Abstract
Microbial processes that mineralize organic carbon and enhance solute production at the bed of polar ice sheets could be of a magnitude sufficient to affect global elemental cycles. To investigate the biogeochemistry of a polar subglacial microbial ecosystem, we analyzed water discharged during the summer of 2012 and 2013 from Russell Glacier, a land-terminating outlet glacier at the western margin of the Greenland Ice Sheet. The molecular data implied that the most abundant and active component of the subglacial microbial community at these marginal locations were bacteria within the order Methylococcales (59–100% of reverse transcribed (RT)-rRNA sequences). mRNA transcripts of the particulate methane monooxygenase (pmoA) from these taxa were also detected, confirming that methanotrophic bacteria were functional members of this subglacial ecosystem. Dissolved methane ranged between 2.7 and 83 μM in the subglacial waters analyzed, and the concentration was inversely correlated with dissolved oxygen while positively correlated with electrical conductivity. Subglacial microbial methane production was supported by δ13C-CH4 values between −64‰ and −62‰ together with the recovery of RT-rRNA sequences that classified within the Methanosarcinales and Methanomicrobiales. Under aerobic conditions, >98% of the methane in the subglacial water was consumed over ∼30 days incubation at ∼4 °C and rates of methane oxidation were estimated at 0.32 μM per day. Our results support the occurrence of active methane cycling beneath this region of the Greenland Ice Sheet, where microbial communities poised in oxygenated subglacial drainage channels could serve as significant methane sinks.
Similar content being viewed by others
Introduction
Environments below inland portions of the Greenland Ice Sheet are dark, cold and poorly ventilated, with basal melt in regions of elevated heat flux providing the main source of liquid water at the bed (Bell, 2008). In the absence of direct exchange with the atmosphere, aerobic respiration and chemical oxygen consumption create anoxia, providing conditions favorable for obligately anaerobic processes such as methanogenesis (Skidmore et al., 2005, 2010; Tranter et al., 2005; Wadham et al., 2008). Whether or not the basal zones of ice sheets are sources of atmospheric methane will depend on the in situ rates of methane production and consumption (that is, methanotrophy) and how effectively gases are advected to the margin. Although it is hypothesized that substantial subglacial methane reservoirs exist (Wadham et al., 2012), the release of which could potentially influence atmospheric methane concentrations during deglaciation (Wadham et al., 2008, 2012), there have been few data available on microbial carbon transformations beneath ice sheets.
Measurements of methane excesses in basal ice (Souchez et al., 1995; Miteva et al., 2009) and frozen water (Christner et al., 2012) have suggested that methanogenesis occurs beneath the Greenland Ice Sheet. It is also possible that the erosion of pre-glacial permafrost through glaciological processes could provide an additional source of legacy methane under the ice (Miteva et al., 2009). Measurements of methane production in laboratory microcosms and molecular analyses have confirmed that methanogenic archaea are active members of the microbial communities in these biomes (Boyd et al., 2010; Stibal et al., 2012a, 2012b). However, data to assess carbon dynamics in these ecosystems have been challenging to obtain, and our understanding about the structure and diversity of microbial communities inhabiting the basal environments of polar ice sheets is still very limited.
Here we present molecular and biogeochemical data from a study that investigated the role of microbes in subglacial carbon transformations in West Greenland. Near the ice sheet margin, surface melt enters the subglacial hydrological system through crevasses and moulins, transporting water, atmospheric gases, nutrients and allochthonous carbon to the base (Tranter et al., 2005). Microbiological and physicochemical characteristics of this aqueous environment were examined by collecting and analyzing subglacial outflows from the western margin of the Greenland Ice Sheet near Kangerlussuaq during the melt season in 2012 and 2013. Our results show the dominance of active methanotrophic bacteria as well as phylotypes related to a diverse assortment of obligately anaerobic bacteria and methanogenic archaea, confirming that a range of oxic to anoxic conditions are present in the subglacial environment at the marginal zone. The implications of unearthing a methanotrophic component in the subglacial ecosystem, and how effluxes and cycling of methane may occur beneath the Greenland Ice Sheet, are discussed.
Materials and methods
Study site description
Sampling at the western margin of the Greenland Ice Sheet was conducted at Russell Glacier (67°7′6′′ N, 50°10′5′′ W), a land-terminating outlet glacier located approximately 25 km east of Kangerlussuaq (Figure 1). Following a recession for about five decades, most of the Russell Glacier margin has recently advanced 30–70 m from 1984–1999 (Knight et al., 2000). The local climate of the area is characteristic of an arctic dessert (Hobbs, 1927) with mean annual and summer air temperatures of −5.1 °C and 9.8 °C (June to August), respectively, between 1948 and 2003 (Aebly and Fritz, 2009). Annual precipitation averaged around 158 mm per year during the same period (Aebly and Fritz, 2009). The geology of the area consists of Archaean granitic, tonalitic gneisses and the Kangâmiut dykes that are composed of metamorphosed to amphibolite facies of the southern Nagssugtoqidian orogen (Mayborn and Lesher, 2006).
Sampling locations on the western margin of the Greenland Ice Sheet. The sites were located east of Kangerlussuaq. (A) Subglacial outflow (SUB-RG) and supraglacial stream (SUPRA-RG) at Russell Glacier. (A1) Early season, (A2) mid-season and (A3) late season discharge at SUB-RG. (B) Moraine stream (MS).
Water sampling
Sampling was conducted between 21 May and 11 September 2012 and on 31 August 2013 at Russell Glacier (designated SUB-RG; Figure 1a). For direct comparison, water samples were also collected from a supraglacial stream (67°7′6′′ N, 50°10′5′′ W; designated SUPRA-RG; Figure 1a) and a proglacial moraine stream (67°9′24′′ N, 50°3′54′′ W; designated MS; Figure 1).
