Abstract
Despite knowledge that viruses are abundant in natural ecosystems, there is limited understanding of which viruses infect which hosts, and how both hosts and viruses respond to those interactions—interactions that ultimately shape community structure and dynamics. In Deep Lake, Antarctica, intergenera gene exchange occurs rampantly within the low complexity, haloarchaea-dominated community, strongly balanced by distinctions in niche adaptation which maintain sympatric speciation. By performing metaproteomics for the first time on haloarchaea, genomic variation of S-layer, archaella and other cell surface proteins was linked to mechanisms of infection evasion. CRISPR defense systems were found to be active, with haloarchaea responding to at least eight distinct types of viruses, including those infecting between genera. The role of BREX systems in defending against viruses was also examined. Although evasion and defense were evident, both hosts and viruses also may benefit from viruses carrying and expressing host genes, thereby potentially enhancing genetic variation and phenotypic differences within populations. The data point to a complex inter-play leading to a dynamic optimization of host–virus interactions. This comprehensive overview was achieved only through the integration of results from metaproteomics, genomics and metagenomics.
Similar content being viewed by others
Introduction
Hypersaline lakes from warm-hot latitudes that harbor halophilic members of the Archaea (haloarchaea) tend to sustain a low abundance of higher trophic predators and support high viral densities (Oren et al., 1997; Santos et al., 2007; Atanasova et al., 2012; Santos et al., 2012; Luk et al., 2014). In polar lakes, metazoan grazers also tend to occur in low abundance, and viruses are hypothesized to have a key role in the microbial loop and to function as drivers of microbial evolution (Kepner et al., 1998; Anesio and Bellas, 2011; Wilkins et al., 2013). Metagenomic studies of Antarctic lakes (freshwater to hypersaline) have revealed high viral diversity, novel viruses, unusual host–virus relationships including virus resistance, and a high level of viral regulation of microbial loop dynamics that appears to be driven by the seasonal polar light regime (López-Bueno et al., 2009; Lauro et al., 2011; Yau et al., 2011; Wilkins et al., 2013). Algal viruses including virophages and bacteriophages have been characterized in hypersaline Organic Lake (~230 g l−1 maximum salinity), but Archaea are rare in this meromictic system (Yau et al., 2011, 2013). Therefore, although it may be expected that viruses would have a particularly important role in aquatic environments that are both polar and sufficiently hypersaline to sustain high abundances of haloarchaea, to date, virus–host interactions in such systems have not been examined.
Deep Lake is a ∼3500-years-old, marine-derived system in the Vestfold Hills region of East Antarctica (68°33’36.8 S, 78°11’48.7 E) that is so saline it remains liquid even when temperatures drop to –20 °C (Campbell 1978; Ferris and Burton 1988; Gibson 1999). By analyzing the metagenome data of biomass taken directly from the lake, and genome sequences of four isolates (Halorubrum lacusprofundi strain ACAM34 (Franzmann et al., 1988); Halohasta litchfieldiae strain tADL (Mou et al., 2012); strain DL31; strain DL1 (Halobacterium sp.)), the lake was found to be dominated by haloarchaea throughout the 36 m-deep water column (DeMaere et al., 2013; Williams et al., 2014). The microbial community in the lake differs greatly to warm-hot latitude hypersaline systems, has overall low complexity and is hierarchically structured with tADL representing ~44%, DL31 ~18%, Hrr. lacusprofundi ~10% and DL1 ~0.3%; in combination, ~72% of the entire lake community (DeMaere et al., 2013).
A striking feature of the genomes of the four isolates is the sharing of long (up to 35 kb) and ~100% conserved, high identity regions (HIRs) (DeMaere et al., 2013). Being indicative of promiscuous gene exchange, such sharing across genus-level boundaries could potentially drive homogenization of the Deep Lake community. However, genomic traits exist that confer ecotype distinctions, demonstrating that niche adaptation counteracts coalescence and helps to maintain sympatric speciation. Indicators of niche adaptation were observed at the genus level in the form of genomic distinctiveness of the four isolates (for example, major differences in resource utilization) and at the strain level as genomic variation (that is, as determined by metagenome fragment recruitment (FR) coverage and de novo assembly) of sub-populations of the lake community (DeMaere et al., 2013; Williams et al., 2014).
The genome of tADL is composed of a single replicon, whereas Hrr. lacusprofundi, DL31 and DL1 each have multiple replicons (DeMaere et al., 2013). The genomes of the four isolates share many mobile genetic elements, particularly insertion sequences and genes associated with viruses such as integrases (DeMaere et al., 2013). In Hrr. lacusprofundi, a putative defective provirus (Hlac-Pro1) is present which has sequence similarity to the head-tailed virus BJ1 (Pagaling et al., 2007; Krupovič et al., 2010; DeMaere et al., 2013). BJ1-like sequences are also present in the genomes of DL31 and DL1, and are associated with regions of genome variation, possibly reflecting virus-infected subpopulations (for example, regions of low metagenome FR) (DeMaere et al., 2013). These data indicate that the Antarctic haloarchaea encode viral genes, possibly representing integrated lysogens and/or virions, and that viruses therefore potentially infect and influence gene transfer, genome composition and community structure in Deep Lake.
Host mechanisms involved in responding to viruses have been proposed for haloarchaea from warm-hot latitudes, including evasion via cell surface proteins which have sequence variation and may therefore reduce rates of viral attachment, and defenses involving CRISPR systems (Breitbart and Rohwer, 2005; Legault et al., 2006; Cuadros-Orellana et al., 2007; Pyatibratov et al., 2008; Rodriguez-Valera et al., 2009; Emerson et al., 2012; Garcia-Heredia et al., 2012; Emerson et al., 2013; Maier et al., 2013; Vestergaard et al., 2014). CRISPR systems potentially provide haloarchaea with a dynamic means to respond and defend against specific haloarchaeoviruses (for example, Andersson and Banfield, 2008; Tyson and Banfield, 2008; Heidelberg et al., 2009; Anderson et al., 2010; Held et al., 2010; Emerson et al., 2013; Maier et al., 2013; Vestergaard et al., 2014). Spacers are incorporated into loci between short direct repeat sequences and, along with the CRISPR-associated (Cas) proteins, are characteristic of the host in which they evolved. Spacers in particular provide a historical record of previous encounters with invading DNA, enabling specific interactions to be inferred about viruses and the hosts they infect. To date, analyses of CRISPR systems of haloarchaeal communities have only been examined using DNA sequence data of environmental samples, and not from functional studies (for example, metaproteomics) which have the added potential to inform about the genes expressed by active members of the communities (both hosts and viruses), and the molecular mechanisms of ongoing interactions. Recently, Bacteriophage Exclusion (BREX) systems were described that function in Bacillus species by preventing viral replication (Goldfarb et al., 2015). BREX gene clusters are present in ~10% of sequenced bacterial and archaeal genomes, including the Type 5 system which has only been identified in haloarchaea, including DL31 and Hrr. lacusprofundi (Goldfarb et al., 2015).
Metaproteomic analyses of hypersaline systems harboring haloarchaea have not been reported, but have been performed on Antarctic marine (Williams et al., 2012, 2013) and lacustrine (Ng et al., 2010; Lauro et al., 2011; Yau et al., 2011) systems. To explore system-wide, virus–host interactions, we used metaproteomics to identify Deep Lake viral proteins, and cellular proteins relevant to viral infection and evasion. Metaproteomics was performed on biomass that was collected in the austral summer of 2008 by sequential size fractionation (20–3 μm, 3–0.8 μm, 0.8–0.1 μm) that directly matched the samples used for metagenomics (DeMaere et al., 2013). Led by the protein identifications, in-depth analyses of metagenome/genome data were performed to assess viral gene/genome content and function, genome variation associated with expressed genes and active host defense systems (for example, CRISPR systems). The data were integrated to obtain a systems-level view of the active host–virus interactions occurring in this novel aquatic Antarctic system.
