Abstract
Cold seeps, located along the Sonora Margin transform fault in the Guaymas Basin, were extensively explored during the ‘BIG’ cruise in June 2010. They present a seafloor mosaic pattern consisting of different faunal assemblages and microbial mats. To investigate this mostly unknown cold and hydrocarbon-rich environment, geochemical and microbiological surveys of the sediments underlying two microbial mats and a surrounding macrofaunal habitat were analyzed in detail. The geochemical measurements suggest biogenic methane production and local advective sulfate-rich fluxes in the sediments. The distributions of archaeal communities, particularly those involved in the methane cycle, were investigated at different depths (surface to 18 cm below the sea floor (cmbsf)) using complementary molecular approaches, such as Automated method of Ribosomal Intergenic Spacer Analysis (ARISA), 16S rRNA libraries, fluorescence in situ hybridization and quantitative polymerase chain reaction with new specific primer sets targeting methanogenic and anaerobic methanotrophic lineages. Molecular results indicate that metabolically active archaeal communities were dominated by known clades of anaerobic methane oxidizers (archaeal anaerobic methanotroph (ANME)-1, -2 and -3), including a novel ‘ANME-2c Sonora’ lineage. ANME-2c were found to be dominant, metabolically active and physically associated with syntrophic Bacteria in sulfate-rich shallow sediment layers. In contrast, ANME-1 were more prevalent in the deepest sediment samples and presented a versatile behavior in terms of syntrophic association, depending on the sulfate concentration. ANME-3 were concentrated in small aggregates without bacterial partners in a restricted sediment horizon below the first centimetres. These niche specificities and syntrophic behaviors, depending on biological surface assemblages and environmental availability of electron donors, acceptors and carbon substrates, suggest that ANME could support alternative metabolic pathways than syntrophic anaerobic oxidation of methane.
Similar content being viewed by others
Introduction
At continental margins, large amounts of sedimented organic matter are progressively degraded and transformed into deeply buried hydrocarbons, such as methane. In localized areas, named cold seeps, these hydrocarbons rise in a fluid to the seafloor and form oases of elevated microbial biomass and various faunal assemblages (Jorgensen and Boetius, 2007). According to recent estimates, more than 90% of the emitted methane is metabolized by chemosynthetic archaeal anaerobic methanotroph (ANME) and bacterial sulfate reducer consortia during sulfate reduction coupled to anaerobic oxidation of methane (AOM) in sediments (see Knittel and Boetius (2009) and references therein). ANME clades are phylogenetically divided into three distinct lineages (ANME-1, ANME-2 and ANME-3) affiliated to the Methanosarcinales and Methanomicrobiales orders (Knittel and Boetius, 2009). Metagenomic and metaproteomic studies of ANME communities support the existence of a reverse methanogenesis metabolic pathway for AOM (Meyerdierks et al., 2005, 2010). The products of these anaerobic microbial metabolisms (sulfide or incompletely oxidized organic compounds) are assimilated by mat-forming giant sulfide-oxidizing Bacteria, such as Beggiatoa spp. (Lloyd et al., 2010; Grünke et al., 2011; McKay et al., 2012) or invertebrate symbiosis-involved microorganisms (Cambon-Bonavita et al., 2009). However, much remains to be understood about the AOM mediated by ANME. For instance, the diversity and habitat preference of involved microorganisms or the environmental parameters that impact on these archaeal groups’ distribution need to be defined.
The methane-rich hydrothermal sediments of Guaymas Basin, with a high sedimentation rate (2.7 mm per year), were the first hydrothermal vent habitats in which AOM was demonstrated and have since been studied extensively (Simoneit et al., 1996; Teske et al., 2002; Dhillon et al., 2003, 2005; Page et al., 2008; Holler et al., 2011; Biddle et al., 2012; McKay et al., 2012). In contrast, a nearby cold seep area, which harbors local patches of microbial mats and various faunal assemblages (clams, tubeworms), was observed few kilometers away from hydrothermal sites on a transform fault located on the Sonora Margin (Simoneit et al., 1990; Paull et al., 2007), but remains unsampled for microbiology.
To characterize for the first time microbial communities on the Sonora Margin and to understand the structure and function of the observed microbial ecosystems (two microbial mats and surrounding macrofauna), phylogenetic, microscopic and quantitative analyses of archaeal communities involved in methane cycling and in cold seep processes were undertaken and completed using geochemical analyses.
Materials and methods
Site description and sampling
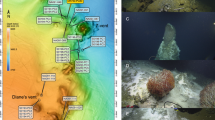
Sediment push core samples were collected in triplicate at cold hydrocarbon-rich seeps from the Sonora Margin using the manned submersible Nautile on dives 1755-11, 1756-12 and 1758-14 at 1574 m depth during the cruise ‘BIG’ (RV L’Atalante, June 2010). The sediments, located near marker BIG18 at ‘Vasconcelos’ site (27°35.5770N; 111°28.9840W) (Supplementary Material and Figure 1), highly supplied by visible sinking organic matter, were covered with patchy thick white Beggiatoa mats occasionally surrounded by grey mat-like macrofauna assemblage. Four different sampling locations were included in this study: (i) White MAT 12 (WM12), a sediment covered with a white microbial mat <1 m wide; (ii) White MAT 14 (WM14), a larger white microbial mat area where gas bubbles and fluids were expelled through a hole; (iii) the Edge of White MAT 14 (EWM14), colonized by dense populations of grey polychaetes and gastropods (M Rabillier and K Olu, personal communications) assumed to be, respectively, deposit feeders and grazers (Fauchald and Jumars, 1979; Waren and Ponder, 1991) (Supplementary Material and Figure 2); and (iv) outside of the active area, that is, the reference site (27°25.483N; 111°30.0759W). At each location, except for WM12 due to its narrow diameter, additional cores were collected on each habitat for methane and sulfide analyses (Supplementary Material and Figure 1). Autonomous temperature sensors (T-Rov; NKE Electronics, Hennebont, France) indicated in situ temperatures around 3 °C from the surface water to the 40 cm below the sea floor (cmbsf) on each habitat before sampling.
