Abstract
Microbes drive myriad ecosystem processes, and their viruses modulate microbial-driven processes through mortality, horizontal gene transfer, and metabolic reprogramming by viral-encoded auxiliary metabolic genes (AMGs). However, our knowledge of viral roles in the oceans is primarily limited to surface waters. Here we assess the depth distribution of protein clusters (PCs) in the first large-scale quantitative viral metagenomic data set that spans much of the pelagic depth continuum (the Pacific Ocean Virome; POV). This established ‘core’ (180 PCs; one-third new to science) and ‘flexible’ (423K PCs) community gene sets, including niche-defining genes in the latter (385 and 170 PCs are exclusive and core to the photic and aphotic zones, respectively). Taxonomic annotation suggested that tailed phages are ubiquitous, but not abundant (<5% of PCs) and revealed depth-related taxonomic patterns. Functional annotation, coupled with extensive analyses to document non-viral DNA contamination, uncovered 32 new AMGs (9 core, 20 photic and 3 aphotic) that introduce ways in which viruses manipulate infected host metabolism, and parallel depth-stratified host adaptations (for example, photic zone genes for iron–sulphur cluster modulation for phage production, and aphotic zone genes for high-pressure deep-sea survival). Finally, significant vertical flux of photic zone viruses to the deep sea was detected, which is critical for interpreting depth-related patterns in nature. Beyond the ecological advances outlined here, this catalog of viral core, flexible and niche-defining genes provides a resource for future investigation into the organization, function and evolution of microbial molecular networks to mechanistically understand and model viral roles in the biosphere.
Similar content being viewed by others
Introduction
Viruses are abundant in marine environments and generally outnumber bacteria, their most numerous hosts, by an order of magnitude (reviewed in Wommack and Colwell (2000)). The estimated 1028 viral infections in the ocean per day (Suttle, 2007) substantially affect marine systems by causing host mortality, facilitating horizontal gene transfer and influencing biogeochemical cycles via production of dissolved organic matter through cell lysis (reviewed in Breitbart (2012)). An emerging paradigm is that viruses also possess auxiliary metabolic genes (AMGs; Breitbart et al., 2007)—‘host’ genes that may be expressed to augment viral-infected host metabolism and facilitate production of new viruses (reviewed in Breitbart (2012) and Rohwer and Thurber (2009)). Due to the availability of cultures and genomes, AMGs are most extensively explored in marine cyanophages (viruses that infect cyanobacteria) and include genes involved in photosynthesis, carbon metabolism, phosphate metabolism and stress response (Mann et al., 2003; Lindell et al., 2004, 2005; Sullivan et al., 2005; Clokie et al., 2006; Sullivan et al., 2006; Weigele et al., 2007; Dammeyer et al., 2008; Millard et al., 2009; Thompson et al., 2011; Zeng and Chisholm, 2012; Frank et al., 2013). AMGs have also been observed in other cultivated viral isolates including genes for sugar metabolism, lipid–fatty acid metabolism and signalling (Derelle et al., 2008). Further, culture-independent metagenomic surveys have identified additional AMGs involved in motility, anti-oxidation, photosystem I, energy metabolism and iron–sulphur clusters (Yooseph et al., 2007; Dinsdale et al., 2008; Sharon et al., 2009, 2011), with recent, focused pathway analysis expanding these ocean virus-encoded AMG lists to include nearly all of central carbon metabolism (Hurwitz et al., 2013).
Given that microorganisms drive global biogeochemical cycles (Falkowski et al., 2008) and that about 2–66% of marine bacteria are infected by viruses at any given time (Wommack and Colwell, 2000), viral-encoded AMGs must alter global biogeochemistry and microbial metabolic evolution. This might best be exemplified by cyanophages and ‘phage photosynthesis’ (Mann et al., 2003). Briefly, cyanophage genomes nearly universally contain the core photosystem II gene psbA (Sullivan et al., 2006) that is expressed during infection (Lindell et al., 2005; Clokie et al., 2006). This gene has been shown to increase cyanophage fitness (Bragg and Chisholm, 2008), and commonly constitutes a large fraction of total psbA genes in marine microbial metagenomes (Sharon et al., 2007). Beyond elevating cyanophage fitness, these viral psbA gene copies alter the evolutionary trajectory of globally distributed cyanobacterial photosystems as the viral versions evolve under different selective pressures than their host versions and have recombined back into the host (Sullivan et al., 2006). Cyanophage AMGs can also evolve to the point that they perform modified function. For example, when discovered, viral pebS was most similar to a cyanobacterial pebA gene, which partners with the pebB product in phycoerythrobilin synthesis in the cyanobacterial host. Subsequent experimental work showed that this highly divergent viral PebS functionally replaces both host gene products (Dammeyer et al., 2008). Thus AMGs may directly influence system productivity and biogeochemical cycling by metabolically reprogramming host cells during infection and accelerating host niche differentiation through horizontal gene transfer of viral-evolved AMGs (Lindell et al., 2004; Ignacio-Espinoza and Sullivan, 2012).
In the oceans, bacterial taxonomy and metabolic potential strongly vary with depth (DeLong et al., 2006; Ghiglione et al., 2008), suggesting that viral taxonomy and AMGs should also have depth-dependent distribution. Indeed, whole-viral genome fingerprinting shows that viral communities change with depth in the ocean (Steward et al., 2000; Brum, 2005), but to our knowledge, this depth-related genetic variability has not been further explored. Although viral metagenomes (viromes) provide community-wide information relatively quickly, studies to date have been limited by non-quantitative methodologies (reviewed in Duhaime and Sullivan (2012); Solonenko et al. (2013); Solonenko and Sullivan (2013)), under sampling due to older sequencing technologies (Breitbart et al., 2002, 2004), and a high percentage (63-93%) of ‘unknown’ reads (reviewed in Hurwitz and Sullivan (2013)). These issues, coupled with few metagenomic investigations of pelagic viral communities below the oceanic mixed layer (Williamson et al., 2008; Steward and Preston, 2011; Cassman et al., 2012), result in depth-related taxonomic and functional variability in marine viral communities remaining relatively unknown.
The Pacific Ocean Virome (POV) data set overcomes many of these limitations. The 32 POV viromes were quantitatively generated from diverse pelagic ocean habitats including many aphotic zone depths, and are processed using an open reading frame-binning strategy to generate protein clusters (PCs) that help organize the dominant unknown sequence space (Hurwitz and Sullivan, 2013). We previously used this data set to show that viral impacts due to metabolic reprogramming extend well beyond cyanophage and photosynthesis, with Pacific Ocean viral communities encoding nearly all of central carbon metabolism (Hurwitz et al., 2013). Here we extend these analyses to define a ‘core’ (shared by all samples) and ‘flexible’ (found in a subset of samples) Pacific Ocean viral community metagenome, because such analyses have proven fundamental to reconstructing biological function and niche differentiation in viral genomes (for example, in T7- and T4-like phages (Labrie et al., 2012; Sullivan et al., 2010)). This first, large-scale genetic survey of viruses in the photic and aphotic ocean regions results in identification of core, flexible and niche-defining genes, taxonomic patterns with depth and evidence of vertical flux of viral genetic material from the upper ocean to the deep.
