Abstract
Diversity has a key role in the dynamics and resilience of communities and both interspecific (species) and intraspecific (genotypic) diversity can have important effects on community structure and function. However, a critical and unresolved question for understanding the ecology of a community is to what extent these two levels of diversity are functionally substitutable? Here we show, for a mixed-species biofilm community composed of Pseudomonas aeruginosa, P. protegens and Klebsiella pneumoniae, that increased interspecific diversity reduces and functionally substitutes for intraspecific diversity in mediating tolerance to stress. Biofilm populations generated high percentages of genotypic variants, which were largely absent in biofilm communities. Biofilms with either high intra- or interspecific diversity were more tolerant to SDS stress than biofilms with no or low diversity. Unexpectedly, genotypic variants decreased the tolerance of biofilm communities when experimentally introduced into the communities. For example, substituting P. protegens wild type with its genotypic variant within biofilm communities decreased SDS tolerance by twofold, apparently due to perturbation of interspecific interactions. A decrease in variant frequency was also observed when biofilm populations were exposed to cell-free effluents from another species, suggesting that extracellular factors have a role in selection against the appearance of intraspecific variants. This work demonstrates the functional substitution of inter- and intraspecific diversity for an emergent property of biofilms. It also provides a potential explanation for a long-standing paradox in microbiology, in which morphotypic variants are common in laboratory grown biofilm populations, but are rare in diverse, environmental biofilm communities.
Similar content being viewed by others
Introduction
Diversity is a fundamental property at all levels of ecological organization. At the community level, species (interspecific) diversity is recognized as one of the key factors governing community function and stability (Elton, 1958; May, 1973; Tilman and Downing, 1994; Naeem and Li, 1997; Cardinale et al., 2002). Similarly, genotypic (intraspecific) diversity is increasingly recognized to have a significant role in enhancing the performance of populations (Hughes et al., 2008; Aguirre and Marshall, 2012; Koh et al., 2012; Latzel et al., 2013). Although inter- and intraspecific diversity are important for communities (multi-species) and populations (single-species), respectively, several studies have also shown that inter- and intraspecific diversity do not operate independently in communities. In fact, intraspecific diversity interacts with interspecific diversity and can profoundly affect community performance (Agrawal, 2003; Vellend and Geber, 2005; Whitham et al., 2006; Hersch-Green et al., 2011; Rowntree et al., 2011). For example, increased intraspecific diversity of Ammophila breviligulata, a dominant plant in some sand dune ecosystems, changed the relationship between interspecific diversity and the above ground biomass of the whole community (Crawford and Rudgers, 2012). Such interactive effects between the two levels of diversity can also be observed in other systems, for example plant associated arthropods and microbes (Dungey et al., 2000; Whitham et al., 2003; Johnson and Agrawal, 2005; Wimp et al., 2005).
One fundamental and critical question in an era of increasingly threatened ecosystems and declining biodiversity is to what extent are these two levels of diversity ‘functionally substitutable’? That is, can enhanced genotypic diversity at the population level enhance community performance and stability when species diversity is low, or conversely does greater species diversity enhance community performance when genotypic diversity is low? In addition, if inter- and intraspecific diversity are ‘functionally substitutable’, which diversity comes to predominate in a community? To our knowledge, these questions have not been investigated together experimentally in any system. Here we address these questions using an experimental, mixed-species biofilm system.
Microorganisms are the most diverse group of organisms on Earth, and any general theory of diversity must apply to microbial communities. Consequently, microbial communities are increasingly being used to test theories of ecological diversity (Davey and O'Toole, 2000; Jessup et al., 2004; Burke et al., 2011). Microbial biofilm communities are ideal for such investigations, given their spatially localized nature on surfaces, the small spatial and temporal scales of their responses, the ability to identify genetic mechanisms that drive ecological and evolutionary processes (Little et al., 2008) and the close proximity of cells, which in biofilm communities facilitates interspecific interactions that enhance their tolerance to stresses such as antibiotics and surfactant (Burmolle et al., 2006; Lee et al., 2014). Furthermore, many biofilm populations produce heritable genetic variants (that is, spontaneous development of intraspecific diversity), which increase the tolerance of biofilm populations to stresses such as hydrogen peroxide, antibiotics and predation (Mah and O'Toole, 2001; Stewart and Costerton, 2001; Boles et al., 2004; Koh et al., 2012).
Pseudomonas aeruginosa PAO1, P. protegens Pf-5 and Klebsiella pneumoniae KP-1 naturally coexist, in environments as diverse as metalworking fluids (Chazal, 1995) and the gut of silk moth, Bombyx mori (Anand et al., 2010). In laboratory experiments, biofilm communities composed of these three species exhibited enhanced tolerance to sodium dodecyl sulfate (SDS) and the antibiotic tobramycin relative to individually grown monospecific populations (Lee et al., 2014). Hence, biofilm populations and communities of P. aeruginosa, P. protegens and K. pneumoniae were used as model biofilms to investigate the functional substitution of inter- and intraspecific diversity in a biofilm community. The data in this study show that interspecific diversity can functionally substitute for intraspecific diversity in mixed-species biofilm communities. Furthermore, intraspecific variation is reduced in biofilm communities with extracellular factors having a role in selection against the appearance of intraspecific variants.