Water samples for the nucleic acid extractions were collected two to three times a week from SUB-RG (n=38) and approximately once a week from SUPRA-RG (n=12). Each sample was approximately 50 l of water, maintained at ∼4 °C after collection, and processed within 4–6 h. Particulates larger than 0.2 μm in the water were concentrated by filtration through a 142 mm Supor-200 0.2 μm pore size membrane filters (Pall; ∼10 l per filter). Separate filters were collected for the RNA and DNA extractions. All samples were maintained at −20 °C while at the Kangerlussuaq International Science Support facility (⩽65 days), and subsequently, were transferred to −80 °C for permanent storage when received at Louisiana State University or the University of Washington.
For dissolved methane measurements in the subglacial discharge, water samples were collected between 8 August 2012 and 9 September 2012 on eight occasions from SUB-RG when electrical conductivity (EC) was above 7 μS cm−1 and dissolved oxygen (DO) was below 14.8 mg l−1, and about once a week from MS (n=6). Duplicate 120 ml serum vials were collected (leaving no headspace), one of which was fixed immediately by the addition of 0.5 ml of 12 N HCl. Fixed and unfixed water samples were stored at ∼4 °C for 4–5 months before analyzed.
Methane oxidation rates were determined from water samples collected on 31 August 2013 at SUB-RG. For a direct comparison to ambient dissolved methane concentrations, water samples (n=3) were also collected from a nearby river. Outflow samples (n=27) were collected in 160 ml serum vials leaving no headspace. Triplicate subglacial and river samples were fixed with 0.5 ml of 12N HCl immediately after collection. The remaining 24 samples were incubated at ∼4 °C for 24, 48, 72 and 96 h before fixation, with the final time point upon return to the laboratory at Louisiana State University 33 days after collection. Triplicate ‘laboratory’ and ‘field’ controls containing methane standards (11 000 p.p.m. CH4; Sigma-Aldrich, St Louis, MO, USA) were included to account for methane losses during transport and storage over time. All samples were stored at ∼4 °C until analyzed.
Molecular biological analysis
The nucleic acid extraction procedure included lysozyme digestion in combination with a bead beating step, followed by phenol and chloroform extraction and isopropanol precipitation. For RNA isolation, genomic DNA was eliminated from the extracts with DNase (TURBO DNA-free kit, Ambion, Austin, TX, USA) according to the manufacturer’s guidelines and total RNA was purified using the MEGAclear kit (Ambion). Complementary DNA was reverse transcribed (RT) from RNA by priming with random hexamers and SuperScript II reverse transcriptase (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s specifications. The Illumina MiSeq system (Covance Genomics Laboratory, Seattle, WA, USA) was used to sequence amplicons of the hypervariable V4 region of the 16S rRNA gene using the primers described by Bates et al. (2011; Supplementary Table S1).
The sequences obtained were analyzed using the Mothur platform v.1.29.1 (Schloss et al., 2009). After quality screening, the data set was divided into rare (that is, <1%) and abundant (that is, >1%) sequences. An operational taxonomic unit (OTU) was defined as all sequences that had ⩾97% identity. OTUs were sorted based on being shared or unique to the SUB-RG and SUPRA-RG environment. Metabolically active OTUs were inferred by analyzing the ratio of the relative abundance of sequences obtained in the 16S RT-rRNA and rRNA gene data sets.
The 16S rRNA from methanogenic archaea and mRNA of particulate methane monooxygenase subunit A (pmoA) were amplified by RT-PCR of total RNA from the SUB-RG and SUPRA-RG samples (Supplementary Table S1). To demonstrate the absence/presence of these sequences, only relatively small clone libraries were generated and sequenced (n=48 clones per library) on an ABI 3130XL Genetic analyzer (Genomics Facility, Louisiana State University). OTUs for the PmoA amino-acid sequences were defined on a ⩾93% similarity level (Degelmann et al., 2010). The V4 sequence data were deposited in the NCBI SRA database under the accession number SRP029752, the 16S rRNA sequences from methanogenic taxa as KF649970–KF649974, and pmoA gene sequences as KF649975–KF649979.
Methane concentration and isotopic analysis
Before analysis, the water samples were equilibrated to 22 °C for 18 h. Subsequently, 4 cm3 of water was replaced with 5 cm3 of ultra-high purity dinitrogen gas. The samples were mixed for 5 min to equilibrate dissolved gases in the water with the headspace. Dissolved methane measurements were conducted via headspace analysis using an SRI 8610C gas chromatograph (SRI instruments, Torrance, CA, USA) equipped with a 1-m porpak Q column and a flame ionization detector. The carrier gas was ultra-high purity helium operating at a flow rate of 20 cm3 min−1. All measurements were conducted at standard temperature (22 °C) and pressure (1 atm). Peak areas were quantified using an eDAQ Powerchrom acquisition and analysis system (Denistone East, NSW, Australia). Methane concentrations were calculated using a certified gas standard (Sigma-Aldrich).
The δ13CH4 measurements were performed by the Stable Isotope Laboratory at the University of Arkansas. Dissolved gases were extracted using a membrane extraction method developed by the Stable Isotope Laboratory and analyzed by G2201-i cavity ring down spectrometry (Picarro, Santa Clara, CA, USA).
Detailed methodology
The Supplementary Material contains explicit detail on the (i) sample collection and in situ measurements; (ii) nucleic acid extraction; (iii) amplification of 16S rRNA genes, RT-16S rRNA molecules and mRNA of functional genes; and (iv) sequencing and analysis.
Results
Physicochemical characteristics of the supra- and subglacial waters
The water samples from the subglacial outflow at SUB-RG encompassed the early melt water that emerged from the glacier, high discharge related to increasing surface melt during summer, and the late season when surface melt water input to the basal environment diminished and the subglacial outflow decreased (Figure 1). At the beginning of July, a slight deviation in the subglacial drainage was observed, and the outflow location migrated ∼50 m down-glacier.
