Abstract
Microbial metabolism in aromatic-contaminated environments has important ecological implications, and obtaining a complete understanding of this process remains a relevant goal. To understand the roles of biodiversity and aromatic-mediated genetic and metabolic rearrangements, we conducted ‘OMIC’ investigations in an anthropogenically influenced and polyaromatic hydrocarbon (PAH)-contaminated soil with (Nbs) or without (N) bio-stimulation with calcium ammonia nitrate, NH4NO3 and KH2PO4 and the commercial surfactant Iveysol, plus two naphthalene-enriched communities derived from both soils (CN2 and CN1, respectively). Using a metagenomic approach, a total of 52, 53, 14 and 12 distinct species (according to operational phylogenetic units (OPU) in our work equivalent to taxonomic species) were identified in the N, Nbs, CN1 and CN2 communities, respectively. Approximately 10 out of 95 distinct species and 238 out of 3293 clusters of orthologous groups (COGs) protein families identified were clearly stimulated under the assayed conditions, whereas only two species and 1465 COGs conformed to the common set in all of the mesocosms. Results indicated distinct biodegradation capabilities for the utilisation of potential growth-supporting aromatics, which results in bio-stimulated communities being extremely fit to naphthalene utilisation and non-stimulated communities exhibiting a greater metabolic window than previously predicted. On the basis of comparing protein expression profiles and metagenome data sets, inter-alia interactions among members were hypothesised. The utilisation of curated databases is discussed and used for first time to reconstruct ‘presumptive’ degradation networks for complex microbial communities.
Similar content being viewed by others
Introduction
Polyaromatic hydrocarbons (PAHs) are widely distributed in the environment owing to their abundance in crude oil and their widespread use in chemical manufacturing (Kästner, 2000). PAHs are pollutants of great concern owing to their toxicity, mutagenicity and carcinogenicity. A number of microorganisms, via the action of Rieske non-haem iron oxygenases, have the ability to grow on PAHs as a sole carbon and energy source (Beil et al., 1998; Roling et al., 2003; Witzig et al., 2006; Peng et al., 2008).
The majority of efforts aimed at understanding microbial responses to aromatic compounds have been focused on genomic (Jiménez et al., 2002; Nogales et al., 2008; Puchałka et al., 2008), transcriptomic (Domínguez-Cuevas et al., 2006; Yuste et al., 2006) and proteomic (Santos et al., 2004; Segura et al., 2005; Kim et al., 2006; Kurbatov et al., 2006; Tomás-Gallardo et al., 2006) analyses conducted in pure cultures. Among PAH-(including naphthalene, fluorene, phenanthrene, pyrene and dibenzofuran, to cite some) mineralising strains, a number of bacteria have received special attention. Those include members of the common genera of Pseudomonas, Sphingomonas, Cycloclasticus, Burkholderia, Rhodococcus, Polaromonas, some novel genera of Neptunomonas and Janibacter, some thermophilic bacteria of Nocardia and Bacillus, some anoxic bacteria of Deltaproteobacteria and Alcaligenes, and high molecular weight PAH-degrading bacteria of Mycobacterium, Stenotrophomonas and Pasteurella; a list with known pure bacterial cultures capable of degrading PAHs can be seen in a recent review by Lu et al. (2011).
However, a large number of culture-independent techniques have shown that pollutant-degrading organisms enriched in the laboratory often do not have an important role in the in situ biodegradation of pollutants and that the diversity of pollutant-degrading organisms in polluted environmental samples is much greater than the diversity determined from cultivation (Abulencia et al., 2006; Liu et al., 2009; Boronin and Kosheleva, 2010; Yagi et al., 2010; Liang et al., 2011). Under this scenario, it is relevant to use molecular microbial tools to define key catabolic players at contaminated sites to predict pollutant degradation networks in the environment and to suggest methods for rational intervention associated with the implementation of bioremediation (Vilchez-Vargas et al., 2010). However, the number of integrative ‘omic’ investigations that have been carried out in PAH-associated microbial communities is limited (Kweon et al., 2007; Powell et al., 2008; Selesi et al., 2010), because of the incomplete genomic information and curated databases available (Pérez-Pantoja et al., 2009, 2012).
Taking all of the above information into consideration, we performed a thorough and holistic (or ecosystems biology approach) phylogenetic, functional and proteomic analysis of the key players in a sample of strongly anthropogenically influenced, PAH-contaminated soil (N) and a naphthalene-enriched community derived from this soil (CN1). This study was carried out using also samples of the same soil bio-stimulated with calcium ammonia nitrate, NH4NO3 and KH2PO4, and the commercial surfactant Iveysol (Ivey International Inc., Campbell River, BC, Canada) (herein referred as Nbs), and the naphthalene-enriched community derived from it (CN2). In all cases, the biodegradation networks of the respective whole communities were reconstructed. This study clarified the genomic and proteomic basis for the purpose of understanding microbial biodiversity, ecology and function in response to both PAHs (represented by naphthalene) and bio-stimulation. It should be noted that our metaproteomic investigations were restricted to naphthalene-enriched communities because of their lower complexity as compared with the soil samples as well as the larger assembled sequences obtained through direct pyro-sequencing for both samples.
Materials and methods
General methods and ‘OMIC’ data analysis and processing
Full descriptions of the materials and methods used for soil characterisation; hydrocarbon analysis; DNA extraction; construction of 16 S RNA gene clone libraries, sequencing and phylogenetic analysis; denaturing gradient gel electrophoresis analysis; metagenomic setup and sequencing, assembly and gene prediction; metaproteomic setup and protein extraction, separation and identification and data processing are available in the Supplementary Materials and methods.
Soil sample collection and preparation of bio-stimulated soil and enrichment cultures
Soil samples (herein referred to as N) were obtained from a parcel on the northern Iberian Peninsula contiguous to a chemical plant (Lugones, Oviedo, Spain; 40°18′33′′N, 3°39′31′′W, at an altitude of 300 m). The chemical plant (Supplementary Figure S1) was used for several decades for the production of naphthalene, phenols and other compounds from coal tar processing as well as for manufacturing resins. A considerable amount of other chemical products (pesticides, solvents, etc.) was stored, although probably not manufactured, in the plant. In 1989 the plant was closed and then used for years to store chemical waste, particularly polychlorinated biphenyls, coolants and other unspecified products. In 2006 most of the buildings were demolished and the characterisation of soil contamination started. Thus, a significant amount of the PAH detected were conceivably formed by the enriched bacterial populations present in the soil through a natural attenuation effect. The top soil was sampled at a depth of 0–30 cm on February 2007 (soil temperature 18 °C). Three replicates (1 kg each) were collected within a 1 m distance, and the samples were kept in open plastic bags in the dark at 4 °C. Vegetation and other non-soil materials, including cobbles, were removed prior to homogenising the samples. Immediately after acquiring the soil samples, they were sieved (2-mm mesh size), followed by mixing of representative subsamples of the triplicates samples, and 100 g aliquots of sample N were used for chemical analysis. In addition, 10 g aliquots were stored at −20 °C in sterile flasks for DNA- and proteome-based analyses.
Bioremediation was performed over approximately 900 m3 of the contaminated soil (N), where 9 tons of dehydrated calcium ammonia nitrate and 3 tons of a mixture of dehydrated NH4NO3 and KH2PO4 combined with 4500 l of the commercial surfactant Iveysol were applied to the homogenised soil. The C:N:P ratio that was applied was 100:10: 1, as recommended for bioremediation purposes (Gallego et al., 2007a). The treatment was performed over 231 days, and the biopile was routinely watered and tilled to maintain humidity (15–20%) and adequate oxygenation. Samples of the top bio-stimulated soil, herein named Nbs, were taken and processed using the same protocols as described above for the original soil.
