Abstract
Patterns of microbial biogeography result from a combination of dispersal, speciation and extinction, yet individual contributions exerted by each of these mechanisms are difficult to isolate and distinguish. The influx of endospores of thermophilic microorganisms to cold marine sediments offers a natural model for investigating passive dispersal in the ocean. We investigated the activity, diversity and abundance of thermophilic endospore-forming sulfate-reducing bacteria (SRB) in Aarhus Bay by incubating pasteurized sediment between 28 and 85 °C, and by subsequent molecular diversity analyses of 16S rRNA and of the dissimilatory (bi)sulfite reductase (dsrAB) genes within the endospore-forming SRB genus Desulfotomaculum. The thermophilic Desulfotomaculum community in Aarhus Bay sediments consisted of at least 23 species-level 16S rRNA sequence phylotypes. In two cases, pairs of identical 16S rRNA and dsrAB sequences in Arctic surface sediment 3000 km away showed that the same phylotypes are present in both locations. Radiotracer-enhanced most probable number analysis revealed that the abundance of endospores of thermophilic SRB in Aarhus Bay sediment was ca. 104 per cm3 at the surface and decreased exponentially to 100 per cm3 at 6.5 m depth, corresponding to 4500 years of sediment age. Thus, a half-life of ca. 300 years was estimated for the thermophilic SRB endospores deposited in Aarhus Bay sediments. These endospores were similarly detected in the overlying water column, indicative of passive dispersal in water masses preceding sedimentation. The sources of these thermophiles remain enigmatic, but at least one source may be common to both Aarhus Bay and Arctic sediments.
Similar content being viewed by others
Introduction
Mounting evidence indicates that microbial populations exhibit complex biogeographical patterns (see Martiny et al., 2006; Falcón et al., 2008; Ionescu et al., 2010; Schauer et al., 2010; Nemergut et al., 2011). Passive dispersal of cells seems to have an important role in shaping these patterns (Green and Bohannan, 2006; Zengler, 2008), but little is known about the extent of dispersal, due to the difficulty of unlinking dispersal and evolutionary history of free-living microorganisms (Martiny et al., 2006; Ramette and Tiedje, 2007). The presence of endospores of thermophiles in permanently cold marine sediments appears to be a result of passive dispersal in the ocean, as these bacteria must originate from warm environments distant from the sediments where they have been discovered (Bartholomew and Paik, 1966; Isaksen et al., 1994; Hubert et al., 2009). Given that these exogenous cells are inactive and thereby less prone to selective pressure leading to speciation and extinction, these particular populations offer a natural model for investigating dispersal processes in the marine environment.
Investigations of the diversity and abundance of such thermophiles have focused on endospores of sulfate-reducing bacteria (SRB) in cold marine sediments of Svalbard. We have estimated that these thermophilic SRB have been supplied to Arctic sediments of western Svalbard at a rate of 108 endospores per square meter per year for at least the past 120 years and have identified them as members of the genus Desulfotomaculum (Hubert et al., 2009), which encompasses the majority of known endospore-forming SRB species. The influx furthermore includes a diverse group of dormant endospores capable of catalyzing anaerobic hydrolysis and fermentation of organic matter upon germination (Hubert et al., 2010). Six different species-level Desulfotomaculum 16S rRNA sequence phylotypes have been enriched and identified in heated Arctic sediments (Vandieken et al., 2006; Hubert et al., 2009, 2010). The studies have included the isolation of pure cultures of some of these SRB by incubating sediment at 28 to 50 °C, which is far above the year-round in situ temperatures of −2 to +4 °C. The Svalbard findings coincide with the earlier results of incubation experiments conducted with sediment from Aarhus Bay, on the east coast of Jutland, Denmark. These showed that the germination of endospores of aerobic (Thamdrup et al., 1998), fermentative and sulfate-reducing thermophiles (Isaksen et al., 1994) can be induced in sediment incubated at up to 70 °C. A thermophilic sulfate-reducing bacterium (strain P60) was isolated and classified as Desulfotomaculum kuznetsovii based on its physiology (Isaksen et al., 1994). However, so far the diversity of thermophilic endospore-forming SRB present in Aarhus Bay or other sediments has not been investigated, hence the extent of dispersal of the thermophilic populations discovered in Svalbard sediments remains unknown. In the present study, we expand the study of thermophilic SRB in cold marine sediments. By incubating sediment with radioactive sulfate tracer, followed by community fingerprinting, cloning and sequencing of 16S rRNA and dsrAB (encoding the two major subunits of the dissimilatory (bi)sulfite reductase; (Zverlov et al., 2005)), we provide evidence of large-scale and long-term passive dispersal of a diverse group of SRB endospores in the marine environment.
Materials and methods
Sampling
Marine sediment was sampled in January 2009 and May 2010 from station M5 (56°06′20′N, 10°27′48′E, 28 m water depth) in Aarhus Bay, located on the transition between the Baltic Sea and North Sea. Surface sediment (0–10 cm) was collected with a box corer (Hessler and Jumars, 1974) and stored at 4 °C in gas-tight plastic bags (Hansen et al., 2000). Sediment from 0 to 650 cm depth was collected using a gravity corer (the top 10 cm were discarded). Cores were sectioned and sediment was stored as described above.
Seawater was sampled in August 2010 at the nearby station M1 (56°09′10′N, 10°19′20′E) in Aarhus Bay, from 1 and 14 m water depth (25 l at either depth). The total water depth at this site is 16 m. Seawater was sequentially filtered through glass fiber (GF) filters (GF-75, Advantec, Knebel, Denmark) and through polycarbonate filters of 0.45 (HVLP, Millipore, Hellerup, Denmark) and 0.2 μm pore size (Cyclopore PC, Whatman, Kent, UK) to collect suspended material. A total of seven GF, three 0.45 and three 0.2 μm pore size filters from each water depth were used for incubation experiments. In all cases, material collected on each filter was derived from 1 l of seawater. All filters were stored in an oxygen-free atmosphere in anaerobic jars at 4 °C for 3 months before experimental incubation.
Sediment slurry preparation
Surface sediment slurries were prepared by homogenizing sediment with sterile anoxic synthetic seawater medium at a 1:2 (w/w) ratio under a constant flow of N2 gas. Synthetic seawater was prepared according to Isaksen et al. (1994). The sulfate concentration was 20 mM. The sediment slurry was divided into two separate master slurries. One slurry was left unamended and the other slurry was amended with organic substrates (formate, lactate, acetate, succinate, propionate, butyrate and ethanol, each to a final concentration of 1 mM).