The pH of the subglacial water at SUB-RG ranged from 5.1 to 7.8 during the 2012 field season, with an average pH of 6.8±0.5 s.d. (Supplementary Figure S1). EC was highest at the beginning and end of the melt season (46.8 and 64.0 μS cm−1, respectively) and decreased to values generally below 10 μS cm−1 from the beginning of June to the end of August. The SUB-RG discharge water temperature ranged between 0.0 and 0.1 °C and contained an average DO concentration of 13.87±1.50 mg l−1. The most significant change in the DO occurred between 3 and 9 September when it decreased from 13.97 to 10.14 mg l−1 (Supplementary Figure S1). There were also occasions in August and September when lower DO values were observed, and collectively the DO data were significantly negatively correlated with the EC data (Pearson correlation; r=−0.939, P<0.001). After a snowfall and subsequent melt on 9 September, extensive melting coincided with an abrupt increase in the DO concentration and decrease in EC on 11 September (Supplementary Figure S1). In contrast, the DO and EC in the SUPRA-RG varied little throughout the melt season and ranged from 14.12 to 14.38 mg l−1and 1.6 to 1.7 μS cm−1, respectively. The pH of SUPRA-RG ranged between 5.4–6.9.
Dissolved methane measurements
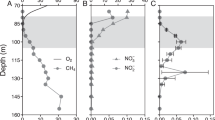
Dissolved methane concentrations were low in the six water samples collected between 8 August and 9 September 2012 from the MS and ranged between 5 and 18 nM. In contrast, dissolved methane in the subglacial outflow from SUB-RG was between 2.7 and 83 μM and was, on average, approximately 3000-fold higher when compared with MS (Figure 2). The high dissolved methane concentrations at SUB-RG were positively correlated with EC (Pearson correlation; r=0.985, P<0.001) and negatively correlated with DO (r=−0.990, P<0.001; Figure 2). The δ13C-CH4 values in the samples from SUB-RG (n=7) ranged between −64‰ and −62‰. The water samples contained 1.7–22 nM of methane after incubation at ∼4 °C for 4–5 months; this is on average 99.9±0.08% less than the concentration observed in samples fixed immediately after collection, indicating methane consumption (Figure 2).
(a) Dissolved methane concentrations in glacial discharge (●) and after 4–5 months of incubation at 4 °C ( ). (b) Correlations between dissolved methane, electrical conductivity (▪) and dissolved oxygen (
). (b) Correlations between dissolved methane, electrical conductivity (▪) and dissolved oxygen ( ) in subglacial discharge at Russell Glacier (SUB-RG) and the moraine stream (MS). Note that different scales are used on the x axes.
) in subglacial discharge at Russell Glacier (SUB-RG) and the moraine stream (MS). Note that different scales are used on the x axes.
A similar decrease in dissolved methane concentrations over time was observed in SUB-RG samples from August 2013, and the water samples contained 1.2±0.73% of the initial methane concentration after 33 days at ∼4 °C. Methane oxidation followed a biphasic decay (Figure 3) with 50% of methane consumed during the first 5 days of incubation. An oxidation rate of 0.32 μM CH4 per day was estimated from the first-order decrease (rate constant=0.138; Figure 3). Methane concentrations in the laboratory and field controls were not statistically different from the initial values (9.1±0.5 μM) after 85 days of storage at ∼4 °C (one-way ANOVA; P=0.528), indicating that methane loss was negligible.
Identification of the autochthonous subglacial microbial community
Based on seasonal changes in EC, DO and select analytes (Supplementary Table S2) in the subglacial discharge, nucleic acids from 19 SUB-RG and 5 SUPRA-RG samples were selected for community structure analysis via 16S RT-rRNA and 16S rRNA gene sequencing. Amplicon sequences of the hypervariable V4 region of the 16S rRNA and 16S rRNA gene were examined using Mothur, and 1 693 724 bacterial sequences passed all quality screening criteria. Subsequently, the analysis focused on the dominant fraction of sequences in each barcoded sample (that is, >1%), which comprised a total of 891 785 bacterial sequence reads. To identify the source of origin, OTUs from the subglacial and supraglacial microbial communities were sorted based on being either shared or unique to the SUB-RG and SUPRA-RG samples (Supplementary Figures S2 and S3). Of all sequences in the 16S RT-rRNA and 16S rRNA gene libraries, 76% and 61% were shared and 3% and 15%, respectively, were unique to the SUPRA-RG samples.
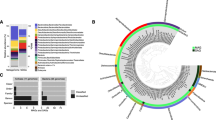
There were 200 650 sequences unique to the SUB-RG samples, accounting for 21% and 24% of the sequences in the 16S RT-rRNA and gene libraries, respectively, and comprising a total of 43 OTUs. Of the 12 OTUs classified in the 16S RT-rRNA data set; only six of these were also present in the 16S rRNA gene library (Supplementary Figures S4A and B). The remaining 31 OTUs were only present in the 16S rRNA gene library (Supplementary Figure S4C). The majority of the sequences unique to the subglacial discharge were phylogenetically related to a single OTU of the methanotrophic order Methylococcales (OTU1). From May to September 2012, OTU1 represented 59–100% of the SUB-RG 16S RT-rRNA and 0–66% of the 16S rRNA gene libraries (Figure 4). The ratio of sequences for each phylotype in the 16S RT-rRNA and rRNA gene libraries was the largest for OTU1 (4:1).
Seasonal distribution of the microbial assemblage unique to the subglacial outflow at Russell Glacier (SUB-RG). The relative abundance of (a) 16S rRNA gene and (b) 16S RT-rRNA sequences. The missing sample on 25 June in the 16S rRNA gene data set was the result of a technical error during sequencing.