Enrichment cultures were performed in Bushnell Haas (Sigma Chemical Co., St Louis, MO, USA) mineral medium that contained naphthalene at a concentration of 0.1% (w/v) as the sole carbon source as described previously (Gallego et al., 2007b). The composition of the medium was as follows: MgSO4·H2O (0.20 g l−1), CaCl2·2 H2O (0.02 g l−1), KH2PO4 (1.0 g l−1), K2HPO4 (1.0 g l−1), NH4NO3 (1.0 g l−1) and FeCl3 (0.05 g l−1). Two different inocula were used. The CN1 enrichment culture was obtained by inoculating 1 g of the polluted soil (N) into a flask containing 100 ml of the culture medium; the CN2 culture was inoculated with 1 g of the same soil that had been subjected to a massive bioremediation (bio-stimulation) process (Nbs). The enrichment cultures were incubated at 30 °C and 250 r.p.m., in which 0.1% (v/v) of the culture was transferred to fresh medium each week.
Results and discussion
Sample characteristics
An agronomic analysis of the soil N revealed a loamy clay soil with a pH of 8.2 and a conductivity of 0.13 dS m−1 clearly containing low amounts of the typically predominant ions (calcium, magnesium, potassium and sodium); the detected natural organic matter, nitrogen and phosphorus levels in the soil (Supplementary Table S1) are characteristic of infertile soils. Gas chromatography–mass spectrometry (GC–MS) was used to quantify the levels of the 16 EPA (Environmental Protection Agency) priority PAHs present in the soil (Supplementary Figure S2). Totally, the soil exhibited an average concentration of 805 mg PAH per kg (Table 1), which is in the range or slightly lower to the level found in previously reported PAH contaminated soils (that is, 1667 mg kg−1 in Ní Chadhain et al. (2006); 589 mg kg−1 in Richardson et al. (2012); 335–8645 mg kg−1 in Thavamani et al. (2012); 3000 mg kg−1 in Lors et al. (2012)). The bio-stimulated soil exhibited a total concentration of 221.6 mg kg−1 (average) of the 16 EPA priority PAHs and the quantified compounds are listed in Table 1. The degradation rate of naphthalene in N was too low and was difficult to be established owing to the long time transcurred in the presence of the contaminants; however, once submitted to bio-stimulation, we calculated the degradation rate of naphthalene in soil from time 0 to 231 days: 1.818 mg kg−1 soil per day.
Denaturing gradient gel electrophoresis was used to survey the development of the microbial communities in the enrichment cultures and to deduce the time point at which a stable community was obtained. We observed that the denaturing gradient gel electrophoresis profiles for CN1 enrichment cultures changed between the 45 and 60 transfers (Supplementary Figure S3); changes in the profiles were also observed for CN2 after 35 and 60 transfers. These changes were subsequently maintained and samples subjected to at least 60 transfers were herein used for further investigations. The major bands were excised and sequenced, and their affiliation revealed the presence of microorganisms identified as members of the genera Achromobacter, Flavobacterium and Acidovorax in both enrichments, and Pseudomonas, Microbacterium, Lysobacter and to an endosymbiont of Acanthomaeba in CN2. Results indicated that consortia CN1 and CN2 were able to degrade 97% naphthalene at an initial concentration of 1000 p.p.m. within 72 and 80 h, respectively.
Bacterial diversity and composition blueprints
DNA isolated from each investigated microbial community was employed for a PCR-based 16S recombinant DNA (rDNA) gene diversity survey of the community structures in the samples. For this purpose, clone libraries were constructed as described in the Supplementary Materials and methods, and the clones were fully sequenced to obtain the almost complete cloned 16S rDNA gene sequence. Additionally, and because of the low yields in the clone libraries of Nbs, we used the 16S rDNA gene partial sequences obtained in the metagenome survey. For this purpose, we used only those sequences that had a length >600 nucleotides. All shorter sequences were discarded. A total of 670 sequences (that is, N: 212; Nbs: 261 (86 clones+175 454 partial sequences); CN1: 90; CN2: 107) were obtained, and analysed. The overall phylogenetic composition in the libraries is shown in Figures 1, 2, 3, in which all operational phylogenetic units (OPUs) affiliated with twelve phyla of the domain Bacteria, namely, Proteobacteria followed to much lower extend by Bacteroidetes, Verrucomicrobia, Firmicutes, Actinobacteria, Chloroflexi, Cyanobacteria, Planctomycetes, Spirochaetes and the candidatus phyla OP8, TM7 and WCHB1 (Figure 4).
A neighbour-joining tree of the proteobacterial 16S rDNA gene sequences from the gene sequences originated from N, Nbs, CN1 and CN2 communities. For the reconstruction, only the almost complete sequences of the clones from the four samples had been used. Besides, partial 454 sequences of >600 nucleotides were inserted in the tree using the Parsimony tool implemented in ARB. The number of sequences comprised in each identity cluster (OPU) is specified for each sample. The colour code for the 16S rDNA sequences is as follows: N (green), Nbs (purple), CN1 (blue), CN2 (red). Whether an OPU contains sequences of more than one sample, the OPU denomination is written in black followed by the code colour and sequence number of each studied sample.
A neighbour-joining tree of non-proteobacterial 16S rDNA gene sequences from the clone libraries established from N, Nbs, CN1 and CN2 community DNA. Layout, reconstruction and colour codes used are the same as in Figure 1.
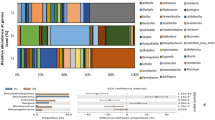
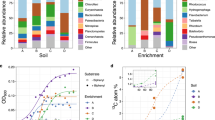
Comparison of bacterial phylotypes (cut-off of >97% sequence identity) based on the 16 S rDNA sequences extracted from N, Nbs, CN1 and CN2 community DNA. The percentages of bacterial phylogenetic lineages detected were based on OPUs. (a) Percentage of 16S rDNA sequences distributed by phyla. (b) Contribution of dominant groups among Gammaproteobacteria. (c) Relative distribution of major bacterial phylotypes (based on 16S rDNA gene sequences (OPUs)) for soil communities CN1 and CN2 at the genus level.
The clones were classified into 124 bacterial operational phylogenic units (OPUs) (López-López et al., 2010) (Table 2 and Supplementary Figure S2). Higher values for Shannon–Wiener’s and Good’s coverage indexes indicated that communities N and Nbs were much more complex (51 OPUs and 53 OPUs were detected, respectively) than the CN1 and CN2 (13 OPUs and 12 OPUs, respectively) communities, which were dominated by few microbial species or strains and exhibited a rather simple structure (Figures 1, 2, 3). It should be recalled that each of the distinct OPUs detected was identified as a putative single species owing to the high sequence identity (Yarza et al., 2008). However, it should also be noted that the sequencing survey covered over 75% of the expected OPU diversity in all cases as it can be deducted from the high Good’s coverage in all samples (Table 2), and the rarefraction curves indicating closeness to saturation (Supplementary Figure S4). A large proportion of the obtained sequences affiliated with known families and clustered with branches represented by cultured microorganisms that have previously been found in contaminated environments (Figures 1, 2, 3 and Supplementary Figures S5 and S6), and thus, they do not appear to be specific to the soil and the enrichment samples that were investigated here. Whatever the case, despite soil characteristics and specific pollutants composition may differ, the biodiversity found in the original soils (N and Nbs) (Shannon indexes of 2.81 and 3.32, respectively) are in the range of that found in other PAH-contaminated soils. For example, Shannon indexes of 1.74–2.78 (Vivas et al., 2008), 2.38–2.96 (Thavamani et al., 2012) and 4.87–5.01 (Martin et al., 2012) were observed in soils that had been historically contaminated with PAHs.