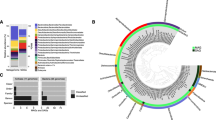
Phylogenetic affiliations of archaeal 16S rRNA genes and 16S rRNA-derived sequences at different depths for cold seep sediments of the Sonora Margin: (a) Reference, (b) White MAT12, (c) White MAT14 areas and (d) the Edge of White MAT 14. Shades of green, brown and yellow denote putative methane-oxidizing or methane-producing groups. ‘Not determined’ means that no amplification was obtained. DHVE8, Deep-Sea Hydrothermal Vent Euryarchaeotal Group 8; MBG-D, -A, -E, -B Marine Benthic Group-D, -A, -E, -B; MCG, Miscellaneous Crenarchaeotic Group; MG II/I, Marine Group II/I; RCV, Rice Cluster V; TMEG, Terrestrial Miscellaneous Euryarcheotal Group; ANME, Anaerobic Methanotrophs; GOM Arc I, Golf of Mexico Archaea group I.
On board, sediment cores were immediately transferred to the cold room. Sediments recovered from WM14 and EWM14 consisted of a dark olive-green ooze in the upper 5 cm and color became lighter with depth. In contrast, sediments color from WM12 was black throughout the core. Gas bubbles were noted immediately after recovery. Sediment cores were subsampled aseptically for molecular analyses in 2- or 5-cm-thick layers using 2 ml cutoff syringes. Sediment subsamples were then frozen at −80 °C for nucleic acid extractions. Sediments (2 g) were collected from each layer and fixed for 4 h in phosphate-buffered saline/formaldehyde (3% final) at 4 °C, washed two times with phosphate-buffered saline and stored in an ethanol/phosphate-buffered saline (1:1, vol vol−1) buffer at −20 °C until analysis by fluorescence in situ hybridization (FISH).
Porewater analyses
Sediment samples were processed on board for porewater chemical analyses. Porewater for sulfate analysis was obtained by spinning down 10 cm3 of sediments per horizon or using Rhizon samplers on closest geochemical cores for hydrogen sulfide and methane (Supplementary Material and Figure 1). Porewater samples were fixed with: zinc chloride (1:1, vol vol−1) for sulfide and 50 μl of sodium azide for methane analyses and nitric acid for sulfate. Sulfate concentrations were measured by ion exchange chromatography as described previously (Lazar et al., 2012). Hydrogen sulfide concentrations were determined by colorimetry (Fonselius et al., 2007). Methane concentrations were quantified by using the headspace technique (HSS 86.50, Dani Instruments, Milan, Italy) and a gas chromatograph (Perichrom 2100, Alpha MOS, Toulouse, France) equipped with flame-ionization detector (Sarradin and Caprais, 1996).
The stable isotope composition of methane was measured by ISOLAB b.v. company (Neerijnen, The Netherlands) in the deepest sediment section (12–17 cmbsf) of the WM14 core CT4 using a MAT Finnigan delta S mass spectrometer (San Jose, CA, USA) coupled to a gas chromatograph by a GC/C II interface.
RNA and DNA extractions from sediments, PCR and RT-PCR amplifications, cloning and sequencing
Total nucleic acids (DNA and RNA) were directly extracted in duplicate from 2.5 g of sediments (Zhou et al., 1996) with modifications, pooled and purified (Lazar et al., 2010). Total RNA were purified using Nucleospin RNA II Kit (Macherey Nagel, Düren, Germany) before reverse transcription-polymerase chain reaction (PCR). Aliquots of rRNA were reverse transcribed using the Quanta qScript Kit according to the manufacturer’s protocol (Quanta Bioscience, Gaithersburg, MD, USA). All PCR reactions were monitored as detailed previously (Lazar et al., 2010), with appropriate annealing temperature for primers listed in Table 1. Gene libraries from 16S rRNA genes (named 16S rDNA in the text) and crDNA templates (named 16S rRNA and used as a proxy for the detection of metabolically active populations) were constructed from pooled and gel-purified triplicate reactions into TOPO XL PCR Cloning Kit (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s recommendations (Ibis Biosciences, Carlsbad, CA, USA). Sequencing was performed on an ABI3730xl Genetic Analyzer (Applied Biosystems, Foster City, CA, USA) using M13 universal primers by GATC Biotech (Constance, Germany). Sequences were analyzed using the NCBI BLASTn search program GenBank (Altschul et al., 1990) and aligned with the closest representative sequences using ClustalW function within BioEdit v.7.1 program (Ibis Biosciences, Carlsbad, CA, USA). Sequences data were analyzed with the MEGA4.0.2 program (Tamura et al., 2007) and checked manually for chimera. Phylogenic trees were calculated by neighbor-joining analysis (Saitou and Nei, 1987) using the Kimura two-parameter correction matrix (Kimura, 1980) and pairwise deletion parameter. The robustness of inferred topology was tested by boostrap resampling (1000) (Felsenstein, 1985). Sequences have been deposited in the EMBL database under the following accession numbers: HE774517–HE774603.
ARISA
An Automated method of Ribosomal Intergenic Spacer Analysis (ARISA) was carried out for a rapid monitoring of microbial diversity and community composition in all sediment samples (Fisher and Triplett, 1999). ARISA-PCR was performed with primers targeting the archaeal 16S–23S intergenic spacer region (Table 1). PCR conditions were as follows: denaturation at 95 °C for 30 s, annealing at 55 °C for 30 s and extension for 1 min 20 s at 72 °C for 35 cycles, followed by a final extension step at 72 °C for 15 min. A measure of 1 μM of each PCR reaction mixture was analyzed according to the manufacturer’s protocol on a DNA 7500 Chip with an Agilent 2100 Bioanalyzer (Agilent Technology, Santa Clara, CA, USA). All data were recovered and analyzed using the R Software with the ade4TkGUI package (Thioulouse and Dray, 2007).
Quantitative real-time PCR
Quantitative measurements were duplicated for each cores. Amplifications were performed with a 7300 Real-Time PCR Systems (Applied Biosystems, Foster City, CA, USA) in a final volume of 25 μl using PerfeCTa SYBR Green SuperMix ROX (Quanta Bioscience), 1 ng of crude DNA template and primers with appropriate concentrations and annealing temperatures (Table 1) according to the manufacturer’s instructions. Primer concentrations were optimized to minimize the secondary structure formations and to maximize the reaction efficiency. Annealing temperatures were adjusted for amplification specificity. As published primer pairs appeared inefficient for quantitative (Q-PCR), new primer sets specific for methanogenic and anaerobic methanotrophic communities, were designed using ARB package (Ludwig et al., 2004) and web-based application Primaclade (Gadberry et al., 2005) (Table 1). Primers were checked for specificity using Oligocheck software (Cardiff University, Cardiff, UK) and tested by PCR. Standard curves were obtained in triplicate with dilutions ranging from 0.001 to 100 nM of DNA extracted from pure cultures of representative groups (Supplementary Materials and Table 1). As no pure culture is available for ANME, standards were made from 10-fold serial dilutions (105–109 copies per μl) of plasmids containing environmental 16S rRNA genes of anaerobic methanotrophs. The R2 of standard curves obtained by real-time PCR were up to 0.997 and efficiency of the reaction up to 90%. Samples were diluted until the crossing point decreased log-linearly with sample dilution, indicating the absence of inhibition effect. The specificity of the primer sets was confirmed by control sequencing of the amplification products. Q-PCR results were expressed in copy number per gram of sediment.