Materials and methods
The data set
PCs defined in the 32-virome POV data set (Hurwitz and Sullivan, 2013) were used in all analyses. All metagenomic sequences used in the analysis are available at CAMERA (Sun et al., 2011) under the project accessions: CAM_P_0000914 and CAM_P_0000915. Metagenomic sequences, assemblies, PCs and annotation are available at iPlant (iPlant, 2014) in the community directory (imicrobe/pov). Briefly, these PCs were generated by clustering ORFs to known PCs from the Global Ocean Survey (Yooseph et al., 2007) and proteins from known phage genomes in NCBI using cd-hit-2d (‘-g 1 -n 4 -d 0 -T 24 -M 45 000’; 60% percent identity and 80% coverage). ORFs that did not map to those data sets were then self-clustered using cd-hit (using the same parameters as above), resulting in a total of 27 685 POV PCs containing ⩾20 ORFs each. These PCs were then annotated using the Similarity Matrix of Proteins (Rattei et al., 2006) to assign taxonomy (NCBI) and function (TIGRFAM), with additional functional information obtained from eggNOG (Powell et al., 2012; 4 March 2012).
Defining core and flexible PCs
A PERL script (create_core_pan_genome_all.pl) was used to find the fraction of PCs that were ‘core’ to all samples. Briefly, the script sequentially adds in PCs from each virome (one by one) and determines by a unique PC identifier (PC ID #) whether the newly added PC is common to all viromes added up to that step. Viromes were added step-wise from the most to least similar, ordered using hierarchical k-means clustering output from the MATLAB clustergram(DataMatrix) function. ‘Core’ PCs were defined as those present in all 32 viromes, whereas ‘flexible’ PCs were found in only a subset of viromes. PCs were then categorized as either ‘photic core’ or ‘aphotic core’ for PCs found in all photic or aphotic zone viromes, respectively. PCs present in the ‘photic core’ and absent in the ‘aphotic core’ were defined as ‘photic core exclusive’ (PCE), and vice versa as ‘aphotic core exclusive’ (ACE). TIGRfam annotations of PCE and ACE PCs were then compared with defined distinct functions in each zone.
Differentiating viral DNA from cellular DNA contamination
All 32 POV metagenomes were purified with both DNase and CsCl density gradients to reduce cellular DNA contamination (Hurwitz and Sullivan, 2013). Extensive BLAST-, kmer- and contig-based analyses of these viromes have suggested low bacterial contamination that compares quite favorably to others mined for AMG-like signals (<0.002% for POV (Hurwitz and Sullivan, 2013); vs <0.1% for the human phageome (Modi et al., 2013)). To more specifically investigate potential bacterial contamination in this study, we compared bacterial taxonomy associated with POV ORFs to 16S ribosomal RNA gene taxonomy in the PCE and ACE PCs, and found minimal parallels between these variables (Supplementary Figure S1). Bacterial taxonomy was assigned at the level of order for PCE and ACE ORFs using the top hits from Similarity Matrix of Proteins (as described above). ORFs were then compared with small subunit 16S ribosomal RNA from the Ribosomal Database Project (release 10_30 (Cole et al., 2009)) using BLASTX. Only the top hit to Ribosomal Database Project was retained with a minimum of 75% coverage to the shortest sequence and 97% nucleotide identity. All hits were normalized by total nucleotide count for each virome to allow for direct comparison across viromes.
Overall, only eight bacterial orders were represented in the PCE and ACE PCs at >50 hits per a given sample. Of these eight bacterial orders, 4 (Bacteroidales, Flavobacteriales, Bacillales and Clostridiales) lacked detectable 16S (<10 top-hits) in any virome and together were responsible for 99.8% of the reads associated with the PCE and ACE PCs. The four other bacterial orders (Rhizobiales, Rhodobacterales, Burholderales, Altermonadales) had trace amounts (0.2%) of the virome reads, and contained >10 hits from 16S reads in only 1–6 of the 32 viromes.
Perhaps the most compelling evidence for the viral origin of AMGs is the co-localization of AMGs with verified viral genes on contigs. To investigate this, POV reads in each virome were assembled using Newbler version 2.5.3 (454 Life Sciences, Branford, CT, USA) with default parameters. ORFs were detected on resulting contigs using Prodigal version 2.5.0 (Hyatt et al., 2010) in metagenomic mode (-meta). Taxonomic and functional annotation of ORFs was then assigned using BLASTP to compare ORFs against the Similarity Matrix of Proteins database (20 June 2013). Contigs containing POV core, PCE and ACE PCs were passed through a secondary filter to find contigs with at least one open reading frame of taxonomically defined viral origin based on superkingdom assignment in Similarity Matrix of Proteins.
In spite of limited assembly across these viromes (Hurwitz and Sullivan, 2013), 19 of the 32 POV core, PCE and ACE AMGs co-localized on contigs with bona fide viral genes (Tables 1, 2, 3) and greater than 5 × read coverage. These 19 confirmed AMGs include six previously observed (gmd, speD, psbA, psbD, grx and trx) and 12 novel (glgA, autotrans_barl, cysK/M, iscA/sufA, iscU, sensory box, rfbB, galE, cyt_trans, BclB, QueA and ydeH). Extrapolating from these findings, we infer that most PCs observed in this data set, including those encoding AMGs are of viral origin and not cellular DNA contamination or gene transfer agent-packaged host DNA (Lang and Beatty, 2007).
Detection and alignment of ORFs encoding the psbA gene
PsbA genes were detected in the POV data set by blasting (TBLASTN) POV ORFs against psbA nucleotide sequences from phage genomes (Sullivan et al., 2006). ORFs that matched the most abundant psbA gene for Synechococcus phage S-SM2 and were extracted for further analysis. Representative bacterial and viral proteins for psbA were then aligned with the ORFs from above using UGene Pro. The alignment was manually curated and trimmed to a region with the most coverage between POV ORFs and representative sequences. Protein structure data for psbA was extracted from the Protein Data Bank, by querying for the bacterial psbA protein sequence.
All scripts and associated documentation for methods are archived at the TMPL google code site (Hurwitz, 2014).