Slurry incubations in a temperature gradient
Aliquots of 11 ml from substrate-amended and unamended master slurries were transferred to Hungate tubes under a constant flow of N2 and stored overnight at 4 °C. The subslurries were pasteurized at 85 °C for 30 min and thereafter immediately incubated between 28 and 85 °C in a temperature gradient block (TGB). Replicate substrate-amended and unamended subslurries (15 each) all received 200 kBq 35S-labeled carrier-free sulfate tracer and were incubated in alternating slots of the TGB for subsequent measurements of sulfate reduction. Additional 15 replicate subslurries without radioactive tracer were incubated in adjacent TGB slots for subsequent RNA extraction. Slurries were subsampled after 24, 72 and 120 h. Three milliliter aliquots were removed from the radioactive (35S-sulfate) slurries, mixed with 6 ml 20% zinc acetate solution and frozen at −20 °C until distillation by the single-step cold chromium method (Kallmeyer et al., 2004). At the same time, 2 ml aliquots were removed from the non-radioactive slurries, pelleted by centrifugation and immediately frozen in liquid N2 and stored at −80 °C until RNA extraction (see below).
T-MPN enumeration of thermophilic SRB endospores
Radiotracer most probable number enumerations (T-MPN, Vester and Ingvorsen, 1998) were performed to estimate the abundance of thermophilic endospore-forming SRB in Aarhus Bay sediment samples. Sediment from seven depths (0–10, 110–120, 210–220, 310–320, 410–420, 510–520 and 640–650 cm below seafloor) were inoculated into sterile sediment medium prepared according to Vester and Ingvorsen (1998) with the following modifications. Aarhus Bay station M5 surface sediment was diluted with synthetic seawater medium (1:1, w/w; composition described above), homogenized and successively passed through 2.0, 1.0 and 0.5 mm mesh sieves to facilitate a homogeneous distribution of slurry medium into tubes. This sediment slurry medium was autoclaved for 1 h at 121 °C and distributed into Hungate tubes (9 ml aliquots) under a constant flow of N2, incubated at 50 °C for 3 days, autoclaved a second time for 1 h, incubated again at 50 °C for 1 day and finally autoclaved a third time for 1 h. Before use, sodium dithionite (200 μM) was injected as a reducing agent and the sterile sediment medium was amended with organic substrates (formate, lactate, acetate, succinate, propionate, butyrate and ethanol, each at a final concentration of 1 mM).
For each sediment depth, triplicate 10-fold serial dilutions were prepared in Hungate tubes from undiluted to a 10−7 dilution. The first tube of each series was inoculated with 1 ml of pristine sediment and pasteurized at 85 °C for 30 min before the subsequent dilution transfers. Each tube was homogenized by vigorous shaking after inoculation, which was facilitated by the presence of three 0.5 cm diameter glass beads in the tubes. Finally, each tube received 400 kBq 35S-labeled carrier-free sulfate tracer and was incubated at 50 °C. In addition, 35 non-inoculated tubes (that is, only the autoclaved sediment medium and radiotracer) were incubated as negative controls. After 30 days of incubation, 3 ml aliquots were withdrawn from each tube and preserved by the addition of 6 ml 20% (w/v) zinc acetate and freezing. Following cold chromium distillation of the total reduced inorganic sulfur (TRIS; Kallmeyer et al., 2004), tubes were scored positive for SRB if 35S-calculated sulfate reduction was above the threshold of TRIS%=0.1% (Vester and Ingvorsen, 1998). TRIS% is calculated as [a/(a+A)] × 100, where a is the total radioactivity of reduced inorganic 35S-labeled sulfur produced (TRIS) per volume and A is the total radioactivity of 35S-labeled sulfate per volume after incubation (Vester and Ingvorsen, 1998). For sediment samples from 0–10 and 640–650 cm depth, tracer-free dilution series were prepared and incubated in parallel to the series that received radiolabeled tracer. From these series, DNA was extracted (see below) for further population studies (2 ml aliquots were removed after 30 days and preserved as described above).
Filter incubations
Filters that collected planktonic biomass from 1 and 14 m water depth were incubated at 50 °C in three times autoclaved sediment medium (described above) that was amended with organic substrates and 400 kBq 35S-labeled carrier-free sulfate tracer in 10 ml Hungate tubes. Five GF filter incubations were unpasteurized and two were pasteurized at 85 °C for 30 min immediately before the incubation for both 1 m and 14 m water depth. The 0.45 and 0.2 μm pore size filters were incubated without pasteurization. In addition, 24 negative controls were incubated: 5 contained pristine GF filters and 19 contained no filter but only medium. After 30 days of incubation, the samples were processed as described above for T-MPN.
RNA extraction and RT-PCR
RNA was extracted using the Power Soil Total RNA Isolation Kit (Mo Bio Laboratories, Carlsbad, CA, USA) from ca. 0.3 g of sediment. Yield and purity of extracts were evaluated using a NanoDrop spectrophotometer (Thermo Scientific, Hvidovre, Denmark). RNA extracts were treated with 2 units per μl Turbo DNase (Ambion/Applied Biosystems, Naerum, Denmark) for 30 min at 37 °C, and purified with RNeasy Mini Kit (Qiagen, Copenhagen, Denmark). RT-PCR was performed using the OneStep RT-PCR Kit (Qiagen). Reactions of 50 μl included 2 μl bovine serum albumin (10 μg μl−1), 3 μl of each of the general bacterial 16S rRNA primers 8F and 1492R (10 pmol μl−1; Loy et al., 2002) and 10 μl of DNase-treated RNA template (between 4 and 14 ng μl−1). Reverse transcription at 50 °C for 30 min was immediately followed by 20 PCR cycles of 94 °C for 40 s, 52 °C for 40 s and 72 °C for 1.5 min, and final elongation at 72 °C for 10 min. As controls for the efficiency of DNase treatment, parallel PCR reactions using non-RT-PCR-treated purified RNA extracts as template were set using the HotStarTaq Master Mix Kit (Qiagen) and did not yield any product. All PCR products were purified using GenElute PCR Clean-Up Kit (Sigma-Aldrich, Copenhagen, Denmark), and used as template for Desulfotomaculum-specific 16S rRNA gene PCR with the primers DEM116F and DEM1164R as described by Stubner and Meuser (2000). PCR mixtures of 50 μl contained 2 μl bovine serum albumin (10 μg μl−1), 25 μl of 2 × Taq Master Mix RED (Ampliqon, Odense, Denmark), 1 μl of each primer (forward and reverse primers at a concentration of 10 μM each) and 2 μl of template cDNA. Thermal cycling was performed as described by Stubner and Meuser (2000), with annealing at 63 °C and 30 cycles.
DNA extraction and PCR
DNA was extracted as described previously (Kjeldsen et al., 2007) from ca. 0.3 g sediment samples before incubation (0 h) and after pasteurization and incubation at 50 °C (120 h). For DNA extracts obtained after incubation at 50 °C, amplification of Desulfotomaculum spp. 16S rRNA genes with the DEM116F-DEM1164R primer pair was performed as described above (with 35 PCR cycles). For the DNA extracted from pristine sediment, nested PCR was performed and consisted of 20 cycles with the universal bacterial primers 8F and 1492R, followed by 35 cycles with the DEM primer pair. PCR products were purified using GenElute PCR Clean Up Kit (Sigma-Aldrich) before cloning.