Fluorescence in situ hybridization
A measure of 20 μl of a 100-fold dilution of fixed sediment subsamples were immobilized on 0.22 μm GTTP polycarbonate filters (Merck Millipore, Darmstadt, Germany). Hybridization conditions were optimized to maximize fluorescence signal and probe specificity. Filters were incubated for 3 h at 44 °C, with hybridization buffer containing 40% formamide and labeled probes (Table 2) before washing at 46 °C for 20 min (Snaidr et al., 1997). Finally, filters were fixed on slides and treated with antifading/ 4′,6-diamidino-2-phenylindole solution (SlowFade Gold; Invitrogen). Observations and imaging were performed using an epifluorescence microscope Apotome with Axio Imager Z2 equipped with a COLIBRI system (Zeiss, Göttingen, Germany)
Results
Geochemical characterization
In WM14 and EWM14 sediments, porewater sulfate concentrations decreased rapidly from 22 mM in the surface layer to 2 and 5 mM respectively, at 5 cmbsf (Figures 1c and d), corresponding to the observed sediment color change. In contrast, WM12 sulfate concentrations decreased slightly from 22 mM at the surface to 12 mM by 18 cmbsf (Figure 1b). Sulfate concentrations in reference sediments remained constant to 28 mM with analyzed depth (Figure 1a).
Porewater sulfide concentrations increased quickly with depth and reached 40 mM in the WM14 core and 20 mM in the EWM14 core at the deepest sediment layers (Figure 1c). Peaks of sulfide (28 and 25 mM) were measured at 2 and 5 cmbsf for WM14 and EWM14, respectively (Figures 1c and d). Reference core sulfide concentrations remained under the detection limit (Figure 1a).
Methane porewater concentrations might have been underestimated owing to outgassing during core retrieval. Therefore, no clear methane profiles representing a ‘sulfate-methane transition’ could be inferred. The highest methane concentrations (almost 1.2 mM) were measured in WM14 cores, probably influenced by proximity to the fluid seep hole (Supplementary Material and Figure 1). Methane concentrations in the EWM14 were lower (0.7 mM) and varied according to depth (Figure 1d). Methane was not detected in the reference sediment core.
At the bottom (15–17 cmbsf) of WM14,  13C-CH4 signature was −63‰, which could be interpreted as a dominant proportion of biogenic methane, mixed with abiotic and thermogenic methane (Damm and Budeus, 2003).
13C-CH4 signature was −63‰, which could be interpreted as a dominant proportion of biogenic methane, mixed with abiotic and thermogenic methane (Damm and Budeus, 2003).
ARISA fingerprints
ARISA was used as a rapid method to compare archaeal community structure across the different samples. ARISA fingerprints displayed up to 10 different archaeal phylotypes throughout WM12, WM14 and EWM14 sediment cores, suggesting a low archaeal diversity. In all the sediment cores, community patterns were similar, with an observable shift in archaeal population composition at 4 cmbsf as shown in Supplementary Material and Figure 3a for WM12. Reconstructed ARISA electrophoregrams, compared using principal component analysis (Supplementary Material and Figure 3b), highlighted a very low archaeal community structure evolution between 6 and 22 cmbsf EWM14 sediment layers and a close archaeal communities composition in the deepest sediment layers of each habitat. Principal component analysis revealed a relatively low variability between replicate cores for each habitat. Therefore, only one sediment core was selected per habitat to construct 16S rRNA gene libraries from RNA and DNA.
Neighbor-joining distance phylogeny of the archaeal 16S rRNA genes and rRNA-derived sequences in sediments of the Sonora Margin cold seeps based on the Kimura 2 parameters method with 1000 replicates, including 825 homologous positions. Only bootstrap values up to 70% are shown. The scale bar indicates five substitutions per 100 nucleotides. Sequences amplified from sections 0 to 4 cmbsf are labeled ‘Top’, sequences from 4 to 6 cmbsf are ‘Middle’ and sequences from 8 cmbsf to end are tagged ‘Bottom’. Only one representative sequence (>97% identical) is shown. Number within parantheses show the number of clones analyzed from RNA or DNA clone libraries. DHVE8, Deep-Sea Hydrothermal Vent Euryarchaeotal Group 8; EWM14, Edge of White MAT14; MBG-D/E/B/C, Marine Benthic Group D/E/B/C; MCG, Miscellaneous Crenarchaeotic Group; MG II/I, Marine Group II/I; RCV, Rice Cluster V; TMEG, Terrestrial Miscellaneous Euryarcheotal Group; ANME, Anearobic Methanotrophs; REF, Reference site; WM12, White MAT12;WM14, White MAT14.
Also based on both ARISA analysis and geochemical data, six 16S rRNA gene libraries (from DNA and cDNA) of pooled sediment layers (top, middle and bottom) were constructed for each habitat.
Phylogenetic diversity of metabolically active Archaea
A total of 1204 partial archaeal 16S rRNA sequences were obtained. Archaeal 16S rRNA gene libraries showed a relative low diversity (that is, 5–6 phylogenetic lineages for rRNA and 7–11 for rDNA) (Figures 2 and 3). The most frequently retrieved sequences were affiliated to uncultured ANME. Most of these sequences belonged to ANME-2 subdivisions a and c; these groups occurred in all sediment layers, with the exception of the reference site where no rRNA amplification was obtained. ANME-1-related sequences increased proportionally with depth in rDNA sequence libraries and were dominant in both rRNA and rDNA sequence libraries obtained from the deepest EWM14 sediments (10–17 cmbsf). ANME-3 cluster-related sequences were retrieved in 16S rRNA gene libraries from all cold seep habitats, except for the deepest layer of EWM14. In contrast, rRNA sequences were only detected in the intermediate layers (4–6 cmbsf; Figure 2). Representative sequences of the ANME clusters matched with sequences currently found at Mediterranean Sea mud volcanoes, Gulf of Mexico and Eel River cold seeps (Beal et al., 2009; Lloyd et al., 2010; Pachiadaki et al., 2010, 2011) (Figure 3). However, phylogenetic analysis of 16S rRNA sequences revealed a monophyletic distinct subgroup of ANME-2c with no previous related sequences (Figure 4) named ‘ANME-2c Sonora’ and was found exclusively in the WM12 sulfate-rich sediments both in rRNA and rDNA libraries.