Phylogenetic analysis indicated that OTU1 had 97% identity to the 16S rRNA gene of Methylobacter tundripaludum and M. psychrophilus (Figure 5a). The clone library of RT-PCR pmoA amplicons from SUB-RG (9 September) confirmed that bacteria affiliated with the Methylococcales were metabolically active in the subglacial samples (Figure 5b); however, pmoA transcripts were not detectable in SUPRA-RG. The most abundant PmoA phylotype (74% of the clones) had an amino-acid sequence 99% similar to M. tundripaludum (PmoA_OTU1; Figure 5b), which agreed well with the most abundant taxon based on 16S rRNA data (Figure 5a). Four additional PmoA OTUs were identified from the library that were most closely related to M. psychrophilus (PmoA_OTU3, OTU5) and Methylovulum miyakonense (PmoA_OTU2, OTU4).
Phylogenetic analysis of (a) 16S RT-rRNA sequences related to type I and type II methanotrophs and (b) PmoA amino-acid sequences from subglacial samples at Russell Glacier (SUB-RG). Filled circles (●) and bold lettering indicate sequences obtained in this study. The 16S RT-rRNA and PmoA amino-acid neighbor joining trees were rooted with the 16S rRNA gene of Clostridium frigoriphilum and AmoA sequences of the family Nitrosomonadaceae, respectively. Evolutionary distance was computed using the Jukes-Cantor method. Gaps and missing data were eliminated from the alignments. Bootstrap values (1000 replications) >50% are shown. The scale bar represents 3% and 10% divergence of nucleic acid sequence and amino-acid sequence, respectively. The numbers in brackets following the OTU indicate the number of sequences represented for each OTU. Environmental clades of methanotrophs were included in the tree (upland soil clade (USC), glacial forefield soils, Greenland and Jasper Ridge clade (JR1)) for reference. Genbank accession numbers are shown in parentheses.
The remaining 11 16S RT-rRNA OTUs unique to subglacial samples were represented by seven bacterial taxonomic orders. OTUs classifying within the orders Sphingobacterales (⩽17%) and Burkholderiales (⩽13%) were found in approximately half of the SUB-RG RT-rRNA sequence libraries (Figure 4b). The Verrucomicrobiales (⩽13%) appeared consistently in the RT-rRNA libraries toward the end of August and the beginning of September, whereas Vibrionales were only present on 9 September. OTUs related to the Actinomycetales, Rhodospirales and Rhizobiales were detected on 28 July in the SUB-RG 16S RT-rRNA libraries (Figure 4b). Sequences obtained from the Rhizobiales contained a single OTU that classified within the methanotrophic genus Methylocella (Figure 5a).
SUB-RG 16S rRNA gene libraries were significantly different compared with the indigenous SUB-RG 16S RT-rRNA data set (Mothur-based AMOVA, P<0.001). Of the 31 OTUs exclusively found in the 16S rRNA gene libraries, OTUs related to strict anaerobes within the orders Anaerolineales (⩽51%) and Holophagales (⩽19%) as well as Coriobacteriales, Syntrophobacterales and Clostridiales were abundant (Figure 4a). Further, OTUs predominately related to bacterial genera known for facultative anaerobic or fermentative metabolisms were identified (Figure 4, Supplementary Figure S4).
Although none of the amplified sequences obtained from the V4 region of the 16S rRNA gene were identified as archaeal, the use of methanogen-specific 16S rRNA gene primers and RT-PCR was successful in amplifying sequences related to species within the Methanosarcinales and Methanomicrobiales in the SUB-RG and SUPRA-RG samples (Figure 6), but identical OTUs were not shared between the two environments. Three OTUs were classified in the SUPRA-RG samples within the order Methanosarcinales, with the most abundant OTU (SUPRA-RG_Methanogen_OTU1; most closely related to Methanosarcina mazei) accounting for 95% of the clone library. The two SUB-RG methanogen OTUs, SUB-RG_Methanogen_OTU1 (88% of the clones) and SUB-RG_Methanogen_OTU2 (12% of the clones) were most closely related to Methanococcoides alaskense and Methanoregula boonei, respectively (Figure 6).
Phylogenetic analysis of methanogen 16S RT-rRNA clones from supraglacial (SUPRA-RG) and subglacial (SUB-RG) water samples at Russell Glacier using the neighbor joining method. Evolutionary distances were computed using the Jukes-Cantor method. Evolutionary analyses were conducted in MEGA v5.2.2. The tree is rooted with the 16S rRNA gene sequence of Clostridium frigoriphilum and bootstrap values (1000 replications) >50% are shown. The scale bar represents 5% divergence of nucleic acid sequence. The sequences obtained are shown in bold: (♦) SUPRA-RG OTUs, (●) SUB-RG OTUs; numbers in parentheses indicate the number of clones grouped with a ⩾97% identity level. Selected 16S rRNA sequences of clone sequences obtained from other subglacial systems and type species were obtained from public databases. Genbank accession numbers are shown in parentheses.
Discussion
This study coincided with the most widespread surface melt event observed in Greenland over the past 120 years, and liquid water was present on 98.6% of the ice sheet surface during 12 July 2012 (Nghiem et al., 2012). This exceptionally high melting also increased bulk runoff from the ice sheet margin by 113% compared with the average melt for 2009 in the study site's hydrological catchment (Tedstone et al., 2013). Given the ubiquity of ephemeral supraglacial aquatic ecosystems (for example, McMillan et al., 2007; Stibal et al., 2010, 2012c; Cameron et al., 2012; Irvine-Fynn et al., 2012; Telling et al., 2012), microorganisms, dissolved gases and nutrients in the runoff inevitably enter the englacial drainage system. These surface runoff contributions alter the hydrological flow path, and affect access to atmospheric oxygen and the redox potential of environments at the glacier bed; processes that have important implications for understanding biogeochemical reactions that occur beneath ice sheets as well as interpreting data obtained from subglacial outflows at the margin.