Approximately 1.9 kbp fragments of dsrAB were amplified using primer variant mixtures DSR1Fabc and DSR4Rabcd (Loy et al., 2004) and 40 PCR cycles with an annealing temperature of 52 °C. Specific primer pairs were designed for two main thermophilic Desulfotomaculum lineages. The primer pair TsporeF2F (5′-CGTTGGTGTTATGGGTTATG-3′) and TsporeF2R (5′-GGAGCTTTTCTGTGRATACC-3′) targets ca. 1.7 kbp fragments of D. alkaliphilum-related dsrAB sequences (that is, 16S rRNA phylotype F; see Figure 2). The primer pair TsporeG2F (5′-CAGGTATTCCGACCTTCCG-3′) and TsporeG2R (5′-GAATTTAGGCAGACCAAAGCG-3′) targets ca. 900 kbp fragments of dsrAB sequences of D. thermosapovorans-related strains isolated from Svalbard sediments (16S rRNA phylotype G; see Figure 2). Both primer pairs were applied at an annealing temperature of 54 °C using 40 PCR cycles. PCR amplicons of the expected size were purified from 1% (w/v) agarose gels (GenElute Gel Extraction Kit, Sigma-Aldrich) before cloning.
Cloning and sequence analyses
PCR products were cloned using the pGEM-T Vector System II cloning kit (Promega Biotech, Nacka, Sweden), according to the manufacturer’s instructions. Plasmids were extracted using GenElute Plasmid Miniprep Kit (Sigma-Aldrich) and inserts were sequenced (Macrogen, Seoul, Korea) using the vector-specific primers M13F and M13R (Messing, 1983). A set of internal sequencing primers (data not shown) was designed to obtain full-length sequences of the cloned dsrAB PCR amplicons. The 16S rRNA sequences were aligned using the SILVA webaligner (Pruesse et al., 2007) and added to the SILVA SSURef version 102 small subunit ribosomal RNA database (Pruesse et al., 2007) using the ARB software package (Ludwig et al., 2004). The dsrAB sequences were added to a dsrAB sequence ARB database and aligned according to their inferred amino-acid sequence (Zverlov et al., 2005). Phylogenetic sequence analyses were performed using ARB except for bootstrap analyses, which were performed using PHYLIP v3.67 (distributed by J Felsenstein, Department of Genome Sciences, University of Washington, Seattle, WA, USA). After manually refining alignments, 16S rRNA gene sequences were grouped into phylotypes using a 99% uncorrected similarity cutoff and the furthest neighbor algorithm of Mothur (Schloss et al., 2009). The obtained 16S rRNA and dsrAB sequences have been deposited in the GenBank database under the accession numbers JQ304523 to JQ304753 and JQ304754 to JQ304785, respectively.
Denaturing gradient gel electrophoresis (DGGE)
For DGGE analyses of Desulfotomaculum-derived 16S rRNA fragments, RT-PCR (from temperature gradient incubations) or PCR (from T-MPN) were conducted with primers DEM116F and DEM1164R for 30 cycles, as described above. The products were purified and used as template for a second PCR reaction with the general bacterial DGGE primers 341F (including a 40 nt 5′-terminal GC-clamp, Muyzer et al., 1993) and 907R (Lane, 1991). For the second PCR reaction, thermocycling consisted of initial denaturation at 94 °C for 1 min, 20 cycles of 94 °C for 40 s, 57 °C for 40 s and 72 °C for 1 min, and a final elongation step at 72 °C for 10 min.
DGGE was performed on 8% acrylamide gels with a 20–80% urea and formamide denaturing gradient (100% denaturant was 7 M urea and 40% (v/v) deionized formamide). Gels were run in 60 °C Tris-acetate-EDTA buffer for ca. 15 h at 100 V, then stained with SybrGold (Invitrogen, Naerum, Denmark) for visualization under UV light (Gel Doc XR, Bio-Rad, Copenhagen, Denmark).
For the samples derived from T-MPN incubations, visible bands were excised and reamplified using the DGGE primer pair (without the GC-clamp on the forward primer), purified (GenElute PCR Clean-Up Kit) and sequenced using the reverse primer. The DGGE banding pattern of the samples from the temperature gradient incubations was subjected to non-metric multidimensional scaling analysis (Legendre and Legendre, 1998) using metaMDS of the vegan package (Oksanen et al., 2010). The analysis was based on the presence/absence of DGGE bands that occurred in more than one sample and used a Jaccard distance measure. The variables of temperature and incubation time were fitted as z-scores.
Results
Identification of distinct communities of thermophilic Desulfotomaculum at different temperatures
Germination of SRB endospores was induced by incubating pasteurized Aarhus Bay sediment between 28 and 85 °C as slurries that were either amended with organic substrates or left unamended (Figures 1a and b, respectively). Under either condition, two temperature optima for sulfate reduction emerged during a 5-day incubation period. Broad temperature-activity distributions with optima between 45 and 60 °C were apparent in both conditions after 24 and 72 h. After 120 h of incubation, a second optimum at ca. 70 °C was observed. Seventy degrees Celsius was also the highest incubation temperature at which sulfate reduction could be measured. Sulfate reduction was detected at 28 °C, the lowest temperature tested, but only after 72 and 120 h in both amended and unamended incubations. The amount of sulfate reduced at temperatures below 40 °C was smaller than at higher temperatures. Organic substrate amendment promoted 5 to 10-fold higher sulfate reduction (Figures 1a and b).
Replicate Aarhus Bay sediment slurries were amended with organic substrates (a) or left unamended (b) and were incubated between 28 and 85 °C in a TGB for up to 120 h. Sulfate reduction (μM sulfate consumed) at each incubation temperature was measured after 24 (triangles), 72 (squares) and 120 h of incubation (circles). RNA was extracted from selected unamended samples (b) and DGGE targeting the 16S rRNA of Desulfotomaculum spp. was performed. DGGE images for samples from 24 and 120 h are shown (c); samples from 72 h are omitted due to their similarity to 120 h. Non-metric multidimensional scaling analysis (d) was conducted to compare the DGGE patterns at each temperature and incubation time. Symbols in (d) correspond to the ones in (a) and (b).
The broad temperature distribution of sulfate reduction, from 28 to 70 °C, and the occurrence of two optima after 120 h of incubation indicate that distinct populations of endospore-forming SRB were induced at different temperatures. This hypothesis was supported by rRNA-based DGGE fingerprinting analysis of the Desulfotomaculum community in unamended sediment incubated at 32, 46, 57, 61, 63 and 69 °C (Figures 1b–d). Non-metric multidimensional scaling analysis of shared DGGE bands indicated that temperature was a significant parameter explaining the difference in the community fingerprints (P=0.001). Samples obtained after 72 and 120 h of incubation were more similar to each other than to those collected after 24 h (Figure 1d), suggesting a community shift as a function of incubation time. However, the effect of incubation time was not statistically significant (P=0.340).
