate with different types or subclades depending on the oceanic region (for example, Mediterranean Sea and South Pacific Ocean; Figure 3). However, up to three different Symbiodinium types can also be found at the same location (for example, station 72 in the south Atlantic Ocean and station 41 in the North Indian Ocean). From station 72, we note that host genotypes 1 and 2 are associated with different but genetically close Symbiodinium types (3 nucleotide differences out of 795 bp of the 28S rDNA sequence), indicating a symbiont specificity within the same location.
Maximum likelihood phylogeny of the genus Symbiodinium inferred from 28S rDNA sequences (D1–D3 regions) and the general time-reversible (GTR) model of nucleotide substitutions. Novel Symbiodinium sequences obtained in this study are colored according to their geographic origin, and the name of their corresponding host is given (for example, TI_363). Bootstrap percentages based on 100 pseudoreplicates are indicated at each node when support values are >50%.
The earliest diverging clade A is found within several benthic hosts (for example, mollusks, corals, anemones and jellyfish) in temperate and tropical waters of the Atlantic, Pacific and Indian Oceans and Red Sea (Goulet et al., 2008; Franklin et al., 2012). Our results raise the question of whether the novel diversity of Symbiodinium associated with a planktonic ciliate in the open ocean can also live within benthic hosts from coastal waters. A complex genetic diversity composed of 15 common subclade types has been recognized in clade A with the highly resolutive ITS2 marker (LaJeunesse et al., 2001, 2009; Franklin et al., 2012; Lee et al., 2015). This marker was therefore used in this study to assign the new Symbiodinium sequences at the subclade level, and thus to provide a better understanding of the identity of the Tiarina symbiont and the specificity of the symbiotic partnership (LaJeunesse, 2001). A statistical parsimony network using the ITS2 sequences (239 bp long) was constructed including Symbiodinium sequences obtained from Tiarina sp. and those of 14 described subclade types from different hosts and oceanic regions (Figure 4 and Supplementary Table S3). Subclade A2 was not included in the network because it was too divergent (over 10 polymorphic sites). Confirming the 28S rDNA phylogeny, ITS2 sequences of Symbiodinium found in Tiarina sp. did not match any previously described clade A sequence types, and instead represented 8 novel divergent subclade types (7 of which have ⩾2 nucleotide differences). Remarkably, this Symbiodinium diversity from a single host taxon is relatively high, and equivalent to the diversity described so far from ∼50 benthic host taxa in reef ecosystems worldwide (Franklin et al., 2012). The ITS2 diversity of Symbiodinium from Tiarina sp., like that of the 28S rRNA gene, is partly structured by geography, whereby sequences from the same oceanic region tend to be similar (South Pacific Ocean, and Mediterranean and Red Seas). However, the converse also occurs in the Indian and South Atlantic oceans, where up to three highly divergent Symbiodinium subclade types (over 10 polymorphic sites) can coexist at the same location. Overall, the ITS2 haplotype network showed a clear genetic separation between Symbiodinium from benthic hosts and Symbiodinium from the pelagic Tiarina sp. For instance, in the Pacific Ocean, Symbiodinium from benthic hosts (subclades A5, A6, A7, A8 and A12, in light blue) are very distinct from Symbiodinium found in Tiarina sp. (TI_377 and TI_378). With two nucleotide differences in the ITS2 sequence, the closest subclade type to the ciliate Symbiodinium is A4. In corals, this subclade type is known for its high photosynthetic efficiency and resistance to photodamage when exposed to temperature stress (Warner et al., 2006). Symbiodinium A4 has also been found in the jellyfish Linuche unguiculata that alternates between the benthic (polyp stage) and planktonic compartments during its life history (Montgomery and Kremer, 1995; Trench and Thinh, 1995; LaJeunesse, 2001). Therefore, subclade A4 could represent an evolutionary and ecological transition between the benthic and pelagic environments, where different selective pressures have caused evolutionary changes within Symbiodinium.
Statistical parsimony network with ITS2 sequences of the different subclades of Symbiodinium clade A. Published sequences and those obtained in this study from the planktonic ciliate Tiarina sp. are represented by a square and a circle, respectively (see Supplementary Table S4 for the GenBank accession numbers). Based on the GeoSymbio database (Franklin et al., 2012), the geographic origin and host spectrum for each Symbiodinium subclade are indicated by a color and a letter, respectively. All of the previously described subclade types (A1–A16 in squares) were found in benthic coastal hosts (∼50 host species).