Results and discussion
Defining the core and flexible PCs in POV communities
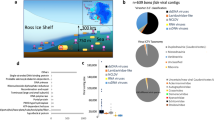
Only 180, 565 or 350 PCs were shared across the entire POV data set (core), the photic zone viromes or the aphotic zone viromes, respectively (Figure 1a). It took ca. 4–6 viromes for core PCs to start to plateau, whereas the flexible PCs took ca. 15 and 8 viromes in the entire POV dataset and photic zone viromes, respectively, reaching ca. 423 and 324K PCs (Figure 1b; Supplementary Figure S2A). In contrast, the flexible PCs in the aphotic viromes did not plateau even after all viromes were added, reaching ca. 215K PCs (Supplementary Figure S2B). Of the core photic and aphotic PCs, 385 and 170 were exclusive to each zone and are termed PCE and ACE, respectively (Figure 1a).
The core and flexible Pacific Ocean viromes. (a) Euler diagram depicting shared and exclusive PCs that are core to the photic and aphotic zone viromes. (b) Core and flexible PCs as a function of the number of viromes in the analysis. Core PCs (squares) are present in all viromes considered, whereas flexible PCs (triangles) are present in only a subset of viromes. Symbols represent the average number of PCs for all combinations of a given number of metagenomes, and error bars represent the range.
To examine the effect of sequencing effort on the development of core PCs, the maximum and minimum number of PCs shared with another virome and unique PCs were determined for each virome were compared with sequencing effort (Supplementary Figure S3). Overall, sequencing effort correlates to the number of PCs unique to any virome based on a Pearson coefficient (0.904, P<0.001), as well as the lowest (0.666, P<0.001) and highest (0.906, P<0.001) number of PCs shared with another virome (Supplementary Figure S3). Thus ‘core PCs’ as defined is a function of sequencing effort.
Taxonomy of POV PCs reveals tailed phages are the minority
Of the 180 core PCs, 64% derived from known viral families, whereas this was true of only 5% in the complete 456K PC data set (Figure 2). The majority of these known families are Myoviridae, Podoviridae and Siphoviridae in both the core (94%) and complete data sets (99%). This shows that tailed viruses (Myoviridae, Podoviridae and Siphoviridae) are the most ubiquitous known virus taxa in POV samples, perhaps explaining why 96% of phage isolates are tailed (Ackermann, 2001). However, they comprise much less of the PCs (ca. 5%) in any given community, which provides genetic evidence consistent with a recent quantitative morphological study with similar taxonomic results for tailed viruses in global ocean samples (Brum et al., 2013b).
This large-scale data set spanning much of the Pacific Ocean depth continuum allows other taxonomic patterns to emerge. These include relatively even distribution of Siphoviridae in the PCE and ACE PCs (Figure 2), although with greater representation of PhiC31-like siphoviruses in the ACE PCs (Supplementary Figure S4). In contrast, PCE PCs had nearly half as many taxonomically unknown PCs as ACE PCs and were enriched for Myoviridae and Podoviridae (Figure 2), primarily comprises T4-like myoviruses, and T7-like and LUZ24-like podoviruses (Supplementary Figure S4). These PC-based data demonstrate vertical zonation of viral taxa, consistent with prior studies of depth-related variation of viral genome size distributions in marine and lacustrine environments (Steward et al., 2000; Jiang et al., 2003a, 2003b; Brum, 2005) and by morphological characterization in lakes (Colombet et al., 2006; Brum and Steward, 2010).
Core PCs for comparative ecology of photic and aphotic zone POV communities
Functional annotation (TIGRfams) was available for 31, 17 and 16% of POV core, PCE and ACE PCs, respectively, and depth-stratified niche-specialization was investigated by examining TIGRfams present in only the POV core (30 TIGRfams; Table 1), PCE (35 TIGRfams; Table 2) or ACE PCs (8 TIGRfams; Table 3). Within these groups, AMGs were defined as metabolic genes not directly involved in viral replication (for example, not including genes involved in DNA packaging, nucleotide transport and metabolism, protein metabolism and assembly or DNA synthesis, replication, recombination and repair). To more explicitly refine this definition, AMGs were subdivided into two classes where Class I AMGs were present in KEGG metabolic pathways (Kanehisa and Goto, 2000), and Class II AMGs were annotatable with only a general metabolic function or entirely absent from KEGG metabolic pathways, presumably due to being peripherally involved in metabolism (for example, transport functions). Given the observation of many AMGs in this data set, we investigated potential cellular DNA contamination as previously discussed in the Methods section, but instead determined that AMGs likely derived from true viral signal.
A potential link between photic and aphotic zone viral communities
Although POV core PCs were, by definition, found in all POV viromes, they had distinct depth-related distribution, with 87 to >99% of ORFs in the photic zone and <1 to 13% in the aphotic zone (normalized to total ORFs in all viromes in each zone; Table 1). Notably, these POV core PCs included the photosystem II reaction center gene psbA, which is widespread among cyanophages (Sullivan et al., 2006). As well, the photosystem gene psbD, also present in cyanophages (Sullivan et al., 2006), was core to the photic zone (Table 2), and present in most (13 of 16) aphotic zone viromes. Given that cyanophage psbA genes are divergent from cyanobacterial host homologs (Sullivan et al., 2006), we constructed a protein alignment, which confirmed that the observed psbA genes derived from T4-like cyanophages rather than cyanobacteria and did not vary with depth (Figure 3).
psbA gene protein sequence assembled from the POV data set. (a) Alignment of psbA gene protein sequences showing amino acids that are specific to viral sequences (noted with red arrows) as compared with bacteria. Virome names follow the convention for the POV data set as described by Hurwitz and Sullivan (2013). Briefly, the initial letter indicates location (L=LineP), season is indicated after the first period (Spr=spring, Sum=summer, Win=winter), proximity to shore is indicated after the second period (C=coastal, I=intermediate, O=open ocean), and depth in meters is indicated after the third period. Sequences in red are from the photic zone and sequences in blue are from the aphotic zone. Representative protein sequences from Synechococcus phage are shown in black and representative bacteria in black and bolded. (b) Protein structure for Thermosynechococcus bacterial protein for Photosystem Q(B) (Uniprot: P0A444). The region from the alignment in (a) is shown with maroon arrows, amino-acid changes that are specific to the viral sequences are denoted in maroon above the bacterial sequence. Amino acids in the bacterial sequence are color-coded based on the following designation (red=protein interaction site, blue=structurally important site, green=accessible site).