Materials and methods
Metaproteomics was performed based on methods described previously (Ng et al., 2010; Williams et al., 2012, 2013), from biomass obtained from Deep Lake (68°33’36.8 S, 78°11’48.7 E), Vestfold Hills, Antarctica between 30 November and 5 December 2008, by filtering water taken from 0, 5, 13, 24 and 36 m depths, through a 20 μm prefilter sequentially onto 293-mm polyethersulfone membrane filters with 3.0, 0.8 and 0.1 μm pore sizes, as described previously (DeMaere et al., 2013; Williams et al., 2014). All CRISPR repeat sequences identified by CRISPR Recognition Tool (Bland et al., 2007) were re-examined using CRISPRFinder (default settings; Grissa et al., 2007) and inspected for motifs associated with functional repeats (Maier et al., 2013). All data were analysed based on approaches developed for studying Antarctic lake microbial communities (Ng et al., 2010; Lauro et al., 2011; Yau et al., 2011; DeMaere et al., 2013; Williams et al., 2014). Full details are provided in Supplementary Information.
Results
Protein identifications were made by matching mass spectra to peptides derived from a composite database that was comprised of assemblies of Deep Lake metagenome data and the genome sequences of the four haloarchaeal isolates, tADL, DL31, Hrr. lacusprofundi and DL1 (DeMaere et al., 2013). All proteins reported were manually verified to contain at least one unique peptide, and were annotated via BLASTP searches on IMG and ExPASy, recording the best matching sequence and percentage identity. Proteins sharing the same set of detected peptides were grouped into protein families and are described in the Supplementary Information.
A notable feature of the metaproteomics was the proportion of proteins (194 from the 1109) which had best matches to the genomes of the four isolates but with <100% sequence identity. These proteins were identified from metagenome contigs and represent genomic variants (that is, phylotypes) of the four isolates that are present within the lake population (Supplementary Table S1, also see Supplementary Information). The term ‘variant’ is used to describe proteins that have <100% amino acid sequence identity to the sequences from the genomes of the four isolates. For all variants, protein matches were assigned based on at least one unique peptide mapping to a region of sequence variation (in the metagenome contig). Percent identity for a variant provides a measure of the extent of protein sequence identity relative to the best matching sequence in the genome of one of the four isolates; the lower the identity, the higher the extent of variation.
Cell surface protein variation
Annotation as a ‘cell surface protein’ (that is, predicted to be exposed to the external environment) required the presence of an N-terminal secretion signal peptide and/or a single C-terminal transmembrane helix and/or homology to experimentally characterized cell surface proteins, including S-layer proteins, archaeal flagella (archaella) and adhesion pili. Cell surface proteins accounted for ~24% of detected spectra and ~10% of protein identifications (Supplementary Table S2).
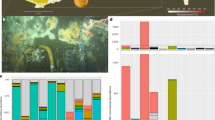
The class of proteins most represented by protein variation was cell surface proteins (Figure 1,Supplementary Table S2). The proportion of variant cell surface proteins relative to the total number of variants detected was: tADL, 18/178; DL31, 3/6; Hrr. lacusprofundi, 5/7 (Supplementary Table S2). The extent of variation ranged from: tADL, 77 to 29%; DL31, 47 to 39%; Hrr. lacusprofundi, 54 to 27%. Cell surface proteins represented a particularly high proportion of variants with high levels of variation (<60% identity) (Figure 1): tADL, 11/13; DL31, 3/4; Hrr. lacusprofundi, 5/5. Moreover, these cell surface variants with high levels of variation were among the most abundant DL31 and Hrr. lacusprofundi variants detected in the metaproteome, representing ~82% and ~99% of all spectra for proteins with variation. For tADL, the proportion was lower (15%).
Relative abundance of cell surface protein variants. All detected protein variants from tADL, DL31 and Hrr. lacusprofundii are shown categorized into cell surface (dark grey) and all other functions (light grey), and grouped according to their identity relative to the best matching sequence in their respective genome. The total number of variant proteins was 191, and for each range of identities, the number of variants was: 90–99%, 109 proteins; 80–89%, 36; 70–79%, 18; 60–69%, 6; 50–59%, 7; 40–49%, 8; 30–39%, 5; 20–29%, 2. Variants with high levels of variation tended to be cell surface proteins.
S-layer proteins
The S-layer is the outer layer of the haloarchaeal cell, and is composed of a glycosylated protein that forms a highly porous, paracrystalline lattice (Albers and Meyer, 2011). tADL, DL31 and DL1 encode one S-layer protein (halTADL_1043, Halar_0829 and HalDL1_0395, respectively), while Hrr. lacusprofundi is conspicuous in encoding two (Hlac_2976, Hlac_0412). Because S-layer proteins encase the cell they are among the most abundant proteins synthesized by the cell (Jarrell et al., 2010), and this is reflected in their high spectral counts (for example, Halar_0829 variant, second highest spectral count) (Supplementary Table S1).
The S-layer proteins are large (834–1063 amino acids), contain a N-terminal surface glycoprotein signal peptide (~35 amino acids) and a C-terminal PGF-CTERM archaeal protein-sorting signal (~25 amino acids) separated by a long extra-cytoplasmic domain (Supplementary Table S5). Multiple tADL, DL31 or Hrr. lacusprofundi S-layer variants were identified, all of which were detected through unique sets of peptides (Supplementary Figure S1,Supplementary Table S3): five tADL proteins, 34–51% identity; two DL31 proteins, 45–47% identity; one Hrr. lacusprofundi protein, 38% identity. The variation occurs primarily within the long extra-cytoplasmic domain of the S-layer protein (Supplementary Figure S1). Non-variants (that is, 100% identity to the sequence from the isolate genomes) were also identified for tADL, DL31, Hrr. lacusprofundi and DL1, but their abundance was much lower than variant forms (Supplementary Table S3). As the metaproteome data revealed that Hrr. lacusprofundi synthesized two types of S-layer proteins, a range of combinations of variant (Hlac_2976, 38%) and non-variant (Hlac_0412, Hlac_2976) S-layer proteins could conceivably be supported within the Hrr. lacusprofundi population growing in the lake.
Archaella proteins
Archaella are type IV pili-like structures that allow cells to swim by ATP-driven rotation (Alam and Oesterhelt, 1984; Streif et al., 2008), and represent an important class of cell surface protein detected in the metaproteome. In contrast to DL31 which does not encode archaella, tADL has seven archaellin genes (halTADL_0078/1543/1544/1810/1811/1812/1813). Proteins corresponding to all the tADL genes were detected except for halTADL_1543, and variant forms (75–77% identity) of four of these were also detected (halTADL_0078/1544/1812/1813). The non-variant form of halTADL_1544 is the single most abundant protein in the metaproteome, with halTADL_1812/1813 being the third and eighth most abundant, respectively. Variant forms tended to be proportionately less abundant. The archaellin genes are overall similar to each other (>65% identity) and the variation observed tended to occur in the N-terminal region and around amino acids 70, 120 and 150 (Supplementary Figure S2), indicating that variation was localized to specific protein domains. Hrr. lacusprofundi has one archaellin gene, Hlac_2557, for which only a variant (54% identity) was detected. DL1 has three genes halDL1_1517/1518/1563, and despite the low abundance of DL1 in the lake, non-variant forms of halDL1_1517/1518 were detected, and a variant of halDL1_1563 (41% identity) (Supplementary Tables S1, S3).
Adhesin and other predicted cell surface proteins
Adhesins are cell surface proteins containing adhesion domains and are involved in cell attachment (for example, pili; Esquivel et al., 2013). Two tADL adhesin variants were detected in the metaproteome with 66% and 33% identity to haltADL_1387 and halTADL_1885, respectively. The non-variant form of halTADL_1387 was also detected (Supplementary Table S3). Six additional tADL cell surface protein variants with 29–77% identity were detected: halTADL_0751/0878/1047/1761/1765 which have predicted N-terminal signal sequences and extra-cytoplasmic domains, and halTADL_1403 which has a C-terminal transmembrane domain and nine extra-cytoplasmic bacterial Ig-like, group 1 domains (Supplementary Table S3). For Hrr. lacusprofundi, two variants (34% and 27% identity) of Hlac_2824 were detected, but not the non-variant form. Both contain multiple PKD domains and a long extra-cytoplasmic structure.
High identity regions
Shared HIRs are notable features of the four isolate genomes, and network analysis established that Hrr. lacusprofundi had the greatest number of links to the other genomes (DeMaere et al., 2013). Metaproteomics identified nine proteins from HIRs (Supplementary Table S4), all of which matched HIRs in Hrr. lacusprofundi as well as at least one other isolate, and in one case, protein 744 matched to genes from all four isolates. In three cases, proteins mapped to two genes from the one HIR. Although the proteins cannot be assigned to a specific host, their presence illustrates that HIR genes are expressed. The annotated functions of the proteins included maltose-binding protein, metallophosphoesterase, nucleotidyltransferase, TATA-binding protein, and hypothetical proteins.