A priori it was reasonable to assume that the bulk composition of microbial assemblages present within the subglacial discharge would represent an amalgam of species that originated from ecosystems on the surface and beneath the ice. For example, phototrophic organisms would not be expected to be active members of a sub-ice ecosystem, and the recovery of 16S rRNA sequences from cyanobacteria (Supplementary Figure S2) suggested these species originated from habitats on the ice surface. To address this issue, we conducted a comparative analysis of the supra- and subglacial data to deduce the source of the phylotypes observed (Supplementary Figures S2 and S3). In addition to cyanobacteria, all OTUs classifying within the Methylophilales and Deinococcales were inferred to be of supraglacial origin as members of these orders were either unique to SUPRA-RG samples or shared between both environments (Supplementary Figure S2). One potential caveat of this approach is that generalist species with ecological roles in both the surface and sub-ice ecosystem were neglected. As such, our data should be viewed as a conservative appraisal of the endogenous subglacial microbial community composition from this region.
In agreement with previous molecular surveys of Russell Glacier basal ice (Yde et al., 2010), sediments (Stibal et al., 2012a) and other subglacial environments (Christner et al., 2001; Skidmore et al., 2005; Lanoil et al., 2009; Hamilton et al., 2013), the phylotypes unique to the subglacial outflow samples were dominated by bacteria in the phyla Bacteroidetes, Actinobacteria, Firmicutes, Acidobacteria and Proteobacteria, the latter of which was the predominant phylum in the 16S RT-rRNA data set. A single phylotype with 97% 16S rRNA sequence identity to the type I methanotrophs M. tundripaludum and M. psychrophilus (OTU1) was the most abundant OTU in the subglacial discharge samples, representing up to 66% and 100% of the 16S rRNA gene and RT-rRNA sequences, respectively (Figure 4). Phylogenetically similar taxa have also been found to be dominant in Arctic soils (for example, Berestovskaya et al., 2002; Liebner et al., 2009; Martineau et al., 2010) and low-temperature lakes (Borrel et al., 2011 and references within), supporting the view that this group of methanotrophs has an important role in a variety of Arctic environments. In addition, two less abundant phylotypes with high sequence identity to alpha-(type II) and gammaproteobacterial (type I) methanotrophs were unique to the subglacial samples (Figure 5).
It is interesting to speculate on the environmental factors that may affect the distribution of type I and type II methanotrophs in the subglacial environment. Berestovskaya et al. (2002) showed that the type II methanotroph Methylocella palustris (related to OTU3; Figure 5a) was more abundant than the type I methanotrophs M. psychrophilus (OTU1; Figure 5a) at low pH (4–6). Furthermore, type II methanotrophs appear to be favored at temperatures >15 °C (Mohanty et al., 2007) and can outcompete type I methanotrophs when oxygen becomes limited and the methane concentration is high (Amaral et al., 1995). Similar to findings obtained from glacial forefield sediments (Bárcena et al., 2010), type II methanotrophs were more abundant in the RG-SUPRA (Broemsen, 2013), where they presumably rely on atmospheric methane concentrations. Hence, the combination of physicochemical conditions (that is, near saturated DO concentrations, circumneutral pH, and temperatures near 0°C) in this subglacial aquatic ecosystem may explain why type I methanotrophs were more abundant during the period of investigation.
The relative abundance of an OTU can be used to assess its activity in the community, and given that rRNA content and growth rate are correlated, the ratio of 16S RT-rRNA/rRNA sequences can be used as an indicator of growth rate for individual taxa (for example, Campbell et al., 2011). Although there are caveats with this approach (for example, Blazewicz et al., 2013), the ratio of 16S RT-rRNA/rRNA gene sequences for OTU1 (that is, 97% sequence similarity with M. tundripaludum and M. psychrophilus) was the largest (4:1) observed for any OTU, suggesting this phylotype was both abundant and metabolically active in the subglacial environment (Supplementary Figure S4A). Further evidence for the in situ activity of methanotrophs was provided through the detection of pmoA mRNA sequences that were closely related to the PmoA amino-acid sequence of M. tundripaludum and M. psychrophilus (OTU1: 99% and OTU2-5: 90–96% sequence similarity, respectively). Direct confirmation of methanotrophic activity was provided by methane consumption in water samples during incubation at ∼4 °C, and methane oxidation rates of 0.32 μM per day indicate that methanotrophs could serve as a biological methane sink in this subglacial ecosystem. This rate of methane oxidation is 2.5 to 46 times higher than rates reported for temperate aquatic systems (Kelley, 2003; Carini et al., 2005) but lower than values from Arctic lakes (Lofton et al., 2014).
A subglacial channelized drainage system is expected to be the major path of water flow near the ice sheet margin during maximum summer melt. Water residence time in these effective drainage systems will be short and microbial activity is more likely confined to basal sediments and the flanking marginal zones. Channel margin habitats would provide conditions favorable for microorganisms capable of aerobic respiration as well as those that generate energy via fermentation or anaerobic respiration (Tranter et al., 2005). Therefore, the abundance of phylotypes related to obligately aerobic species (Figure 5) together with methanogens and other known anaerobes in the orders Anaerolineales (⩽51%) and Holophagales (⩽19%; Figure 4a) was consistent with a range of redox conditions existing in the environment beneath the ice sheet. Previous studies have provided evidence for methanogenesis in subglacial sediments from alpine valley glaciers (Boyd et al., 2010) and beneath polar ice sheets (Christner et al., 2012) including Russell Glacier (Stibal et al., 2012b). The dissolved methane concentrations we documented in subglacial outflow waters ranged between 2.7 and 83 μM and were similar in magnitude to values reported in freshwater wetlands (Devol et al., 1988) and temperate swamps (Amaral and Knowles, 1994). The δ13C-CH4 data (range of −62‰ to −64‰) were slightly heavier than values observed in recently deglaciated lakes near Kangerlussuaq (−63‰ to −67; Walter Anthony et al., 2012) and were consistent with expectations for methane produced from a mixture of methyl type-fermentation and CO2 reduction (Whiticar, 1999). For comparison, the δ13C-CH4 values from the basal ice of GRIP (−81 to −84‰; Souchez et al., 2006) and GISP2 (−76‰; Miteva et al., 2009) were isotopically lighter and characteristic of methane produced by CO2 reduction (Whiticar, 1999). It is important to note that we lack an endmember source and alteration of the methane isotopic signature would have occurred in situ as a result of methane oxidation. Hence, a strict interpretation of our isotopic data should be avoided.