Materials and methods
Strains and growth conditions
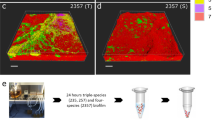
Bacterial strains used in this study were P. aeruginosa PAO1, P. protegens Pf-5 and K. pneumoniae KP-1 (Lee et al., 2013). These strains were labeled with different fluorescent proteins as described (Lee et al., 2014). Biofilms were cultivated in three-channel flow chambers and set up as described (Lee et al., 2014) with M9 minimal medium (48 mm Na2HPO4; 22 mm KH2PO4; 9 mm NaCl; 19 mm NH4Cl; 2 mm MgSO4; 0.1 mm CaCl2 and 0.04% w/v glucose) supplemented with 0.2% w/v casamino acids. Biofilm effluents were collected from 3 day biofilm populations, centrifuged (5000 g, 10 min) and filtered (0.22 μm). During SDS treatment, the biofilms were treated with 0.1% w/v SDS in casamino acids supplemented M9 minimal medium for 2 h. P. aeruginosa biofilms were treated with SDS after 4 days of growth, while P. protegens and K. pneumoniae biofilms were treated with SDS after 3 days of growth.
Isolation and enumeration of morphotypic variants from biofilms and planktonic cultures
Samples (approximately 1 ml) of biofilm effluent and planktonic culture were collected daily. The samples were sonicated (37 kHz, 100% power) in a water bath sonicator (Elmansonic P, Elma, Singen, Germany) for 5 min, vortexed for three periods of 6 s, serially diluted, plated onto LB10 agar plates (10 g l−1 NaCl; 10 g l−1 tryptone; 5 g l−1 yeast extract; 1.5% w/v agar) as well as Pseudomonas isolation agar plates (Becton Dickinson, Franklin Lakes, NJ, USA) in triplicate and incubated at room temperature (25±1 °C) for 2 days. The morphotypic variants were identified and classified by their colony morphologies.
Microscopy, image and statistical analysis
Microscopic observation and image acquisition were performed using a confocal laser scanning microscope (LSM 780, Carl Zeiss, Oberkochen, Germany) as described (Lee et al., 2014). For image analysis, three biological replicates (flow chambers) with a total of 45 image stacks (15 from each flow chamber) were quantified for each biofilm using IMARIS (Bitplane, Belfast, UK). Tolerance to SDS was determined as the percentage of biomass remaining post-treatment. Statistical analysis was performed using IBM SPSS Statistics 21 (IBM, Armonk, NY, USA). Analysis of variance (ANOVA) was performed to test the significant difference between groups. When more than two groups were compared, Tukey's honest significance test was performed post ANOVA to compare the means for all possible pairs.
Whole genome sequencing and genetic analysis
The genomic DNA of each morphotypic variant, wild-type biofilm isolate and wild-type strain was extracted using QIAamp DNA Mini Kit (Qiagen, Venlo, The Netherlands) according to the manufacturer's protocol. Sequencing libraries were prepared using TruSeq DNA Sample Preparation Kit (Illumina, San Diego, CA, USA) and sequenced on the MiSeq bench top sequencer (Illumina) according to the manufacturer's protocols. The insertion deletion (INDELs) and single-nucleotide polymorphisms (SNPs) present in the morphotypic variants and wild-type biofilm isolates were identified using the probabilistic variant detection analysis in CLC genomic workbench 6 (CLC bio, Aarhus, Denmark). First, the paired-end reads were mapped to the reference genomes of P. aeruginosa PAO1, P. protegens Pf-5 and K. pneumoniae NTUH-K2044 (GenBank accession number: AE004091 (PAO1), CP000076 (Pf-5), AP006725 (NTUH-K2044)). Second, all the INDELs and SNPs were identified with the probabilistic variant detection analysis. Third, the identified INDELs and SNPs were filtered, where paired-ends reads from the wild-type strains were used to remove INDELs and SNPs already present in the wild-type strains due to genetic drift.
Site-specific amplification and sequencing
PCR was performed for the additional morphotypic variant isolates using the primers in Supplementary Table S1. The amplicons were sequenced using sanger sequencing.
Growth assay
The morphotypic variants and their respective wild-type strains were grown in casamino acids supplemented M9 minimal medium (200 r.p.m. at 25±1 °C). Growth was monitored (OD600) over a 12 h period using an UV spectrophotometer (UV-1800, Shimadzu, Kyoto, Japan). At 3 h intervals, the colony forming units per milliliter (c.f.u. ml−1) was also quantified as described (Miles et al., 1938) and used to calculate the doubling time.
Attachment assay
An aliquot of a 24 h culture was distributed into a 96-well Nunclon Delta-treated polystyrene microtiter plate (Thermo Scientific, Waltham, MA, USA) in triplicate, incubated at room temperature without agitation for 2 h and assessed by crystal violet staining. Briefly, the culture was removed from each well, the wells were washed twice with phosphate buffer saline and stained with 0.1% w/v crystal violet for 20 min. The crystal violet was subsequently removed, the wells were washed thrice with phosphate buffer saline before 95% v/v ethanol was added to each well, mixed and transferred into a clean microtiter plate for quantification (OD600) using a microplate reader (Infinite M200 Pro, Tecan, Männedorf, Switzerland).