The dominant group among the Gammaproteobacteria consisted of members of the genus Pseudomonas (Figures 2 and 4). The N sequences were distributed rather evenly within this genus, with approximately 40% of the total gammaproteobacterial sequences branching in the P. fluorescens, P. frederiksbergensis and P. amygdale lineages, followed by those affiliated with the Pseudoxanthomonas spp. (20%) and P. putida (14%) lineages (Supplementary Table S2). A similar even distribution was observed in the sample Nbs, being as well the members of Pseudomonadaceae the most abundant. However, in this case, the Pseudomonas species composition shifted towards the major representatives P. tuomourensis, P. pertucinogena, P. stutzeri genomovar 1, and yet undescribed Pseudomonas species. These results clearly contrast with the distribution of the CN2 sequences, where just representatives of P. stutzeri (46%) lineages (Supplementary Table S2) were observed in accordance with this being one of the major represented lineages in Nbs. Finally, in CN1, no representation of the Pseudomonas genus was found, but six sequences representing Pseudoxanthomonas were identified (6% of the total) (Figures 1 and 4 and Supplementary Table S2); this is in agreement with previous observations made in PAH microcosm experiments that suggest that, although, Pseudomonas spp. can be easily enriched and use a variety of organic substrates, they may be repressed by other Proteobacteria (Rhee et al., 2004).
The alphaproteobacterial sequences detected in N were widely distributed among Rhizobium, Devosia, Novosphingobium, Sphingomonas and Brevundimonas, which were the most represented genera (Figure 1 and Supplementary Table S2). On the other hand, members of this lineage were barely represented in Nbs with just seven sequences, being Brevundimonas the most represented. None of these genera were detected in any of the enrichment cultures (Figure 1). In contrast, most of the alphaproteobacterial clones in CN1 (73%) were affiliated with Azospirillum species, that is, Azospirillum oryzae, whereas only three alphaproteobacterial sequences (out of a total of 108) were found in CN2, and only one was affiliated with Azospirillum (Figure 5 and Supplementary Table S2).
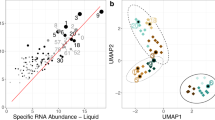
Relative abundances and distributions of gene coding sequences in the CN1 and CN2 communities based on taxonomic bins for bacterial 16S rDNA gene sequences (OPUs) (blue), taxonomic bins for metagenome-derived genes encoding proteins that could be assigned a taxonomic annotation (red) and taxonomic categories of proteins that were identified in the metaproteomes (green). As shown, the metagenome and the 16S rDNA analyses yielded congruent results. Further, only a minor number of bacterial members seemed to be metabolically active in both communities, by meaning of expressed proteins binned to them.
Finally, the two N clones (out of 214) representing Betaproteobacteria affiliated with Acidovorax and Achromobacter spp. (Supplementary Table S2). On the other hand, Nbs sequences were importantly represented in this lineage with 27 sequences (10%), affiliated with Tetrathiobacter and Acidovorax. In this regard, the percentage of Betaproteobacteria found in the enrichment cultures was much higher (Figures 1 and 4); we observed an overrepresentation of sequences affiliated with the genera Comamonas and Achromobacter in CN1 and Achromobacter in CN2 (Figures 1 and 5). In addition to Delta- and Epsilon-proteobacteria, members of Tenericutes, Verrucomicrobia, Firmicutes and Cyanobacteria were only detected in samples N and Nbs (Figures 3 and 4).
The above data demonstrated that the CN1 and CN2 communities displayed considerably different phylogenetic structures (Figures 4 and 5), sharing only two OPUs: Beta- and Gamma-proteobacteria representatives of the denitrifying Achromobacter (that is, Achromobacter spanius) and Azospirillum (that is, Azospirillum doebereinerae) genera, respectively. Whereas the first one was most abundant in CN2 (29 vs 7 sequences), the second was particularly enriched in CN1 (35 vs 1 sequences) (Supplementary Table S2). In addition to supplying biosurfactants, the addition of different nitrate compounds may have stimulated the growth of denitrifying Achromobacter spp. and P. stutzeri (Song and Ward, 2003) in the bio-stimulated community, whereas depletion of nitrate may have favoured the presence of the nitrogen-fixing Azospirillum species in the non-stimulated community. In addition to denitrification, P. stutzeri is well known as naphthalene degrader, which has been isolated in such hydrocarbon enrichments from a wide range of environmental samples (Rosselló-Mora et al., 1994), and was already an important component of the original Nbs soil. However, the possibility that the selective desorption and liberation of recalcitrant compounds from soils during the bio-stimulation process may have key functions in shaping the genomes and community structures of the associated microorganisms should not be ruled out. The findings of previous studies using BTEX biodegradation co-cultures (Shim et al., 2005) agree with this hypothesis. In any case, although bacteria related to Pseudomonas, Achromobacter and Comamonas spp. have been widely associated with PAH degradation in a number of contaminated ecosystems and microbial consortia (Goyal and Zylstra, 1997), the presence of the Azospirillum species in contaminated soils and PAH-degrading bacterial consortia has only previously been reported in a heavily creosote-contaminated soil (Viñas et al., 2005). This is of special interest because it is known that the recently sequenced Azospirillum sp. B510 harbours gene sets for degradation pathways, though their functions have not yet been experimentally elucidated (Kaneko et al., 2010).
Due to the clear shifts in community composition detected among the investigated conditions, it is important to establish the roles and degradation capabilities of the individual microbial members of the communities to understand the overall function of each community and its members.
Genomic signatures associated with the aerobic degradation of aromatics
DNA isolated from each microbial community was sequenced using a Roche GS FLX DNA sequencer, at Lifesequencing SL (Valencia, Spain) which produced 1 471 821 reads (N: 165 791; Nbs: 438 821; CN1: 448 713; CN2: 418 293) with an average length per read of 395 bp for CN1 and CN2, 336 bp for N and 531 for Nbs. Accordingly, a total of 55.7, 233.3, 177.2 and 165.2 Mbp of raw DNA sequences for N, Nbs, CN1 and CN2 were obtained, which were assembled into 2.6 Mbp (4335 contigs), 17.9 Mbp (16 032 contigs), 20.0 Mbp (20 809 contigs) and 13.0 Mbp (9915 contigs), respectively. CN1 (266) and CN2 (256) contained a higher number of contigs with lengths greater than 10 kbp compared with Nbs (68) and to major extent N (only 1); this may be due to the high diversity of the N sample, which resulted in a large pool of reads that could not be assembled (Supplementary Figure S7). Further information regarding the obtained metagenome characteristics is given in Supplementary Table S3. With a threshold of higher than 95% identity and an aligned length of more than 150 bp, 91.7% (for N), 95.2% (for Nbs), 92.3% (for CN1) and 89.9% (for CN2) of the predicted genes were assigned to particular genera, the analysis of which showed results comparable to those found in the 16S rDNA assignments (see Figure 5 for CN1 and CN2 comparison). Approximately 89% of the predicted genes (63 974 in total) were assigned to COG protein families, and 65% were assigned to KEGG pathways (Supplementary Table S3). Among the 1538 (for N), 2751 (for Nbs), 2775 (for CN1) and 2507 (for CN2) KEGG orthologs detected, 891 were shared between the two enrichment cultures, and 66% of these were putatively attributed to Achromobacter species.