Neighbor-joining distance phylogeny of the archaeal 16S rRNA sequences and rRNA-derived sequences for ANME-methanogens clusters from sediment of Sonora Margin cold seeps based on the Kimura 2 parameters method with 1000 replicates, including 825 homologous positions. Only bootstrap values up to 70% are shown. Only one representative sequence (>97% identical) is shown. Number within parantheses show the number of clones analyzed from RNA or DNA clone libraries. ANME, Anaerobic Methanotrophs; GOM Arc I, Golf of Mexico Archaea group I; WM12, White MAT12;WM14, White MAT14; EWM14, Edge of White MAT14; REF, reference site.
Also, a significant number of sequences was affiliated to other uncultivated Archaea belonging to members of the Deep Sea Hydrothermal Vent Euryarchaeotal Group 8 (DHVE-8) detected in hydrocarbon-rich cold seeps (Lloyd et al., 2010). Other sequences closely related to the Marine Benthic Groups D and B, Terrestrial Miscellaneous Euryarchaeotal Group and Marine Group II, commonly observed in deep marine sediments (Teske and Sorensen, 2008), were found not only in sediment cores underlying microbial mat but also in the reference core. Sequences affiliated to Marine Benthic Group E and Marine Group I dominated in samples from the reference site but were rarely detected in the cold seep habitats (Figures 2 and 3). Finally, only seven sequences affiliated to the Methanoarchaea genus Methanosarcina were found in the deepest layers of WM14 and EWM14, where sequences related to the emergent group of GOM Arc I were also detected (Figure 4).
Quantitative real-time PCR
Owing to the very few data available on microorganisms from cold seeps (pure culture or metagenomic analysis), the number of 16S rRNA genes (rDNA) copy per cell remains undetermined. To compare microbial populations abundance in our samples, we postulated a single 16S rRNA gene copy number for each quantified population.
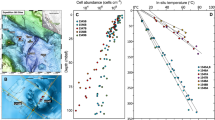
Archaeal and bacterial abundance obtained by Q-PCR suggested an overall dominance of bacterial communities in sediment cores excepted in EWM14 sediments (Figure 5). In the first WM (WM12 and WM14) sediment layers (0–2 cmbsf), Bacteria were almost 20-fold more abundant than Archaea with ∼5 × 108 and ∼9 × 109 16S rDNA copies per g of sediment, respectively. In WM14, archaeal 16S rDNA copy numbers increased with depth (∼3.4 × 109 copies per g at the WM14 deepest layers), while Bacteria 16S rDNA copy numbers decreased (∼3.8 × 109 copies per g). In contrast, in EWM14 sediments, Archaea gene copies remained constant (∼2 × 108 copies per g) and Bacteria concentration decreased rapidly to non-active site bacterial concentration. Archaea abundance was twofold higher in WM12 than in other habitats and increased to 7.8 × 109 16S rDNA copies per g with depth. At the reference site, Bacteria 16S rDNA copies were 10-fold more abundant than archaeal gene copies throughout the entire sediment core and never exceed 2.2 × 109 and 2.9 × 108 16S rDNA copies per g for Bacteria and Archaea, respectively.
DNA copy numbers of the 16S rDNA gene per gram of sediment for Bacteria, Archaea, methanogens and anaerobic methanotrophs, from cores of (a) White MAT12, (b) White MAT14 and (c) Edge White MAT14 in function of depth (0–17 cmbsf) in cold seep sediments of the Sonora Margin. Quantification of specific DNA sequences of methanogenic populations are in logarithmic scale.
The Methanosarcinales, which include the ANME-2 and -3 phylogenetic clusters, dominated the archaeal community in WM sediments by rRNA gene copy abundance (109 16S rDNA copies per g). In EWM14 sediments, the Methanosarcinales remained the most abundant methanogenic order, representing 75% of Archaea in the upper layer (6.7 × 108 copies per g) and 30% below 8 cmbsf (2 × 108 copies per g). 16S rDNA genes copy numbers of Methanococcales and Methanomicrobiales orders were close to 107 copies per g without significant changes. 16S rDNA copy numbers of Methanobacteriales, close to the Q-PCR detection limit in the upper sediment section, increased with depth but remained low (4.8 × 106 copies per g). Methanopyrales representatives, previously isolated in the Guaymas Basin hydrothermal fields, were not detected in cold seep sediments.
Owing to the difference in the standard calibration process (plasmid and/or genomic DNA dilutions), 16S rRNA gene copy numbers of ANMEs could not be directly compared with previous quantifications. However, we were able to compare depth distribution of ANME representative sequences throughout the sediment cores. Quantifications highlighted ANME community concentrations fivefold more elevated in WM12 sediment cores, strongly dominated by ANME-2c (4.9 × 1011 16S rDNA copies per g). ANME-2c representatives also dominated WM14 surface and intermediate sediment layers (0–8 cmbsf) and EWM14 surface layers (0–4 cmbsf) with a maximum of 1.5 × 1011 and 8.5 × 1010 16S rDNA copies per g, respectively (Figure 5). ANME-1 16S copy numbers significantly increased with depth (∼1.3 × 1011 copies per g for WM14 and EWM14 and 2.35 × 1011 for WM12), dominating the WM14 bottom sediment ANME communities and appeared to be the unique ANME population below 8 cmbsf in EWM14. ANME-2a quantifications showed a similar depth distribution as those of ANME-2c but with lower concentrations. Specific DNA sequences of ANME-3 were also present in the intermediate sediment layers of all cold seep samples with a maximum of 5 × 1010 and 4 × 1010 16S rDNA copies per g for WM14 and EWM14 sediments and 9.5 × 1010 16S rDNA copies per g for WM12 cores.