Metagenome data supportive of variation
By mapping metagenome reads to the genomes of the four isolates, previous analyses identified specific regions of the genomes that had low FR coverage and were therefore markers of genome variation that corresponded to phylotypes (DeMaere et al., 2013). The FR coverage for genes of cell surface variants was examined, and generally found to be low (Supplementary Table S3). For example, the average rpkm (reads per kilobase per 1 000 000 recruited reads) for the whole replicon compared with just the S-layer genes was 460/105 for halTADL_1043, 474/141 for Halar_0829, 3114/122 for Hlac_2976 and 452/67 for Hlac_0412. For halTADL_1043, rpkm values are low from the beginning of the gene through to the end of the region that encodes the extra-cytoplasmic portion (17 rpkm), whereupon the coverage increases to 518 rpkm through the end of the gene which encodes the conserved PGF-CTERM archaeal sorting signal (Figure 2). The low coverage extends into halTADL_1042, a putative membrane-bound sialidase. A similar pattern of low FR coverage throughout the region encoding the extra-cytoplasmic domain of the S-layer genes was observed for DL31 and Hrr. lacusprofundi.
Metagenome data indicative of genome variation. Upper panel: Region of low fragment recruitment (FR) for tADL S-layer gene, halTADL_1043. The upper portion of the panel shows FR coverage obtained from Deep Lake metagenome reads using the tADL genome (DeMaere et al., 2013) viewed using Artemis (Carver et al., 2012). The main tADL S-layer protein with six detected variants (gene 1043); cell surface protein genes (1040, 1047), including putative membrane-bound sialidase (1042); proteins detected with variation (1042, 1047). Lower panel: Expressed S-layer gene from Hrr. lacusprofundi (Hlac_2976) neighboring viral gene, transposases and HIR. Metagenome contig number ctg7180000444255. Nucleotide sequence identity between metagenome and genome sequences is shown as %. Both panels: Locus tags (numbers); genes encoding proteins detected in the metaproteome (purple); viral genes (green); transposases (yellow); HIR (red).
The metagenome contigs were also useful for identifying changes in gene context for variants (by comparing with the isolate genome sequences). For example, the Hrr. lacusprofundi S-layer gene Hlac_2976 appears to be located in a mobile region of the genome near two insertion sequences and a HIR region, with the metagenome contig of the variant containing different neighboring genes, including a putative BJ1 virus gene adjacent to the S-layer gene (Figure 2).
Viruses
Eight distinct major capsid proteins (MCPs) were identified in the metaproteome indicative of the detection of eight different head-tailed viruses (Caudovirales) (Supplementary Table S5). Five of the MCPs best matched to MCPs of isolated haloarchaeal siphoviruses: two to members of the HCTV-1-like subgroup (for example, HCTV-1 and HVTV-1), and one each to HCTV-2, HHTV-1 and Halorubrum virus CGΦ46 (Kukkaro and Bamford, 2009; Atanasova et al., 2012; Pietilä et al., 2013; Senčilo and Roine, 2014). However, the latter was encoded on a metagenome contig (deg7180000401097) that also contained genes homologous to Halorubrum virus BJ1, a possible lysogen (Pagaling et al., 2007), as well as to CGΦ46. Another metaproteome MCP matched to a contig (deg7180000409203) that contained genes homologous to the Hlac-Pro1 provirus of Hrr. lacusprofundi (Hlac_0736-0775), which has extensive sequence similarity with BJ1 (Krupovič et al., 2010), although the MCP encoded on the contig was homologous to MCPs from the HCTV-1-like subgroup. This underscores the mosaic nature of haloarchaeoviruses, with individual genes showing sequence identities to different viruses and cellular genes (Krupovič et al., 2010). Another MCP in the metaproteome matched to putative lytic head-tailed viruses (eHP-12 and eHP-6) that were assembled from metagenome data (Garcia-Heredia et al., 2012). The last MCP matched to the bacteriophage myovirus VBM1 from the marine bacterium Vibrio parahaemolyticus (YP_007674343.1). Metaproteome proteins that best matched to VBM1 (MCP and a hypothetical) may originate from a virus that infected a Halomonas sp. as this genus also belongs to the Gammaproteobacteria, and Halomonas sp. represents one of the most abundant bacterial taxa in Deep Lake (~0.8% of the lake community) (DeMaere et al., 2013).
On the basis of spectral counts, the HCTV-1-like viruses were the most abundant, and a prohead protease was detected for two of the HCTV-1-like viruses, in addition to MCPs (Supplementary Table S5). Typical of HCTV-1-like viruses, the MCP and prohead protease were encoded within the same gene cluster. As the prohead protease is involved in maturation of the capsid (Cheng et al., 2004; Pietilä et al., 2013), the detection of these proteases is indicative of active lytic cycle gene expression. The diameter of heads of haloarchaeoviruses is typically ⩽70 nm (Kukkaro and Bamford, 2009; Pietilä et al., 2013) and the minimum filter pore size during sampling was 0.1 μm. As a result, it is likely that rather than planktonic haloarchaeoviruses, the filters would tend to retain cells infected by viruses, viruses attached to cells, proviruses and any viruses with a diameter larger than 0.1 μm (Emerson et al., 2013). The detection of a scaffold protein best matching to HRTV-7 (Pietilä et al., 2013) is also indicative of the expression of myovirus genes involved in the lytic cycle (Supplementary Table S5). These data indicate that the detected viral proteins arose during active viral life cycles.
Six other proteins matched to five contigs that all contained at least one gene of putative viral origin, although it cannot be determined whether these particular contigs originate from viruses. The inferred functions of these proteins were: a hypothetical protein, a pilin (pilus protein–halTADL1885 homolog) (Supplementary Figure S3), two predicted cell surface proteins and two linocin M18 type bacteriocins (Valdés-Stauber and Scherer, 1994) (Supplementary Table S5). Bacteriocins are often derived from proviruses that are subsequently ‘domesticated’ by their hosts (Bobay et al., 2014). One of the cell surface proteins was encoded on a contig that includes an open reading frame from Halorubrum virus GNf2. The linocin-containing contigs each possess orthologs of virus and plasmid related elements present in certain Halorubrum spp., and one contig contained an open reading frame matching a prohead protease from HGTV-1 myovirus.
Overall, a total of 25 viral proteins were identified in the metaproteome illustrating that proteins of viral origin are expressed in the Deep Lake community. However, because metaproteome matches were made to metagenome contigs and not assembled virus genomes, it is not possible to unambiguously determine whether the proteins were derived from extracellular virions, viruses undergoing a lytic cycle, integrated active viruses (proviruses) or integrated inactivated viruses (viral remnants).
Defense against invading DNA
The metaproteome provided evidence for functional CRISPR systems. Cas proteins were identified in the metaproteome that best matched to tADL (Cas7, Cas8b), Hrr. lacusprofundi (Cas7, Cas10d, Csc2) and an unknown taxon of the Halobacteriaceae (Cas7), but none to DL31 or DL1 (Supplementary Figure S4,Supplementary Table S6). Cas7 forms the ‘backbone’ of the CRISPR RNA (crRNA) guided Cascade (CRISPR-associated complex for antiviral defense) in type I-B systems and protects the crRNA from degradation (Sorek et al., 2013). In type I-D systems, Csc2 appears to perform a similar function to Cas7 (Haft et al., 2005; Makarova et al., 2011a and b). The Cas8b and Cas10d proteins constitute the large subunit of the Cascade complex in I-B and I-D systems, respectively, and may be involved in binding the 5’ end of the crRNA and engaging invading DNA (Makarova et al., 2011a, 2011b; Sorek et al., 2013; Brendel et al., 2014). The detection of protein subunits of the Cascade complex in the metaproteome is indicative of Deep Lake haloarchaea CRISPR systems actively scanning and targeting invading DNA.