We first calculated the overrepresentation of functions between metagenomes; for that, we applied a z-test for independent proportions proposed by Li (2009) to statistically analyse the changes of the functional categories between samples. It should be noticed that the comparison was restricted to samples Nbs, CN1 and CN2 because the lower number of pyro sequences and COGs in sample N may have impact on the comparative result. We found a rather stable distribution of the functional categories composition between the samples with significant different contribution of only 238 out of 3293 COGs (Supplementary Table S4). Among them, 56/23 did show a significant increase in the bio-stimulated soil as compared with the CN1/CN2 enrichments, 58/26 increased in CN1 as compared with Nbs/CN2 and 41/55 were overrepresented in CN2 as compared with Nbs/CN1. Overall, we observed that both bio-stimulated communities were particularly enriched in COGs related to ‘Replication and repair’ and ‘Translation’ (29 vs 4 distinct COGs in total in CN1). Overrepresentation of such functions is typical for microbial communities developed under very dynamic environmental conditions, such as bio-stimulation; the addition of different nitrate compounds may have stimulated a competition between fast-growing organisms and organisms capable of metabolising poly-aromatics. By contrast, it is noteworthy that COGs related to ‘Transcription’ were most likely characteristics of enrichment cultures (13 and 7 COG enriched in CN1 and CN2, respectively), whereas only two (COG0789 and COG1974) were enriched in Nbs (Supplementary Table S4). This suggests a plausible scenario in which the stress caused by aromatic compounds during the naphthalene cultivation led to an enrichment of genomes containing transcriptional elements that could be required for stress endurance (Domínguez-Cuevas et al., 2006) and/or activation of functions required for substrate uptake. Ten distinct COGs within the ‘Cell wall, membrane, envelop biogenesis’ category were found overrepresented in CN1, a number much higher that those found in Nbs (four COGs) and in CN2 (two COGs) (Supplementary Table S4). In addition, CN1 was also particularly enriched in ‘ABC Transport Systems’ by meaning of genes distributed in 11 distinct overrepresented COGs, for which only three and two were found in Nbs and CN2, respectively. COG0318 acetyl-CoA synthethase was also associated to CN1 community; enzymes of the COG0318 catalyse the formation of acetyl-CoA from acetate, suggesting that acetate may be a major end product used to produce acetyl-CoA feeding the Krebs pathway in members of CN1 (in agreement with proteomic data that will be discussed below). By contrast, CN2 was particularly enriched in genes coding proteins containing Fe–S clusters, namely COG3210 (large exo-proteins involved in haem utilisation or adhesion), COG1049 (aconitase B), COG0348 (polyferredoxin) as well as COG4773 (outer membrane receptor for ferric coprogen and ferric-rhodotorulic acid) (Supplementary Table S4), indicating iron being particularly relevant for CN2 community members. Taken together, the most notable observations drawn from all these data sets are that bio-stimulated communities appear to be extensive reservoirs of genes involved in replication, repair and translation and iron adhesion and utilisation whereas bacterial components of non-stimulated communities seems to be most active in cell wall, membrane and envelop biogenesis; in addition, enrichment most likely favoured transcriptional events.
The obtained coverage was sufficient to produce a further partial theoretical metabolic reconstruction (including both exclusive and common capabilities) of the aerobic aromatic catabolic routes in the investigated communities; although these reconstructions were incomplete and likely represent composite cell networks, the information obtained may be sufficient to achieve a better understanding of how the four communities behave regarding their biodegradation capabilities on a genomic scale. To do that, a proper functional assignment of the predicted genes was performed using an in-house database containing protein sequences with biochemical functions shown to be involved in biodegradation (for details see Supplementary Materials and methods). According to this protocol, 428 (N: 38; Nbs: 115; CN1: 132; CN2: 143) open-reading-frame fragments were identified as showing close sequence similarity to genes that encode enzymes known to be involved in the aerobic metabolism of aromatics via di- and tri-hydroxylated intermediates and, accordingly, presumptive functions were assigned. The overall gene features and presumptive catabolic routes most likely associated to them, are shown in Supplementary Table S5 and Figure 6, the results of which are described in details below. It should be noticed that the lower number of hits in the original community (N) may be due to the broad spectrum of carbon sources that are present in the soil and may have been biased by the lower number of assembled sequences due to the high diversity and the corresponding low coverage of the metagenome in this community.
Potential aerobic degradation networks of aromatics via di- and tri-hydroxylated intermediates in the four investigated communities. The colour code used for the respective pathways is as follows: black, all samples; green, N and Nbs (specific for soil communities); blue, CN1 (N) (specific for non bio-stimulated communities]; red, CN2 (Nbs) (specific for bio-stimulated communities). Illustrations were created by ChemDraw graphic programme Chem Draw Ultra 8.0 (CambridgeSoft; http://www.cambridgesoft.com/) based on substrate specificity of enzymes listed in Supplementary Table S5, and the corresponding metabolic pathways were established based on bibliographic records. It should be noticed that the identification of a particular activity in CN1 and CN2 implies its presence in N and Nbs, respectively, although their signatures were not identified in the later most likely owing to their higher biodiversity and low coverage of the metagenome. Accordingly, those activities were indicated as CN1 (N) and CN2 (Nbs).
Eighteen/thirteen types of oxygenases (including the component systems reductase and ferredoxin and large and small subunits of the terminal oxygenase component) and 3/13 types of accessory enzymes involved in subsequent degradation or transport steps were identified in the N/Nbs soil samples, with the majority (∼86% and 53%, respectively) being binned to strain(s) of the Pseudomonas genus (Supplementary Table S5). Among the most significant enzymes were those involved in potential degradation pathways for naphthalene, biphenyl, phenol and carbazole (Figure 6). The absence of genomic signatures for carbazole degradation in the enriched communities further suggests that either the carbazole-degrading bacterium (or its plasmid) containing such genes was lost, or was present at a very low abundance, during the enrichment process.
A total of 275 (CN1: 132; CN2: 143) open-reading-frame fragments covering 186 (CN1: 83; CN2: 103) unique proteins were identified in the enriched cultures, of which 152 (CN1: 62; CN2: 90) were characterised as oxygenase components, 43 (CN1: 11; CN2: 32) as accessory enzymes and 9 (CN1: 5; CN2: 4) as probable regulators. The functional analysis of these gene sets (Supplementary Table S5) suggested that microbial communities from enrichment samples and, in turn their corresponding soil samples, possessed a number of common metabolic degradation signatures (Figure 6), whereas their phylogenetic composition is significantly different; however, a number of community-specific genomic determinants were also found, which may be of particular interest regarding the possible energy sources available to community members (Eaton, 1996).
We first obtained genomic evidence of the presence of genes belonging to the naphthalene upper pathway leading to the formation of salicylate (Eaton and Chapman, 1992) in both enrichment cultures, including naphthalene dioxygenases (CN1: 1; CN2: 2) and accessory enzymes (CN1: 6; CN2: 8) (Supplementary Table S5 and Figure 6). Additionally, the identification of salicylate-5-hydroxylases (CN1: 5; CN2: 5) and salicylate-1-hydroxylases (CN1: 0; CN2: 2) provided evidence that although the salicylate-to-gentisate pathway operated in both communities, the salicylate-to-catechol pathway most likely occurs only in CN2. Both communities contained genomic signatures suggesting their ability to transform phenol to catechol (Supplementary Table S5 and Figure 6). In CN2, a multicomponent phenol hydroxylase (⩾69% sequence identity to lapKLMNOP of P. alkylphenolia) (Jeong et al., 2003) was encoded in a single contig (contig01289) that also contained enzymes involved in transforming catechol to Krebs cycle intermediates (LapBCEHGIR, an acetylaldehyde dehydrogenase and a transporter); in CN1, a phenol oxygenase component was detected, although neither a clear indication of a complete pathway nor a clear phylogenetic affiliation could be established. The CN1 metagenome did not contain genomic determinants associated with the salicylate-to-catechol pathway, but did contain genes encoding enzymes involved in the ortho-cleavage of catechol (Supplementary Table S5); however, this is not surprising, considering that the catechol pathway is widespread among Proteobacteria (Pérez-Pantoja et al., 2008). Catechol is a central metabolite in the biphenyl/benzoate degradation pathways, and six benzoate dioxygenases (CN1: 2; CN2: 4), four 2,3-dihydroxybiphenyl dioxygenases (CN1: 1; CN2: 3) and two benzoate dihydrodiol dehydrogenases (CN1:0; CN2:2) were identified in the two communities (Supplementary Table S5). Other common degradation capabilities observed in both communities included the potential to degrade 4-hydroxyphenylacetate via homoprotocatechuate, which may be transformed by homoprotocatechuate 2,3-dioxygenases (CN1: 2; CN2: 2); furthermore, 4-hydroxyphenylpyruvate may be transformed by 4-hydroxyphenylpyruvate dioxygenases (CN1: 4; CN2: 5) to homogentisate, which can be cleaved by homogentisate 1,2-dioxygenases (CN1: 5; CN2: 3) (Supplementary Table S5 and Figure 6). The identification of a 2-amino-1,6-dioxygenase in both communities suggests that aminophenol may be also a potential common carbon source. It was recently reported that nicotinic acid, an n-heterocyclic carboxylic derivative of pyridine, that is essential for microorganisms, can also be used as a carbon and nitrogen source (Jiménez et al., 2008); this substrate enters the Krebs cycle via the action of a number of enzymes, and degradation is initiated by a 6-hydroxynicotinate 3-monooxygenase (Jiménez et al., 2008). Accordingly, two putative 6-hydroxynicotinate 3-monooxygenases (CN1: 1; CN2: 1) were identified (Supplementary Table S5). Further genomic evidence suggested the presence of metabolic routes associated with vanillate (vanillate monooxygenase; CN1: 2; CN2: 2) and p-hydroxybenzoate (4-hydroxybenzoate 3-hydroxylase; CN1: 3; CN2: 2) (Supplementary Table S5 and Figure 6).