Structure and morphologies of ANME aggregates
FISH observations using archaeal and bacterial probes revealed a high diversity of cell aggregates in size (5–50 μm in diameter), form (regular and irregular balls, chains, and so on) and organization (proportion and/or localization of Archaea and Bacteria) (Figure 6). Using more specific probes, ANME-2/Bacteria-labeled aggregates were found to be dominant in the surface and middle sediment horizons of WM14 and throughout WM12. Microscopic observations of ANME-2/bacterial aggregates showed a wide range of spatial organization, including mixed, concentric or intermediate aggregates (Figure 6D1–D3). Archaeal cells hybridized with ANME-1 probes were detected in the deeper sediment layers of WMs and seemed to be widely dominant in the deeper sediment layers of EWM14. ANME-1 were observed in EWM14 and WM14 as rod chains without detectable bacterial partner either as a single chain or in unstructured heaps (Figure 6B1). In contrast, in WM12 deepest sediments, ANME-1 have also been observed in straight aggregates with bacterial partners (Figure 6B2). Cells labeled with ANME-3 probes were detected in small aggregates (<10 μm) without any detectable bacterial partner in intermediate sediment layers (Figure 6C).
Individual cells and cell aggregates of Archaea, Bacteria and ANMEs visualized with fluorescent-labeled oligonucleotide probes. Each aggregates shown were taken from different pictures. (A, B1 and B2) are maximal image projection pictures, which superposed several stacks in a same representation. (A) Large Archaeal/Bacterial aggregate labeled with ARC915 (yellow) and EUB338 (green) probes observed in the deepest sediment layer of the White MAT14. (B1) Heap of monospecific and single ANME-1 archaeal rods in chain labeled with ANME-1-350 (yellow) probe visualized at the deepest sediment layer of the Edge White MAT14. (B2) Straight aggregate of ANME-1 (ANME-1-350 (yellow)) and Bacteria (EUB338 (green)). (C) Tightly associated cluster of ANME-3 cells, stained with the ANME-3-1249 (orange) obtained from White MAT14. (D1–D3) Various ANME-2/Bacteria aggregates labeled with ANME-2-712 (red) and EUB338 (green) probes. Scale is 10 μm.
Discussion
The occurrence of dominant populations of ANME clusters, detected by FISH, Q-PCR and gene libraries, combined with geochemical gradients, is the first evidence that AOM represented a major microbial process on the Sonora Margin cold seeps. ANME-1, -2 and -3, anaerobic methane-oxidizing populations commonly retrieved in the active microbial cold seep communities (see Knittel and Boetius (2009) and reference therein), were metabolically active in our samples, as indicated by their 16S rRNA detection, and represented almost the whole active archaeal diversity. This co-occurrence of the three ANME clades has only been reported in few studies (Heijs et al., 2005; Pernthaler et al., 2008; Lloyd et al., 2010). However, ANME populations presented different distributions according to the visual and geochemical patterns of the sampled habitats.
Diversity and distribution of ANMEs
ANME-2c dominated both 16S libraries (rRNA and rDNA), quantifications and FISH observations in sediments characterized by higher sulfate concentrations (first sediment layers of WM14 and EWM14; the entire sulfate-rich WM12 sediment cores), suggesting a major role of these ANME representatives in AOM processes at the Sonora Margin. All ANME-2 observed by FISH formed morphologically diverse (size, form and spatial organization) aggregates with Bacteria, as previously reported for AOM (Knittel and Boetius, 2009). This apparently obligatory association of ANME-2c with presumably sulfate-reducing Bacteria gains metabolic energy for both partners (metabolic exchanges improvement due to cell proximity; Stams and Plugge, 2009). However, this obligatory relationship would also restrict those consortia to a specific ecological niche where both electron acceptors and carbon sources for consortia partners were available. If methane was available throughout the sediment cores, sulfate could be limiting (only detected in the WM14 and EWM14 upper sediments layers and in the entire WM12 cores). The observed ANME-2c consortia distribution in WMs could thereby be explained by the bacterial partner requirement for sulfate. Thus, the dense populations of sulfate-dependent ANME-2c consortia, observed throughout WM12 sediments, could be linked to the advective import of seawater sulfate as previously suggested for the Guaymas Basin (Biddle et al., 2012). These particular environmental conditions also seemed to allow the development of the new ‘ANME-2c Sonora’ representatives below 4 cmbsf. However, in EWM14 sediments, ANME-2c were restricted to the upper sediment layers despite a slightly deeper penetration of sulfate. This ANME-2c sulfate-independent distribution in EWM14 sediments indicated that other parameters, such as non-methane hydrocarbon degradation (Orcutt et al., 2008), fluid flow regimen (Niemann et al., 2006; Losekann et al., 2007) or a sediment perturbation by surface assemblages could be involved in ANME-2 distribution. Here, surface colonizers (Beggiatoa mats and macraufona) could be considered as influent factor. Beggiatoa mats, by their gliding mobility through the sediment, could micro-oxygenate the first sediment layers and then inhibit the ANME-2 consortia or be competitive, such as DHVE-8 with ANME-2 bacterial partners, for substrates, such as acetate in the shallow sediment layers. Otherwise, the EWM14 surface deposit feeders probably modified the structure and/or the composition of the underlying sediments (organic matter degradation, substrate availabilities, for instance) and then vertically changed the ANME-2 ecological niche.
Molecular data also highlighted the presence and activity of ANME-1 relatives in each core, in increasing proportion with depth as observed previously (Knittel et al., 2005; Yanagawa et al., 2011). ANME-1 were detected in larger proportion in WM12 sediments and as unique ANME community in EWM14 deepest sediment layers. Moreover, FISH observations highlighted a versatile ANME-1 behavior in terms of bacterial association. The EWM14 and WM14 ANME-1 communities were detected as single or heaps of rods-in-chain. In contrast, ANME-1 were also observed in straight aggregates with bacteria in WM12 deepest sediments. Combined with geochemical data, these results confirmed that ANME-1 preferred completely anoxic and highly sulfidic sediments (Biddle et al., 2012) and suggested that ANME-1 ecophysiology could depend on environmental conditions and more particularly on sulfate concentrations. In sulfate-poor and methane-rich EWM14 sediments, ANME-1 could support a bacterial-independent AOM as suggested previously (Orphan et al., 2002; Maignien et al., 2012), using a different metabolic pathway providing terminal electron acceptors for AOM, such as the use of extracellular proteins, as mentioned by recent metaproteomic analyses (Stokke et al., 2012). Such metabolism, independent of bacterial association and then metabolic bacterial requirements, could occur and be more competitive than syntrophic AOM in EWM14 sediments. ANME-1 metabolism could also persist in sulfate-depleted WM14 sediments, perhaps by switching to methanogenesis, as suggested previously (Meyerdierks et al., 2010; Lloyd et al., 2011; Stokke et al., 2012). In contrast, advective sulfate inputs in WM12 deepest sediments could induce another ANME-1 metabolic shift to a bacterial-dependent AOM in straight consortia as previously observed in sulfate-rich cold seeps (Orcutt et al., 2005), which could be more energetically effective.