There is neither a priori reason to expect that photosynthesis genes confer an adaptive advantage for viruses below sunlit waters, nor are cyanobacteria expected to produce extracellular phage in the aphotic zone. Thus, we hypothesize that the presence of POV core PCs, including psbA, in the aphotic zone result from photic zone viruses transported to the deep ocean on sinking particles, either intra- or extracellularly, where they are released from the cell or particulate matter. This hypothesis has some support from previous studies as follows. First, viruses are known to adsorb to sinking particles. which facilitates their transport to deeper waters (Hewson and Fuhrman, 2003). Second, cyanophages have previously been observed in deep-sea microbial metagenomes (DeLong et al., 2006) and in deep-sea viral communities using a cyanophage gene marker (Short and Suttle, 2005). Although the decay rate of marine viral communities is extremely variable (1–54% per hour; reviewed in Wommack and Colwell (2000)), cyanophages are generally considered stable with measurements of persistence ranging from several years in unfixed samples to at least 100 years in sediments reviewed in (Suttle (2000)). We therefore suggest that these observed core PCs may derive from viruses with low decay rates that can survive transport to the deep sea on sinking particles. This suggested transport of upper ocean viruses to the deep sea represents a revival of an intriguing avenue of research initiated over a decade ago (Hewson and Fuhrman, 2003) and has serious implications for how we think about viral depth distribution in the sea. These results indicate that the viral genetic signal at depth may integrate the vertical flux of upper ocean viruses plus viruses adapted to infecting deep sea organisms. Thus, the analysis of genes found exclusively in deep sea samples (as reported in this study using ACE genes) should result in the most accurate description of viral deep sea adaptive genes.
Although acknowledging that some or all of the POV core PCs may, in fact, be derived from the photic zone, we restrict the rest of our analysis of functional niche specialization to PCs that are exclusive to either the photic (PCE) or aphotic (ACE) regions. In this way, we hope to avoid confounding depth signals with this potential surface-to-deep ocean genetic link, while exploring functional roles of unique genes in viruses in the photic and aphotic ocean.
PCE PCs highlight the importance of Fe-S clusters, DNA metabolism and host resuscitation to photic zone Pacific Ocean viruses
Of the 36 unique TIGRfam functions present in the PCE PCs, 12 represent proteins related to iron-sulphur (Fe-S) clusters, of which 9 are AMGs as strictly defined here (Table 2). Fe-S cluster proteins participate in a wide array of essential physiological pathways including electron transfer, catalysis and regulatory processes that are conserved across the tree of life (Rouault and Tong, 2005; Rouault, 2012). Here, viral-encoded PCE AMGs encode genes for Fe-S protein biogenesis and Fe-S proteins that suggest critical functions in phages linked to electron transfer and enzyme catalysis, but not regulatory functions. Fe-S cluster biogenesis may be enabled by six AMGs likely associated with the ISC and SUF machinery (Fontecave and Ollagnier-de-Choudens, 2008; Shepard et al., 2011). These include the Fe-S cluster assembly protein (iscA), two components of SufABCD Fe-S cluster scaffold complex (sufA and sufB), cysteine synthase (cysK/M), Fe-S assembly scaffold protein (iscU) and the chaperone HscB (hscB). Given that sufA is homologous to iscA (Ollagnier-de-Choudens et al., 2004), it cannot be confirmed whether one or both genes are present; however, both function as scaffold proteins in Fe-S cluster assembly indicating that this is likely important in viruses. Another such biogenesis gene (sufE) was also detected and has been previously documented in viromes (Sharon et al., 2011), but occurred in only 12 of the 16 POV photic zone viromes. In addition, three PCE genes suggest that Pacific Ocean viruses also augment Fe-S protein folding (ATP-dependent molecular chaperone, (clpX)) and degradation of Fe-S cluster proteins (serine protease (clpP) and DNA-binding ATP-dependent protease La (lon) (Rouault and Tong, 2005)), although their functions may be generalizable to other proteins.
Ecologically, the ability to modulate synthesis and degradation of Fe-S cluster proteins in photic zone viral communities may be important as a means to create Fe-S clusters that drive phage production and reduce host stress while preserving environmentally limited iron in regions with high primary productivity, as follows. First, glutaredoxin, the most abundant Fe-S cluster protein in the POV data set, reduces ribonucleotide reductase and may augment the conversion of RNA to DNA to produce genomes of viral progeny (Dwivedi et al., 2013; Holmfeldt et al., 2013). Second, additional POV-encoded genes suggest that viruses mediate host stress response through (i) producing Fe-S cluster containing polyamines including adenosylmethionine decarboxylase (speD) or methionine adenosyltransferase (metK) (Imai et al., 2004; Igarashi and Kashiwagi, 2010), or (ii) degrading sigma factors via Fe-S protein degradation genes clpX and clpP (Hengge, 2008; Calhoun and Kwon, 2011). Among these, only speD has been described in phage genomes (Sullivan et al., 2005).
Viruses are primarily nucleic acids and proteins, so it is not surprising that 9 of 36 PCE TIGRfams were involved with dNTP and protein biosynthesis and repair (Table 2), similar to recent work showing host metabolism shifts to nucleotide biosynthesis (Enav et al., 2014). Seven of these were observed previously in viral genomes and metagenomes including thyA, def (PDF), dnaB, dnaG, xerC, ligD and dam (Scherzinger et al., 1977; Yonesaki, 1994; Subramanya et al., 1996; Huber and Waldor, 2002; Pitcher et al., 2006; Sullivan et al., 2010; Sharon et al., 2011). The two that are newly described for viruses include genes encoding an endonuclease (uvdE) and a phosphate transport protein (phoU). Photorepair of ultraviolet-damaged viral DNA has been well documented (for example, Wilhelm et al., 1998) and uvdE may provide a genetic mechanism for how some viruses achieve such repair. Although phoU has not been observed in viruses, phosphate stress and acquisition genes (for example, pstS, phoA, phoH) are common in T4-like cyanophages isolated from low-phosphate waters (Sullivan et al., 2010). Here, phoU may enable rapid uptake of free phosphate for use in DNA synthesis (Muda et al., 1992), and adds to the paradigm that phosphate scavenging is critical to phosphate-intensive marine viral reproduction (Bratbak, 1993; Clasen and Elser, 2007).
Finally, a PCE gene appears to encode an exosporium leader peptide. In Bacillus, an exosporium leader peptide targets collagen-like proteins to the exosporium, which is a structure that protects the inner spore and can modulate germination (Henriques and Moran, 2007). Perhaps, in viruses, this gene product revives sporulating bacterium by compromising the exosporium. Such a viral-encoded ‘wake up’ strategy is not unprecedented, and in fact, may be common in nature, as 10 soil mycobacteriophages also encode a ‘resuscitation factor’ in their genomes (Pedulla et al., 2003).