Genomic evidence of the capability to potentially transform phthalate (four phthalate dioxygenases) and terephthalate (one terephthalate dioxygenase) into protocatechuate was only observed in CN1 (Supplementary Table S5 and Figure 6). Other potential growth-supporting aromatic compounds found to be CN1 specific were gallate (through a gallate dioxygenase), 2,3-dihydroxyphenylpropionate (through a 2,3-dihydroxy-phenylpropionate dioxygenase), 2,3-dihydroxy-p-cumate (through a 2,3-dihydroxy-p-cumate dioxygenase), indole carboxylate, polycyclic arene diols (PhnC-like polycyclic arene diol extradiol dioxygenase (PhnC-like)) and diterpenoids, such as abietic acid and its intermediate 7-oxo-11,12-dihydroxy-8,13-abietadiene acid (through an abietane diterpenoid dioxygenase and the corresponding 7-oxo-11,12-dihydroxydehydroabietate dioxygenase, which has an unclear phylogenetic affiliation) (Supplementary Table S5 and Figure 6). By contrast, genomic signatures for the potential utilisation of ibuprofen, through an ibuprofen-CoA dioxygenase, were only found in bio-stimulated derived community (Supplementary Table S5).
The functional phylogenetic assignments further indicated potential major ‘food’ webs that were primarily composed of Pseudomonas gammaproteobacterial species (most likely operating in both soils and CN2 communities, but not in CN1), Azospirillum alphaproteobacterial species (operating in non-stimulated communities) and betaproteobacterial Achromobacter (operating in all communities) and Comamonas (operating in non-stimulated communities) species. Based on the phylogenetic analysis provided in Supplementary Table S5, different interactions among these members can be hypothesised. Achromobacter spp. may supply the genetic potential for transforming naphthalene to salicylate, benzoate/biphenyl to catechol, 4-hydroxybenzoate, 4-hydroxyphenylpyruvate, ibuprofen, vanillate, homogentisate, (homo-) protocatechuate and cumate. Potential activities of Azospirillum may be responsible for the ability to transform salicylate to gentisate, 4-hydroxybenzoate and 4-hydroxyphenylpyruvate. The potential roles of Pseudomonas may cover the abilities to transform carbazole to catechol, naphthalene to salicylate to gentisate and catechol, benzoate/biphenyl/phenol to catechol, 4-hydroxybenzoate, 4-hydroxyphenylpyruvate, vanillate and protocatechuate. The potential activities of Comamonas may include the transformation of 4-hydroxybenzoate, 4-hydroxyphenylpyruvate, vanillate, gallate, nicotinate, 2,3-dihydroxyphenylpropionate and (tere-) phthalate.
Taken together, the data may have a significant ecological impact while it is plausible that bio-stimulation per se favours ‘specialists’ for naphthalene-degradation, non-stimulated communities may display a higher metabolic plasticity to access pollutants with complex chemical structures and physical properties. This pattern was clearly demonstrated as bio-stimulated community members presented two different routes for salicylate utilisation, in contrast to the non-stimulated ones, which were only capable of metabolising naphthalene via gentisate but have the genetic potential to degrade a range of carboxylated aromatics (Figure 6), for which no genomic signatures were found in the bio-stimulated communities. This should be of special interest in defining future strategies for implementing enrichment cultures in settings associated with biodegradation based on the pollutant profile and thus producing a-la-carte degrading communities.
Proteomic blueprints of CN1 and CN2 naphthalene-enriched communities
A total of 1116 proteins were unambiguously identified and quantified from cells harvested from enrichment cultures CN1 and CN2 (60 transfers) used also for DNA isolation, following the protocol and criteria described in Supplementary Materials and Methods (Supplementary Table S6). Because the analysis was performed using a draft metagenome, the metaproteome size is within common ranges that have been observed for other communities and is only three times lower than that observed for cultivable organisms (Benndorf et al., 2007; Passalacqua et al., 2009; Abu Laban et al., 2010; Selesi et al., 2010). As the majority of the spectra could be assigned to a taxonomic and functional annotation based on highly similar homologues, the metaproteomic approach applied here allowed us to compare taxonomic annotations to evaluate the differences between the contributions of particular groups of organisms to the global community, as well as to predict the importance of particular sets of proteins for the overall functioning of the community.
A total of 582 (or 52%) proteins were identified in both samples, with the relative intensities of 123 and 227 proteins being significantly higher in CN1 and CN2, respectively (Supplementary Table S6). Additionally, a total of 132 proteins were specific to CN1, most of which were affiliated with the Azospirillum (∼71%) and Comamonas (∼17%) species, whereas 402 proteins (∼94% associated with Gammaproteobacteria of the Pseudomonas genus) were found exclusively in CN2. This clearly indicated that the CN1 and CN2 communities displayed considerable heterogeneity, in accordance with their distinct phylogenetic compositions (Figures 4 and 5). On the basis of their probable functions, the proteins were sorted into five major groups, which were associated with roles in aerobic degradation, electron acceptors, energy metabolism, transport and folding/stress.
(i) Aerobic degradation proteins: As expected for the communities utilising aromatics as their sole carbon source, peptides from 27 unique proteins directly assigned to different aromatic degradation pathways were found in the metaproteomes, albeit at different intensities (Supplementary Table S7). The peptides included 19 proteins belonging to the naphthalene degradation pathway with gentisate as an intermediate, with abundance levels up to a relative concentration (rel. conc.) of 2.1%. Seventeen proteins were observed at higher abundances in CN1 compared with CN2, whereas two proteins were CN1 and CN2-specific. Additionally, four proteins involved in the catechol meta-cleavage pathway were found to be most expressed in CN2, with remarkable expression levels up to a 0.53% rel. conc. Expression of a glutathione S transferase (NagJ-like) and a membrane protein of unknown function was observed and may be implicated in naphthalene degradation. It should be noted that the most abundant protein in CN1 showed a relative concentration of ∼3.6%, coding for the naphthalene dioxygenase alpha and beta subunits (Supplementary Table S6); therefore, proteins involved in naphthalene degradation were among those that were most highly expressed (that is, CN1/23999 gentisate dioxygenase, with a 2.0% rel. conc.), as expected for a community utilising naphthalene as the only carbon source. Similarly, in the case of CN2, the most abundant protein was a salicylaldehyde dehydrogenase (1.7% rel. conc.) (Supplementary Table S6).