CRISPR systems present in the isolate genomes were analyzed (Supplementary Figure S4 and Supplementary Table S6) and cas genes were identified in discrete gene clusters. tADL and DL1 each possessed a single I-B system, DL31 a single I-D system, and Hrr. lacusprofundi one I-B and one I-D system located on a secondary replicon with each flanked by transposases. A CRISPR locus was situated adjacent to each of the cas gene clusters, and additional CRISPR loci were present in the genomes, bringing the total to four CRISPR loci for tADL, two each for Hrr. lacusprofundi and DL31, and one for DL1 (Supplementary Table S7). Hrr. lacusprofundi also encodes a putative three-gene V-1 CRISPR interference module (Vestergaard et al., 2014) encoded on a different replicon to the I-B and I-D systems (Hlac_2813–2815). Contigs that included cas gene clusters for the four isolate genomes were identified in the metagenome. Also, a number of additional cas gene clusters from other organisms were identified on metagenome contigs (Supplementary Table S8).
Complete BREX systems are present in DL31 and Hrr. lacusprofundi (Goldfarb et al., 2015), with a portion of the BREX genes contained on a ~14 kb HIR present only in DL31 and Hrr. lacusprofundi, and FR coverage was low for the region containing the pglX gene (Supplementary Figure S7,Supplementary Table S9 and Supplementary Information). A methylase of the Hrr. lacusprofundi BREX system (Hlac_3189, PglX) was detected in the metaproteome. The genome sequence of the Hrr. lacusprofundi isolate contains a transposon within the pglX open reading frame. However, a contig representing the uninterrupted gene was present in assembled metagenome data, indicating members of the Hrr. lacusprofundi population synthesize a functional PglX.
Within the BREX gene clusters of DL31 and Hrr. lacusprofundi are two genes for a type II toxin-antitoxin system (VapBC; Yamaguchi et al., 2011): Halar_0259/0258, Hlac_3192/3193, respectively (Supplementary Figure S7). The respective VapB and VapC proteins for DL31 and Hrr. lacusprofundi each have very high sequence identities (99–100%). Additionally, adjacent to the CRISPR I-D locus in Hrr. lacusprofundi are two genes that encode another VapBC toxin-antitoxin system (Hlac_3586/3585), and the VapC (toxin) protein was detected in the metaproteome. VapC is a ribonuclease, and to offset toxicity of VapC to the cell it is co-expressed with VapB, a tight-binding inhibitor that forms a complex with VapC (Gerdes, 2000; Arcus et al., 2011). The toxin is stable, whereas the cognate antitoxin is unstable and prone to degradation (Arcus et al., 2011; You et al., 2011). Genes for VapBC systems were also identified in the genomes of tADL and DL1 (for example, halTADL_1260/1261, HalDL1_0971/0970).
Targets of the CRISPR systems—Deep Lake viruses and virus–host interactions
Because CRISPR spacers are derived from invading DNA, they were used to identify the source by matching spacers from the four isolate genomes and metagenome contigs to the genome sequences and metagenome contigs. This served to identify spacers derived from any invading source of DNA, or ‘self’ DNA. Head-tailed viruses were the major source, and a small fraction were pleolipoviruses, haloarchaeal plasmids and chromosomes, with the latter often corresponding to genomic islands inferred to be of viral origin (proviruses or viral remnants) (Figure 3,Supplementary Table S10). No genomic or metagenomic spacers matched contigs assigned to the two most abundant viruses in the metaproteome (based on spectral counts), both of which were HCTV-1-like viruses; this is consistent with data from another hypersaline environment (Lake Tyrrell) in which the most targeted viruses are at relatively low abundance (Emerson et al., 2013). Thus, the high-abundance viruses in Deep Lake might be viruses that have yet to be kept in check by CRISPR systems.
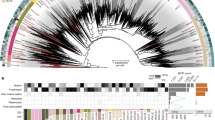
CRISPR systems in Deep Lake haloarchaea. Cas proteins detected in the metaproteome (yellow), and Cas protein gene clusters for tADL, Hrr. lacusprofundi, DL31 and DL1 are shown with their associated CRISPR loci containing repeats (black) and spacers (white). Spacers identified in metagenome contigs are shown separately, linked to specific CRISPR loci based on their repeat sequences. tADL locus 1 and 4 cannot be differentiated in metagenome data because they have identical repeats. For both genomic and metagenomic spacers, spacers that matched to sequences in metagenome contigs or isolate genomes sequences are in red. Spacer number is shown for repeats (red) from genomic loci, numbered relative to the first spacer in the locus. The contig (red text) matching the spacer (red) includes a description (taxa and/or function; black text) of relevant genes that were able to be annotated (no description indicates insufficient level of match to provide annotation). Full details of each contig are given in Supplementary Tables S10 and S11. Cdc6, cell division control protein 6; IF-2, translation initiation factor IF-2; IgA, immunoglobulin A; IgB, immunoglobulin B; MCP, major capsid protein; PadR, PadR transcriptional regulator; PLD, Phospholipase D active site domain; PQQ/WD-40, pyrrolo-quinoline quinone repeat; TFIIB, transcription initiation factor IIB; UspA, universal stress protein A; VP, virion protein; VWA, von Willebrand factor type A domain; wHTH, winged helix-turn-helix; XRE, XRE transcriptional regulator.
Of the eight head-tail viruses represented by MCPs in the metaproteome, three were targeted by the CRISPR system of tADL: one virus with genetic similarities to BJ1 and CGΦ46; another to HHTV-1; and a third to eHP-12 and eHP-6 (see Viruses above). A fourth head-tail virus, related to HCTV-2, was targeted by the DL31 CRISPR system. One pleolipovirus-derived (HRPV-1-like) contig (ctg7180000266221) was targeted solely by Hrr. lacusprofundi (Figure 3, Supplementary Table S10). Pleolipoviruses are pleomorphic viruses that are released from the cell by budding and do not cause host lysis (Pietilä et al., 2009, 2012; Roine et al., 2010; Atanasova et al., 2012). CRISPR spacers from Hrr. lacusprofundi also targeted a linocin gene, and spacers from a metagenome contig derived from an uncharacterized Deep Lake haloarchaeon targeted a contig containing a linocin gene (Figure 3,Supplementary Table S10); both linocin proteins were identified in the metaproteome (Supplementary Table S5) (see Viruses above).
Some spacers from different haloarchaea matched the same viral contig indicating that the virus infected multiple Deep Lake genera (Figure 3,Supplementary Table S10). Most of the broad host range viruses had similarity to myoviruses HSTV-2 and HRTV-7. This is consistent with evidence that, within head-tailed viruses, myoviruses generally have a broader host range (including members of different genera) than siphoviruses (Nuttall and Dyall-Smith, 1993; Sullivan et al., 2003; Atanasova et al., 2012). Some viruses appear to have prompted strong CRISPR responses as they were targeted by multiple spacers to different regions of the virus, including tADL and Hrr. lacusprofundi spacers to a virus of the HRTV-7-like subgroup of haloarchaeal myoviruses (Supplementary Table S10).
There are also instances of the two Hrr. lacusprofundi CRISPR systems targeting the same element represented by a viral contig (for example, ctg7180000271717, ctg7180000396713; Figure 3,Supplementary Table S10). CRISPR spacers from Hrr. lacusprofundi also matched to sequences in the genomes of tADL, DL31 and DL1. These spacer data are consistent with the HIR metaproteome data and network analyses indicating that Hrr. lacusprofundi is a HIR distribution hub (see High identity region s above; DeMaere et al., 2013). In particular, the spacer data support Hrr. lacusprofundi being a net recipient of HIRs within the exchange group.
A number of CRISPR spacers from Hrr. lacusprofundi matched to one gene each from tADL, DL31 and DL1. This included a cdc6-like gene present in the DL31 primary replicon and a N4/N6 DNA methylase gene from the tADL replicon, both of which possibly originated from archaeal plasmids or viruses (Krupovič et al., 2010). A DL1 spacer matched to a putative DL1 peptidase gene (HalDL1_0747), and a spacer from a metagenome contig representing a tADL phylotype (that is, repeats matched tADL CRISPR loci 1 and 4 but the spacer was not in any CRISPR locus of the tADL isolate genome) matched to a tADL gene for a hypothetical protein (halTADL_1427). The tADL gene is encoded within a region (halTADL_1376 to 1454) containing multiple transposases and two integrase genes and may therefore be of viral origin. These spacer data provide evidence that CRISPR systems respond to DNA derived from other haloarchaeal hosts, and self DNA, much of which appears to be of viral and/or plasmid origin.