ACE PCs suggest viral co-evolution with bacterial hosts under high pressure
Bacterial adaptations for high pressure and deep sea survival are poorly understood, but likely include significant modifications to pressure-sensitive biological processes (reviewed in Bartlett (2002)). Comparative genomics of the deep sea bacterium Photobacterium profundum SS9 (Bartlett, 2002; Eloe et al., 2008; El-Hajj et al., 2009) suggests genes especially tuned to deep sea living include the following: (i) DNA replication initiation (DnaA initiator-associating factor for replication initiation (diaA), chromosomal replication initiator (dnaA), negative modulator of initiation of replication (seqA)), (ii) DNA repair (a component of the RecBCD helicase/nuclease complex (recD)) and (iii) motility (flagellar MS-ring protein (flab) and proton conductor component of the flagellar motor complex (motA)).
Striking functional parallels were observed in the POV ACE PCs (Table 3) as follows. First, DNA replication initiation may be augmented by POV-encoded DnaA. Second, DNA repair may be augmented by two ACE genes (deoxyuridine triphosphatase (dut) and DNA recombination protein (radA)). Third, virus-directed augmentation of motility is suggested by ACE POV-encoded pseudaminic synthase (pseI), which likely glycosylates flagellins (Hopf et al., 2011). Flagellar genes (flaB and motA) are critical for maintaining mobility and particle association for deep sea bacteria (Eloe et al., 2008; Qin et al., 2011). Such genes for ‘chemotaxis and motility’ have been observed previously in viromes (Dinsdale et al., 2008) and may boost their host’s motility to improve nutrient acquisition in the deep sea.
In addition to these more readily identifiable deep sea microbial adaptations, succinate semialdehyde dehydrogenase was core to all aphotic POV samples. This enzyme provides a source of energy within the TCA cycle by converting the carbon backbone of gamma aminobutyric acid to succinic acid. succinate semialdehyde dehydrogenase is just one of 35 AMGs that were recently documented with pathway analyses suggesting that Pacific Ocean viruses reprogram most of host central carbon metabolism to incease nucleotide and energy production during infection (Hurwitz et al., 2013).
Ecological and evolutionary implications of diverse viral-encoded AMGs
Phages have long been suggested to evolve through ‘moron accretion’ (Hendrix et al., 2000) whereby randomly sampled, transcriptionally autonomous host genomic DNA (‘morons’) can accumulate in phage genomes with those conferring a fitness advantage for the phage being selected for and fixed in the populations. Molecular evolutionary studies of viral-encoded photosynthesis AMGs showed that such ‘host’ genes are obtained from within their known host ranges, evolve independently in the viruses and can recombine back into their host cells to alter the evolutionary trajectory of the particular host metabolism (Sullivan et al., 2006), while also improving phage fitness (Bragg and Chisholm, 2008). Although viruses obtain and maintain such AMGs for their own fitness advantage, it is probable—as in the psbA and psbD cases (Sullivan et al., 2006)—that independently evolving viral-encoded AMGs also serve as a source of genetic novelty that periodically alters host metabolic function and evolution. Given so many newly discovered AMGs (this study and Hurwitz et al. (2013); Sharon et al. (2011)) and the fact that a large fraction of microbes are infected by viruses at any given time (Wommack and Colwell, 2000), suggests that cells should be modeled separately as ‘infected’ and ‘uninfected’ as their metabolic output undoubtedly sharply differs.
For example, mutations in the viral protein sequences as compared with bacterial resulted in amino acids with lower energy costs for metabolic biosynthesis (Figure 3, in 5 of 6 amino-acid changes in the viral gene copy (Akashi and Gojobori, 2002)) that may result in energy use efficiency and benefit lytic phages such as T4-like cyanophages. Specifically, by using lower ‘cost’ amino acids phage-driven protein production could be sped up during infection, which in turn, benefits short-term viral replication, whereas the host copy may be evolutionarily tuned for longer term use and slower protein degradation. Further, these mutations occurred in accessible sites in the protein (alpha helices and bends) indicating that the mutations may be non-random and evolutionarily conserved through interaction with other viral specific proteins. In addition, all core T4-like cyanophage genes combined (Ignacio-Espinoza and Sullivan, 2012) showed similar depth-related distribution as psbA, with an average of 73 and 27% of their ORFs in the photic and aphotic zones, respectively.
Conclusions
Viruses are emerging as fundamental drivers of ecosystems ranging from oceans to humans. Yet, the functional diversity of viruses across environments remains largely unexplored due to the lack of sufficient tools available to identify the extent of their roles in driving the Earth’s ecosystems. New tools are rapidly increasing our ability to ‘see’ viral diversity and roles in nature (Hurwitz et al., 2014). The ‘core’ and ‘flexible’ genomic repertoire documented here in the Pacific Ocean offers new biological insight into spatial patterns of viral-encoded, niche-defining functions that are fundamental to viral and host ecology. Linking niche-defining AMGs identified in these ‘gene ecology’ observations to their viral ‘owners’ will help in elucidating which viruses drive specific metabolic pathways in the sunlit and dark oceans, and can be accomplished through screening for particular AMGs in large-insert fosmid libraries (Beja et al., 2000; Mizuno et al., 2013), novel model phage–host systems (for example, phages for the abundant bacterial phyla SAR11 (Zhao et al., 2013), SAR116 (Kang et al., 2013) and Bacteriodetes (Holmfeldt et al., 2012)) or simplified/enriched viral metagenomes (Brum et al., 2013a; Deng et al., 2014). With such a refined toolkit in-hand and a metagenome-inferred roadmap of hypotheses to test, we can now more comprehensively develop experiments to untangle viral–host interactions in nature.
References
Ackermann HW . (2001). Frequency of morphological phage descriptions in the year 2000. Arch Virol 146: 843–857.
Akashi H, Gojobori T . (2002). Metabolic efficiency and amino acid composition in the proteomes of Escherichia coli and Bacillus subtilis. Proc Natl Acad Sci USA 99: 3695–3700.
Bartlett DH . (2002). Pressure effects on in vivo microbial processes. Biochim Biophys Acta 1595: 367–381.
Beja O, Aravind L, Koonin EV, Suzuki MT, Hadd A, Nguyen LP et al. (2000). Bacterial rhodopsin: evidence for a new type of phototrophy in the sea. Science 289: 1902–1906.
Bragg JG, Chisholm SW . (2008). Modelling the fitness consequences of a cyanophage-encoded photosynthesis gene. PLoS One 3: e3550.
Bratbak G . (1993). Viral mortality of the marine alga Emiliania huxleyi (Haptophyceae) and termination of algal blooms. Mar Ecol Prog Ser 93: 39–48.