ANME-3-related sequences were present throughout the entire sediment cores with a slight increase of 16S copy numbers restricted to the 4–8 cmbsf sediment horizon. Furthermore, ANME-3 communities were shown to be active only in this sediment layer. This suggests that ANME-3 might be restricted to this sediment horizon, as previously reported in submarine mud volcano (Losekann et al., 2007), just below the surface layers influenced by surface communities. ANME-3 were previously detected in syntrophic consortia with sulfate-reducer Bacteria (Niemann et al., 2006; Losekann et al., 2007). However, our FISH observations with the same probe highlighted some unstructured aggregates with no direct contact with bacterial partners, which is consistent with the support of another metabolism independent of bacterial association for ANME-3 (Losekann et al., 2007; Omoregie et al., 2008). Again, another metabolic pathway other than syntrophic AOM, such as methanogenesis or use of metabolic products from the surface layers as substrates or electrons acceptors, could be considered for ANME-3 aggregates in Sonora Margin sediments.
Methane hypothetical origins
As we mentioned above, methanogenesis was often suggested for single ANME, due to their phylogenetic proximity with methanogens and recent metagenomic studies. In cold seep sediments, methane is a preponderant factor, fueling the AOM aggregates, and its origin in the sediments also remained misunderstood.
Methane isotopic ratio suggested a dominant biogenic methane origin, involving methanogens populations. However, abiotic and thermogenic methane origin from deeper sediment layers could not be excluded. Confirming a biogenic origin, sequences relatives to methanogenic Archaea were detected in gene libraries. Methanosarcinales gene copy numbers were high, but anaerobic methanotrophs (ANME-2 and ANME-3) are affiliated to the Methanosarcinales group and could represent most of this population rather than the true methanogens. Nevertheless, Methanococcales, Methanomicrobiales and Methanobacteriales 16S rDNA gene numbers were detected by Q-PCR in the sediment. Even if methanogen 16S rDNA concentrations were low, methanogenic enrichments (culture conditions as described in Lazar et al., 2011b) also confirmed the presence of active methanogen communities (data not shown). Taken together, these results suggest that AOM fueling methane could be produced by small phylogenetically diverse populations of methanogens, eventually supplemented by other unknown or poorly understood microbial populations involved in methane production, such as ANME-1 in the shallow sediment layers. Alternatively, methane could also be produced by larger methanogenic populations in much deeper sediments.
Conclusion
In the anoxic sediments of cold seep environments, the geochemically rich emitted fluids coupled with surface inputs (sulfate and organic carbon) allow the development of complex microbial ecosystems mainly based on methane and sulfur cycles (methanogenesis, AOM, sulfate reduction). If ANME communities were frequently detected and appeared to be ubiquitous in cold seep environments (Knittel and Boetius, 2009), their distributions and their precise functions remained misunderstood. The Sonora Margin cold seeps, by their visually and/or geochemically different habitats, presented a unique opportunity to access the environmental parameters and key factors involved in archaeal distribution and ecophysiology. On the basis of our sampling strategy and detailed microbiological characterizations, we initially confirmed that known methanogens represent only a small proportion of the microbial biomass in the shallow sediments (Lazar et al., 2011a). Indeed, such as for other sedimentary cold seep environments (Gulf of Mexico mats, Mediterranean mud volcanoes or gas hydrate mats), archaeal communities were dominated by ANMEs in the shallow sediment layers (Knittel et al., 2005; Niemann et al., 2006; Lloyd et al., 2010). ANME groups were commonly linked to AOM metabolism; however, the previously reported and the presently observed ANME differences in vertical niche distribution (Knittel et al., 2005; Niemann et al., 2006; Losekann et al., 2007; Yanagawa et al., 2011) and in consortia formation (Orphan et al., 2002; Knittel et al., 2005; Orcutt et al., 2005; Losekann et al., 2007) clearly indicate different ecophysiology. These data suggest that ANME distributions and metabolisms could be directly or indirectly (through bacterial partner requirements) related to flux rate-dependent geochemical gradients (methane, sulfate and sulfide) and substrate availabilities (organic carbon). In addition, these microbiological processes lead to various surface community settlements, such as microbial mats and diverse macrofauna, depending on fluid flux and composition, probably affecting in return the underlying microbial communities by modifying the sediment structure and composition. Depending on these environmental factors, ANME subgroups could have different functions in cold seep environments, such as methanogenesis, and could not be restricted to methane oxidation. Interestingly, ANMEs were also detected in hydrothermal underlying Beggiatoa mat sediments of the Guaymas Basin, monitoring AOM from 15 °C to 95 °C; comparative analyses between those geographically close ecosystems would be needed to discuss on AOM mechanisms and settlement worldwide in sediments.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ . (1990). Basic local alignment search tool. J Mol Biol 215: 403–410.
Amann RI, Binder BJ, Olson RJ, Chisholm SW, Devereux R, Stahl DA . (1990). Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analyzing mixed microbial populations. Appl Environ Microbiol 56: 1919–1925.
Beal EJ, House CH, Orphan VJ . (2009). Manganese- and iron-dependent marine methane oxidation. Science 325: 184–187.
Biddle JF, Cardman Z, Mendlovitz H, Albert DB, Lloyd KG, Boetius A et al (2012). Anaerobic oxidation of methane at different temperature regimes in Guaymas Basin hydrothermal sediments. ISME J 6: 1018–1031.
Boetius A, Ravenschlag K, Schubert CJ, Rickert D, Widdel F, Gieseke A et al (2000). A marine microbial consortium apparently mediating anaerobic oxidation of methane. Nature 407: 623–626.
Cambon-Bonavita MA, Nadalig T, Roussel E, Delage E, Duperron S, Caprais JC et al (2009). Diversity and distribution of methane-oxidizing microbial communities associated with different faunal assemblages in a giant pockmark of the Gabon continental margin. Deep-Sea Res Part I 56: 2248–2258.
Casamayor EO, Massana R, Benlloch S, Ovreas L, Diez B, Goddard VJ et al (2002). Changes in archaeal, bacterial and eukaryal assemblages along a salinity gradient by comparison of genetic fingerprinting methods in a multipond solar saltern. Environ Microbiol 4: 338–348.
Damm E, Budeus G . (2003). Fate of vent-derived methane in seawater above the Hakon Mosby mud volcano (Norwegian Sea). Mar Chem 82: 1–11.