Motility assays
A tryptone swimming plate (10 g l−1 tryptone; 5 g l−1 NaCl; 0.3% w/v agar) and a swarming plate (8 g l−1 nutrient broth; 5 g l−1 glucose; 0.5% w/v agar) were point inoculated with a 24 h culture, and incubated at 30 °C (P. protegens) and 37 °C (P. aeruginosa and K. pneumoniae) for 16 h. Swimming and swarming motilities were assessed quantitatively by measuring the diameters of the turbid circular zone and the swarm zone, respectively. Twitching motility for P. aeruginosa was assessed using a LB10 agar (1% w/v agar) stab inoculated with a 24 h culture of P. aeruginosa. After incubation at 37 °C for 24 h, twitching motility was quantified by removing the agar, washing the unattached cells with water, staining the cells that attached on the polystyrene plate with crystal violet as described above and measuring the diameter of the twitch zone.
Production of pyoverdine and pyocyanin
Bacteria were cultured in 10 ml of King's B medium (20 g l−1 protease peptone; 1.5 g l−1 K2HPO4; 1.5 g l−1 MgSO4·7H2O; 10 ml l−1 glycerol) at 30 °C (P. protegens) and 37 °C (P. aeruginosa) for 24 h. Pyoverdine in the supernatant was semi-quantified by measuring the emission fluorescence at 460 nm after excitation at 400 nm. Similarly, bacteria were cultured in 10 ml of King’s A medium (20 g l−1 Bacto peptone; 1.4 g l−1 MgCl2; 10 g l−1 K2SO4; 10 ml l−1 glycerol) and pyocyanin in the supernatant was semi-quantified by measuring the absorbance (OD695).
Colanic acid assay
Colanic acid was estimated by measuring l-fucose. Briefly, bacteria were cultured for 24 h and the OD600 measured. The culture was boiled (100 °C, 15 min), cooled to room temperature and centrifuged (14 000 g, 30 min, 4 °C). Subsequently, 120 ml of 70% v/v ethanol was added to 40 ml of the supernatant. The mixture was kept at 4 °C overnight and centrifuged (14 000 g, 30 min, 4 °C). The pellet was dissolved in 1 ml of sterile water and subsequently diluted 10 times. To quantify l-fucose, 4.5 ml of H2SO4/H2O (6:1 v/v) was mixed with 1 ml of the sample, heated (100 °C, 20 min) and cooled to room temperature. The absorbance at 396 and 427 nm were measured before and after the addition of 100 μl of 1 m cysteine hydrochloride. Fucose concentration in a sample was determined by comparing the differences in measurements (OD396-OD427) against a standard curve.
Results
Morphotypic variants and their occurrence
In this study, the frequency of morphotypic variants in biofilm populations and communities were used as a measure of intraspecific diversity. A small colony variant (P. aeruginosa SCV) and a non-mucoid colony variant (K. pneumoniae NMV) were isolated based on colony morphology from P. aeruginosa and K. pneumoniae biofilms, respectively. Four morphotypic variants were also isolated from P. protegens biofilms, the small colony variant (P. protegens SCV), cauliflower colony variant (P. protegens CCV), rough colony variant (P. protegens RCV) and wrinkled colony variant (P. protegens WCV) (Table 1). Although these morphotypic variants made up as much as 44%, 62% and 36% of the P. aeruginosa, P. protegens and K. pneumoniae biofilms respectively, they were not found in planktonic cultures (Figure 1), suggesting that these morphotypic variants were specific to growth as a biofilm.
Morphotypic variants were genotypically distinct
Morphotypic variants can result from either genetic mutation that leads to heritable variants (Koh et al., 2007; McElroy et al., 2014), or phase variation that leads to transient variants (Drenkard and Ausubel, 2002; van der Woude and Baumler, 2004). Here, the variant morphologies and phenotypes were stable after multiple rounds of culturing (data not shown), indicating that the morphotypic variants were heritable. Additionally, whole genome shotgun sequencing (>46-fold coverage) was performed for each morphotypic variant and revealed that each had between one to two mutations across the entire genome (Table 1) that could be linked to its morphotypic change. For example, P. aeruginosa SCV has a deletion at position 628 (relative to the start site of the gene) on the pilT gene that results in a frame shift mutation at Thr210 of the twitching motility protein PilT. This mutation impairs twitching motility and can account for the reduced motility and, thus the small colony size of P. aeruginosa SCV. Similarly, the other morphotypic variants have mutations in genes encoding proteins with functions that can be related to their colony morphotypes. These proteins include the UDP-glucose lipid carrier transferase which is responsible for colanic acid biosynthesis in K. pneumoniae, the flagellar motor switch protein (FliM) which is essential for swimming motility of P. protegens, the chemotaxis-specific methylesterase which is required for chemotaxis in P. protegens, the cell division protein (FtsW) which is important for regulating cell shape of P. protegens and the group 1 family glycosyltransferase, involved in the biogenesis of cell envelope for P. protegens (Table 1).
These mutations were further verified by site-specific amplification and sequencing of three additional isolates for each type of morphotypic variant. The results showed that most of the additional isolates also possess mutations in the same gene, although these may be at different positions within that gene (Supplementary Table S2). Hence, morphotypic variants with identical colony morphology largely possessed mutations in the same gene, indicating that the frequency of morphotypic variants can be used as a proxy for intraspecific (genotypic) diversity within biofilms in this study.
Morphotypic variants differed phenotypically and were more competitive than their wild-type strains
Morphotypic variants differed from their wild-type strains in various functional traits ranging from motility, attachment, siderophore production as well as the synthesis of colanic acid (Table 2). These phenotypic differences have been associated with different niche utilization relative to the wild-type strains, as well as enhanced stress tolerance and better competitive abilities (Workentine et al., 2013). In fact, the morphotypic variants outcompeted their wild-type strains when co-cultured as biofilms, making up 67–99% of the total biomass in these polytypic co-cultures after 1 day of growth (Figure 2a).