Some of the genes on viral contigs that were targeted by CRISPR spacers are homologous to haloarchaeal cellular genes. Hrr. lacusprofundi and/or tADL spacers matched to contigs (ctg7180000446992 and ctg7180000403008) with genes homologous to virus GNf2 and haloarchaeal genes encoding hypothetical proteins (Figure 3,Supplementary Table S10). Cell surface protein genes (from both contigs) were detected in the metaproteome, and had the highest identities to genes in haloarchaea, but were also homologous to GNf2 gene HAPG_00095. Different spacers from Hrr. lacusprofundi and tADL targeted this gene. Overall, the data for CRISPR spacers describe a complex and intricate series of host-defense responses to viruses and the genes they carry (Figure 4).
Proposed host–virus interactions occurring in Deep Lake. Interactions between Deep Lake haloarchaea and haloarchaeoviruses and other mobile elements in Deep Lake were inferred from metaproteome data for viruses, cellular proteins relevant to viral infection and evasion, and CRISPR spacers linking previous encounters of hosts to invading DNA. Virus designation is based on sequence similarity to known viruses as described in the text. Certain viruses (for example viruses matching to HRTV-7 or HSTV-1) are shown to have multiple haloarchaeal hosts, infecting distinct lake genera. All featured viruses are inferred to have haloarchaea as hosts except for VBM1 (vibriophage). Proteins detected in the metaproteome that were inferred to be of viral origin are indicated. The color scheme for the S-layer reflects variation in the dominant S-layer protein for tADL, Hrr. lacusprofundi and DL31 observed in the metaproteome: different colors represent the relative abundance of each variant protein as quantified by normalized spectral counts, highlighting distinct phylotypes within populations of each organism. Viruses detected in the metaproteome that were not able to be assigned to a specific host are drawn in the center of the figure. BREX, Bacteriophage Exclusion system; CRISPR, clustered regularly interspaced short palindromic repeats system (type I-B and type I-D shown); CSP, cell surface protein; hyp., hypothetical protein; MCP, major capsid protein; S-layer, surface layer. CSP (black) and pilin (black) illustrate cell surface proteins and pilins, respectively, which were inferred to be carried and expressed by viruses in hosts.
Discussion
The metaproteome for Deep Lake illustrates that genomic variation known to exist within the community (DeMaere et al., 2013), manifests at the protein level. A high extent of variation (ranging from 27 to 77% identity) is particularly apparent in abundant cell surface proteins (Figure 1), including the major S-layer proteins, archaella and adhesins. This variation likely provides the lake population of haloarchaea with a diverse range of cell envelope structures that functions to help host cells evade viral infection. Prevention of attachment and therefore entry of a virus into a host cell is potentially the most effective line of defense against viruses (Avrani et al., 2011). Consistent with a need to evade viruses, the metaproteome revealed the presence of at least eight haloarchaeoviruses, and the expression of a wide range of proteins of viral origin (Supplementary Table S5). The head-tail viruses detected in the metaproteome include those with genetic similarities to lytic viruses (for example, HCTV-1-like and BJ1-like siphoviruses), suggesting that virus infection contributes to cell lysis and nutrient remobilization in Deep Lake. The metaproteome data also support the occurrence of active virus life cycles (for example, prohead proteases and scaffold proteins).
Metaproteomics has not previously been performed on haloarchaeal communities, and our study shows its value for determining mechanisms of virus–host interactions; mechanisms which have previously only been speculated about based upon inferences of genome structure and genetic potential of haloarchaea (Breitbart and Rohwer, 2005; Legault et al., 2006; Cuadros-Orellana et al., 2007; Rodriguez-Valera et al., 2009; Emerson et al., 2012; Garcia-Heredia et al., 2012; Emerson et al., 2013). Although it is theoretically possible that some of the variants we observed may derive from haloarchaea other than tADL, Hrr. lacusprofundi, DL31 or DL1, it is unlikely. Small subunit rRNA gene pyrotag sequencing has shown that tADL, Hrr. lacusprofundi, DL31 and DL1 account for ~72% of the entire lake community, and the relative proportions of each was verified from FR read depth of 454 and Illumina data (DeMaere et al., 2013). Moreover, only five other members of the Halobacteriaceae were identified in the small subunit rRNA gene sequence data with an abundance of 0.2–2% of the lake community (DeMaere et al., 2013). This is consistent with the detection of only 93 proteins in the metaproteome (including 29 cell surface proteins) out of a total of 1109 proteins that had their best BLAST match to Halobacteriaceae other than the four isolates. As many of the detected cell surface variants are highly abundant, it is also very unlikely they could have been produced by rare members. Finally, the FR coverage of the specific cell surface genes also supports the existence of sequence variation that we see manifested as protein variation.
Cost-benefit of CRISPR and BREX systems
Metaproteomics supports the functioning of CRISPR defense systems to counter virus infection (Figure 4). By mapping CRISPR repeats and spacers between host and source sequences, the history of invasion and defense within the lake community was evaluated, indicating that both specific and broad host-range viruses infect the haloarchaea (Figure 3). A broad host-range capacity to infect the three major haloarchaeal genera in the lake, tADL, DL31 and Hrr. lacusprofundi, as well as less abundant members such as DL1, illustrates that viruses have great potential to redistribute DNA within the lake population. However, despite the presence of haloarchaeal genes (including cell surface proteins) in contigs of viral origin and many potential viral genes in HIRs, the data do not clearly reveal whether viruses can be vehicles for HIR transmission.
In general, the CRISPR systems of the haloarchaea were found to be deployed against a broad range of genetic elements, including diverse viruses, plasmids, mobile elements with uncertain identities, as well as haloarchaeal genomic DNA (Figure 3). It has been reported that archaeal CRISPR loci are likely to be constitutively expressed, but may require further activation to target and eliminate invading DNA, such as a sufficient level of replication of the invading element (Garrett et al., 2011). It was previously speculated that gene transfer among haloarchaea in Deep Lake could be mediated by cell-cell contact (for example, cell fusion, conjugation), viruses (transduction) and/or transformation (naked DNA) (DeMaere et al., 2013). Conceivably, DNA transferred by these mechanisms could evade the CRISPR system, recombine and be stably inherited.
BREX systems in DL31 and Hrr. lacusprofundi also potentially provide an additional mechanism for modulating the impact of viruses (Goldfarb et al., 2015), but much remains to be learned about the activity, viral specificity and dissemination of this system. Surveys of bacterial and archaeal genomes have shown that the pglX gene is prone to variation, including gene disruption, duplication and possible phase variation (Goldfarb et al., 2015). The metagenome FR data reveal that a high level of within-population variation exists for pglX in Hrr. lacusprofundi and DL31 (Supplementary Figure S8,Supplementary Table S9).
Cells carrying virus defense systems such as CRISPR and BREX can incur a fitness cost, especially if they synthesize proteins that are toxic to the cell (Makarova et al., 2012), and this can lead to inactivation or outright loss of these systems (Stern and Sorek 2011; Jiang et al., 2013; Goldfarb et al., 2015). During stress, such as viral attack, the antitoxin of the toxin-antitoxin system is inactivated and the toxin serves to induce a dormant state or a suicide response in the infected cell to restrict the extent of infection (Gerdes et al., 2005; Makarova et al., 2012). By promoting their own maintenance the toxin-antitoxin systems may also help to retain chromosomal regions that are themselves prone to loss, including CRISPR systems (You et al., 2011) and BREX systems (Goldfarb et al., 2015). Toxin-antitoxin systems also appear to be subject to horizontal gene transfer (Gerdes, 2000; You et al., 2011). The defense (CRISPR/BREX) and suicide (VapBC) systems tend to be co-localized (You et al., 2011; Makarova et al., 2011c; Jaubert et al., 2013; Makarova et al., 2013; Goldfarb et al., 2015) and we predict this genetic module is capable of being disseminated and selected within the Deep Lake haloarchaea community to mediate virus–host interactions.
Archaella—infection vs swimming
Indicative of being motile in the lake, archaella were detected for tADL, Hrr. lacusprofundi and DL1, including variants for four archaellin proteins from tADL, and one protein each from Hrr. lacusprofundi and DL1. Archaella variation may reduce infection rates for viruses which attach to these appendages, as was suggested for Halorubrum marismortui where the composition of archaella is known to vary (Pyatibratov et al., 2008). However, DL31 is conspicuous in not possessing archaella genes, and also synthesizing Cas proteins below the level of detection in the metaproteome, in contrast to tADL and Hrr. lacusprofundi. The apparent reduced deployment of CRISPR defense by DL31 may indicate it is less subject to infection as a result of lacking archaella. Achieving reduced rates of infection may be a trade-off with the lack of ability to swim and perform chemo/phototaxis. Niche adaptation of DL31 is inferred to involve an association with particulate organic matter containing proteinaceous compounds, in contrast to tADL which is highly saccharolytic (Williams et al., 2014). Although there is evidence of viral attack on DL31, lack of motility may contribute to reduced susceptibility to infection by DL31, and therefore provides an additional explanation for how it achieves the status of being the second most abundant species in the lake.