Phylogenetic analysis suggests that in CN1 the primary proteins that are active are identical to those described in the pWWU2 plasmid of Ralstonia sp. U2, and most likely binned to several species or genomes of Achromobacter (Supplementary Table S7). In contrast, the degradation of naphthalene in CN2 may be ‘presumptively’ carried out both by organisms that utilise a catechol pathway similar to the pathway encoded by the plasmid pND6-1 of Pseudomonas sp. strain ND6 and those that employ gentisate pathways; however, in the latter case, at least two different type of organisms may be involved, as proteins identical (similarity ranging from 57 to 100%) to those encoded by the pWWU2 and pND6-1 plasmids from Ralstonia sp. strain U2 (in CN1 and CN2) and Pseudomonas sp. strain ND6 (in CN2), respectively, were found (Supplementary Tables S6 and S7). Additional organisms may have minor roles in the naphthalene degradation network in both cultures, as indicated by the expression of CN1/3872 gentisate dioxygenase, which has a sequence identity of only 53% to one of Ralstonia sp. U2 (Supplementary Table S7). In any case, it should be noted that among a total of 62 unique proteins taking part in the putative naphthalene degradation network based on the obtained metagenome sequence data (Supplementary Table S5), only approximately 43% of the proteins passed the strict ⩾2 unique peptides filtering criterion (Supplementary Table S7). As the physico-chemical features of peptides that define their suitability of detection by mass spectrometry are more or less equally distributed the likeliness for detecting a protein is mostly a matter of abundance. Thus, the set of label-free detected proteins is by itself indicative for both the abundance of the producing species and the protein of interest.
(ii) Proteins involved in electron acceptors: A total of seven genes (covered by five distinct orthologs: K00370, K00371, K00373, K00374 and K00376) involved in nitrate (NO3-) and nitric/nitrous oxide (NxO) metabolism were found among the genes that were exclusive to the CN2 metagenome, the majority (∼90%) of which were associated with a single (or several) species or with a Pseudomonas genome (most likely P. stutzeri). Consistent with this finding, significant expression of a nitric-oxide reductase (CN2/16494; 0.25% rel. conc.), a nitrous-oxide reductase (CN2/13702; 0.30% rel. conc.), and four nitrate reductases (CN2/1030, CN2/16257, CN2/15634 and CN2/1029; from 0.01% to 0.20% rel. conc.) was found in the CN2 metaproteome (Supplementary Table S6). The presence of such enzymes suggests that NxO may be used as an alternative (or dominant) electron acceptor (Hemme et al., 2010) by members of the CN2 community, most likely by Pseudomonas stutzeri strain(s) that are capable of denitrification (Chen et al., 2011).
(iii) Proteins involved in energy metabolism: Enzymes belonging to the naphthalene degradation pathway convert the naphthalene to pyruvate and fumarate, which feed the Krebs cycle. A total of 72 central metabolic enzymes were found in both the CN1 and CN2 metaproteomes during growth using naphthalene, including 41 proteins involved in the Krebs and glyoxylate cycles (Supplementary Table S6), which binned majoritarily to several species or genomes of Achromobacter, Comamonas and Azospirillum in CN1 and Achromobacter and Pseudomonas in CN2. The enzyme diversity and taxonomic classification of the expressed proteins suggested that during growth with naphthalene, acetyl-CoA was metabolised in CN1 and CN2 via both the Krebs and the glyoxylate cycles. However, it is noteworthy that the expression of citrate synthase (for example, CN2/0977; 0.37% rel. conc.) in CN2 was approximately threefold higher than the expression of malate synthase (for example, CN2/15317; 0.11% rel. conc.), whereas in CN1, an opposite scenario was observed, as malate synthase (for example, CN1/16907; 0.07% rel. conc.) was expressed at a higher level than citrate synthase (0.0096% rel. conc.) (Supplementary Table S6). This suggests that acetyl-CoA may be preferentially metabolised via the Krebs cycle in CN2, whereas it may be preferentially metabolised via the glyoxylate cycle in CN1. Kurbatov et al. (2006) observed that during the growth of P. putida KT2440 on phenol, the glyoxylate cycle was preferred, suggesting that bio-stimulation prior to the enrichment may also have a direct influence on the overall preferred mode of energetic metabolism in settings associated with biodegradation.
Binning analysis suggests Pseudomonas species (in CN2) most likely use both pyruvate and fumarate (generated from naphthalene) as substrates to feed the Krebs and glyoxylate pathways, whereas members of CN1 primarily use pyruvate. This can be seen clearly in the abundances of fumarate hydratases and succinate dehydrogenases, which were observed at significant levels only in the CN2 proteome (seven proteins in total, all binned to the Pseudomonas genus: CN2/0973, CN2/0975, CN2/0974, CN2/7221, CN2/17464, CN2/17714 and CN2/17817, showing up to a 0.46% rel. conc.) (Supplementary Table S6).
The metaproteomic analysis revealed the presence of five acetyl-CoA synthases (3 binned to Azospirillum in CN1 and 1 to Achromobacter and 1 to Pseudomonas in CN2; up to a 0.51% rel. conc.; Supplementary Table S6), which suggests acetate, in combination with pyruvate (produced from naphthalene), is a major end product used to produce acetyl-CoA feeding the Krebs and glyoxylate pathways. In CN1, acetyl-CoA may additionally be produced from acetate via the action of an Azospirillum-derived formate c-acetyltransferase (CN1/4225), which was detected at a low expression level (0.02% rel. conc.) (Supplementary Table S6); moreover, in CN1, acetate may be used as an energy source via an acetate kinase (0.04% rel. conc.), which uses ATP to produce acetyl phosphate. Taking these findings together, we suggest that the production and utilisation of acetate yielding maximal energy while avoiding acidification of the environment owing to the accumulation of acetate is potentially more efficient in CN1. Together with the previous observations related to the Krebs and glyoxylate cycles, these results suggest that distinct energetic networks operate in both communities during naphthalene utilisation.
(iv) Proteins involved in transport. Both enrichment cultures contained a large number of cytoplasmic membrane transport proteins, which may facilitate the uptake of a multitude of organic compounds: strong production of 69 (up to a 2.28% rel. conc.) and 24 (up to a 1.55% rel. conc.) proteins associated with (in)organic molecule uptake systems (including transporters, TolB proteins and porins) was evident in the CN1 and CN2 metaproteomes, respectively (Supplementary Table S6). The large number of significantly expressed transporters in CN1 is in agreement with the profiling analysis based on the COG-enrichment values calculated for each microbiomes (Supplementary Table S4), which showed an overrepresentation of COGs classified into the ‘ABC transport systems’ in the non-stimulated enriched community.
(v) Proteins involved in folding and stress: Proteins involved in detoxification and oxidative stress response mechanisms were identified only in CN1 (22 hits) (Supplementary Table S6). Interestingly, all of these proteins were binned to Azospirillum, which may suggest a major level of radical stress on these organisms compared with others within the community. Among the stress proteins detected, one peroxidase and one catalase (CN1/2218 and CN1/18517) and two putative xenobiotic reductases (CN1/14685 and CN1/16278) were identified, with relative abundance levels ranging from 0.09 to 0.19%. Expression of these types of stress proteins has been reported in several studies related to the responses of organisms to different sources of carbon and energy, for example, as a response to phenol (Kurbatov et al., 2006). However, to the best of our knowledge, no experimental (for example, proteomic) evidences, suggesting higher stress responses of Azospirillum strains compared with other degraders in contaminated or aromatic-enriched cultures, have been obtained. In contrast, high abundances (21 hits) of Pseudomonas- and Achromobacter-binned stress proteins potentially involved in protein folding mechanisms were identified in CN2, which included CspA, Clp, GroEL, DnaJ and thiol-disulphide interchange-like proteins (Supplementary Table S6). Taken together, these data suggest the action of distinct coping mechanisms in response to naphthalene-derived stress during the enrichment process. Members of CN1 appear to be more affected by radical oxygen species most likely produced during naphthalene degradation (for example, via enzyme inefficiencies and/or dioxygenase inhibition), whereas stress factors derived from naphthalene itself appear to be the major determinants affecting individual members, and thus, modulating the structure of the CN2 community.