Growth competitiveness of wild-type and morphotypic variants. (a) Relative biomass of the morphotypic variants in co-cultured biofilm populations with their respective wild-type strains at 0 h (white bars), 24 h (gray bars) and 72 h (black bars). (b) Relative biomass of the wild-type strains and the morphotypic variants in the triple-species biofilm communities. Each bar represents the average of three biological replicates. Error bars represent standard deviations.
In the triple-species biofilm communities, all of the morphotypic variants also performed better than their wild-type strains. For example, the relative biomass of P. aeruginosa wild type in the triple-species biofilm communities was typically 3.5% in contrast to the 29.1% achieved by P. aeruginosa SCV (Figure 2b). Similarly, the morphotypic variants of P. protegens achieved at least three times more biomass compared with the P. protegens wild type (Figure 2b). K. pneumoniae wild type and variant performed similarly (Figure 2b).
Interspecific diversity dominated biofilm communities
Given the results in Figure 2, and previous studies demonstrating that morphotypic variants were generally more tolerant to stresses relative to the wild-type strains (Mah and O'Toole, 2001; Stewart and William Costerton, 2001; Boles et al., 2004; Koh et al., 2012), we hypothesized that these morphotypic variants would dominate biofilm communities and play a crucial role in defending them against stresses.
Surprisingly, we found that while P. aeruginosa and K. pneumoniae biofilms produced morphotypic variants that represented up to 44% and 36% of their populations, respectively, neither species generated morphotypic variants when cultivated as biofilm communities (Figures 3a and b, respectively). The frequency of P. protegens morphotypic variants also decreased with increasing species richness, albeit to a lesser degree than for P. aeruginosa and K. pneumoniae (Figure 3c). Whole genome shotgun sequencing performed for 30 isolates (10 isolates for each species from the triple-species biofilm communities) with wild-type colony morphology revealed that only one of the P. aeruginosa isolates carried (two) SNPs, resulting in amino acid substitutions in the flhA gene involved in flagella biosynthesis. Similarly, two of the K. pneumoniae isolates, each had a single SNP, resulting in amino acid changes in genes involved in either lactate dehydrogenase or glutathione reductase synthesis. No mutation was found in the P. protegens isolates (Supplementary Table S3). Thus, overwhelmingly, the population of each species within the triple-species biofilm communities lacked or had low level of intraspecific variation, in contrast to each of the species when grown individually as biofilm populations.
Frequency of morphotypic variants in biofilm populations and communities. (a) P. aeruginosa, (b) P. protegens and (c) K. pneumoniae in biofilm populations (white bars), biofilm communities with two species (light and dark gray bars), and biofilm communities with all three species (black bars). Each bar represents the average from three biological replicates. Error bars represent s.d.
Interspecific diversity functionally substituted for intraspecific diversity
As intraspecific diversity decreased while interspecific diversity increased in the biofilm communities, it was compelling to ask whether interspecific diversity could functionally substitute for intraspecific diversity in mediating the performance of biofilm communities.
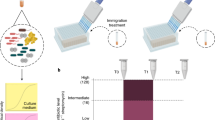
To investigate this, biofilms with different levels and types of diversity were established and treated with the anionic surfactant SDS, a general stressor that has been widely used to test the performance of biofilms (Barraud et al., 2006; Nett et al., 2008; Li et al., 2013; Lee et al., 2014). The biofilms tested include monotypic biofilm populations established with isolates (P. aeruginosa SCV, K. pneumoniae NMV, P. protegens SCV and CCV) that do not self-generate additional morphotypic variants (Supplementary Figure S1), polytypic biofilm populations established with wild-type strains that self-generate morphotypic variants (P. aeruginosa, K. pneumoniae and P. protegens) and triple-species biofilm communities with high interspecific, but low intraspecific diversity.
Monotypic P. aeruginosa SCV biofilms were sensitive to SDS, with <30% of the biomass remaining after SDS stress. In comparison, polytypic P. aeruginosa biofilms (high intraspecific diversity) and P. aeruginosa within the triple-species biofilm communities (communities have high inter-, but low intraspecific diversity) were significantly more tolerant to SDS stress, with more than 60% of their biomass remaining after treatment (Figure 4a). In addition, no significant difference was observed between the biomass remaining for polytypic P. aeruginosa biofilms and for P. aeruginosa within the triple-species biofilm communities. Similar findings were also observed for K. pneumoniae (Figure 4b), suggesting that interspecific diversity was functionally substituting for intraspecific diversity within the triple-species biofilm communities in mediating biofilm tolerance to SDS stress.
Sensitivity of biofilms with different levels of diversity to SDS stress. Biomass remaining after SDS stress for (a) P. aeruginosa; (b) K. pneumoniae and (c) P. protegens found in monotypic biofilm populations, polytypic biofilm populations and triple-species biofilm communities. P. aeruginosa biofilms were treated on day 4, while K. pneumoniae and P. protegens biofilms were treated on day 3. These time points were chosen based on when variants appear in the biofilms. Each bar represents the average from three biological replicates. Error bars represent s.d. (a–c) Denote significant differences (Tukey's honest significance test: P<0.1).