Significant quantities of organic carbon from vegetation, soils and lacustrine and marine sediments are thought to have been overridden by ice masses and sequestered globally (Wadham et al., 2008, 2012). If mineralized to methane, modeled estimates suggest an equivalent of 1.31–7.28 × 1014 m3 of methane gas could be stored beneath the East Antarctic Ice Sheet alone (Wadham et al., 2012). Methane reservoirs of this magnitude could have significant climatic implications; however, the net flux of methane from these regions will ultimately depend on the sources and sinks for methane beneath the ice sheet. The recovery of RT-rRNA molecules related to species of the Methanosarcinales and Methanomicrobiales (Figure 6) supports that viable methanogens were present beneath this portion of the ice sheet. The phylotypes observed formed clades with clones previously observed in sediments from the Russell Glacier (Stibal et al., 2012b) and alpine glaciers (Boyd et al., 2010), suggesting that specific lineages of methanogens may be common to subglacial environments. The nearest neighbors to these OTUs have the capacity to produce methane from CO2 reduction (Bräuer et al., 2011) and methylated compounds (Singh et al., 2005). Although methanogens would not be metabolically active in the oxygenated subglacial waters we sampled (Supplementary Figure S1), methanogenesis would be favorable in anoxic regions beneath the ice sheet. As such, methane produced in anoxic regions of the subglacial aquifer could diffuse or advect to the oxygenated, channelized drainage system at the ice sheet margin (Hewitt, 2011), providing conditions favorable for aerobic methanotrophy. Our data support the working hypothesis that anoxic–oxic transitions in the subglacial environment affect the nature of microbial communities and carbon cycling at the ice sheet bed. Whether or not the prominence of methane oxidation in the marginal zones is seasonally limited to periods of increased inputs of surface runoff (for example, Chandler et al., 2013) or occurs throughout the year (for example, anaerobic methanotrophy; Strous, 2010) merits further study.
The dissolved methane concentrations we observed in subglacial waters (Figure 2) indicate that this region of the ice sheet was a source of atmospheric methane. We used cumulative discharge measured between May and August 2012 from Leverett Glacier (2.2 km3; Tedstone et al., 2013) to provide an estimate of seasonal methane fluxes from this region. Leverett Glacier is the southern lobe of the Russell-Leveritt Glacier catchment, draining approximately half (∼1200 km2; Lawson et al., 2013) of the entire hydrological catchment area investigated in the present study (∼2400 km2; Fitzpatrick et al., 2013). When the one month average methane concentration in SUB-RG (n=8) is scaled to the Leverett Glacier discharge, the total dissolved methane flux during the 4 months of summer melt season is estimated at 990 Mg CH4. Considering that the Russell-Leverett Glacier is representative of many land-terminating Greenland outlet glaciers (Lawson et al., 2013), the Greenland Ice Sheet as a whole may represent a significant global methane source.
Conclusion
Our molecular and biogeochemical data indicate that methanotrophic bacteria were both abundant and active members of the microbial communities found in regions beneath the western margin of the Greenland Ice Sheet. These data have expanded our understanding of the subglacial carbon cycle and point toward the existence of a complex subglacial ecosystem at the marginal zones driven by organic carbon mineralization, methanogenesis and methanotrophy. Stibal et al. (2012b) and Wadham et al. (2012) predict that the degradation of organic carbon beneath the world’s ice sheets would lead to a substantial subglacial methane reservoir that could have significant climatic implications if released to the atmosphere during deglaciation. In the absence of empirical data, such modeling efforts have been unable to account for the potential of aerobic methanotrophy as a sink for methane in ventilated regions near the ice sheet margin. Our findings make clear the need for further research to constrain methane reservoirs beneath ice masses and assess how microbial activities in subglacial ecosystems influence methane cycling and its flux to the atmosphere.
Accession codes
References
Aebly FA, Fritz SC . (2009). Palaeohydrology of Kangerlussuaq (Søndre Strømfjord), West Greenland during the last ∼8000 years. Holocene 19: 91–104.
Amaral JA, Knowles R . (1994). Methane metabolism in a temperate swamp. Appl Environ Microbiol 60: 3945–3951.
Amaral JA, Archambault C, Richards SR, Knowles R . (1995). Denitrification associated with Groups I and II methanotrophs in a gradient enrichment system. FEMS Microbiol Ecol 18: 289–298.
Bárcena TG, Yde JC, Finster KW . (2010). Methane flux and high-affinity methanotrophic diversity along the chronosequence of a receding glacier in Greenland. Ann Glaciol 51: 23–31.
Bates ST, Berg-Lyons D, Caporaso JG, Walters WA, Knight R, Fierer N . (2011). Examining the global distribution of dominant archaeal populations in soil. ISME J 5: 08–917.
Bell RE . (2008). The role of subglaical water in ice-sheet mass balance. Nat Geosci 1: 297–304.
Berestovskaya YY, Vasil'eva LV, Chestnykh OV, Zavarzin GA . (2002). Methanotrophs of the psychrophilic microbial community of the Russian Arctic Tundra. Microbiol 71: 538–544.