Dhillon A, Lever M, Lloyd KG, Albert DB, Sogin ML, Teske A . (2005). Methanogen diversity evidenced by molecular characterization of methyl coenzyme M reductase A (mcrA) genes in hydrothermal sediments of the Guaymas Basin. Appl Environ Microbiol 71: 4592–4601.
Dhillon A, Teske A, Dillon J, Stahl DA, Sogin ML . (2003). Molecular characterization of sulfate-reducing bacteria in the Guaymas Basin. Appl Environ Microbiol 69: 2765–2772.
Fauchald K, Jumars PA . (1979). The diet of worms: a study of polychaete feeding guilds. Oceanogr Mar Biol Annu Rev 17: 193–284.
Felsenstein J . (1985). Confidence-limits on phylogenies—an approach using the bootstrap. Evolution 39: 783–791.
Fisher MM, Triplett EW . (1999). Automated approach for ribosomal intergenic spacer analysis of microbial diversity and its application to freshwater bacterial communities. Appl Environ Microbiol 65: 4630–4636.
Fonselius S, Dyrssen D, Yhlen B . (2007). Determination of hydrogen sulphide. In: Grasshoff K, Kremling K and Ehrhardt M (eds). Methods of Seawater Analysis, 3rd edition. Wiley-VCH Verlag GmbH: Weinheim, Germany, pp 91–100.
Gadberry MD, Malcomber ST, Doust AN, Kellogg EA . (2005). Primaclade—a flexible tool to find conserved PCR primers across multiple species. Bioinformatics 21: 1263–1264.
Grünke S, Felden J, Lichtschlag A, Girnth AC, De Beer D, WenzhÖFer F et al (2011). Niche differentiation among mat-forming, sulfide-oxidizing bacteria at cold seeps of the Nile Deep Sea Fan (Eastern Mediterranean Sea). Geobiology 9: 330–348.
Heijs SK, Damste JS, Forney LJ . (2005). Characterization of a deep-sea microbial mat from an active cold seep at the Milano mud volcano in the Eastern Mediterranean Sea. FEMS Microbiol Ecol 54: 47–56.
Holler T, Widdel F, Knittel K, Amann R, Kellermann MY, Hinrichs KU et al (2011). Thermophilic anaerobic oxidation of methane by marine microbial consortia. ISME J 5: 1946–1956.
Jorgensen BB, Boetius A . (2007). Feast and famine–microbial life in the deep-sea bed. Nat Rev Microbiol 5: 770–781.
Kimura M . (1980). A Simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide-sequences. J Mol Evol 16: 111–120.
Knittel K, Boetius A . (2009). Anaerobic oxidation of methane: progress with an unknown process. Annu Rev Microbiol 63: 311–334.
Knittel K, Losekann T, Boetius A, Kort R, Amann R . (2005). Diversity and distribution of methanotrophic archaea at cold seeps. Appl Environ Microbiol 71: 467–479.
Lazar CS, Dinasquet J, Pignet P, Prieur D, Toffin L . (2010). Active archaeal communities at cold seep sediments populated by Siboglinidae tubeworms from the Storegga Slide. Microb Ecol 60: 516–527.
Lazar CS, John Parkes R, Cragg BA, L’Haridon S, Toffin L . (2012). Methanogenic activity and diversity in the centre of the Amsterdam Mud Volcano, Eastern Mediterranean Sea. FEMS Microbiol Ecol 81: 243–254.
Lazar CS, L’Haridon S, Pignet P, Toffin L . (2011a). Archaeal populations in hypersaline sediments underlying orange microbial mats in the Napoli mud volcano. Appl Environ Microbiol 77: 3120–3131.
Lazar CS, Parkes RJ, Cragg BA, L'Haridon S, Toffin L . (2011b). Methanogenic diversity and activity in hypersaline sediments of the centre of the Napoli mud volcano, Eastern Mediterranean Sea. Environ Microbiol 13: 2078–2091.
Lloyd KG, Albert DB, Biddle JF, Chanton JP, Pizarro O, Teske A . (2010). Spatial structure and activity of sedimentary microbial communities underlying a Beggiatoa spp. mat in a Gulf of Mexico hydrocarbon seep. PLoS One 5: e8738.
Lloyd KG, Alperin MJ, Teske A . (2011). Environmental evidence for net methane production and oxidation in putative ANaerobic MEthanotrophic (ANME) archaea. Environ Microbiol 13: 2548–2564.
Losekann T, Knittel K, Nadalig T, Fuchs B, Niemann H, Boetius A et al (2007). Diversity and abundance of aerobic and anaerobic methane oxidizers at the Haakon Mosby Mud Volcano, Barents Sea. Appl Environ Microbiol 73: 3348–3362.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: 1363–1371.
Maignien L, Parkes RJ, Cragg B, Niemann H, Knittel K, Coulon S et al (2012). Anaerobic oxidation of methane in hypersaline cold seep sediments. FEMS Microbiol Ecol 83: 214–231.
McKay LJ, MacGregor BJ, Biddle JF, Albert DB, Mendlovitz HP, Hoer DR et al (2012). Spatial heterogeneity and underlying geochemistry of phylogenetically diverse orange and white Beggiatoa mats in Guaymas Basin hydrothermal sediments. Deep Sea Res Part I 67: 21–31.
Meyerdierks A, Kube M, Kostadinov I, Teeling H, Glockner FO, Reinhardt R et al (2010). Metagenome and mRNA expression analyses of anaerobic methanotrophic archaea of the ANME-1 group. Environ Microbiol 12: 422–439.
Meyerdierks A, Kube M, Lombardot T, Knittel K, Bauer M, Glockner FO et al (2005). Insights into the genomes of archaea mediating the anaerobic oxidation of methane. Environ Microbiol 7: 1937–1951.
Niemann H, Losekann T, de Beer D, Elvert M, Nadalig T, Knittel K et al (2006). Novel microbial communities of the Haakon Mosby mud volcano and their role as a methane sink. Nature 443: 854–858.
Omoregie EO, Mastalerz V, de Lange G, Straub KL, Kappler A, Roy H et al (2008). Biogeochemistry and community composition of iron- and sulfur-precipitating microbial mats at the Chefren mud volcano (Nile Deep Sea Fan, Eastern Mediterranean). Appl Environ Microbiol 74: 3198–3215.
Orcutt B, Boetius A, Elvert M, Samarkin V, Joye SB . (2005). Molecular biogeochemistry of sulfate reduction, methanogenesis and the anaerobic oxidation of methane at Gulf of Mexico cold seeps (vol 69, pg 4267, 2005). Geochim Cosmochim Acta 69: 5633–5633.