On the basis of the initial DGGE results (Figures 1c and d), cDNA clone libraries were constructed from unamended samples incubated at 46, 61 and 69 °C to identify active Desulfotomaculum populations. To focus on the effect of temperature, sequences that were retrieved from samples collected after 24 and 120 h of incubation were pooled before the analysis. The largest diversity, of 11 Desulfotomaculum phylotypes, was observed in clone libraries derived from sediment samples that were incubated at 46 °C. At 61 and 69 °C, five and one phylotype were retrieved, respectively (Figure 2 and Table 1). None of the 17 phylotypes occurred in more than one of the three clone libraries, indicating that different temperatures induced different populations of the overall community of thermophilic SRB endospores.
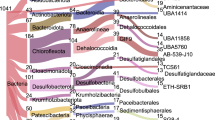
Phylogenetic positions of retrieved 16S rRNA sequences affiliated with the endospore-forming Desulfotomaculum lineage. Phylotypes of putative thermophilic Desulfotomaculum spp. were defined based on 99% sequence similarity. A total of 128 clone sequences were derived from RNA extracted from pasteurized sediment slurries incubated at 46 ( ), 61 (
), 61 ( ) and 69 °C (
) and 69 °C ( ). A total of 63 sequences were derived from DNA extracted from pasteurized sediment slurries incubated at 50 °C for 120 h (
). A total of 63 sequences were derived from DNA extracted from pasteurized sediment slurries incubated at 50 °C for 120 h ( : unamended slurry;
: unamended slurry;  : organic substrate-amended slurry). Additionally, 36 clone sequences were derived from DNA extracted from sediment not incubated at high temperature (DNA 0 h,
: organic substrate-amended slurry). Additionally, 36 clone sequences were derived from DNA extracted from sediment not incubated at high temperature (DNA 0 h,  ). Also shown (
). Also shown ( ) are DNA sequences originating from two studies of thermophilic Desulfotomaculum in permanently cold Arctic sediment from Svalbard (Hubert et al., 2009, 2010). Filled and open squares at branching nodes indicate lineages with >90% and 70 to 90% parsimony bootstrap support, respectively (1000 resamplings). The scale bar indicates 5% sequence divergence as estimated from maximum-likelihood analysis. A complete list of sequences and GenBank accession numbers is provided in Supplementary Table S1 (Supplementary Information).
) are DNA sequences originating from two studies of thermophilic Desulfotomaculum in permanently cold Arctic sediment from Svalbard (Hubert et al., 2009, 2010). Filled and open squares at branching nodes indicate lineages with >90% and 70 to 90% parsimony bootstrap support, respectively (1000 resamplings). The scale bar indicates 5% sequence divergence as estimated from maximum-likelihood analysis. A complete list of sequences and GenBank accession numbers is provided in Supplementary Table S1 (Supplementary Information).
To assess the effect of organic substrates on the composition of induced thermophilic Desulfotomaculum communities, DNA clone libraries of 16S rRNA genes were constructed from substrate-amended and unamended sediments following 50 °C incubation for 120 h. The 31 sequences obtained from the unamended incubation clustered into seven phylotypes, whereas the 32 sequences obtained from the amended incubation clustered into four phylotypes, of which two (G and R) were also present in the unamended incubation (Table 1 and Figure 2). This suggests that amendment with the chosen substrates promoted higher sulfate reduction rates by stimulating the growth of a limited subset of populations among the thermophilic SRB that are present in Aarhus Bay sediment, rather than by evenly stimulating several SRB populations that would otherwise remain undetected.
To investigate whether the detected Desulfotomaculum phylotypes represent vegetative populations in the sediment surviving pasteurization, a Desulfotomaculum-specific 16S rRNA gene library was constructed from pristine sediment (that is, not incubated at high temperature). Only 1 out of the 36 sequences obtained from this preincubation library clustered with sequences from heated sediment (phylotype D, Figure 2). Detecting this sequence in the preincubation library is most likely due to DNA extraction from endospores of this taxon present in pristine sediment and due to the increased sensitivity of the nested PCR approach, as there have been no indications of vegetative cells of thermophiles in previous studies of pristine Aarhus Bay sediments (Thomsen et al., 2001; Leloup et al., 2009; Webster et al., 2011). The fact that phylotype D increased in relative abundance in clone libraries from 3 to 52% when the sediment was incubated at 50 °C (unamended; Figure 2, Table 1) suggests that an enrichment of this particular organism occurred at high temperature. Not detecting any other putative thermophilic endospore-forming Desulfotomaculum spp. in the pristine sample and the pasteurization procedure support the assumption that these SRB are present as endospores. Incubation at temperatures above those in situ unveils the phylogenetic diversity of these thermophiles in cold sediments.
Genotypic identity between thermophilic Desulfotomaculum present in both Aarhus Bay and Svalbard sediments
Three of the thermophilic Desulfotomaculum phylotypes present in Aarhus Bay (D, F and G) were previously discovered in heated sediment from an Arctic fjord in Northwest Svalbard (Figure 2 and Table 1; Hubert et al., 2010). Alignments of Svalbard and Aarhus Bay 16S rRNA sequences revealed up to 99.9% sequence identity for both phylotypes D and F, and 100.0% for phylotype G. These close genetic relationships are also observed when comparing dsrAB sequences. Our clone libraries based on the general dsrAB primer pair DSR1Fabc–DSR4Rabcd revealed that a cloned sequence related to D. thermosapovorans (corresponding to phylotype G), derived from heated Aarhus Bay sediment, and the dsrAB sequence of a thermophilic Desulfotomaculum isolated from heated Svalbard sediment have only one mismatch over 1937 nucleotides (that is, 99.9% sequence identity; Figure 3, accession numbers JQ304758 and JQ304755, respectively). This single mismatch could be due to random errors introduced by Taq polymerase during PCR amplification, as Taq misincorporation rates are estimated to be ca. 2 × 10−5 nucleotides per cycle (Klepac-Ceraj et al., 2004). The general dsrAB primer pair failed to target dsrAB sequences related to D. halophilum and D. alkaliphilum, which correspond to 16S rRNA phylotype F (Figure 3). Therefore, a new dsrAB primer pair was designed to target members of this lineage, which enabled cloning dsrAB sequences derived from Aarhus Bay heated sediment sharing 99.7% similarity to a cloned sequence from Svalbard heated sediment; that is, with five mismatches over 1654 nucleotides (Figure 3; accession numbers JQ304768 and JQ304772, respectively). Due to the lack of close cultured relatives, organisms belonging to phylotype D (Figure 2) currently remain unidentified at the dsrAB level and thus the relatedness between dsrAB sequences of this third Desulfotomaculum sp. found in both Aarhus Bay and Svalbard sediments could not be analyzed.