Breitbart M, Salamon P, Andresen B, Mahaffy JM, Segall AM, Mead D et al. (2002). Genomic analysis of uncultured marine viral communities. Proc Natl Acad Sci USA 99: 14250–14255.
Breitbart M, Felts B, Kelley S, Mahaffy JM, Nulton J, Salamon P et al. (2004). Diversity and population structure of a near-shore marine-sediment viral community. Proc Biol Sci 271: 565–574.
Breitbart M, Thompson LR, Suttle CA, Sullivan MB . (2007). Exploring the vast diversity of marine viruses. Oceanography 20: 135–139.
Breitbart M . (2012). Marine viruses: truth or dare. Ann Rev Mar Sci 4: 425–448.
Brum J . (2005). Concentration, production, and turnover of viruses and dissolved DNA pools at Station ALOHA, North Pacific Subtropical Gyre. Aquat Microb Ecol 41: 103–113.
Brum J, Culley A, Steward G . (2013a). Assembly of a marine viral metagenome after physical fractionation. PLoS ONE 8: e60604.
Brum JR, Steward GF . (2010). Morphological characterization of viruses in the stratified water column of alkaline, hypersaline mono lake. Microb Ecol 60: 636–643.
Brum JR, Schenck RO, Sullivan MB . (2013b). Global morphological analysis of marine viruses shows minimal regional variation and dominance of non-tailed viruses. ISME J 7: 1738–1751.
Calhoun LN, Kwon YM . (2011). Structure, function and regulation of the DNA-binding protein Dps and its role in acid and oxidative stress resistance in Escherichia coli: a review. J Appl Microbiol 110: 375–386.
Cassman N, Prieto-Davó A, Walsh K, Silva GG, Angly F, Akhter S et al. (2012). Oxygen minimum zones harbour novel viral communities with low diversity. Environ Microbiol 14: 3043–3065.
Clasen J, Elser J . (2007). The effect of host Chlorella NC64A carbon: phosphorus ratio on the production of Paramecium bursaria Chlorella Virus-1. Freshwater Biol 52: 112–122.
Clokie MRJ, Shan J, Bailey S, Jia Y, Krisch HM . (2006). Transcription of a 'photosynthetic' T4-type phage during infection of a marine cyanobacterium. Environ Microbiol 8: 827–835.
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ et al. (2009). The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37: D141–D145.
Colombet J, Sime-Ngando T, Cauchie HM, Fonty G, Hoffmann L, Demeure G . (2006). Depth-related gradients of viral activity in Lake Pavin. Appl Environ Microbiol 72: 4440–4445.
Dammeyer T, Bagby SC, Sullivan MB, Chisholm SW, Frankenberg-Dinkel N . (2008). Efficient phage-mediated pigment biosynthesis in oceanic cyanobacteria. Curr Biol 18: 442–448.
DeLong EF, Preston CM, Mincer T, Rich V, Hallam SJ, Frigaard NU et al. (2006). Community genomics among stratified microbial assemblages in the ocean's interior. Science 311: 496–503.
Deng L, Ignacio-Espinoza JC, Gregory AC, Poulos BT, Weitz JS, Hugenholtz P et al. (2014). Viral tagging reveals discrete populations in Synechococcus viral genome sequence space. Nature ; e-pub ahead of print 13 July 2014 doi:10.1038/nature13459.
Derelle E, Ferraz C, Escande ML, Eychenié S, Cooke R, Piganeau G et al. (2008). Life-cycle and genome of OtV5, a large DNA virus of the pelagic marine unicellular green alga Ostreococcus tauri. PLoS One 3: e2250.
Dinsdale EA, Edwards RA, Hall D, Angly F, Breitbart M, Brulc JM et al. (2008). Functional metagenomic profiling of nine biomes. Nature 452: 629–632.
Duhaime MB, Sullivan MB . (2012). Ocean viruses: rigorously evaluating the metagenomic sample-to-sequence pipeline. Virology 434: 181–186.
Dwivedi B, Xue B, Lundin D, Edwards RA, Breitbart M . (2013). A bioinformatic analysis of ribonucleotide reductase genes in phage genomes and metagenomes. BMC Evol Biol 13: 33.
El-Hajj ZW, Tryfona T, Allcock DJ, Hasan F, Lauro FM, Sawyer L et al. (2009). Importance of proteins controlling initiation of DNA replication in the growth of the high-pressure-loving bacterium Photobacterium profundum SS9. J Bacteriol 191: 6383–6393.
Eloe EA, Lauro FM, Vogel RF, Bartlett DH . (2008). The deep-sea bacterium Photobacterium profundum SS9 utilizes separate flagellar systems for swimming and swarming under high-pressure conditions. Appl Environ Microbiol 74: 6298–6305.
Enav H, Mandel-Gutfreund Y, Béjà O . (2014). Comparative metagenomic analyses reveal viral-induced shifts of host metabolism towards nucleotide biosynthesis. Microbiome 2: 1–12.
Falkowski PG, Fenchel T, Delong EF . (2008). The microbial engines that drive Earth's biogeochemical cycles. Science 320: 1034–1039.
Fontecave M, Ollagnier-de-Choudens S . (2008). Iron-sulfur cluster biosynthesis in bacteria: mechanisms of cluster assembly and transfer. Arch Biochem Biophys 474: 226–237.
Frank JA, Lorimer D, Youle M, Witte P, Craig T, Abendroth J et al. (2013). Structure and function of a cyanophage-encoded peptide deformylase. ISME J 7: 1150–1160.
Ghiglione J, Palacios C, Marty J, Mevel G, Labrune C, Conan P et al. (2008). Role of environmental factors for the vertical distribution (0-1000 m) of marine bacterial communities in the NW Mediterranean Sea. Biogeosciences 5: 1751–1764.
Hendrix RW, Lawrence JG, Hatfull GF, Casjens S . (2000). The origins and ongoing evolution of viruses. Trends Microbiol 8: 504–508.
Hengge R . (2008). The two-component network and the general stress sigma factor RpoS (sigma S) in Escherichia coli. Adv Exp Med Biol 631: 40–53.
Henriques AO, Moran CP Jr . (2007). Structure, assembly, and function of the spore surface layers. Annu Rev Microbiol 61: 555–588.
Hewson I, Fuhrman JA . (2003). Viriobenthos production and virioplankton sorptive scavenging by suspended sediment particles in coastal and pelagic waters. Microb Ecol 46: 337–347.
Holmfeldt K, Odić D, Sullivan MB, Middelboe M, Riemann L . (2012). Cultivated single-stranded DNA phages that infect marine Bacteroidetes prove difficult to detect with DNA-binding stains. Appl Environ Microbiol 78: 892–894.