Blazewicz SJ, Barnard RL, Daly RA, Firestone MK . (2013). Evaluating rRNA as an indicator of microbial activity in environmental communities: limitations and uses. ISME J 7: 2061–2068.
Borrel G, Jézéquel D, Biderre-Petit C, Morel-Desrosiers N, Morel J-P, Peyret P et al. (2011). Production and consumption of methane in freshwater lake ecosystems. Res Microbiol 162: 832–847.
Boyd ES, Skidmore M, Mitchell AC, Bakermans C, Peters JW . (2010). Methanogenesis in subglacial sediments. Environ Microbiol Rep 2: 685–692.
Bräuer SL, Cadillo-Quiroz H, Ward RJ, Yavitt JB, Zinder SH . (2011). Methanoregula boonei gen. nov., sp. nov., an acidiphilic methanogen isolated from an acidic peat bog. Int J Syst Evol Microbiol 61: 45–52.
Broemsen ELJE . (2013). Evidence of methane cycling beneath the western margin of the Greenland Ice Sheet. Master's Thesis. Louisiana State University. http://etd.lsu.edu/docs/available/etd-01142014-185208/.
Cameron KA, Hodson AJ, Osborn AM . (2012). Structure and diversity of bacterial, eukaryotic and archaeal communities in glacial cryoconite holes from the Arctic and the Antarctic. FEMS Microbiol Ecol 82: 254–267.
Carini S, Bano N, LeCleir G, Joye SB . (2005). Aerobic methane oxidation and methanotroph community composition during seasonal stratification in Mono Lake, California (USA). Environ Microbiol 7: 127–1138.
Campbell BJ, Yu L, Heidelberg JF, Kirchman DL . (2011). Activity of abundant and rare bacteria in a coastal ocean. Proc Natl Acad Sci USA 108: 12776–112781.
Chandler DM, Wadham JL, Lis GP, Cowton T, Sole A, Bartholomew I et al. (2013). Evolution of the subglacial drainage system beneath the Greenland Ice Sheet revealed by tracers. Nat Geosci 6: 195–198.
Christner BC, Mosley-Thompson E, Thompson LG, Reeve JN . (2001). Isolation of bacteria and 16S rDNAs from Lake Vostok accretion ice. Environ Microbiol 3: 570–577.
Christner BC, Montross GG, Priscu JC . (2012). Dissolved gases in frozen basal water from the NGRIP borehole: implications for biogeochemical processes beneath the Greenland ice sheet. Polar Biol 35: 1735–1741.
Degelmann DM, Borken W, Drake HL, Kolb S . (2010). Different atmospheric methane-oxidizing communities in European beech and Norway spruce soils. Appl Environ Microbiol 76: 3228–3235.
Devol AH, Richey JE, Clark WA, King SL, Martinelli LA . (1988). Methane emissions to the troposphere from the Amazon floodplain. J Geophys Res Atmos 93: 1583–1592.
Fitzpatrick AAW, Hubbard A, Joughin I, Quincey DJ, van As D, Mikkelsen APB et al. (2013). Ice flow dynamics and surface meltwater flux at a land-terminating sector of the Greenland ice sheet. J Glaciol 59: 687–696.
Hamilton TL, Peters JW, Skidmore ML, Boyd ES . (2013). Molecular evidence for an active endogenous microbiome beneath glacial ice. ISME J 7: 1402–1412.
Hewitt IJ . (2011). Modelling distributed and channelized subglacial drainage: the spacing of channels. J Glaciol 57: 302–314.
Hobbs WH . (1927). The first Greenland expedition of the University of Michigan. Geographical Rev 17: 1–35 American Geographical Society of New York.
Irvine-Fynn TDL, Edwards A, Newton S, Langford H, Rassner SM, Telling J et al. (2012). Microbial cell budgets of an Arctic glacier surface quantified using flow cytometry. Environ Microbiol 14: 2998–3012.
Kelley C . (2003). Methane oxidation potential in the water column of two diverse coastal marine sites. Biogeochemistry 65: 05–120.
Knight PG, Waller RI, Patterson CJ, Jones AP, Robinson ZP . (2000). Glacier advance, ice-marginal lakes and routing of meltwater and sediment: Russell Glacier, Greenland. J Glaciol 46: 423–426.
Lanoil B, Skidmore M, Priscu JC, Han S, Foo W, Vogel SW et al. (2009). Bacteria beneath the West Antarctic Ice Sheet. Environ Microbiol 11: 609–615.
Lawson EC, Wadham JL, Tranter M, Stibal M, Lis GP, Butler CEH et al. (2013). Greenland Ice Sheet exports labile organic carbon to the Arctic oceans. Biogeosci Discuss 10: 19311–19345.
Liebner S, Rublack K, Stuehrmann T, Wagner D . (2009). Diversity of aerobic methanotrophic bacteria in a permafrost active layer soil of the Lena Delta, Siberia. Microb Ecol 57: 25–35.
Lofton DD, Whalen SC, Hershey AE . (2014). Effect of temperature on methane dynamics and evaluation of methane oxidation kinetics in shallow Arctic Alaskan lakes. Hydrobiology 721: 209–222.
Martineau C, Whyte LG, Greer CW . (2010). Stable isotope probing analysis of the diversity and activity of methanotrophic bacteria in soils from the Canadian High Arctic. Appl Environ Microbiol 76: 5773–5784.
Mayborn KR, Lesher CE . (2006). Origin and evolution of the Kangâmiut mafic dyke swarm, West Greenland. GEUS 11: 61–86.
McMillan M, Nienow P, Shepherd A, Benham T, Sole A . (2007). Seasonal evolution of supra-glacial lakes on the Greenland Ice Sheet. Earth Planet Sci Lett 262: 484–492.