Ecological significance of the symbiosis between the ciliate Tiarina and Symbiodinium
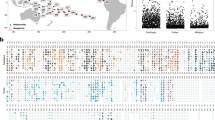
To explore the geographical distribution of the symbiotic relationship over large spatial scales, the two V9 sequences of the host ciliate (genotypes 1 and 2, differentiated by a single nucleotide) were used to interrogate the V9 rDNA Tara Oceans metabarcoding data set (De Vargas et al., 2015). V9 metabarcodes that were 100% identical to the V9 of genotypes 1 and 2 were detected in 32 stations across the oceans, mainly at equatorial, tropical and subtropical latitudes (Figures 5a and b). The highest abundances of V9 reads occurred in the Red Sea (station 32), and the Indian (station 41) and South Atlantic (station 72) Oceans, and they were not detected in the Southern Ocean. Both genotypes coexisted in 21 stations in the Mediterranean and Red Seas, and the Indian, Atlantic and Pacific Oceans. Genotype 1 was more cosmopolitan and abundant (10 to 100 times more, in terms of the relative proportion of reads in a given sample) as compared with genotype 2 that was detected in fewer stations (Figures 5a and b).
Geographic distribution and ecology of the photosymbiosis between the host ciliate Tiarina and the microalga Symbiodinium. (a, b) Mapping of the relative abundance of rDNA V9 reads identical to the V9 sequences of Tiarina genotype 1 (a) and Tiarina genotype 2 (b) in Tara Oceans stations (surface samples in the 20–180 μm size fraction). The color gradient indicates the relative abundance of V9 reads from green to red for low to high values, respectively. Black dots indicate that no V9 reads were detected at the given station. (c) Network showing the different correlations (statistically significant; P<0.05) between the V9 read abundances of Symbiodinium (represented by five distinct V9 sequences), Tiarina genotypes 1 and 2, T. fusus and several physicochemical parameters (see Supplementary Tables S6 and S8 for the data used and the P-values of the different correlations, respectively).
The ecological preferences of the symbiosis between Tiarina sp. and Symbiodinium were investigated through correlation analyses including different contextual oceanographic parameters and the relative abundance of V9 reads of Symbiodinium (representing five distinct V9 sequences), Tiarina sp. (genotypes 1 and 2) and T. fusus (Figure 5c,Supplementary Tables S6 and S8). This analysis revealed a positive correlation between Symbiodinium and both genotypes of Tiarina sp., demonstrating that there is an intimate ecological interaction between these taxa in surface ocean waters. In addition, Symbiodinium V9 reads correlated negatively to the concentration of some nutrients (NO2, PO4) and chlorophyll a. Reads of genotypes 1 and 2 of the ciliate Tiarina sp. significantly co-occurred and displayed negative correlation with chlorophyll a. Reads of genotype 2 also displayed negative correlations with some nutrients (NO2, PO4, Si), whereas those of genotype 1 had positive correlations with the depth of the deep chlorophyll maximum. Thus, the correlation patterns between the host, its symbiont and their abiotic environment clearly indicate that the symbiotic partnership predominantly occurs in oligotrophic conditions (typically characterized by deep chlorophyll maximum and low chlorophyll and nutrient concentrations), mirroring the ecology of other known planktonic photosymbioses (Shaked and de Vargas, 2006; Stoecker et al., 2009; Decelle et al., 2012). More specifically, we note that genotype 2 positively correlated with salinity, very likely because of its prevalence in the Mediterranean Sea. No significant correlations were detected between T. fusus and Symbiodinium, or between T. fusus and different oceanographic parameters, showing that the life mode and the ecology of T. fusus and the novel Tiarina sp. are distinct. To date, T. fusus has been reported essentially from coastal and productive marine waters, such as European fjords (Smetacek, 1981; Dale and Dahl, 1987; Lynn, 2008; Chen et al., 2012), where it can dominate ciliate abundance and/or biomass (maximum density reported: 34 000 cells m−l). T. fusus is particularly known for its significant grazing activity on a variety of toxic and/or red-tide microalgae (Jeong et al., 2002). In this study, V9 reads assigned to T. fusus (100% identical to the reference sequence) were only detected in low abundance in coastal stations in the Adriatic and Red Seas, and in the Indian and Atlantic Oceans (Supplementary Figure S4). According to the ecological species concept (Boenigk et al., 2012), these results therefore provide additional evidence that the symbiotic Tiarina sp. is a novel species distinct from T. fusus.