Members of a polytypic biofilm were equally protected
Both monotypic P. protegens SCV and CCV biofilms were more sensitive to SDS than the polytypic biofilms formed by wild-type P. protegens. However, when the two variants were grown together (P. protegens SCV+CCV) to establish polytypic biofilm populations, they were significantly more tolerant to SDS stress than when they were tested alone (Figure 4c), suggesting mutualistic interactions between the morphotypic variants. In addition, while individual variants were more sensitive to SDS stress than their wild type, SDS treatment did not significantly affect the percentages of morphotypic variants within polytypic biofilm populations (Supplementary Figure S2). This indicates that the morphotypic variants were equally protected from SDS stress in polytypic biofilms. These results collectively suggest that while the individual variants may show reduced tolerance to SDS, the presence of multiple variants or wild-type strains enhances tolerance of all members within the polytypic biofilms.
Morphotypic variants disrupted community tolerance
Given that both high intraspecific and interspecific diversity enhanced SDS tolerance (Figure 4), the reduction in morphotypic variant frequency in biofilm communities was surprising. We recently reported that the communal tolerance to SDS and tobramycin stresses for the triple-species biofilm community was achieved through balanced interactions between the three organisms (Lee et al., 2014). Since the morphotypic variants were found to outcompete other species in the biofilm communities (Figure 2b), it was hypothesized that the variants could disturb the species equilibrium in the community and thereby disrupt the communal tolerance. To test this hypothesis, the P. protegens SCV, was substituted for its wild-type strain in the establishment of triple-species biofilm communities. The results showed that P. protegens SCV made up 45% of the triple-species biofilm communities, which was significantly (P<0.05, one-way ANOVA) higher than the 18% achieved by the P. protegens wild-type strain. The substitution also led to significant reductions (P<0.05, one-way ANOVA) in the relative biomass of P. aeruginosa (6% to 0.05%) and K. pneumoniae (76% to 54%) in the triple-species biofilm communities (Figure 5a). These changes in species composition and potentially interactions between community members resulted in a twofold increase in the sensitivity of the variant substituted biofilm when exposed to SDS stress (P<0.05, one-way ANOVA) (Figure 5b). Thus, the morphotypic variants, which were more competitive than their wild-type strains in co-cultured biofilms, disrupted the communal tolerance observed for the triple-species biofilm communities. As a result, there is selection against the morphotypic variants in the presence of interspecific diversity and SDS stress.
Differences in composition and communal resistance to SDS stress for the biofilm communities. (a) Relative biomass of the different species within the biofilm communities composed of P. aeruginosa, P. protegens and K. pneumoniae (white bars) and biofilm communities composed of P. aeruginosa, P. protegens SCV and K. pneumoniae (black bar). (b) Biomass remaining after SDS stress for the two different biofilm communities. Each bar represents the average from three biological replicates. Error bars represent s.d.
Cell-free effluents-reduced frequency of morphotypic variants
The frequency of morphotypic variants was very low in biofilm communities (Figure 3) and this may be a consequence of selection against the less competitive variants in the communities. However, the results above (Figure 2b) show that the morphotypic variants in fact display a competitive advantage over their wild-type strains, based on increased biomass in the biofilm communities. Alternatively, the frequency of morphotypic variants in the biofilm communities may be influenced by other factors.
As bacteria secrete a wide range of extracellular factors including cues, signals and metabolites (McDougald et al., 2007; Garbeva et al., 2011), we added 10% v/v cell-free effluents from P. protegens and K. pneumoniae biofilms to P. aeruginosa biofilms. The frequency of the P. aeruginosa SCV was significantly reduced (P<0.05, Tukey's honest significance test) when cultured in the presence of cell-free effluents from both P. protegens and K. pneumoniae biofilms (Figure 6a). Similar results were also observed when P. protegens and K. pneumoniae (Figures 6b and c, respectively) were exposed to cell-free effluents from biofilms of another species, indicating that extracellular factors that were present in the cell-free effluents played a role in reducing the frequency of morphotypic variants.
Frequency of morphotypic variants grown with cell-free biofilm effluents. Frequency of (a) P. aeruginosa SCV, (b) K. pneumoniae NMV and (c) P. protegens morphotypic variants in biofilm populations grown with 10% v/v cell-free effluents of biofilm populations from the other two species. Each bar represents the average from three biological replicates. Error bars represent s.d.
Discussion
A range of ecological phenomena such as niche adaptation (Jousset et al., 2011) and tolerance to stresses (Mah and O'Toole, 2001) are facilitated by genotypic variation and the generation of morphotypic variants. This study, as well as those of others (Koh et al., 2007; Singh et al., 2009; Workentine et al., 2010; Cooper et al., 2014; Wang et al., 2015), have shown that the production of morphotypic variants is commonly associated with biofilm formation, where mutations are induced by stresses such as starvation and oxidative stress (Bjedov et al., 2003).
As a result, the frequency of morphotypic variants was used to measure intraspecific diversity within biofilms in this study. Together, this work and those of others (Koh et al., 2007; Cooper et al., 2014; McElroy et al., 2014) have shown that morphotypic variants are often associated with a number of genetic modifications that are specific to genes responsible for the changes in colony morphology. Moreover, these subsets of genes are also found to facilitate adaptation to the biofilm life cycle (McElroy et al., 2014), which is supported by the presence of morphotypic variants only in biofilms and not planktonic cultures. It was also determined in this study that wild-type biofilm isolates exhibited no significant mutations as demonstrated by whole genome sequencing (Supplementary Table 3). Furthermore, previous studies have also shown that wild-type biofilm isolates are similar to their wild-type strains for various functional traits (Koh et al., 2012; Woo et al., 2012), suggesting that they do not carry significant mutations. Therefore, as a consequence of specific selection pressures, mutations within biofilms might be restricted to a few genes that cause morphological changes and adaptations to the biofilm life cycle. Hence, the frequency of morphotypic variants is a good proxy to quantify intraspecific diversity within biofilms for this study.