Holmfeldt K, Solonenko N, Shah M, Corrier K, Riemann L, Verberkmoes NC et al. (2013). Twelve previously unknown phage genera are ubiquitous in global oceans. Proc Natl Acad Sci USA 110: 12798–12803.
Hopf PS, Ford RS, Zebian N, Merkx-Jacques A, Vijayakumar S, Ratnayake D et al. (2011). Protein glycosylation in Helicobacter pylori: beyond the flagellins? PLoS One 6: e25722.
Huber KE, Waldor MK . (2002). Filamentous phage integration requires the host recombinases XerC and XerD. Nature 417: 656–659.
Hurwitz BL . (2014). TMPL source code. p http://code.google.com/p/tmpl.
Hurwitz BL, Hallam SJ, Sullivan MB . (2013). Metabolic reprogramming by viruses in the sunlit and dark ocean. Genome Biol 14: R123.
Hurwitz BL, Sullivan MB . (2013). The Pacific Ocean Virome (POV): a marine viral metagenomic dataset and associated protein clusters for quantitative viral ecology. PLoS One 8: e57355.
Hurwitz BL, Westveld AH, Brum JR, Sullivan MB . (2014). Modeling ecological drivers in marine viral communities using comparative metagenomics and network analyses. Proc Natl Acad Sci USA 111: 10714–10719.
Hyatt D, Chen GL, LoCascio PF, Land ML, Larimer FW, Hauser LJ . (2010). Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11: 119.
Igarashi K, Kashiwagi K . (2010). Modulation of cellular function by polyamines. Int J Biochem Cell Biol 42: 39–51.
Ignacio-Espinoza JC, Sullivan MB . (2012). Phylogenomics of T4 cyanophages: lateral gene transfer in the 'core' and origins of host genes. Environ Microbiol 14: 2113–2126.
Imai R, Ali A, Pramanik HR, Nakaminami K, Sentoku N, Kato H . (2004). A distinctive class of spermidine synthase is involved in chilling response in rice. J Plant Physiol 161: 883–886.
iPlant. (2014). iPlant Collaborative https://www.iplantcollaborative.org/.
Jiang S, Fu W, Chu W, Fuhrman JA . (2003a). The vertical distribution and diversity of marine bacteriophage at a station off Southern California. Microb Ecol 45: 399–410.
Jiang S, Steward G, Jellison R, Chu W, Choi S . (2003b). Abundance, distribution, and diversity of viruses in alkaline, hypersaline Mono Lake, California. Microb Ecol 47: 9–17.
Kanehisa M, Goto S . (2000). KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28: 27–30.
Kang I, Oh HM, Kang D, Cho JC . (2013). Genome of a SAR116 bacteriophage shows the prevalence of this phage type in the oceans. Proc Natl Acad Sci USA 110: 12343–12348.
Labrie SJ, Frois-Moniz K, Osburne MS, Kelly L, Roggensack SE, Sullivan MB et al. (2012). Genomes of marine cyanopodoviruses reveal multiple origins of diversity. Environ Microbiol 155: 1356–1376.
Lang AS, Beatty JT . (2007). Importance of widespread gene transfer agent genes in alpha-proteobacteria. Trends Microbiol 15: 54–62.
Lindell D, Sullivan MB, Johnson ZI, Tolonen AC, Rohwer F, Chisholm SW . (2004). Transfer of photosynthesis genes to and from Prochlorococcus viruses. Proc Natl Acad Sci USA 101: 11013–11018.
Lindell D, Jaffe JD, Johnson ZI, Church GM, Chisholm SW . (2005). Photosynthesis genes in marine viruses yield proteins during host infection. Nature 438: 86–89.
Mann NH, Cook A, Millard A, Bailey S, Clokie M . (2003). Bacterial photosynthesis genes in a virus. Nature 424: 741.
Millard AD, Zwirglmaier K, Downey MJ, Mann NH, Scanlan DJ . (2009). Comparative genomics of marine cyanomyoviruses reveals the widespread occurrence of Synechococcus host genes localized to a hyperplastic region: implications for mechanisms of cyanophage evolution. Environ Microbiol 11: 2370–2387.
Mizuno CM, Rodriguez-Valera F, Kimes NE, Ghai R . (2013). Expanding the marine virosphere using metagenomics. PLoS Genet 9: e1003987.
Modi SR, Lee HH, Spina CS, Collins JJ . (2013). Antibiotic treatment expands the resistance reservoir and ecological network of the phage metagenome. Nature 499: 219–222.
Muda M, Rao NN, Torriani A . (1992). Role of PhoU in phosphate transport and alkaline phosphatase regulation. J Bacteriol 174: 8057–8064.
Ollagnier-de-Choudens S, Sanakis Y, Fontecave M . (2004). SufA/IscA: reactivity studies of a class of scaffold proteins involved in [Fe-S] cluster assembly. J Biol Inorg Chem 9: 828–838.
Pedulla ML, Ford ME, Houtz JM, Karthikeyan T, Wadsworth C, Lewis JA et al. (2003). Origins of highly mosaic mycobacteriophage genomes. Cell 113: 171–182.
Pitcher RS, Tonkin LM, Daley JM, Palmbos PL, Green AJ, Velting TL et al. (2006). Mycobacteriophage exploit NHEJ to facilitate genome circularization. Mol Cell 23: 743–748.
Powell S, Szklarczyk D, Trachana K, Roth A, Kuhn M, Muller J et al. (2012). eggNOG v3.0: orthologous groups covering 1133 organisms at 41 different taxonomic ranges. Nucleic Acids Res 40: D284–D289.
Qin Q, Li Y, Zhang Y, Zhou Z, Zhang W, Chen X et al. (2011). Comparative genomics reveals a deep-sea sediment-adapted life style of Pseudoalteromonas sp. SM9913. ISME J 5: 274–284.
Rattei T, Arnold R, Tischler P, Lindner D, Stumpflen V, Mewes HW . (2006). SIMAP: the similarity matrix of proteins. Nucleic Acids Res 34: D252.
Rohwer F, Thurber RV . (2009). Viruses manipulate the marine environment. Nature 459: 207–212.
Rouault TA, Tong WH . (2005). Iron-sulphur cluster biogenesis and mitochondrial iron homeostasis. Nat Rev Mol Cell Biol 6: 345–351.
Rouault TA . (2012). Biogenesis of iron-sulfur clusters in mammalian cells: new insights and relevance to human disease. Dis Model Mech 5: 155–164.
Scherzinger E, Lanka E, Hillenbrand G . (1977). Role of bacteriophage T7 DNA primase in the initiation of DNA strand synthesis. Nucleic Acids Res 4: 4151–4163.