All together, the present study provided a comprehensive and comparative investigation of four microbial communities containing several OPUs associated with putative metabolism related to degradation capabilities. Their inter-alia interactions were unrevealed, which further demonstrated that a minor fraction of active genomes (and the proteins they contain) is responsible for the degradation of naphthalene both in soil samples and enrichment cultures (Figure 5). It is interesting to mention with respect to the aspects discussed here, that the enrichment serial batch cultures used for CN1 and CN2 could select for the fast growers species, which could also have some impact in the functional phylogenetic structure found in this investigation. Whatever the case, the present study revealed that the soil ecosystem (that is, a PAH-contaminated soil) investigated here was quite delicate and that its metabolic capability may be easily lost as a result of bio-stimulation, which could have a significant ecological role. Finally, the curated database containing protein sequences with biochemical functions, shown to be involved in biodegradation of aromatics via di- and tri-hydroxylated intermediates and are created and applied for first time in this study, could be used in future investigation related to microbial biodiversity, ecology and function as response to aromatics.
References
Abu Laban N, Selesi D, Rattei T, Tischler P, Meckenstock RU . (2010). Identification of enzymes involved in anaerobic benzene degradation by a strictly anaerobic iron-reducing enrichment culture. Environ Microbiol 12: 2783–2796.
Abulencia CB, Wyborski DL, Garcia JA, Podar M, Chen W, Chang SH et al (2006). Environmental whole-genome amplification to access microbial populations in contaminated sediments. Appl Environ Microbiol 72: 3291–3301.
Beil S, Mason JR, Timmis KN, Pieper DH . (1998). Identification of chlorobenzene dioxygenase sequence elements involved in dechlorination of 1,2,4,5-tetrachlorobenzene. J Bacteriol 180: 5520–5528.
Benndorf D, Balcke GU, Harms H, von Bergen M . (2007). Functional (meta-) proteome analysis of protein extracts from contaminated soil and groundwater. ISME J 1: 224–234.
Boronin AM, Kosheleva IA . (2010). Biodiversity of naphthalene biodegradation systems in soil bacteria. In: Timmis KN (ed) Handbook of Hydrocarbon and Lipid Microbiology. Springer-Verlag: Berlin, pp 1155–1163.
Chen M, Yan Y, Zhang W, Lu W, Wang J, Ping S et al (2011). Complete genome sequence of the type strain Pseudomonas stutzeri CGMCC 1.1803. J Bacteriol 193: 6095.
Domínguez-Cuevas P, Gonzalez-Pastor JE, Marques S, Ramos JL, de Lorenzo V . (2006). Transcriptional tradeoff between metabolic and stress-response programs in Pseudomonas putida KT2440 cells exposed to toluene. J Biol Chem 281: 11981–11991.
Eaton RW . (1996). p-Cumate catabolic pathway in Pseudomonas putida Fl: cloning and characterization of DNA carrying the cmt operon. J Bacteriol 178: 1351–1362.
Eaton RW, Chapman PJ . (1992). Bacterial metabolism of naphthalene: construction and use of recombinant bacteria to study ring cleavage of 1,2-dihydroxynaphthalene and subsequent reactions. J Bacteriol 174: 7542–7554.
Gallego JR, Fernández JR, Díez-Sanz F, Ordoñez S, Sastre H, González-Rojas E et al (2007a). Bioremediation for shoreline cleanup: in situ vs. on-site treatments. Environ Engineer Sci 24: 493–504.
Gallego JR, García-Martínez MJ, Llamas JF, Belloch C, Peláez AI, Sánchez J . (2007b). Biodegradation of oil tank bottom sludge using microbial consortia. Biodegradation 18: 269–281.
Goyal AK, Zylstra GJ . (1997). Genetics of naphthalene and phenanthrene degradation by Comamonas testosteroni. J Ind Microbiol Biotechnol 19: 401–407.
Hammer Ø, Harper D, Ryan P . (2001). PAST: Paleontological statistics software package for education and data analysis. Palaeontologia Electronica 4: 1–9.
Hemme CL, Deng Y, Gentry TJ, Fields MW, Wu L, Barua S et al (2010). Metagenomic insights into evolution of a heavy metal-contaminated groundwater microbial community. ISME J 4: 660–672.
Jeong JJ, Kim JH, Kim CK, Hwang I, Lee K . (2003). 3- and 4-alkylphenol degradation pathway in Pseudomonas sp. strain KL28: genetic organization of the lap gene cluster and substrate specificities of phenol hydroxylase and catechol 2,3-dioxygenase. Microbiology 149: 3265–3277.
Jiménez JI, Canales A, Jiménez-Barbero J, Ginalski K, Rychlewski L, García JL et al (2008). Deciphering the genomic determinants for aerobic nicotinic acid degradation: the nic cluster from Pseudomonas putida KT2440. Proc Natl Acad Sci USA 105: 11329–11334.
Jiménez JI, Minambres B, Garcia JL, Diaz E . (2002). Genomic analysis of the aromatic catabolic pathways from Pseudomonas putida KT2440. Environ Microbiol 4: 824–841.
Kaneko T, Minamisawa K, Isawa T, Nakatsukasa H, Mitsui H, Kawaharada Y et al (2010). Complete genomic structure of the cultivated rice endophyte Azospirillum sp. B510. DNA Res 17: 37–50.
Kästner M . (2000). Degradation of aromatic and polyaromatic compounds. Biotechnology 11: 211.
Kim YH, Cho K, Yun SH, Kim JY, Kwon KH, Yoo JS et al (2006). Analysis of aromatic catabolic pathways in Pseudomonas putida KT 2440 using a combined proteomic approach: 2-DE/MS and cleavable isotope-coded affinity tag analysis. Proteomics 6: 1301–1318.
Kurbatov L, Albrecht D, Herrmann H, Petruschka L . (2006). Analysis of the proteome of Pseudomonas putida KT2440 grown on different sources of carbon and energy. Environ Microbiol 8: 466–478.
Kweon O, Kim SJ, Jones RC, Freeman JP, Adjei MD, Edmondson RD et al (2007). A polyomic approach to elucidate the fluoranthene-degradative pathway in Mycobacterium vanbaalenii PYR-1. J Bacteriol 189: 4635–4647.
Li W . (2009). Analysis and comparison of very large metagenomes with fast clustering and functional annotation. BMC Bioinformatics 10: 359.
Liang Y, Van Nostrand JD, Deng Y, He Z, Wu L, Zhang X et al (2011). Functional gene diversity of soil microbial communities from five oil-contaminated fields in China. ISME J 5: 403–413.
Liu R, Zhang Y, Ding R, Li D, Gao Y, Yang M . (2009). Comparison of archaeal and bacterial community structures in heavily oil-contaminated and pristine soils. J Biosci Bioeng 108: 400–407.
López-López A, Yarza P, Richter M, Suárez-Suárez A, Antón J, Niemann H et al (2010). Extremely halophilic microbial communities in anaerobic sediments from a solar saltern. Environ Microbiol Rep 2: 258–271.
Lors C, Ryngaert A, Périé F, Diels L, Damidot D . (2010). Evolution of bacterial community during bioremediation of PAHs in a coal tar contaminated soil. Chemosphere 81: 1263–1271.
Lu XY, Zhang T, Fang HH . (2011). Bacteria-mediated PAH degradation in soil and sediment. Appl Microbiol Biotechnol 89: 1357–1371.
Martin F, Torelli S, Le Paslier D, Barbance A, Martin-Laurent F, Bru D et al (2012). Betaproteobacteria dominance and diversity shifts in the bacterial community of a PAH-contaminated soil exposed to phenanthrene. Environ Pollut 162: 345–353.
Ní Chadhain SM, Norman RS, Pesce KV, Kukor JJ, Zylstra GJ . (2006). Microbial dioxygenase gene population shifts during polycyclic aromatic hydrocarbon biodegradation. Appl Environ Microbiol 72: 4078–4087.
Nogales J, Palsson BO, Thiele I . (2008). A genome-scale metabolic reconstruction of Pseudomonas putida KT2440: iJN746 as a cell factory. BMC Syst Biol 2: 79.
Passalacqua KD, Varadarajan A, Ondov BD, Okou DT, Zwick ME, Bergman NH . (2009). Structure and complexity of a bacterial transcriptome. J Bacteriol 191: 3203–3211.