Although there are many observations of genetic variation in biofilm populations, there is no information on the relationship between intraspecific and interspecific diversity, and whether they play complementary, antagonistic or substitutable roles in biofilm communities. Our data indicate a surprising interplay between these two types of diversity and their roles in an emergent property (stress tolerance) of biofilm communities. Intraspecific diversity can enhanced the tolerance of polytypic biofilm populations to SDS stress as a consequence of mutualistic interactions, such as metabolic cooperation between the different morphotypic variants and wild-type strains. This was clearly demonstrated by P. protegens since neither P. protegens SCV nor CCV was tolerant to SDS stress when challenged alone, but their co-cultures were more tolerant to SDS stress (Figure 4c). On the other hand, biofilm communities with low intraspecific diversity were tolerant to SDS stress, suggesting that individual tolerance mechanisms offered by the more tolerant species, such as secreted public goods including enzymes or matrix, protected the communities (Lee et al., 2014). When experimentally substituted into the biofilm communities, the morphotypic variants were found to lower the SDS tolerance of biofilm communities due to a change in species equilibrium and interspecies interactions (Figure 5). These results therefore suggest that interspecific diversity dominates biofilm communities and can functionally substitute for intraspecific diversity in mediating SDS tolerance.
This study has shown that intraspecific diversity was greatly reduced in biofilm communities, and that this reduction may be due to chemical factors, for example, extracellular factors present in the cell-free biofilm effluents (Figure 6). The mechanisms by which the biofilm communities and effluents affect the frequency of morphotypic variants are not known, but at least two general mechanisms are possible. First, morphotypic variants are generated in biofilm communities at similar rates as in biofilm populations, but there is selection against morphotypic variants (relative to wild types) in the multi-specific environment. Thus, the frequency of morphotypic variants remains low. The selection hypothesis is congruent with classical evolutionary theory, and the introduction of morphotypic variants into biofilm communities made the biofilms less tolerant to a particular selective regime, that is, SDS stress. However, in the absence of SDS, the morphotypic variants outgrew the wild-type strains in biofilm communities (Figure 2b), indicating that there is no selection against morphotypic variants in biofilm communities under other selective environments. This suggests that classical natural selection against variants in biofilm communities may not be the only explanation for their reductions.
As indicated above, variant formation can be attributed to mutations, and mutations can be induced by stressors such as starvation and oxidative stress (Bjedov et al., 2003). As one possible scenario, DNA damaging reactive oxygen and nitrogen species can build up in biofilm microcolonies and trigger the SOS response (Barraud et al., 2006). During the SOS response, the error prone Y-family DNA polymerases (Pol IV and PolV) are induced and led to frequent mutations, which result in the accumulation of mutations within microcolonies (Foster, 2005; Conibear et al., 2009). Therefore, a second, and not mutually exclusive, explanation for the reduction in frequency of morphotypic variants in mixed-species biofilms may be that the reduction or elimination of stressors results in less DNA damage. Factors present in the cell-free biofilm effluents might quench reactive oxygen and nitrogen species within the biofilms and result in a decrease in mutations and consequent morphotypic variant formation. To date, no specific molecular mechanism has been identified that could be responsible for a decrease in mutation rate, and thus a reduction in the generation (as opposed to the proliferation) of variants remains speculative. Further work is required to define the molecular mechanisms by which the generation and/or proliferation of intraspecific variation is reduced in the mixed-species biofilms.
Our results provide an explanation for the apparent rarity of morphotypic variants in environmental multi-species biofilm communities, in contrast to their common appearance in laboratory biofilm populations. From our results, we predict that the frequency of morphotypic variants in environmental multi-species biofilms would be negligible due to their reduction in the presence of interspecific diversity. However, morphotypic variants could still be found in biofilms when interspecific diversity is low and when they are selected for within specialized environments, such as in the lungs of cystic fibrosis patients (Folkesson et al., 2012; Woo et al., 2012). This further highlights the need to understand how variants respond to the selective regime in mono- versus multi-species biofilms, or biofilms in different environments.
References
Agrawal AA . (2003). Community genetics: new insights into community ecology by integrating population genetics. Ecology 84: 543–544.
Aguirre JD, Marshall DJ . (2012). Genetic diversity increases population productivity in a sessile marine invertebrate. Ecology 93: 1134–1142.
Anand AA, Vennison SJ, Sankar SG, Prabhu DI, Vasan PT, Raghuraman T et al. (2010). Isolation and characterization of bacteria from the gut of Bombyx mori that degrade cellulose, xylan, pectin and starch and their impact on digestion. J Insect Sci 10: 107.
Barraud N, Hassett DJ, Hwang SH, Rice SA, Kjelleberg S, Webb JS . (2006). Involvement of nitric oxide in biofilm dispersal of Pseudomonas aeruginosa. J Bacteriol 188: 7344–7353.