Phylogenetic positions of representative dsrAB clones retrieved from slurries of pasteurized sediment (0–10 cm depth) incubated at 50 °C for 120 h with and without organic growth substrate amendment, and of dsrAB clones retrieved from similar incubations of sediment from 5 and 650 cm below the seafloor (cmbsf) with a primer pair designed to target dsrAB sequences affiliated with D. alkaliphilum and D. halophilum. Numbers in parentheses show the relative abundance of the respective clones within a given clone library. The tree was inferred from FITCH distance matrix analysis of 532 aligned DsrAB amino-acid sequence positions with JTT distance correction, global rearrangements, randomized input order of sequences and one category of substitution rates. Closed circles indicate nodes receiving >70% neighbor joining analysis-based bootstrap support (JTT distance correction, 500 resamplings). The scale bar represents 10% estimated sequence divergence. §Short sequence manually drawn in to the tree (the sequence shares 100% identity with its closest relative in the tree). Specific details of the incubation condition of the sediments from which the dsrAB clone sequences were obtained are provided in Supplementary Table S2 (Supplementary Information). A full version of the tree in shown in Supplementary Figure S1.
Thermophilic SRB endospores in deep sediments of Aarhus Bay
The quantitative distribution of thermophilic SRB in deep sediments was investigated by T-MPN (Vester and Ingvorsen, 1998). MPN-estimated cell density decreased exponentially from 4 to 9 × 103 thermophilic SRB endospores per cm3 of sediment at the surface to 1 × 100 at 650 cm depth (Figure 4a). The dominant thermophilic SRB phylotype in the uppermost and deepest sediments were identified via Desulfotomaculum-targeted 16S rRNA gene DGGE using DNA extracted from positive MPN dilutions (Figure 4b). Prominent bands were excised and sequenced (Table 1). Phylotypes G and R were detected in undiluted surface sediment after 30 days of incubation at 50 °C, whereas in the 10−3 MPN dilution of surface sediment only phylotype F was detected, indicating its relative abundance in situ. Furthermore, only phylotype F was detected following incubation of undiluted sediment (the only positive MPN dilution) from 650 cm depth (Figure 4b). The 16S rRNA DGGE results were supported by dsrAB clone library results. Using group-specific dsrAB primer pairs, phylotype F was detected in both the deepest and uppermost sediment, whereas phylotype G was detected in the uppermost sediment but not at 650 cm depth (Figure 3). The cloned dsrAB sequences from 650 cm sediment depth were 99.7% identical (1650 out of 1655 nucleotides) to one of the sublineages of phylotype F detected at the surface sediment (Figure 3, accession numbers JQ304785 and JQ304770, respectively).
The depth distribution of thermophilic SRB endospores in Aarhus Bay sediment was assessed by T-MPN and correlated to sediment age (a) according to C-14 dating of shells (JB Jensen, unpublished). From the 35 non-inoculated controls, only 3 yielded TRIS% above the background threshold of 0.10 (maximum TRIS%=0.47, data not shown). Error bars show 95% confidence intervals of T-MPN results. DGGE targeting the 16S rRNA of Desulfotomaculum spp. was performed (b) with DNA extracted from serial dilutions of sediment from 0–10 and 640–650 cm depth. Bands were excised, sequenced and identified as the phylotypes shown in the figure.
Thermophilic SRB endospores in the water column of Aarhus Bay
Thermophilic SRB were detected at 1 and 14 m water depth, in both pasteurized and unpasteurized incubations that were inoculated with biomass concentrated by filtration of water. For 1 m water depth thermophilic sulfate reduction was observed in three out of the seven filter incubations, whereas at 14 m water depth sulfate reduction was measured in all filter incubations (Figure 5). The detection of greater sulfate reduction for filters from 14 m may indicate higher abundance of endospores in deeper water; a larger initial community of endospores likely corresponds with a greater diversity of endospores, hence the ability to use a wider variety of substrates and increase the total amount of sulfate reduced (Figure 5).
Seawater from 1 and 14 m water depth in Aarhus Bay was sequentially filtered through GF filters, 0.45 and 0.2 μm filters, which were then incubated in sterile sediment medium at 50 °C for 30 days. Thermophilic sulfate reduction was detected in incubations of GF filters from both water depths (TRIS% from 2 to 55), as shown in the figure. No sulfate reduction was detected in incubations of 0.45 and 0.2 μm filters (TRIS%=0.00, data not shown). From the 24 negative control incubations, TRIS% was above background in 1 out of the 5 unused GF filters and 3 out of the 19 non-inoculated controls (maximum TRIS%=0.47, data not shown). BD, below detection limit.
Discussion
Long-distance dispersal of thermophilic endospores
This study shows that the community of dormant thermophilic SRB in cold marine sediments in general and Aarhus Bay in particular is much more diverse than previously recognized (Isaksen et al., 1994; Hubert et al., 2010). In total, 23 phylotypes were identified in incubation experiments conducted under different conditions with respect to temperature and organic substrate availability. A total of 19 of these 23 phylotypes share less than 97% 16S rRNA sequence identity with cultured Desulfotomaculum type strains and thus probably represent novel species within this genus (Table 1). Given that thermophiles cannot, and presumably could not, grow in cold Aarhus Bay sediments, this large diversity must be due to passive influx of SRB endospores from external sources rather than growth and diversification of thermophilic Desulfotomaculum at this location. Of particular interest are phylotypes D, F and G that were also discovered in the Arctic sediment of Smeerenburgfjorden, Svalbard (Table 1; Hubert et al., 2009, 2010). Classification of phylotypes based on a 16S rRNA sequence identity threshold of 99% cannot exclude that sequences from different species are regarded as the same phylotype (Tourova et al., 2001; Stackebrandt and Ebers, 2006). However, the detection of endospores with both highly similar 16S rRNA and dsrAB gene sequences in Svalbard and Aarhus Bay sediments shows that very closely related thermophilic Desulfotomaculum species are found in these two geographically distant locations (Figures 2 and 3). This supports the hypothesis that these bacteria originate from a common warm, anoxic source environment. Such an environment may be theoretically represented by a single-source habitat with a strong efflux that has resulted in an yearly deposition of ca. 108 thermophilic SRB per m2 in the surface sediments of Svalbard and ca. 107 per m2 in Aarhus Bay for the last hundred years (Hubert et al., 2009; Figure 4a), despite more than 3000 km separating these two locations.
Although a common source may supply thermophiles to distant locations, the detection of several thermophilic SRB following incubation of Aarhus Bay sediment at different temperatures suggests that multiple source habitats could contribute to the supply of thermophilic endospores to Aarhus Bay. The diversity of thermophilic Desulfotomaculum spp. reported here may arise from a combination of different source environments in distinct geographic locations. Alternatively, different source environments may be at the same geographic location but associated with a habitat or habitats where temperature varies between at least 40 and 70 °C. These temperature gradients are common in geothermal aquifers, for example, the Great Artesian Basin, Australia, from where D. australicum was isolated (Love et al., 1993). D. australicum is the closest cultivated relative of phylotype AB, the only phylotype identified in our incubations at 69 °C (99% sequence identity, Table 1). Similarly, advection of geofluids (for example, hydrocarbons, mud volcanoes) from warm deep sediments could provide a transport pathway from warm depths up through the seabed to cold surface environments (Judd and Hovland, 2007; Hubert and Judd, 2010). Alternatively, magma-driven hydrothermal circulation in oceanic crust environments, such as ridge flanks and seamount-associated systems, maintains parts of these anaerobic environments at temperatures that favor thermophilic microbial communities, and seawater circulation through the ocean crust may facilitate the transport of microorganisms in and out of these environments (Schrenk et al., 2010). Hydrothermal circulation with diffuse fluid venting, in particular at the Lost City hydrothermal field off the Mid Atlantic Ridge, is also subjected to such temperature gradients and provides conditions that could host abundant thermophilic microbial communities (Früh-Green et al., 2003). A large diversity of endospore-forming Firmicutes, including Desulfotomaculum-related 16S rRNA gene sequences, have been found in carbonate chimneys at this site (Brazelton et al., 2006) and are closely related to phylotype F (Hubert et al., 2010).