Peng RH, Xiong AS, Xue Y, Fu XY, Gao F, Zhao W et al (2008). Microbial biodegradation of polyaromatic hydrocarbons. FEMS Microbiol Rev 32: 927–955.
Pérez-Pantoja D, De la Iglesia R, Pieper DH, González B . (2008). Metabolic reconstruction of aromatic compounds degradation from the genome of the amazing pollutant-degrading bacterium Cupriavidus necator JMP134. FEMS Microbiol Rev 32: 736–794.
Pérez-Pantoja D, Donoso R, Agulló L, Córdova M, Seeger M, Pieper DH et al (2012). Genomic analysis of the potential for aromatic compounds degradation in Burkholderiales. Environ Microbiol 14: 1091–1117.
Pérez-Pantoja D, Donoso R, Junca H, Gonzalez B, Pieper DH . (2009). Phylogenomics of aerobic bacterial degradation of aromatics. In: Timmis KN (ed) Handbook of Hydrocarbon and Lipid Microbiology. Springer-Verlag: Berlin, pp 1356–1397.
Powell SN, Singleton DR, Aitken MD . (2008). Effects of enrichment with salicylate on bacterial selection and PAH mineralization in a microbial community from a bioreactor treating contaminated soil. Environ Sci Technol 42: 4099–4105.
Puchałka J, Oberhardt MA, Godinho M, Bielecka A, Regenhardt D, Timmis KN et al (2008). Genome-scale reconstruction and analysis of the Pseudomonas putida KT2440 metabolic network facilitates applications in biotechnology. PLoS Comput Biol 4: e1000210.
Rhee SK, Liu X, Wu L, Chong SC, Wan X, Zhou J . (2004). Detection of genes involved in biodegradation and biotransformation in microbial communities by using 50-mer oligonucleotide microarrays. Appl Environ Microbiol 70: 4303–4317.
Richardson SD, Jones MD, Singleton DR, Aitken MD . (2012). Long-term simulation of in situ biostimulation of polycyclic aromatic hydrocarbon-contaminated soil. Biodegradation 23: 621–633.
Roling WFM, Head IM, Larter SR . (2003). The microbiology of hydrocarbon degradation in subsurface petroleum reservoirs: perspectives and prospects. Res Microbiol 154: 321–328.
Rosselló-Mora R, Lalucat J, García-Valdés E . (1994). Biochemical and genetic analysis of naphthalene degradation among Pseudomonas stutzeri strains. Environ Microbiol 60: 966–972.
Santos PM, Benndorf D, Sa-Correia I . (2004). Insights into Pseudomonas putida KT2440 response to phenol-induced stress by quantitative proteomics. Proteomics 4: 2640–2652.
Segura A, Godoy P, van Dillewijn P, Hurtado A, Arroyo N, Santacruz S et al (2005). Proteomic analysis reveals the participation of energy- and stress-related proteins in the response of Pseudomonas putida DOT-T1E to toluene. J Bacteriol 187: 5937–5945.
Selesi D, Jehmlich N, von Bergen M, Schmidt F, Rattei T, Tischler P et al (2010). Combined genomic and proteomic approaches indentify gene clusters involved in anaerobic 2-methylnaphthalene degradation in the sulphate-reducing enrichment culture N47. J Bacteriol 192: 295–306.
Shim H, Hwang B, Lee S-S, Kong S-H . (2005). Kinetcis of BTEX biodegradation by a coculture of Pseudomonas putida and Pseudomonas fluorescens under hypoxic conditions. Biodegradation 16: 319–327.
Song B, Ward BB . (2003). Nitrite reductase genes in halobenzoate degrading denitrifying bacteria. FEMS Microbiol Ecol 43: 349–357.
Thavamani P, Malik S, Beer M, Megharaj M, Naidu R . (2012). Microbial activity and diversity in long-term mixed contaminated soils with respect to polyaromatic hydrocarbons and heavy metals. J Environ Manage 99C: 10–17.
Tomás-Gallardo L, Canosa I, Santero E, Camafeita E, Calvo E, López JA et al (2006). Proteomic and transcriptional characterization of aromatic degradation pathways in Rhodoccocus sp. strain TFB. Proteomics 1: S119–S132.
Vilchez-Vargas R, Junca H, Pieper DH . (2010). Metabolic networks, microbial ecology and 'omics' technologies: towards understanding in situ biodegradation processes. Environ Microbiol 12: 3089–3104.
Viñas M, Sabaté J, Espuny MJ, Solanas AM . (2005). Bacterial community dynamics and polycyclic aromatic hydrocarbon degradation during bioremediation of heavily creosote-contaminated soil. Appl Environ Microbiol 71: 7008–7018.
Vivas A, Moreno B, del Val C, Macci C, Masciandaro G, Benitez E . (2008). Metabolic and bacterial diversity in soils historically contaminated by heavy metals and hydrocarbons. J Environ Monit 10: 1287–1296.
Witzig R, Junca H, Hecht HJ, Pieper DH . (2006). Assessment of toluene/biphenyl dioxygenase gene diversity in benzene-polluted soils: links between benzene biodegradation and genes similar to those encoding isopropylbenzene dioxygenases. Appl Environ Microbiol 72: 3504–3514.
Yagi JM, Neuhauser EF, Ripp JA, Mauro DM, Madsen EL . (2010). Subsurface ecosystem resilience: long-term attenuation of subsurface contaminants supports a dynamic microbial community. ISME J 4: 131–143.
Yarza P, Richter M, Peplies J, Euzéby J, Amann R, Schleifer K-H et al (2008). The All-Species Living Tree Project: a 16S rDNA-based phylogenetic tree of all sequenced type strains. Syst Appl Microbiol 31: 241–250.
Yuste L, Hervás AB, Canosa I, Tobes R, Jiménez JI, Nogales J et al (2006). Growth phase-dependent expression of the Pseudomonas putida KT2440 transcriptional machinery analysed with a genome-wide DNA microarray. Environ Microbiol 8: 165–177.
Acknowledgements
This research was supported by the Spanish CSD2007-00005 and CENIT-07-CLEAM projects and by FEDER funds, ERANET (GEN2006-27750-C5-5-E/SYS), and the European Community Projects MAGICPAH (FP7-KBBE-2009-245226) and ULIXES (FP7-KBBE-2010-266473). M-EG thanks the CSIC for a JAE fellowship, and IL thanks FICYT, Principado de Asturias for a Severo Ochoa fellowship. We also thank Professor Michail M Yakimov, Jesús Tornés and Michael Richter for assistance in the bacterial diversity analyses.
Project has been registered as umbrella BioProject at NCBI with the ID PRJNA51851. Metagenomic sequences are deposited in Short Read Archive (http://www.ncbi.nlm.nih.gov/sra) with accession number SRA053109. The GenBank accession numbers for nucleotide sequences are HQ218445 to HQ218870 and JX310869 to JX311048.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Guazzaroni, ME., Herbst, FA., Lores, I. et al. Metaproteogenomic insights beyond bacterial response to naphthalene exposure and bio-stimulation. ISME J 7, 122–136 (2013). https://doi.org/10.1038/ismej.2012.82
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2012.82
Keywords
This article is cited by
-
Advances in metaproteomic profiling of molecular microbiology and environmental responses
Systems Microbiology and Biomanufacturing (2024)
-
Biochemical characterization, microbial diversity and biodegradability of coastal sediments in the Gulf of Gabès, Southern Mediterranean Sea
International Journal of Environmental Science and Technology (2022)
-
Implementation of Genetic Engineering and Novel Omics Approaches to Enhance Bioremediation: A Focused Review
Bulletin of Environmental Contamination and Toxicology (2022)
-
Soil metaproteomics as a tool for monitoring functional microbial communities: promises and challenges
Reviews in Environmental Science and Bio/Technology (2020)
-
Whole metagenome sequencing reveals links between mosquito microbiota and insecticide resistance in malaria vectors
Scientific Reports (2018)