Selection and possible mechanisms of generating sequence variation
Exposed regions of cell surface proteins are under strong selective pressure to diversify to avoid virus predation, which may arise by mechanisms including mutation, horizontal gene transfer and recombination (Rodriguez-Valera et al., 2009; Avrani et al., 2011; Samson et al., 2013). The use of alternative S-layer genes expressed at different growth stages has been reported for Bacillus anthracis (Mignot et al., 2002; Fagan and Fairweather, 2014), and this type of mechanism involving the two S-layer genes Hlac_0412 and Hlac_2976 (both detected in the metaproteome) could possibly facilitate cell surface variation in Hrr. lacusprofundi. Intragenomic recombination between distinct gene clusters that encode cell surface proteins has been proposed for generating variable cell surface proteins of Haloquadratum walsbyi (Cuadros-Orellana et al., 2007), and specific mechanisms for antigenic switching in S-layer proteins and pilus proteins have been reported for certain pathogenic bacteria (Tu et al., 2003; Dingle et al., 2013; Fagan and Fairweather, 2014; Rotman and Seifert, 2015). However, we found no evidence from the Deep Lake haloarchaeal genomes or the Deep Lake metagenome data (for example, duplicated genes or gene fragments) that would enable comparable intraspecies or intragenomic recombination processes to occur.
Do viruses generate variation of host genes?
Prochlorococcus cyanophage encoding cell-surface-related proteins have been reported to generate microdiversity in the host community through the exchange of genes involved in viral attachment (Avrani et al., 2011). Haloarchaeoviruses containing cell surface genes speculated to be involved in virus evasion have previously been described (Legault et al., 2006; Dyall-Smith et al., 2011); and haloarchaeovirus ΦCh1, which infects Natrialba magadii, has been shown to exhibit phase variation in tail fiber proteins, likely as a mechanism to improve attachment to hosts (Rössler et al., 2004). These examples of host-derived cell surface genes carried on viruses illustrate how Deep Lake haloarchaea and haloarchaeoviruses could have been selected to express genes that overall improve their chance to evade or infect, respectively. The expression of alternate host cell surface proteins after virus infection might render the cell more resistant to subsequent infections by other viruses, akin to superinfection exclusion (Labrie et al., 2010). The type of modular variability present in the Deep Lake haloarchaea S-layer genes (for example, the low FR coverage region of halTADL_1042 and 1043; Figure 2) is consistent with the exchange and recombination of specific segments of DNA, as has been described for cyanobacteria and cyanophage genes (Sullivan et al., 2003; Zeidner et al., 2005).
Novel haloarchaeal pili may enhance avenues for attachment and infection of specific advantageous viruses. Moreover, both tADL and Hrr. lacusprofundi form aggregates (Fröls et al., 2012; Mou et al., 2012; Fröls, 2013), so novel pili and cell surface proteins may facilitate aggregation, and possibly attachment to surfaces and lake particulate matter. Pilin and other cell surface proteins were detected that were encoded on contigs that appear to be of viral origin (Supplementary Table S5), and genes on the contigs were targeted by CRISPR systems of tADL and Hrr. lacusprofundi, indicating that host cells mounted a defense response against these infecting viruses. Contig ctg7180000459513 contains the pilin protein 224467999, a pleolipovirus gene (unknown function) and an integrase (Supplementary Table S11). This pilin protein has high sequence identity to the tADL pilin protein halTADL_1885, and clusters with halTADL_1885 in a phylogenetic tree, away from other tADL pilin proteins (Supplementary Figure S9). These data are consistent with a virus carrying and possibly mutating the tADL pilin gene.
Viruses have been postulated to be beneficial to hosts by having high rates of mutation and causing rapid evolution of genes (Santos et al., 2010; Samson et al., 2013), including host genes captured by viruses (Zeidner et al., 2005). As such, haloarchaeoviruses in Deep Lake could potentially provide haloarchaeal hosts a mechanistic ‘short cut’ for acquiring novel genes, or portions of genes. In a hypersaline environment where viruses are the dominant predators (Oren et al., 1997; Kepner et al., 1998; Santos et al., 2007; Anesio and Bellas, 2011; Atanasova et al., 2012; Santos et al., 2012; Wilkins et al., 2013; Luk et al., 2014), and our data indicate in Deep Lake they can have broad host range, shuffling and mutating cell surface genes could bestow protection against more destructive haloarchaeoviruses (for example, virulent forms) by altering and/or replacing the receptors that the more harmful viruses require for attachment. The complexity of such interactions in Deep Lake would be consistent with the constant-diversity dynamics model which proposes that patterns of diversity in microbial populations arise through viral predation (Rodriguez-Valera et al., 2009).
References
Alam M, Oesterhelt D . (1984). Morphology, function and isolation of halobacterial flagella. J Mol Biol 176: 459–475.
Albers S-J, Meyer BH . (2011). The archaeal cell envelope. Nat Rev Microbiol 9: 414–426.
Anderson RE, Brazelton WJ, Baross JA . (2010). Using CRISPRs as a metagenomic tool to identify microbial hosts of a diffuse flow hydrothermal vent viral assemblage. FEMS Microbiol Ecol 77: 120–133.
Andersson A, Banfield JF . (2008). Virus population dynamics and acquired virus resistance in natural microbial communities. Science 230: 1047–1050.
Anesio AM, Bellas CM . (2011). Are low temperature habitats hot spots of microbial evolution driven by viruses? Trends Microbiol 19: 52–57.
Arcus VL, McKenzie JL, Robson J, Cook GM . (2011). The PINdomain ribonucleases and the prokaryotic VapBC toxin–antitoxin array. Prot Engin Design Select 24: 33–40.
Atanasova NS, Roine E, Oren A, Bamford DH, Oksanen HM . (2012). Global network of specific virus-host interactions in hypersaline environments. Environ Microbiol 14: 426–440.
Avrani S, Wurtzel O, Sharon I, Sorek R, Lindell D . (2011). Genomic island variability facilitates Prochlorococcus-virus coexistence. Nature 474: 604–608.
Bland C, Ramsey TL, Sabree F, Lowe M, Brown K, Kyrpides NC et al. (2007). CRISPR recognition tool (CRT): a tool for automatic detection of clustered regularly interspaced palindromic repeats. BMC Bioinformatics 8: 209.
Bobay LM, Touchon M, Rocha EP . (2014). Pervasive domestication of defective prophages by bacteria. Proc Natl Acad Sci USA 111: 12127–12132.
Breitbart M, Rohwer F . (2005). Here a virus, there a virus, everywhere the same virus? Trends Microbiol 13: 278–284.
Brendel J, Stoll B, Lange SJ, Sharma K, Lenz C, Stachler AE et al. (2014). A complex of Cas proteins 5, 6, and 7 is required for the biogenesis and stability of clustered regularly interspaced short palindromic repeats (CRISPR)-derived RNAs (crRNAs) in Haloferax volcanii. J Biol Chem 289: 7164–7177.
Campbell PJ . (1978). Primary productivity of a hypersaline Antarctic lake. Aust J Mar Freshw Res 29: 717–724.
Carver T, Harris SR, Berriman M, Parkhill J, McQuillan JA . (2012). Artemis: an integrated platform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics 28: 464–469.
Cheng H, Shen N, Pei J, Grishin NV . (2004). Double-stranded DNA bacteriophage prohead protease is homologous to herpesvirus protease. Protein Sci 13: 2260–2269.
Cuadros-Orellana S, Martin-Cuadrado AB, Legault B, D'Auria G, Zhaxybayeva O, Papke RT et al. (2007). Genomic plasticity in prokaryotes: the case of the square haloarchaeon. ISME J 1: 235–245.
DeMaere MZ, Williams TJ, Allen MA, Brown MV, Gibson JA, Rich J et al. (2013). High level of intergenera gene exchange shapes the evolution of haloarchaea in an isolated Antarctic lake. Proc Natl Acad Sci USA 110: 16939–16944.