Anthropogenic sources may also be relevant for explaining the occurrence of thermophilic SRB in Aarhus Bay sediment. Local coal-fired power plants and industrial wastewater treatment facilities around the city of Aarhus were established in recent decades, and were previously proposed as the source of thermophilic endospore-forming SRB to Aarhus Bay sediment (Isaksen et al., 1994). Another possible source may be North Sea oil production systems that include different high temperature components and harbor diverse thermophilic SRB (see Stetter et al., 1993; Beeder et al., 1994; Nilsen et al., 1996; Dahle et al., 2008). The Dan and Halfdan oil fields located ca. 200 km west of Denmark are characterized by temperatures from ca. 40 to 80 °C and host a diverse microbiota (Gittel et al., 2009). Desulfotomaculum 16S rRNA gene sequences originating from these oil fields are the closest known relatives of phylotypes R and S (95%; Figure 2), which became highly enriched in substrate-amended 50 °C incubations. Indeed, phylotypes R and S were detected in surface sediment but not in deeper layers (Figure 4b), which is suggestive of a recent, possibly anthropogenic source for these taxa, assuming a similar death rate of different Desulfotomaculum spp. endospores. However, the detection of other thermophilic Desulfotomaculum spp. at 650 cm depth, which corresponds to ca. 4500 years of sedimentation (see below), highlights that the presence of thermophiles in this cold sediment is not exclusively due to human industry. As discussed above, outflow from natural, non-ephemeral environments must contribute to the long-term influx of thermophilic endospores into Aarhus Bay sediment. Whatever the primary origin of these thermophiles is, the presence of thermophilic SRB in the water column of Aarhus Bay (Figure 5) confirms that they are being dispersed in this region by the water currents, and remain viable despite the cold temperatures and oxic surroundings. The detection of thermophilic SRB in coarse GF filters and not in the downstream filters of smaller pore size suggest that these thermophiles are transported not as single cells, but attached to larger particles or aggregates.
Long-term deposition and survival of thermophilic SRB endospores in Aarhus Bay sediments
The abundance of thermophilic SRB endospores as detected by our T-MPN method decreases exponentially from ca. 104 cells per cm3 in surface sediment to 1 cell per cm3 at 650 cm depth (Figure 4a). Migration of cells through sediment layers is not expected, as endospores are not motile and the sediment is composed by fine-grained clay with no evidence of advective fluid flow. Therefore, the profile of thermophilic SRB endospore abundance in Aarhus Bay can be related to sediment age. This profile could result from an increasing influx of thermophilic endospores to Aarhus Bay sediments over the past 4500 years, perhaps due to an accelerating efflux of SRB from a given source or a growing number of sources supplying thermophilic endospores to this location. However, the sedimentation rate at this site has not changed considerably in the last 4500 years (JB Jensen, unpublished); furthermore, there is no evidence of changes in the ocean currents that might be associated with a gradual increase in the influx of thermophilic SRB endospores. It therefore seems unlikely that an increased supply of endospores alone can be responsible for the change of four orders of magnitude in the observed abundance profile in Aarhus Bay (Figure 4a). Therefore, we propose an alternative hypothesis that endospore death on millennial time scales explains the observed decrease in endospore concentrations.
The decrease in thermophilic SRB endospore numbers with depth shown in Figure 4a is three orders of magnitude greater than the decrease in cell counts over the same 650 cm depth interval (4 × 109 to 1 × 108 cells per cm3, personal communication, MA Lever; Langerhuus et al. submitted). Rejuvenation of thermophilic endospore populations through germination, repair and sporulation cycles (Yung et al., 2007) is not expected at the cold in situ temperatures of 0 to 15 °C (Thamdrup et al., 1994), hence their death and decay-dependent decrease in abundance is not counterbalanced by growth and multiplication. Therefore, unlike vegetative cells, the decrease of thermophilic endospores does not relate to factors such as electron donor and electron acceptor availability. Given that the T-MPN method only counts viable cells that are induced to grow, the observed depth profile may be interpreted as a decrease in the overall viability of endospore populations over time, for example, as a result of cellular damage (Johnson et al., 2007). Alternatively, populations of endospores may become reduced following stochastic germination events (Paidhungat and Setlow, 2000; Epstein, 2009; Buerger et al., 2012a, 2012b), which would in this case lead to death of these thermophilic organisms due to the unfavorably low temperature in the Aarhus Bay sediment environment.
The exponential decline in the abundance of viable thermophilic SRB endospores (Figure 4a) and the assumed lack of reproduction and sporulation in situ allow the estimation of a half-life for this dormant community. Using a sediment age model based on C-14 dating of shells from a replicate sediment core (JB Jensen, unpublished), endospore abundance (N) can be related to sediment age (t) according to N(t)=8620 × e−0.002t (Figure 4a). This results in a half-life of thermophilic SRB endospores in Aarhus Bay sediment between 250 and 440 years (95% confidence interval). Mukamolova et al. (2003) speculated that the survival half-life of a microbial cell under optimal conditions, in the absence of nutrient influx, turnover and regeneration, is between 10 and 100 years. The longer half-life calculated in our study may be explained by the higher stress resistance of endospores and lower maintenance requirements as compared with the vegetative cells. Rothfuss et al. (1997) calculated death rates in a study of deep lake sediments dominated by endospore populations and their data result in similar population half-lives, of 525 years for aerobically grown microorganisms and 575 years for anaerobically grown microorganisms. This longer half-life than what we estimate here (Figure 4a) may be explained by the fact that the endospores studied by Rothfuss et al. (1997) are mesophiles and thus can potentially cycle through reproduction and sporulation in situ. To our knowledge, these are the only attempts so far to estimate the half-life of dormant microbial populations in natural environments.