Bjedov I, Tenaillon O, Gérard B, Souza V, Denamur E, Radman M et al. (2003). Stress-induced mutagenesis in bacteria. Science 300: 1404–1409.
Boles BR, Thoendel M, Singh PK . (2004). Self-generated diversity produces ‘insurance effects’ in biofilm communities. Proc Natl Acad Sci USA 101: 16630–16635.
Burke C, Steinberg P, Rusch D, Kjelleberg S, Thomas T . (2011). Bacterial community assembly based on functional genes rather than species. Proc Natl Acad Sci USA 108: 14288–14293.
Burmolle M, Webb JS, Rao D, Hansen LH, Sorensen SJ, Kjelleberg S . (2006). Enhanced biofilm formation and increased resistance to antimicrobial agents and bacterial invasion are caused by synergistic interactions in multispecies biofilms. Appl Environ Microbiol 72: 3916–3923.
Cardinale BJ, Palmer MA, Collins SL . (2002). Species diversity enhances ecosystem functioning through interspecific facilitation. Nature 415: 426–429.
Chazal PM . (1995). Pollution of modern metalworking fluids containing biocides by pathogenic bacteria in France. Reexamination of chemical treatments accuracy. Eur J Epidemiol 11: 1–7.
Conibear TC, Collins SL, Webb JS . (2009). Role of mutation in Pseudomonas aeruginosa biofilm development. PLoS One 4: e6289.
Cooper VS, Staples RK, Traverse CC, Ellis CN . (2014). Parallel evolution of small colony variants in Burkholderia cenocepacia biofilms. Genomics 104: 447–452.
Crawford KM, Rudgers JA . (2012). Plant species diversity and genetic diversity within a dominant species interactively affect plant community biomass. J Ecol 100: 1512–1521.
Davey ME, O'Toole GA . (2000). Microbial biofilms: from ecology to molecular genetics. Microbiol Mol Biol Rev 64: 847–867.
Drenkard E, Ausubel FM . (2002). Pseudomonas biofilm formation and antibiotic resistance are linked to phenotypic variation. Nature 416: 740–743.
Dungey HS, Potts BM, Whitham TG, Li HF . (2000). Plant genetics affects arthropod community richness and composition: evidence from a synthetic eucalypt hybrid population. Evolution 54: 1938–1946.
Elton CS . (1958) The Ecology of Invasion by Plants and Animals. Chicago, IL, USA: University of Chicago Press.
Folkesson A, Jelsbak L, Yang L, Johansen HK, Ciofu O, Hoiby N et al. (2012). Adaptation of Pseudomonas aeruginosa to the cystic fibrosis airway: an evolutionary perspective. Nat Rev Microbiol 10: 841–851.
Foster PL . (2005). Stress responses and genetic variation in bacteria. Mutat Res 569: 3–11.
Garbeva P, Silby MW, Raaijmakers JM, Levy SB, Boer Wd . (2011). Transcriptional and antagonistic responses of Pseudomonas fluorescens Pf0-1 to phylogenetically different bacterial competitors. ISME J. 5: 973–985.
Hersch-Green EI, Turley NE, Johnson MT . (2011). Community genetics: what have we accomplished and where should we be going? Philos Trans R Soc Lond B Biol Sci 366: 1453–1460.
Hughes AR, Inouye BD, Johnson MT, Underwood N, Vellend M . (2008). Ecological consequences of genetic diversity. Ecol Lett 11: 609–623.
Jessup CM, Kassen R, Forde SE, Kerr B, Buckling A, Rainey PB et al. (2004). Big questions, small worlds: microbial model systems in ecology. Trends Ecol Evol 19: 189–197.
Johnson MTJ, Agrawal AA . (2005). Plant genotype and environment interact to shape a diverse arthropod community on evening primrose (Oenothera biennis. Ecology 86: 874–885.
Jousset A, Schulz W, Scheu S, Eisenhauer N . (2011). Intraspecific genotypic richness and relatedness predict the invasibility of microbial communities. ISME J 5: 1108–1114.
Koh KS, Lam KW, Alhede M, Queck SY, Labbate M, Kjelleberg S et al. (2007). Phenotypic diversification and adaptation of Serratia marcescens MG1 biofilm-derived morphotypes. J Bacteriol 189: 119–130.
Koh KS, Matz C, Tan CH, Le HL, Rice SA, Marshall DJ et al. (2012). Minimal increase in genetic diversity enhances predation resistance. Mol Ecol 21: 1741–1753.
Latzel V, Allan E, Bortolini Silveira A, Colot V, Fischer M, Bossdorf O . (2013). Epigenetic diversity increases the productivity and stability of plant populations. Nat Commun 4: 2875.
Lee KW, Arumugam K, Purbojati RW, Tay QX, Williams RB, Kjelleberg S et al. (2013). Draft Genome Sequence of Klebsiella pneumoniae Strain KP-1. Genome Announc 1: e01082-13.
Lee KW, Periasamy S, Mukherjee M, Xie C, Kjelleberg S, Rice SA . (2014). Biofilm development and enhanced stress resistance of a model, mixed-species community biofilm. ISME J 8: 894–907.
Li L, Molin S, Yang L, Ndoni S . (2013). Sodium dodecyl sulfate (SDS)-loaded nanoporous polymer as anti-biofilm surface coating material. Int J Mol Sci 14: 3050–3064.
Little AE, Robinson CJ, Peterson SB, Raffa KF, Handelsman J . (2008). Rules of engagement: interspecies interactions that regulate microbial communities. Annu Rev Microbiol 62: 375–401.