Dingle KE, Didelot X, Ansari MA, Eyre DW, Vaughan A, Griffiths D et al. (2013). Recombinational switching of the Clostridium difficile S-layer and a novel glycosylation gene cluster revealed by large-scale whole-genome sequencing. J Infect Dis 207: 675–686.
Dyall-Smith ML, Pfeiffer F, Klee K, Palm P, Gross K, Schuster SC et al. (2011). Haloquadratum walsbyi: limited diversity in a global pond. PLoS One 6: e20968.
Emerson JB, Andrade K, Thomas BC, Norman A, Allen EE, Heidelberg KB et al. (2013). Virus-host and CRISPR dynamics in Archaea-dominated hypersaline Lake Tyrrell, Victoria, Australia. Archaea 2013: 370871.
Emerson JB, Thomas BC, Andrade K, Allen EE, Heidelberg KB, Banfield JF . (2012). Dynamic viral populations in hypersaline systems as revealed by metagenomic assembly. Appl Environ Microbiol 78: 6309–6320.
Esquivel RN, Xu R, Pohlschroder M . (2013). Novel archaeal adhesion pilins with a conserved N terminus. J Bacteriol 195: 3808–3818.
Fagan RP, Fairweather NF . (2014). Biogenesis and functions of bacterial S-layers. Nat Rev Microbiol 12: 211–222.
Ferris JM, Burton HR . (1988). The annual cycle of heat content and mechanical stability of hypersaline Deep Lake, Vestfold Hills, Antarctica. Hydrobiologia 165: 115–128.
Franzmann PD, Stackebrandt E, Sanderson K, Volkman JK, Cameron DE, Stevenson PL et al. (1988). Halobacterium lacusprofundi, sp. nov., a halophilic bacterium isolated from Deep Lake, Antarctica. Syst Appl Microbiol 11: 20–27.
Fröls S . (2013). Archaeal biofilms: widespread and complex. Biochem Soc Trans 41: 393–398.
Fröls S, Dyall-Smith M, Pfeifer F . (2012). Biofilm formation by haloarchaea. Environ Microbiol 14: 3159–3174.
Garcia-Heredia I, Martin-Cuadrado AB, Mojica FJM, Santos F, Mira A, Antón J et al. (2012). Reconstructing viral genomes from the environment using fosmid clones: The case of haloviruses. PLoS One 7: e33802.
Garrett RA, Vestergaard G, Shah SA . (2011). Archaeal CRISPR-based immune systems: exchangeable functional modules. Trends Microbiol 19: 549–556.
Gerdes K . (2000). Toxin-antitoxin modules may regulate synthesis of macromolecules during nutritional stress. J Bacteriol 182: 561–572.
Gerdes K, Christensen SK, Lobner-Olesen A . (2005). Prokaryotic toxin-antitoxin stress response loci. Nat Rev Microbiol 3: 371–382.
Gibson JAE . (1999). The meromictic lakes and stratified marine basins of the Vestfold Hills, East Antarctica. Antarct Sci 11: 175–192.
Goldfarb T, Sberro H, Weinstock E, Cohen O, Doron S, Charpak-Amikam Y et al. (2015). BREX is a novel phage resistance system widespread in microbial genomes. EMBO J 34: 169–183.
Grissa I, Vergnaud G, Pourcel C . (2007). CRISPRFinder: a web tool to identify clustered regularly interspaced short palindromic repeats. Nucleic Acids Res 35: W52–W57.
Haft DH, Selengut J, Mongodin EF, Nelson KE . (2005). A guild of 45 CRISPR-associated (Cas) protein families and multiple CRISPR/Cas subtypes exist in prokaryotic genomes. PLoS Comput Biol 1: e60.
Heidelberg JF, Nelson WC, Schoenfeld T, Bhaya D . (2009). Germ warfare in a microbial mat community: CRISPRs provide insights into the co-evolution of host and viral genomes. PLoS One 4: e4169.
Held NL, Herrera A, Quiroz NC, Whitaker RJ . (2010). CRISPR associated diversity within a population of Sulfolobus islandicus. PLoS One 5: e12988.
Jarrell KF, Jones GM, Kandiba L, Nair DB, Eichler J . (2010). S-layer glycoproteins and flagellins: reporters of archaeal posttranslational modifications. Archaea pii: 612948.
Jaubert C, Danioux C, Oberto J, Cortez D, Bize A, Krupovič M et al. (2013). Genomics and genetics of Sulfolobus islandicus LAL14/1, a model hyperthermophilic archaeon. Open Biol 3: 130010.
Jiang W, Maniv I, Arain F, Wang Y, Levin BR, Marraffini LA . (2013). Dealing with the evolutionary downside of CRISPR immunity: bacteria and beneficial plasmids. PLoS Genet 9: e1003844.
Kepner RL, Wharton RA, Suttle CA . (1998). Viruses in Antarctic lakes. Limnol Oceanogr 43: 1754–1761.
Krupovič M, Forterre P, Bamford DH . (2010). Comparative analysis of the mosaic genomes of tailed archaeal viruses and proviruses suggests common themes for virion architecture and assembly with tailed viruses of bacteria. J Mol Biol 397: 144–160.
Kukkaro P, Bamford DH . (2009). Virus-host interactions in environments with a wide range of ionic strengths. Environ Microbiol Rep 1: 71–77.
Labrie SJ, Samson JE, Moineau S . (2010). Bacteriophage resistance mechanisms. Nat Rev Microbiol 8: 317–327.
Lauro FM, DeMaere MZ, Yau S, Brown M, Ng C, Wilkins D et al. (2011). An integrative study of a meromictic lake ecosystem in Antarctica. ISME J 5: 879–895.
Legault BA, Lopez-Lopez A, Alba-Casado JC, Doolittle WF, Bolhuis H, Rodriguez-Valera F et al. (2006). Environmental genomics of "Haloquadratum walsbyi" in a saltern crystallizer indicates a large pool of accessory genes in an otherwise coherent species. BMC Genomics 7: 171.
López-Bueno A, Tamames J, Velázquez D, Moya A, Quesada A, Alcamí A . (2009). High diversity of the viral community from an Antarctic lake. Science 326: 858–861.
Luk AW, Williams TJ, Erdmann S, Papke RT, Cavicchioli R . (2014). Viruses of haloarchaea. Life 4: 681–715.
Maier L-K, Lange SJ, Stoll B, Haas KA, Fischer S, Fischer E et al. (2013). Essential requirements for the detection and degradation of invaders by the Haloferax volcanii CRISPR/Cas system I-B. RNA Biol 10: 865–874.
Makarova KS, Anantharaman V, Aravind L, Koonin EV . (2012). Live virus-free or die: coupling of antivirus immunity and programmed suicide or dormancy in prokaryotes. Biol Direct 7: 40.
Makarova KS, Aravind L, Wolf YI, Koonin EV . (2011a). Unification of Cas protein families and a simple scenario for the origin and evolution of CRISPR-Cas systems. Biol Direct 6: 38.
Makarova KS, Haft DH, Barrangou R, Brouns SJ, Charpentier E, Horvath P et al. (2011b). Evolution and classification of the CRISPR-Cas systems. Nat Rev Microbiol 9: 467–477.
Makarova KS, Wolf YI, Koonin EV . (2013). Comparative genomics of defense systems in archaea and bacteria. Nucleic Acids Res 41: 4360–4377.
Makarova KS, Wolf YI, Snir S, Koonin EV . (2011c). Defense islands in bacterial and archaeal genomes and prediction of novel defense systems. J Bacteriol 193: 6039–6056.
Mignot T, Mesnage S, Couture-Tosi E, Mock M, Fouet A . (2002). Developmental switch of S-layer protein synthesis in Bacillus anthracis. Mol Microbiol 43: 1615–1627.
Mou YZ, Qiu XX, Zhao ML, Cui HL, Oh D, Dyall-Smith ML . (2012). Halohasta litorea gen. nov. sp. nov., and Halohasta litchfieldiae sp. nov., isolated from the Daliang aquaculture farm, China and from Deep Lake, Antarctica, respectively. Extremophiles 16: 895–8901.
Ng C, DeMaere MZ, Williams TJ, Lauro FM, Raftery M, Gibson JA et al. (2010). Metaproteogenomic analysis of a dominant green sulfur bacterium from Ace Lake, Antarctica. ISME J 4: 1002–1019.
Nuttall SD, Dyall-Smith ML . (1993). HF1 and HF2: Novel bacteriophages of halophilic Archaea. Virology 197: 678–684.