Conclusion
Thermophilic endospores in cold sediments have been proposed as models for investigating the dispersal of marine microorganisms, as demonstrated for sulfate-reducing Desulfotomaculum spp. supplied to Aarhus Bay. The presence of a Desulfotomaculum species (phylotype F) throughout a 650-cm sediment core indicates that this thermophilic endospore population has been deposited in Aarhus Bay sediments for thousands of years. The presence of this phylotype, as well as two others (Desulfotomaculum phylotypes D and G), in the surface sediments of both Aarhus Bay and a Svalbard fjord suggests that these results are explained by global scale passive dispersal and that these particular taxa might share common source environment(s), whereas on the other hand the molecular and physiological diversity we have uncovered (23 phylotypes in Aarhus Bay alone) indicate that multiple source environments may supply a single location. The presence of approximately 104 thermophilic SRB endospores per cm3 of surface sediment in both Aarhus Bay and Svalbard sediments indicates a large flux of these cells, and their detection in the water column confirms that this passive dispersal is an ongoing process involving seawater circulation. As our analyses have only targeted thermophilic SRB endospores able to germinate under specific incubation conditions, both the diversity and the abundance of these endospore populations in Aarhus Bay sediments may be larger than what has been uncovered by this study.
References
Bartholomew JW, Paik G . (1966). Isolation and identification of obligate thermophilic sporeforming bacilli from ocean basin cores. J Bacteriol 92: 635–638.
Beeder J, Nilsen RK, Rosnes JT, Torsvik T, Lien T . (1994). Archaeoglobus fulgidus isolated from hot North Sea oil field waters. Appl Environ Microbiol 60: 1227–1231.
Brazelton WJ, Schrenk MO, Kelley DS, Baross JA . (2006). Methane- and sulfur-metabolizing microbial communities dominate the Lost City hydrothermal field ecosystem. Appl Environ Microbiol 72: 6257–6270.
Buerger S, Spoering A, Gavrish E, Leslin C, Ling L, Epstein SS . (2012a). Microbial scout hypothesis, stochastic exit from dormancy, and the nature of slow growers. Appl Environ Microbiol 78: 3221–3228.
Buerger S, Spoering A, Gavrish E, Leslin C, Ling L, Epstein SS . (2012b). Microbial scout hypothesis and microbial discovery. Appl Environ Microbiol 78: 3229–3233.
Dahle H, Garshol F, Madsen M, Birkeland NK . (2008). Microbial community structure analysis of produced water from a high-temperature North Sea oil-field. A Van Leeuw 93: 37–49.
Epstein SS . (2009). Microbial awakenings. Nature 457: 1083–1083.
Falcón LI, Noguez AM, Espinosa-Asuar L, Eguiarte LE, Souza V . (2008). Evidence of biogeography in surface ocean bacterioplankton assemblages. Marine Genomics 1: 55–61.
Früh-Green GL, Kelley DS, Bernasconi SM, Karson JA, Ludwig KA, Butterfield DA et al (2003). 30000 years of hydrothermal activity at the Lost City vent field. Science 301: 495–498.
Gittel A, Sørensen KB, Skovhus TL, Ingvorsen K, Schramm A . (2009). Prokaryotic community structure and sulfate reducer activity in water from high-temperature oil reservoirs with and without nitrate treatment. Appl Environ Microbiol 75: 7086–7096.
Green J, Bohannan BJM . (2006). Spatial scaling of microbial biodiversity. Trends Ecol Evolution 21: 501–507.
Hansen JW, Thamdrup B, Jørgensen BB . (2000). Anoxic incubation of sediment in gas-tight plastic bags: a method for biogeochemical process studies. Mar Ecol Prog Ser 208: 273–282.
Hessler RR, Jumars PA . (1974). Abyssal community analysis from replicate box cores in the central North Pacific. Deep Sea Res Oceanogr Abstr 21: 185–209.
Hubert C, Loy A, Nickel M, Arnosti C, Baranyi C, Brüchert V et al (2009). A constant flux of diverse thermophilic bacteria into the cold arctic seabed. Science 325: 1541–1544.
Hubert C, Arnosti C, Brüchert V, Loy A, Vandieken V, Jørgensen BB . (2010). Thermophilic anaerobes in Arctic marine sediments induced to mineralize complex organic matter at high temperature. Environ Microbiol 12: 1089–1104.
Hubert C, Judd A . (2010). Using microorganisms as prospecting agents in oil and gas exploration. In: Timmis KN, (ed). Handbook of Hydrocarbon and Lipid Microbiology. Springer-Verlag: Berlin, Heidelberg.
Ionescu D, Hindiyeh M, Malkawi H, Oren A . (2010). Biogeography of thermophilic cyanobacteria: insights from the Zerka Ma'in hot springs (Jordan). FEMS Microbiol Ecol 72: 103–113.
Isaksen MF, Bak F, Jørgensen BB . (1994). Thermophilic sulfate-reducing bacteria in cold marine sediment. FEMS Microbiol Ecol 14: 1–8.
Johnson SS, Hebsgaard MB, Christensen TR, Mastepanov M, Nielsen R, Munch K et al (2007). Ancient bacteria show evidence of DNA repair. Proc Natl Acad Sci USA 104: 14401–14405.
Judd A, Hovland M . (2007). Seabed fluid flow. Impact on Geology, Biology and the Marine Environment. Cambridge University Press: Cambridge, UK.
Kallmeyer J, Ferdelman TG, Weber A, Fossing H, Jørgensen BB . (2004). A cold chromium distillation procedure for radiolabeled sulfide applied to sulfate reduction measurements. Limnol Oceanogr Methods 2: 171–180.
Kjeldsen KU, Loy A, Jakobsen TF, Thomsen TR, Wagner M, Ingvorsen K . (2007). Diversity of sulfate-reducing bacteria from an extreme hypersaline sediment, Great Salt Lake (Utah). FEMS Microbiol Ecol 60: 287–298.
Klepac-Ceraj V, Bahr M, Crump BC, Teske AP, Hobbie JE, Polz MF . (2004). High overall diversity and dominance of microdiverse relationships in salt marsh sulphate-reducing bacteria. Environ Microbiol 6: 686–698.
Lane DJ . (1991). 16/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M, (eds) Nucleic Acid Techniques in Bacterial Systematics. Wiley: New York, pp 115–175.
Langerhuus AT, Røy H, Lever MA, Morono Y, Inagaki F, Jørgensen BB et al. Endospore abundance and D:L-amino acid modeling of bacterial turnover in holocene marine sediment (Aarhus Bay). Submitted for publication.
Legendre P, Legendre L . (1998). Numerical Ecology, 2nd English edn. Elsevier Science BV. Amsterdam, The Netherlands.
Leloup J, Fossing H, Kohls K, Holmkvist L, Borowski C, Jørgensen BB . (2009). Sulfate-reducing bacteria in marine sediment (Aarhus Bay, Denmark): abundance and diversity related to geochemical zonation. Environ Microbiol 11: 1278–1291.
Love CA, Patel BKC, Nichols PD, Stackebrandt E . (1993). Desulfotomaculum australicum, sp. nov., a thermophilic sulfate-reducing bacterium isolated from the Great Artesian Basin of Australia. Syst Appl Microbiol 16: 244–251.
Loy A, Lehner A, Lee N, Adamczyk J, Meier H, Ernst J et al (2002). Oligonucleotide microarray for 16S rRNA gene-based detection of all recognized lineages of sulfate-reducing prokaryotes in the environment. Appl Environ Microbiol 68: 5064–5081.