Conclusion and perspectives
Our study identified and characterized a novel and widespread pelagic photosymbiosis between an undescribed calcifying ciliate (Tiarina sp.) and the dinoflagellate Symbiodinium. Consistent microscopy observations on multiple specimens, systematic PCR detection of Symbiodinium within the ciliate and the geographic distribution in surface oceans together provide evidence in favor of a long-term mutualistic symbiosis rather than predation or kleptoplastidy. This newly described partnership is relevant not only because the host is a previously unknown planktonic protistan species, but also because the occurrence of Symbiodinium as a symbiont in the open ocean was previously unknown, despite the fact that Symbiodinium is one of the most extensively studied microalgal genera. We revealed a novel and highly diverse assemblage within Symbiodinium clade A composed of eight subclade types that appear to be endemic to pelagic waters as they have not been reported from hosts in benthic coastal habitats so far (for example, tropical reefs, temperate coastal areas). Selective pressures inherent to the pelagic realm and host specialization might have created a distinct ecological niche and driven the tempo and mode of Symbiodinium diversification in the open ocean. Given that clade A is the earliest branching lineage and colepid fossils date back to the Triassic (220 million year ago, Schmidt et al., 2006), it is possible that the Tiarina–Symbiodinium symbiosis occurred relatively early in the evolution of the two lineages.
This cosmopolitan symbiosis likely plays multifaceted biogeochemical roles in pelagic ecosystems by contributing to primary production and calcium cycling through the Symbiodinium photosynthesis and host calcification, respectively. Automatic and in situ imaging techniques could be used in the future to properly quantify this interaction, and thus to determine its ecological impact in the environment. The active motility of the cilia-bearing host, which is absent in the ‘passive’ photosymbiotic rhizarian Radiolaria and Foraminifera, makes the partnership highly original and may indicate a chemotactic behavior for finding symbiotic partners, suitable light conditions, and food or nutrient patches, as hypothesized in the photosymbiotic ciliate Mesodinium rubrum (Wilkerson and Grunseich, 1990; Fenchel and Blackburn, 1999). Given the metabolic capacities of Symbiodinium in reef ecosystems (Yellowlees et al., 2008; Godinot et al., 2009; Pernice et al., 2012), Tiarina sp. very likely benefits from a significant source of carbon, nitrogen, phosphate and other key nutrients for growth and reproduction that are otherwise limiting in the open ocean. In addition to its nutritional role, Symbiodinium could also play a critical role not only in the calcification of the ciliate’s skeleton, as shown in scleractinian corals (Allemand et al., 2011; Davy et al., 2012), but also in ultraviolet radiation protection in transparent open ocean waters, as clade A is known to produce a significant amount of ultraviolet-absorbing mycosporine-like amino acids in comparison with other clades (Banaszak et al., 2000). Clade A is also known for its tolerance for high irradiance and high temperatures, likely explaining why it is commonly found in shallow-water benthic hosts (Warner et al., 2006; Kemp et al., 2014, 2015) or in the free-living environment (Pochon et al., 2014). Thus, the physiological features of clade A would seem to make this Symbiodinium lineage particularly well suited to be a symbiont within single-celled hosts in oligotrophic transparent waters. Future studies should aim to better understand the nature, ecological role and life cycle of this new photosymbiosis, and explore whether other Symbiodinium clades can be found in the open ocean.
References
Allemand D, Tambutté E, Zoccola D, Tambutté S . (2011) Coral calcification, cells to reefs. In: Dubinsky Z, Stambler N (eds). Coral Reefs: An Ecosystem in Transition. Springer: Berlin, Germany.
Amaral-Zettler LA, McCliment EA, Ducklow HW, Huse SM . (2009). A method for studying protistan diversity using massively parallel sequencing of V9 hypervariable regions of small-subunit ribosomal RNA genes. PLoS One 4: e6372.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ . (1990). Basic local alignment search tool. J Mol Biol 215: 403–410.