Mah T-FC, O'Toole GA . (2001). Mechanisms of biofilm resistance to antimicrobial agents. Trends Microbiol 9: 34–39.
May RM . (1973) Stability and complexity in model ecosystems. Princeton University Press: Princeton, NJ, USA.
McDougald D, Rice SA, Kjelleberg S . (2007). Bacterial quorum sensing and interference by naturally occurring biomimics. Anal Bioanal Chem 387: 445–453.
McElroy KE, Hui JG, Woo JK, Luk AW, Webb JS, Kjelleberg S et al. (2014). Strain-specific parallel evolution drives short-term diversification during Pseudomonas aeruginosa biofilm formation. Proc Natl Acad Sci USA 111: E1419–E1427.
Miles AA, Misra SS, Irwin JO . (1938). The estimation of the bactericidal power of the blood. J Hyg (Lond) 38: 732–749.
Naeem S, Li S . (1997). Biodiversity enhances ecosystem reliability. Nature 390: 507–509.
Nett JE, Guite KM, Ringeisen A, Holoyda KA, Andes DR . (2008). Reduced biocide susceptibility in Candida albicans biofilms. Antimicrob Agents Chemother 52: 3411–3413.
Rowntree JK, Shuker DM, Preziosi RF . (2011). Forward from the crossroads of ecology and evolution. Philos Trans R Soc Lond B Biol Sci 366: 1322–1328.
Singh R, Ray P, Das A, Sharma M . (2009). Role of persisters and small-colony variants in antibiotic resistance of planktonic and biofilm-associated Staphylococcus aureus: an in vitro study. J Med Microbiol 58: 1067–1073.
Stewart PS, Costerton J . (2001). Antibiotic resistance of bacteria in biofilms. The Lancet 358: 135–138.
Tilman D, Downing JA . (1994). Biodiversity and stability in graslands. Nature 367: 363–365.
van der Woude MW, Baumler AJ . (2004). Phase and antigenic variation in bacteria. Clin Microbiol Rev 17: 581–611.
Vellend M, Geber MA . (2005). Connections between species diversity and genetic diversity. Ecol Lett 8: 767–781.
Wang D, Dorosky RJ, Han CS, Lo CC, Dichosa AE, Chain PS et al. (2015). Adaptation genomics of a small-colony variant in a Pseudomonas chlororaphis 30-84 biofilm. Appl Environ Microbiol 81: 890–899.
Whitham TG, Bailey JK, Schweitzer JA, Shuster SM, Bangert RK, LeRoy CJ et al. (2006). A framework for community and ecosystem genetics: from genes to ecosystems. Nat Rev Genet 7: 510–523.
Whitham TG, Young WP, Martinsen GD, Gehring CA, Schweitzer JA, Shuster SM et al. (2003). Community and ecosystem genetics: a consequence of the extended phenotype. Ecology 84: 559–573.
Wimp GM, Martinsen GD, Floate KD, Bangert RK, Whitham TG . (2005). Plant genetic determinants of arthropod community structure and diversity. Evolution 59: 61–69.
Woo JK, Webb JS, Kirov SM, Kjelleberg S, Rice SA . (2012). Biofilm dispersal cells of a cystic fibrosis Pseudomonas aeruginosa isolate exhibit variability in functional traits likely to contribute to persistent infection. FEMS Immunol Med Microbiol 66: 251–264.
Workentine ML, Harrison JJ, Weljie AM, Tran VA, Stenroos PU, Tremaroli V et al. (2010). Phenotypic and metabolic profiling of colony morphology variants evolved from Pseudomonas fluorescens biofilms. Environ Microbiol 12: 1565–1577.
Workentine ML, Wang S, Ceri H, Turner RJ . (2013). Spatial distributions of Pseudomonas fluorescens colony variants in mixed-culture biofilms. BMC Microbiol 13: 175.
Acknowledgements
KWKL was supported by the National Research Foundation Singapore under the National Research Foundation Environmental and Water Technologies PhD Scholarship Programme administered by the Environment and Water Industry Programme Office. We acknowledge financial support from the Singapore Centre on Environment Life Sciences Engineering, whose research is supported by the National Research Foundation Singapore, Ministry of Education, Nanyang Technological University and National University of Singapore, under its Research Centre of Excellence Programme. We also acknowledge the work done by the genomics sequencing facility in Singapore Centre on Environment Life Sciences Engineering.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Kelvin Lee, K., Hoong Yam, J., Mukherjee, M. et al. Interspecific diversity reduces and functionally substitutes for intraspecific variation in biofilm communities. ISME J 10, 846–857 (2016). https://doi.org/10.1038/ismej.2015.159
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.159
This article is cited by
-
Carbon starvation of Pseudomonas aeruginosa biofilms selects for dispersal insensitive mutants
BMC Microbiology (2021)
-
rpoS-mutation variants are selected in Pseudomonas aeruginosa biofilms under imipenem pressure
Cell & Bioscience (2021)
-
The initial inoculation ratio regulates bacterial coculture interactions and metabolic capacity
The ISME Journal (2021)
-
Emergence of a Synergistic Diversity as a Response to Competition in Pseudomonas putida Biofilms
Microbial Ecology (2020)
-
Transcriptional changes involved in inhibition of biofilm formation by ε-polylysine in Salmonella Typhimurium
Applied Microbiology and Biotechnology (2020)