Orcutt B, Samarkin V, Boetius A, Joye S . (2008). On the relationship between methane production and oxidation by anaerobic methanotrophic communities from cold seeps of the Gulf of Mexico. Environ Microbiol 10: 1108–1117.
Orphan VJ, House CH, Hinrichs KU, McKeegan KD, DeLong EF . (2002). Multiple archaeal groups mediate methane oxidation in anoxic cold seep sediments. Proc Natl Acad Sci USAm 99: 7663–7668.
Pachiadaki MG, Kallionaki A, Dahlmann A, De Lange GJ, Kormas KA . (2011). Diversity and spatial distribution of prokaryotic communities along a sediment vertical profile of a deep-sea mud volcano. Microb Ecol 62: 655–668.
Pachiadaki MG, Lykousis V, Stefanou EG, Kormas KA . (2010). Prokaryotic community structure and diversity in the sediments of an active submarine mud volcano (Kazan mud volcano, East Mediterranean Sea). FEMS Microbiol Ecol 72: 429–444.
Page A, Tivey MK, Stakes DS, Reysenbach AL . (2008). Temporal and spatial archaeal colonization of hydrothermal vent deposits. Environ Microbiol 10: 874–884.
Paull CK, Ussler W, Peltzer ET, Brewer PG, Keaten R, Mitts PJ et al (2007). Authigenic carbon entombed in methane-soaked sediments from the northeastern transform margin of the Guaymas Basin, Gulf of California. Deep Sea Res Part I 54: 1240–1267.
Pernthaler A, Dekas AE, Brown CT, Goffredi SK, Embaye T, Orphan VJ . (2008). Diverse syntrophic partnerships from deep-sea methane vents revealed by direct cell capture and metagenomics. Proc Natl Acad Sci USA 105: 7052–7057.
Saitou N, Nei M . (1987). The neighbor-joining method—a new method for reconstructing phylogenetic trees. Mol Biol Evol 4: 406–425.
Sarradin P-M, Caprais J-C . (1996). Analysis of dissolved gases by headspace sampling gas chromatography with column and detector switching. Preliminary results. Anal Commun 33: 371–373.
Simoneit BR, Leif RN, Ishiwatari R . (1996). Phenols in hydrothermal petroleums and sediment bitumen from Guaymas Basin, Gulf of California. Organic Geochem 24: 377–388.
Simoneit BRT, Lonsdale PF, Edmond JM, Shanks WC . (1990). Deep-water hydrocarbon seeps in Guaymas Basin, Gulf of California. Appl Geochem 5: 41–49.
Snaidr J, Amann R, Huber I, Ludwig W, Schleifer KH . (1997). Phylogenetic analysis and in situ identification of bacteria in activated sludge. Appl Environ Microbiol 63: 2884–2896.
Stams AJM, Plugge CM . (2009). Electron transfer in syntrophic communities of anaerobic bacteria and archaea. Nat Rev Microbiol 7: 568–577.
Stokke R, Roalkvam I, Lanzen A, Haflidason H, Steen IH . (2012). Integrated metagenomic and metaproteomic analyses of an ANME-1-dominated community in marine cold seep sediments. Environ Microbiol 14: 1333–1346.
Suzuki MT, Taylor LT, DeLong EF . (2000). Quantitative analysis of small-subunit rRNA genes in mixed microbial populations via 5′-nuclease assays. Appl Environ Microbiol 66: 4605–4614.
Tamura K, Dudley J, Nei M, Kumar S . (2007). MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24: 1596–1599.
Teske A, Hinrichs KU, Edgcomb V, Gomez AD, Kysela D, Sylva SP et al (2002). Microbial diversity of hydrothermal sediments in the Guaymas Basin: evidence for anaerobic methanotrophic communities. Appl Environ Microbiol 68: 1994–2007.
Teske A, Sorensen KB . (2008). Uncultured archaea in deep marine subsurface sediments: have we caught them all? ISME J 2: 3–18.
Thioulouse J, Dray S . (2007). Interactive multivariate data analysis in R with the ade4 and ade4TkGUI packages. J Statist Softw 22: 1–14.
Waren A, Ponder WF . (1991). New species, anatomy, and systematic position of the hydrothermal vent and hydrocarbon seep Gastropod Family Provannidae Famn (Caenogastropoda). Zool Scr 20: 27–56.
Yanagawa K, Sunamura M, Lever MA, Morono Y, Hiruta A, Ishizaki O et al (2011). Niche separation of methanotrophic archaea (ANME-1 and -2) in methane-seep sediments of the Eastern Japan Sea Offshore Joetsu. Geomicrobiol J 28: 118–129.
Yu Y, Lee C, Kim J, Hwang S . (2005). Group-specific primer and probe sets to detect methanogenic communities using quantitative real-time polymerase chain reaction. Biotechnol Bioeng 89: 670–679.
Zhou J, Bruns MA, Tiedje JM . (1996). DNA recovery from soils of diverse composition. Appl Environ Microbiol 62: 316–322.
Acknowledgements
We are indebted to the crews of the research vessel L'Atalante and the submersible Nautile of the cruise ‘BIG’ and the scientific team for their help on board. This cruise was funded by IFREMER (France) and has benefited from a work permit in Mexican waters (DAPA/2/281009/3803, 28 October 2009). We thank Stéphane L'Haridon, Gordon Webster and Catherine Larose for very helpful scientific discussions and comments. This study was supported by an IFREMER PhD grant.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Vigneron, A., Cruaud, P., Pignet, P. et al. Archaeal and anaerobic methane oxidizer communities in the Sonora Margin cold seeps, Guaymas Basin (Gulf of California). ISME J 7, 1595–1608 (2013). https://doi.org/10.1038/ismej.2013.18
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2013.18
Keywords
This article is cited by
-
Thermogenic hydrocarbon biodegradation by diverse depth-stratified microbial populations at a Scotian Basin cold seep
Nature Communications (2020)
-
Discriminative biogeochemical signatures of methanotrophs in different chemosynthetic habitats at an active mud volcano in the Canadian Beaufort Sea
Scientific Reports (2019)
-
Effect of pressure and temperature on anaerobic methanotrophic activities of a highly enriched ANME-2a community
Environmental Science and Pollution Research (2018)
-
The vertical distribution of prokaryotes in the surface sediment of Jiaolong cold seep at the northern South China Sea
Extremophiles (2018)
-
Comparative metagenomics of hydrocarbon and methane seeps of the Gulf of Mexico
Scientific Reports (2017)