Banaszak AT, LaJeunesse TC, Trench RK . (2000). The synthesis of mycosporine-like amino acids (MAAs) by cultured, symbiotic dinoflagellates. J Exp Mar Biol Ecol 249: 219–233.
de Bary AH . (1878) Vortrag: Uber Symbiose. Tagblatt der 51 Versammlung Deutscher Naturforscher und Aerzte in Cassel. Baier & Lewalter: Kassel, pp 121–126.
Benjamini Y, Hochberg Y . (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B 57: 289–300.
Benson DA, Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW . (2014). GenBank. Nucleic Acids Res 42: D32–D37.
Bernstein RE, Betzer PR, Feely RA, Byrne RH, Lamb MF, Michaels AF . (1987). Acantharian fluxes and strontium to chlorinity ratios in the North Pacific Ocean. Science 237: 1490–1494.
Boenigk J, Ereshefsky M, Hoef-Emden K, Mallet J, Bass D . (2012). Concepts in protistology: species definitions and boundaries. Eur J Protistol 48: 96–102.
Casado-Amezúa P, Machordom A, Bernardo J, Gonzáles-Wangüemert M . (2014). New insights into the genetic diversity of zooxanthellae in Mediterranean anthozoans. Symbiosis 63: 41–46.
Chen X, Gao S, Liu W, Song W, Al-Rasheid KAS, Warren A . (2012). Taxonomic descriptions of three marine colepid ciliates, Nolandia sinica spec. nov., Apocoleps caoi spec. nov. and Tiarina fusa (Claparède & Lachmann, 1858) Bergh, 1881 (Ciliophora, Prorodontida). Int J Syst Evol Microbiol 62: 735–744.
Clement M, Posada D, Crandall KA . (2000). TCS: a computer program to estimate gene genealogies. Mol Ecol 9: 1657–1660.
Coffroth MA, Santos SR . (2005). Genetic diversity of symbiotic dinoflagellates in the genus Symbiodinium. Protist 156: 19–34.
Costanza R, d’Arge R, de Groot R, Farber S, Grasso M, Hannon B et al. (1997). The value of the world’s ecosystem services and natural capital. Nature 387: 253–260.
Dale T, Dahl E . (1987). Mass occurrence of planktonic oligotrichous ciliates in a bay in southern Norway. J Plankton Res 9: 871–879.
Daugbjerg N, Jensen MH, Hansen PJ . (2013). Using nuclear-encoded LSU and SSU rDNA sequences to identify the eukaryotic endosymbiont in Amphisolenia bidentata (Dinophyceae). Protist 164: 411–422.
Davy SK, Allemand D, Weis VM . (2012). Cell biology of cnidarian-dinoflagellate symbiosis. Microbiol Mol Biol Rev 76: 229–261.
Decelle J, Colin S, Foster RA . (2015) Photosymbiosis in marine planktonic protists. In: Ohtsuka S, Suzaki T, Horiguchi T, Suzuki N, Not F(eds) Marine Protists: Diversity and Dynamics. Springer: Tokyo, Japan, pp 465–500.
Decelle J, Probert I, Bittner L, Desdevises Y, Colin S, de Vargas C et al. (2012). An original mode of symbiosis in open ocean plankton. Proc Natl Acad Sci USA 109: 18000–18005.
De Vargas C, Audic S, Henry N, Mahé F, Decelle J, Logares R et al. (2015). Eukaryotic plankton diversity in the sunlit ocean. Science 348: 1261605-1-11.
Dopheide A, Lear G, Stott R, Lewis G . (2008). Molecular characterization of ciliate diversity in stream biofilms. Appl Environ Microbiol 74: 1740–1747.
Dziallas C, Allgaier M, Monaghan MT, Grossart HP . (2012). Act together—implications of symbioses in aquatic ciliates. Front Microbiol 3: 288.
Edgar RC . (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32: 1792–1797.
Esteban GF, Fenchel T, Finlay BJ . (2010). Mixotrophy in ciliates. Protist 161: 621–641.
Fenchel T, Blackburn N . (1999). Motile chemosensory behaviour of phagotrophic protists: mechanisms for and efficiency in congregating at food patches. Protist 150: 325–336.
Foissner W, Kusuoka Y, Shimano S . (2008). Morphology and gene sequence of Levicoleps biwae n. gen., n. sp. (Ciliophora, Prostomatida), a proposed endemic from the ancient Lake Biwa, Japan. J Euk Microbiol 55: 185–200.