Miteva V, Teacher C, Sowers T, Brenchley J . (2009). Comparison of the microbial diversity at different depths of the GISP2 Greenland ice core in relationship to deposition climates. Environ Microbiol 11: 640–656.
Mohanty SR, Bodelier PLE, Conrad R . (2007). Effect of temperature on composition of the methanotrophic community in rice field and forest soil. FEMS Microbiol Ecol 62: 24–31.
Nghiem SV, Hall DK, Mote TL, Tedesco M, Albert MR, Keegan K et al. (2012). The extreme melt across the Greenland ice sheet in 2012. Geophys Res Lett 39: L20502.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB et al. (2009). Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75: 7537–7541.
Singh N, Kendall MM, Liu Y, Boone DR . (2005). Isolation and characterization of methylotrophic methanogens from anoxic marine sediments in Skan Bay, Alaska: description of Methanococcoides alaskense sp. nov., and emended description of Methanosarcina baltica. Int J Syst Evol Microbiol 55: 2531–2538.
Skidmore M, Anderson SP, Sharp M, Foght J, Lanoil BD . (2005). Comparison of microbial community compositions of two subglacial environments reveals a possible role for microbes in chemical weathering processes. Appl Environ Microbiol 71: 6986–6997.
Skidmore M, Tranter M, Tulaczyk S, Lanoil B . (2010). Hydrochemistry of ice stream beds-evaporitic or microbial effects? Hydrol Process 24: 517–523.
Souchez R, Lemmens M, Chappellaz J . (1995). Flow-induced mixing in the GRIP basal ice deduced from the CO2 and CH4 records. Geophys Res Lett 22: 41–44.
Souchez R, Jouzel J, Landais A, Chappellaz J, Lorrain R, Tison JL . (2006). Gas isotopes in ice reveal a vegetated central Greenland during ice sheet invasion. Geophys Res Lett 33: L24503.
Stibal M, Lawson EC, Lis GP, Mak KM, Wadham JL, Anesio AM . (2010). Organic matter content and quality in supraglacial debris across the ablation zone of the Greenland Ice Sheet. Ann Glaciol 51: 1–8.
Stibal M, Hasan F, Wadham JL, Sharp MJ, Anesio AM . (2012a). Prokaryotic diversity in sediments beneath two polar glaciers with contrasting organic carbon substrates. Extremophiles 16: 255–265.
Stibal M, Wadham JL, Lis GP, Telling J, Pancost RD, Dubnick A et al. (2012b). Methanogenic potential of Arctic and Antarctic subglacial environments with contrasting organic carbon sources. Global Change Biol 18: 3332–3345.
Stibal M, Šabacká M, Žárský J . (2012c). Biological processes on glacier and ice sheet surfaces. Nat Geosci 5: 771–774.
Strous M . (2010). Global consequences of anaerobic methane oxidation. In: Timmis KN, (ed) Handbook of Hydrocarbon and Lipid Microbiology. Springer: Berlin Heidelberg, pp 3077–3085.
Tedstone AJ, Nienow PW, Sole AJ, Mair DWF, Cowton TR, Bartholomew ID et al. (2013). Greenland ice sheet motion insensitive to exceptional meltwater forcing. Proc Natl Acad Sci USA 49: 19719–19724.
Telling J, Stibal M, Anesio AM, Tranter M, Nias I, Cook J et al. (2012). Microbial nitrogen cycling on the Greenland Ice Sheet. Biogeoscience 9: 2431–2442.
Tranter M, Skidmore M, Wadham J . (2005). Hydrological controls on microbial communities in subglacial environments. Hydrol Process 19: 995–998.
Walter Anthony KM, Anthony P, Grosse G, Chanton J . (2012). Geologic methane seeps along boundaries of Arctic permafrost thaw and melting glaciers. Nat Geosci 5: 419–426.
Wadham JL, Tranter M, Tulaczyk S, Sharp M . (2008). Subglacial methanogenesis: a potential climatic amplifier? Global Biogeochem Cycles 22: 1–16.
Wadham JL, Arndt S, Tulaczyk S, Stibal M, Tranter M, Telling J et al. (2012). Potential methane reservoirs beneath Antarctica. Nature 488: 633–637.
Whiticar J . (1999). Carbon and hydrogen isotope systematics of bacterial formation and oxidation of methane. Chem Geol 161: 291–314.
Yde JC, Finster KW, Raiswell R, Steffensen JP, Heinemeier J, Olsen J et al. (2010). Basal ice microbiology at the margin of the Greenland ice sheet. Ann Glaciol 51: 71–79.
Acknowledgements
This collaborative study was funded by National Science Foundation grants (1023462, 1023233 and 1023351) from the Division of Polar Programs. BCC and AMA were also partially supported by funds from NSF award 0838941 and the NSF’s Graduate Research Fellowship Program (1247192), respectively. Logistical support was provided by Polar Field Services and Kangerlussuaq International Science Support. We thank L Liu, Z Harrold, S Foredyce and S Schoenemann for assistance in the field.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Dieser, M., Broemsen, E., Cameron, K. et al. Molecular and biogeochemical evidence for methane cycling beneath the western margin of the Greenland Ice Sheet. ISME J 8, 2305–2316 (2014). https://doi.org/10.1038/ismej.2014.59
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2014.59
Keywords
This article is cited by
-
Thermal acclimation of methanotrophs from the genus Methylobacter
The ISME Journal (2023)
-
Groundwater springs formed during glacial retreat are a large source of methane in the high Arctic
Nature Geoscience (2023)
-
Methylotrophic Communities Associated with a Greenland Ice Sheet Methane Release Hotspot
Microbial Ecology (2023)
-
Biogeochemical and historical drivers of microbial community composition and structure in sediments from Mercer Subglacial Lake, West Antarctica
ISME Communications (2023)
-
Subglacial lakes and their changing role in a warming climate
Nature Reviews Earth & Environment (2022)