Loy A, Kusel K, Lehner A, Drake HL, Wagner M . (2004). Microarray and functional gene analyses of sulfate-reducing prokaryotes in low-sulfate, acidic fens reveal cooccurrence of recognized genera and novel lineages. Appl Environ Microbiol 70: 6998–7009.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: 1363–1371.
Martiny JBH, Bohannan BJM, Brown JH, Colwell RK, Fuhrman JA, Green JL et al (2006). Microbial biogeography: putting microorganisms on the map. Nat Rev Microbiol 4: 102–112.
Messing J . (1983). New M13 vectors for cloning. Methods Enzymol 101: 20–78.
Mukamolova GV, Kaprelyants AS, Kell DB, Young M . (2003). Adoption of the transiently non-culturable state - a bacterial survival strategy? Advances in Microbial Physiology Vol 47. Academic Press Ltd: London, pp 65–129.
Muyzer G, de Waal EC, Uitterlinden AG . (1993). Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59: 695–700.
Nemergut DR, Costello EK, Hamady M, Lozupone C, Jiang L, Schmidt SK et al (2011). Global patterns in the biogeography of bacterial taxa. Environ Microbiol 13: 135–144.
Nilsen RK, Beeder J, Thorstenson T, Torsvik T . (1996). Distribution of thermophilic marine sulfate reducers in North Sea oil field waters and oil reservoirs. Appl Environ Microbiol 62: 1793–1798.
Oksanen J, Blanchet FG, Kindt R, Legendre P, O’Hara RB, Simpson GL et al (2010). Vegan: Community Ecology Package. 2010; R package version 1: 17–3.
Paidhungat M, Setlow P . (2000). Role of Ger proteins in nutrient and nonnutrient triggering of spore germination in Bacillus subtilis. J Bacteriol 182: 2513–2519.
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J et al (2007). SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35: 7188–7196.
Ramette A, Tiedje JM . (2007). Biogeography: an emerging cornerstone for understanding prokaryotic diversity, ecology, and evolution. Microbial Ecol 53: 197–207.
Rothfuss F, Bender M, Conrad R . (1997). Survival and activity of bacteria in a deep, aged lake sediment (Lake Constance). Microbial Ecol 33: 69–77.
Schauer R, Bienhold C, Ramette A, Harder J . (2010). Bacterial diversity and biogeography in deep-sea surface sediments of the South Atlantic Ocean. ISME J 4: 159–170.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB et al (2009). Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75: 7537–7541.
Schrenk MO, Huber JA, Edwards KJ . (2010). Microbial Provinces in the Subseafloor. Annu Rev Marine Sci 2: 279–304.
Stackebrandt E, Ebers J . (2006). Taxonomic parameters revisited: tarnished gold standards. Microbiol Today 33: 152–155.
Stetter KO, Huber R, Blochl E, Kurr M, Eden RD, Fielder M et al (1993). Hyperthermophilic archaea are thriving in deep North Sea and Alaskan oil reservoirs. Nature 365: 743–745.
Stubner S, Meuser K . (2000). Detection of Desulfotomaculum in an Italian rice paddy soil by 16S ribosomal nucleic acid analyses. FEMS Microbiol Ecol 34: 73–80.
Thamdrup B, Fossing H, Jorgensen B . (1994). Manganese, iron and sulfur cycling in a coastal marine sediment, Aarhus Bay, Denmark. Geochim Cosmochim Ac 58: 5115–5129.
Thamdrup B, Hansen JW, Jørgensen BB . (1998). Temperature dependence of aerobic respiration in a coastal sediment. FEMS Microbiol Ecol 25: 189–200.
Thomsen TR, Finster KW, Ramsing NB . (2001). Biogeochemical and Molecular Signatures of Anaerobic Methane Oxidation in a Marine Sediment. Appl Environ Microbiol 67: 1646–1656.
Tourova TP, Kuznetzov BB, Novikova EV, Poltaraus AB, Nazina TN . (2001). Heterogeneity of the Nucleotide Sequences of the 16S rRNA Genes of the Type Strain of Desulfotomaculum kuznetsovii. Microbiology 70: 678–684.
Vandieken V, Knoblauch C, Jørgensen BB . (2006). Desulfotomaculum arcticum sp. nov., a novel spore-forming, moderately thermophilic, sulfate-reducing bacterium isolated from a permanently cold fjord sediment of Svalbard. Int J Syst Evol Microbiol 56: 687–690.
Vester F, Ingvorsen K . (1998). Improved most-probable-number method to detect sulfate-reducing bacteria with natural media and a radiotracer. Appl Environ Microbiol 64: 1700–1707.
Webster G, Sass H, Cragg BA, Gorra R, Knab NJ, Green CJ et al (2011). Enrichment and cultivation of prokaryotes associated with the sulphate–methane transition zone of diffusion-controlled sediments of Aarhus Bay, Denmark, under heterotrophic conditions. FEMS Microbiol Ecol 77: 248–263.
Yung PT, Shafaat HS, Connon SA, Ponce A . (2007). Quantification of viable endospores from a Greenland ice core. FEMS Microbiol Ecol 59: 300–306.
Zverlov V, Klein M, Lucker S, Friedrich MW, Kellermann J, Stahl DA et al (2005). Lateral gene transfer of dissimilatory (bi)sulfite reductase revisited. J Bacteriol 187: 2203–2208.
Zengler K (ed). Accessing Uncultivated Microorganisms: From the Environment to Organisms and Genomes and Back. ASM Press: Washington, DC, 2008, pp 95–115.
Acknowledgements
We thank the crew aboard R/V Tyra and Susanne A for assistance in sampling campaigns, and Jørn Bo Jensen, Mark A Lever, Flemming C Mønsted, Kristoffer Piil, Hans Røy and Cecilia Wentrup for their valuable input on data analysis. We are grateful to Mohamoud Hashi, Britta Poulsen and Trine B. Søgaard for their technical assistance. This work was funded by the Danish National Research Foundation, the Max Planck Society, the Austrian Science Fund (P20185-B17) and an EU Marie Curie fellowship to CH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
de Rezende, J., Kjeldsen, K., Hubert, C. et al. Dispersal of thermophilic Desulfotomaculum endospores into Baltic Sea sediments over thousands of years. ISME J 7, 72–84 (2013). https://doi.org/10.1038/ismej.2012.83
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2012.83
Keywords
This article is cited by
-
Extremophilic microbial metabolism and radioactive waste disposal
Extremophiles (2023)
-
Assessing the Diversity of Benthic Sulfate-Reducing Microorganisms in Northwestern Gulf of Mexico by Illumina Sequencing of dsrB Gene
Microbial Ecology (2021)
-
The sponge microbiome within the greater coral reef microbial metacommunity
Nature Communications (2019)
-
Microbial Community Composition and Putative Biogeochemical Functions in the Sediment and Water of Tropical Granite Quarry Lakes
Microbial Ecology (2019)
-
Beyond the tip of the iceberg; a new view of the diversity of sulfite- and sulfate-reducing microorganisms
The ISME Journal (2018)