Franklin EC, Stat M, Pochon X, Putnam HM, Gates RD . (2012). GeoSymbio: a hybrid, cloud-based web application of global geospatial bioinformatics and ecoinformatics for Symbiodinium-host symbioses. Mol Ecol Resources 12: 369–373.
Gast RJ, Caron DA . (2001). Photosymbiotic associations in planktonic foraminifera and radiolaria. Hydrobiologia 461: 1–7.
Godinot C, Ferrier-Pagès C, Grover R . (2009). Control of phosphate uptake by zooxanthellae and host cells in the scleractinian coral Stylophora pistillata. Limnol Oceanogr 54: 1627–1633.
Gómez F, López‐García P, Moreira D . (2011). Molecular phylogeny of dinophysoid dinoflagellates: the systematic position of Oxyphysis oxytoxoides and the Dinophysis hastata group (Dinophysales, Dinophyceae) 1: Molecular phylogeny of Dinophysales. J Phycol 47: 393–406.
Goulet T, Simmons C, Goulet D . (2008). Worldwide biogeography of Symbiodinium in tropical octocorals. Mar Ecol Prog Ser 355: 45–58.
Gouy M, Guindon S, Gascuel O . (2010). SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27: 221–224.
Guindon S, Gascuel O . (2003). A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52: 696–704.
Jeong JH, Yoon JY, Kim JS, Yoo YD, Seong KA . (2002). Growth and grazing rates of the prostomatid ciliate Tiarina fusus on red-tide and toxic algae. Aquat Microb Ecol 28: 289–297.
Johnson MD . (2011). Acquired phototrophy in ciliates: a review of cellular interactions and structural adaptations. J Euk Microbiol 58: 185–195.
Kahl A . (1930). Urtiere oder Protozoa I: Wimpertiere oder Ciliata (Infusoria) 1. Allgemeiner Teil und Prostomata. Tierwelt Dtl 18: 1–180.
Karsenti E, Acinas S G, Bork P, Bowler C, De Vargas C, Raes J et al. (2011). A holistic approach to marine eco-systems biology. PLoS Biol 9: e1001177.
Keeling PJ . (2010). The endosymbiotic origin, diversification and fate of plastids. Philos Trans R Soc London [Biol] 365: 729–748.
Kemp DW, Hernandez-Pech X, Iglesias-Prieto R, Fitt WK, Schmidt GW . (2014). Community dynamics and physiology of Symbiodinium spp. before, during and after a coral bleaching event. Limnol Oceanogr 59: 788–797.
Kemp DW, Thornhill DJ, Rotjan RD, Iglesias-Prieto R, Fitt WK, Schmidt GW . (2015). Spatially distinct and regionally endemic Symbiodinium assemblages in the threatened Caribbean reef-building coral Orbicella faveolata. Coral Reefs 34: 535–547.
LaJeunesse TC . (2001). Investigating the biodiversity, ecology, and phylogeny of endosymbiotic dinoflagellates in the genus Symbiodinium using the ITS region: in search of a ‘species’ level marker. J Phycol 37: 866–880.
LaJeunesse TC, Loh W, Trench RK . (2009). Do introduced endosymbiotic dinoflagellates ‘take’ to new hosts? Biol Invasions 11: 995–1003.
Lee SY, Jeong HJ, Kang NS, Jang TY, Jang SH, LaJeunesse TC . (2015). Symbiodinium tridacnidorum sp. nov., a dinoflagellate common to Indo-Pacific giant clams, and a revised morphological description of Symbiodinium microadriaticum Freudenthal, emended Trench & Blank. Eur J Phycol 50: 155–172.
Lepère C, Demura M, Kawachi M, Romac S, Probert I, Vaulot D . (2011). Whole-genome amplification (WGA) of marine photosynthetic eukaryote populations. FEMS Microbiol Ecol 76: 513–523.
Lobban CS, Schefter M, Simpson AGB, Pochon X, Pawlowski J, Foissner W . (2002). Maristentor dinoferus n. gen., n. sp., a giant heterotrich ciliate (Spirotrichea: Heterotrichida) with zooxanthellae, from coral reefs on Guam, Mariana Islands. Mar Biol 141: 207–208.
Lobban CS, Modeo L, Verni F, Rosati G . (2005). Euplotes uncinatus (Ciliophora, Hypotrichia), a new species with zooxanthellae. Mar Biol 147: 1055–1061.