Oren A, Bratbak G, Heldal M . (1997). Occurrence of virus-like particles in the Dead Sea. Extremophiles 1: 143–149.
Pagaling E, Haigh RD, Grant WD, Cowan DA, Jones BE, Ma Y et al. (2007). Sequence analysis of an archaeal virus isolated from a hypersaline lake in inner Mongolia, China. BMC Genomics 8: 410.
Pietilä MK, Atanasova NS, Manole V, Liljeroos L, Butcher SJ, Oksanen HM et al. (2012). Virion architecture unifies globally distributed pleolipoviruses infecting halophilic archaea. J Virology 86: 5067–5078.
Pietilä MK, Laurinmäki P, Russell DA, Ko CC, Jacobs-Sera D, Butcher SJ et al. (2013). Insights into head-tailed viruses infecting extremely halophilic Archaea. J Virol 87: 3248–3260.
Pietilä MK, Roine E, Paulin L, Kalkkinen N, Bamford DH . (2009). An ssDNA virus infecting archaea: A new lineage of viruses with a membrane envelope. Mol Microbiol 72: 307–319.
Pyatibratov MG, Beznosov SN, Rachel R, Tiktopulo EI, Surin AK, Syutkin AS et al. (2008). Alternative flagellar filament types in the haloarchaeon Haloarcula marismortui. Can J Microbiol 54: 835–844.
Rodriguez-Valera F, Martin-Cuadrado AB, Rodriguez-Brito B, Pasić L, Thingstad TF, Rohwer F et al. (2009). Explaining microbial population genomics through phage predation. Nat Rev Microbiol 7: 828–836.
Roine E, Kukkaro P, Paulin L, Laurinavičius S, Domanska A, Somerharju P et al. (2010). New, closely related haloarchaeal viral elements with different nucleic acid types. J Virol 84: 3682–3689.
Rössler N, Klein R, Scholz H, Witte A . (2004). Inversion within the haloalkaliphilic virus ΦCh1 DNA results in differential expression of structural proteins. Mol Microbiol 52: 413–426.
Rotman E, Seifert HS . (2015). Neisseria gonorrhoeae MutS affects pilin antigenic variation through mismatch correction and not by pilE G4 binding. J Bacteriol 197: 1828–1838.
Samson JE, Magadán AH, Sabri M, Moineau S . (2013). Revenge of the phages: defeating bacterial defences. Nat Rev Microbiol 11: 675–687.
Santos F, Meyerdierks A, Peña A, Rosselló-Mora R, Amann R, Antón J . (2007). Metagenomic approach to the study of halophages: the environmental halophage 1. Environ Microbiol 9: 1711–1723.
Santos F, Yarza P, Parro V, Briones C, Antón J . (2010). The metavirome of a hypersaline environment. Environ Microbiol 12: 2965–2976.
Santos F, Yarza P, Parro V, Meseguer I, Rosselló-Móra R, Antón J . (2012). Culture-independent approaches for studying viruses from hypersaline environments. Appl Environ Microbiol 78: 1635–1643.
Senčilo A, Roine E . (2014). A glimpse of the genomic diversity of haloarchaeal tailed viruses. Front Microbiol 5: 84.
Sorek R, Lawrence CM, Wiedenheft B . (2013). CRISPR-mediated adaptive immune systems in bacteria and archaea. Annu Rev Biochem 82: 237–266.
Stern A, Sorek R . (2011). The phage-host arms race: shaping the evolution of microbes. BioEssays 33: 43–51.
Streif S, Staudinger WF, Marwan W, Oesterhelt D . (2008). Flagellar rotation in the archaeon Halobacterium salinarum depends on ATP. J Mol Biol 384: 1–8.
Sullivan MB, Waterbury JB, Chisholm SW . (2003). Cyanophages infecting the oceanic cyanobacterium. Prochlorococcus Nature 424: 1047–1051.
Tu ZC, Wassenaar TM, Thompson SA, Blaser MJ . (2003). Structure and genotypic plasticity of the Campylobacter fetus sap locus. Mol Microbiol 48: 685–698.
Tyson GW, Banfield JF . (2008). Rapidly evolving CRISPRs implicated in acquired resistance of microorganisms to viruses. Environ Microbiol 10: 200–207.
Valdés-Stauber N, Scherer S . (1994). Isolation and characterization of Linocin M18, a bacteriocin produced by Brevibacterium linens. Appl Environ Microbiol 60: 3809–3814.
Vestergaard G, Garrett RA, Shah SA . (2014). CRISPR adaptive immune systems of Archaea. RNA Biol 11: 156–167.
Wilkins D, Yau S, Williams TJ, Allen MA, Brown MV, DeMaere MZ et al. (2013). Key microbial drivers in Antarctic aquatic environments. FEMS Microbiol Rev 37: 303–335.
Williams TJ, Allen MA, DeMaere MZ, Kyrpides NC, Tringe SG, Woyke T et al. (2014). Microbial ecology of an Antarctic hypersaline lake: genomic assessment of ecophysiology among dominant haloarchaea. ISME J 8: 1645–1658.
Williams TJ, Long E, Evans F, DeMaere MZ, Lauro FM, Raftery MJ et al. (2012). A metaproteomic assessment of winter and summer bacterioplankton from Antarctic Peninsula coastal surface waters. ISME J 6: 1883–1900.
Williams TJ, Wilkins D, Long E, Evans F, DeMaere MZ, Raftery MJ et al. (2013). The role of planktonic Flavobacteria in processing algal organic matter in coastal East Antarctica revealed using metagenomics and metaproteomics. Environ Microbiol 15: 1302–1317.
Yamaguchi Y, Park JH, Inouye M . (2011). Toxin-antitoxin systems in bacteria and archaea. Annu Rev Genet 45: 61–79.
Yau S, Lauro FM, DeMaere MZ, Brown MV, Thomas T, Raftery MJ et al. (2011). Virophage control of Antarctic algal host–virus dynamics. Proc Natl Acad Sci USA 108: 6163–6168.
Yau S, Lauro FM, Williams TJ, DeMaere MZ, Brown MV, Rich J et al. (2013). Metagenomic insights into strategies of carbon conservation and unusual sulfur biogeochemistry in a hypersaline Antarctic lake. ISME J 7: 1944–1961.
You XY, Liu C, Wang SY, Jiang CY, Shah SA, Prangishvili D et al. (2011). Genomic analysis of Acidianus hospitalis W1 a host for studying crenarchaeal virus and plasmid life cycles. Extremophiles 15: 487–497.
Zeidner G, Bielawski JP, Shmoish M, Scanlan DJ, Sabehi G, Béjà O . (2005). Potential photosynthesis gene recombination between Prochlorococcus and Synechococcus via viral intermediates. Environ Microbiol 7: 1505–1513.
Acknowledgements
This work was supported by the Australian Research Council and the Australian Antarctic Science program. Mass spectrometric results were obtained at the Bioanalytical Mass Spectrometry Facility within the Analytical Centre of the University of New South Wales. This work was undertaken using infrastructure provided by NSW Government co-investment in the National Collaborative Research Infrastructure Scheme. Subsidized access to this facility is gratefully acknowledged. The work conducted by the U. S. Department of Energy Joint Genome Institute is supported by the Office of Science of the U. S. Department of Energy under Contract No. DE-AC02-05CH11231. We thank Matthew DeMaere for assistance with Deep Lake databases, Sheree Yau and Susanne Erdmann for valuable discussion about viruses and CRISPRs, and the PRIDE team and ProteomeXchange for efficiently processing and hosting the mass spectrometry data. We warmly acknowledge the positive and constructive comments made during the review process.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Tschitschko, B., Williams, T., Allen, M. et al. Antarctic archaea–virus interactions: metaproteome-led analysis of invasion, evasion and adaptation. ISME J 9, 2094–2107 (2015). https://doi.org/10.1038/ismej.2015.110
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.110
This article is cited by
-
Population structure of an Antarctic aquatic cyanobacterium
Microbiome (2022)
-
Microbial diversity in extreme environments
Nature Reviews Microbiology (2022)
-
Remarkably coherent population structure for a dominant Antarctic Chlorobium species
Microbiome (2021)
-
Influence of the polar light cycle on seasonal dynamics of an Antarctic lake microbial community
Microbiome (2020)
-
Genomic variation and biogeography of Antarctic haloarchaea
Microbiome (2018)