Sharon I, Tzahor S, Williamson S, Shmoish M, Man-Aharonovich D, Rusch DB et al. (2007). Viral photosynthetic reaction center genes and transcripts in the marine environment. ISME J 1: 492–501.
Sharon I, Alperovitch A, Rohwer F, Haynes M, Glaser F, Atamna-Ismaeel N et al. (2009). Photosystem I gene cassettes are present in marine virus genomes. Nature 461: 258–262.
Sharon I, Battchikova N, Aro EM, Giglione C, Meinnel T, Glaser F et al. (2011). Comparative metagenomics of microbial traits within oceanic viral communities. ISME J 5: 1178–1190.
Shepard EM, Boyd ES, Broderick JB, Peters JW . (2011). Biosynthesis of complex iron-sulfur enzymes. Curr Opin Chem Biol 15: 319–327.
Short CM, Suttle CA . (2005). Nearly identical bacteriophage structural gene sequences are widely distributed in both marine and freshwater environments. Appl Environ Microbiol 71: 480.
Solonenko SA, Ignacio-Espinoza JC, Alberti A, Cruaud C, Hallam S, Konstantinidis K et al. (2013). Sequencing platform and library preparation choices impact viral metagenomes. BMC Genomics 14: 320.
Solonenko SA, Sullivan MB . (2013) Preparation of Metagenomic Libraries from Naturally Occurring Marine Viruses Vol 531 Elsevier: Burlington.
Steward G, Montiel JL, Azam F . (2000). Genome size distributions indicate variability and similarities among marine viral assemblages from diverse environments. Limnol Oceanogr 45: 1697–1706.
Steward GF, Preston CM . (2011). Analysis of a viral metagenomic library from 200 m depth in Monterey Bay, California constructed by direct shotgun cloning. Virol J 8: 287.
Subramanya HS, Doherty AJ, Ashford SR, Wigley DB . (1996). Crystal structure of an ATP-dependent DNA ligase from bacteriophage T7. Cell 85: 607–615.
Sullivan MB, Coleman ML, Weigele P, Rohwer F, Chisholm SW . (2005). Three Prochlorococcus cyanophage genomes: signature features and ecological interpretations. PLoS Biol 3: e144.
Sullivan MB, Lindell D, Lee JA, Thompson LR, Bielawski JP, Chisholm SW . (2006). Prevalence and evolution of core photosystem II genes in marine cyanobacterial viruses and their hosts. PLoS Biol 4: e234.
Sullivan MB, Huang KH, Ignacio-Espinoza JC, Berlin AM, Kelly L, Weigele PR et al. (2010). Genomic analysis of oceanic cyanobacterial myoviruses compared with T4-like myoviruses from diverse hosts and environments. Environ Microbiol 12: 3035–3056.
Sun S, Chen J, Li W, Altintas I, Lin A, Peltier S et al. (2011). Community cyberinfrastructure for advanced microbial ecology research and analysis: the CAMERA resource. Nucleic Acids Res 39: D546–D551.
Suttle CA . (2000). Cyanophages and their role in the ecology of cyanobacteria. In Whitton BA, Potts M, (eds) The Ecology of Cyanobacteria. Kluwer Academic Publishers: Netherlands, pp 563–589.
Suttle CA . (2007). Marine viruses—major players in the global ecosystem. Nat Rev Microbiol 5: 801–812.
Thompson LR, Zeng Q, Kelly L, Huang KH, Singer AU, Stubbe J et al. (2011). Phage auxiliary metabolic genes and the redirection of cyanobacterial host carbon metabolism. Proc Natl Acad Sci USA 108: E757–E764.
Weigele PR, Pope WH, Pedulla ML, Houtz JM, Smith AL, Conway JF et al. (2007). Genomic and structural analysis of Syn9, a cyanophage infecting marine Prochlorococcus and Synechococcus. Environ Microbiol 9: 1675–1695.
Wilhelm SW, Weinbauer MG, Suttle CA, Pledger RJ, Mitchell DL . (1998). Measurements of DNA damage and photoreactivation imply that most viruses in marine surface waters are infective. Aq Microb Ecol 14: 215–222.
Williamson SJ, Cary SC, Williamson KE, Helton RR, Bench SR, Winget D et al. (2008). Lysogenic virus-host interactions predominate at deep-sea diffuse-flow hydrothermal vents. ISME J 2: 1112–1121.
Wommack KE, Colwell RR . (2000). Virioplankton: viruses in aquatic ecosystems. Microbiol Mol Biol Rev 64: 69–114.
Yonesaki T . (1994). Involvement of a replicative DNA helicase of bacteriophage T4 in DNA recombination. Genetics 138: 247–252.
Yooseph S, Sutton G, Rusch DB, Halpern AL, Williamson SJ, Remington K et al. (2007). The Sorcerer II Global Ocean Sampling expedition: expanding the universe of protein families. PLoS Biol 5: e16.
Zeng Q, Chisholm SW . (2012). Marine viruses exploit their host's two-component regulatory system in response to resource limitation. Curr Biol 22: 124–128.
Zhao Y, Temperton B, Thrash JC, Schwalbach MS, Vergin KL, Landry ZC et al. (2013). Abundant SAR11 viruses in the ocean. Nature 494: 357–360.
Acknowledgements
We thank Tucson Marine Phage Lab members for comments on the manuscript; Matt Kane for suggesting the designation of two classes of AMGs; UITS Research Computing Group and the ARL Biotechnology Computing for HPCC access and support. Funding was provided by NSF (DBI-0850105 and OCE-0961947), BIO5 and Gordon and Betty Moore Foundation grants (GBMF2631 and GBMF3790) to MBS, and an NSF Integrative Graduate Education and Research Training Fellowship and NSF Graduate Research Fellowship to BLH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Hurwitz, B., Brum, J. & Sullivan, M. Depth-stratified functional and taxonomic niche specialization in the ‘core’ and ‘flexible’ Pacific Ocean Virome. ISME J 9, 472–484 (2015). https://doi.org/10.1038/ismej.2014.143
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2014.143
This article is cited by
-
Water mass age structures the auxiliary metabolic gene content of free-living and particle-attached deep ocean viral communities
Microbiome (2023)
-
Genomic and transcriptomic insights into complex virus–prokaryote interactions in marine biofilms
The ISME Journal (2023)
-
Integrative omics framework for characterization of coral reef ecosystems from the Tara Pacific expedition
Scientific Data (2023)
-
Infection strategy and biogeography distinguish cosmopolitan groups of marine jumbo bacteriophages
The ISME Journal (2022)
-
Cyanophage Distribution Across European Lakes of the Temperate-Humid Continental Climate Zone Assessed Using PCR-Based Genetic Markers
Microbial Ecology (2022)