Lynn DH . (2008) The Ciliated Protozoa: Characterization, Classification, and Guide to the Literature, third edn. Springer: Dordrecht.
Margulis L, Fester R . (1991) Symbiosis as a Source of Evolutionary Innovation: Speciation and Morphogenesis. MIT Press: Cambridge, MA, USA.
Maynard Smith J . (1989). Generating novelty by symbiosis. Nature 341: 284–285.
Meunier A . (1910) Microplankton des Mers de Barents et de Kara. Campagne Arctique de 1907. C. Bulens: Bruxelles.
Michaels AF, Caron DA, Swanberg NR, Howse FA, Michaels CM . (1995). Planktonic sarcodines (acantharia, radiolaria, foraminifera) in surface waters near Bermuda–abundance, biomass and vertical flux. J Plankton Res 17: 131–163.
Milne I, Lindner D, Bayer M, Husmeier D, McGuire G, Marshall DF et al. (2009). TOPALi v2: a rich graphical interface for evolutionary analyses of multiple alignments on HPC clusters and multi-core desktops. Bioinformatics 25: 126–127.
Montgomery M, Kremer P . (1995). Transmission of symbiotic dinoflagellates through the sexual cycle of the host scyphozoan Linuche unguiculata. Mar Biol 124: 147–155.
Muscatine L, Falkowski PG, Porter JW, Dubinsky Z . (1984). Fate of photosynthetically fixed carbon in light and shade-adapted colonies of the symbiotic coral, Stylophora pistillata. Proc R Soc Lond B 222: 181–202.
Pernice M, Meibom A, Van Den Heuvel A, Kopp C, Domart-Coulon I, Hoegh-Guldberg O et al. (2012). A single-cell view of ammonium assimilation in coral-dinoflagellate symbiosis. ISME J 6: 1314–1324.
Pesant S, Not F, Picheral M, Kandels-Lewis S, Le Bescot N, Gorsky G et al. (2015). Open science resources for the discovery and analysis of Tara Oceans data. Available at http://biorxiv.org/content/early/2015/05/08/019117.
Pochon X, Putnam HM, Gates RD . (2014). Multi-gene analysis of the symbiotic and free-living dinoflagellate genus Symbiodinium. Peer J 2: e394.
Pochon X, Putnam HM, Burki F, Gates RD . (2012). Identifying and characterizing alternative molecular markers for the symbiotic and free-Living dinoflagellate genus Symbiodinium. PLoS One 7: e29816.
Pochon X, Gates RD . (2010). A new Symbiodinium clade (Dinophyceae) from soritid foraminifera in Hawaii. Mol Phylogenet Evol 56: 492–497.
Pochon X, Pawlowski J, Zaninetti L, Rowan R . (2001). High genetic diversity and relative specificity among Symbiodinium-like endosymbiotic dinoflagellates in soritid foraminiferans. Mar Biol 139: 1069–1078.
Posada D . (2008). jModelTest: phylogenetic model averaging. Mol Bio Evol 25: 1253–1256.
Probert I, Siano R, Poirier C, Decelle J, Biard T, Akihiro T et al. (2014). Brandtodinium gen. nov. and B. nutricula comb. Nov. (Dinophyceae), a dinoflagellate commonly found in symbiosis with polycystine radiolarians. J Phycol 50: 388–399.
R Core Team. (2008) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria, Available from https://www.R-project.org/.
Rowan R . (2004). Thermal adaptations in reef coral symbionts. Nature 430: 742.
Savage AM, Goodson MS, Visram S, Trapido-Rosenthal H, Wiedenmann J, Douglas AE . (2002). Molecular diversity of symbiotic algae at the latitudinal margins of their distribution: dinoflagellates of the genus Symbiodinium in corals and sea anemones. Mar Ecol Prog Ser 244: 17–26.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T et al. (2012). Fiji: an open-source platform for biological-image analysis. Nat Methods 9: 676–682.
Schmidt AR, Ragazzi E, Coppellotti O, Roghi G . (2006). A microworld in Triassic amber. Nature 444: 14.
Scholin CA, Herzog M, Sogin M, Anderson DM . (1994). Identification of group‐and strain‐specific genetic markers for globally distributed Alexandrium (Dinophyceae). II. Sequence analysis of a fragment of the LSU rRNA gene. J Phycol 30: 999–1011.
Shaked Y, de Vargas C . (2006). Pelagic photosymbiosis: rDNA assessment of diversity and evolution of dinoflagellate symbionts and planktonic foraminiferal hosts. Mar Ecol Prog Ser 325: 59–71.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al. (2003). CytoScape a software environment for integrated models of biomolecular interaction networks. Genome Res 13: 2498–2504.
Smetacek V . (1981). The annual cycle of protozooplankton in the Kiel Bight. Mar Biol 63: 1–11.
Stabell T, Andersen T, Klaveness D . (2002). Ecological significance of endosymbionts in a mixotrophic ciliate - an experimental test of a simple model of growth coordination between host and symbiont. J Plankton Res 24: 889–899.
Stanley GD Jr . (2006). Ecology, photosymbiosis and the evolution of modern coral reefs. Science 312: 857–858.
Stoecker DK, Johnson MD, de Vargas C, Not F . (2009). Acquired phototrophy in aquatic protists. Aquat Microb Ecol 57: 279–310.
Stoecker DK, Silver MW, Michaels AE, Davis LH . (1988–1989). Enslavement of algal chloroplasts by four Strombidium spp. (Ciliophora, Oligotrichida). Mar Microb Food Webs 3: 79–100.
Swanberg NR, Caron DA . (1991). Patterns of sarcodine feeding in epipelagic oceanic plankton. J Plankton Res 13: 287–322.
Sweeney BM . (1976). Pedinomonas noctilucae (Prasinophyceae), the flagellate symbiotic in Noctiluca (Dinophyceae) in Southeast Asia. J Phycol 12: 460–464.
Taylor FJR . (1982). Symbioses in marine microplankton. Ann Inst Oceanogr Paris (Nouv Ser) 58: 61–90.
Trench RK, Thinh L . (1995). Gymnodinium linucheae sp. nov.: the dinoflagellate symbiont of the jellyfish Linuche unguiculata. Eur J Phycol 30: 149–154.
Warner ME, LaJeunesse TC, Robison JD, Thur RM . (2006). The ecological distribution and comparative photobiology of symbiotic dinoflagellates from reef corals in Belize: potential implications for coral bleaching. Limnol Oceanogr 51: 1887–1897.
Wilkerson FP, Grunseich G . (1990). Formation of blooms by the symbiotic ciliate Mesodinium rubrum: the significance of nitrogen uptake. J Plankton Res 12: 973–989.
Yi Z, Dunthorn M, Song W, Stoeck T . (2010). Increasing taxon sampling using both unidentified environmental sequences and identified cultures improves phylogenetic inference in the Prorodontida (Ciliophora, Prostomatea). Mol Phylogenet Evol 57: 937–941.
Yellowlees D, Rees TAV, Leggat W . (2008). Metabolic interactions between algal symbionts and invertebrate hosts. Plant Cell Environ 31: 679–694.
Acknowledgements
This work was supported by the project OCEANOMICS that has received funding from the French government, managed by the Agence Nationale de la Recherche, under the grant agreement ‘Investissement d′Avenir’ ANR-11-BTBR-0008. We thank the European Molecular Biology Laboratory (EMBL, Heidelberg) and more specifically Rainer Pepperkok for providing access to the advanced light microscopy facility (ALMF). We also thank the coordinators and members of the Tara Oceans expedition, Diana Sarno from the Stazione Zoologica Anton Dohrn of Naples for providing bright-field images-, and the marine station LOV from Villefranche-sur-Mer for field sampling. We are grateful to Ian Probert and Jan Slapeta for valuable comments on the manuscript. This article is contribution number 33 of Tara Oceans.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Mordret, S., Romac, S., Henry, N. et al. The symbiotic life of Symbiodinium in the open ocean within a new species of calcifying ciliate (Tiarina sp.). ISME J 10, 1424–1436 (2016). https://doi.org/10.1038/ismej.2015.211
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.211
This article is cited by
-
Metagenome-assembled genomes of phytoplankton microbiomes from the Arctic and Atlantic Oceans
Microbiome (2022)
-
An integrative approach sheds new light onto the systematics and ecology of the widespread ciliate genus Coleps (Ciliophora, Prostomatea)
Scientific Reports (2021)
-
Tara Oceans: towards global ocean ecosystems biology
Nature Reviews Microbiology (2020)
-
A de novo approach to disentangle partner identity and function in holobiont systems
Microbiome (2018)