Abstract
The human intestinal tract is colonized by a myriad of microbes that have developed intimate interactions with the host. In healthy individuals, this complex ecosystem remains stable and resilient to stressors. There is significant attention on the understanding of the composition and function of this intestinal microbiota in health and disease. Current developments in metaomics and systems biology approaches allow to probe the functional potential and activity of the intestinal microbiota. However, all these approaches inherently suffer from the fact that the information on macromolecules (DNA, RNA and protein) is collected at the ecosystem level. Similarly, all physiological and other information collected from isolated strains relates to pure cultures grown in vitro or in gnotobiotic systems. It is essential to integrate these two worlds of predominantly chemistry and biology by linking the molecules to the cells. Here, we will address the integration of omics- and culture-based approaches with the complexity of the human intestinal microbiota in mind and the mucus-degrading bacteria Akkermansia spp. as a paradigm.
Similar content being viewed by others
Editor's note
Willem M de Vos is Professor and Chair of Microbiology at Wageningen University in the Netherlands and Academy Professor at Helsinki University in Finland. He is a leading expert in gut microbiome research and was one of the first microbial ecologists to venture into this field, following his work since the 80s with lactic acid bacteria in the dairy industry and archaea and other extremophiles in the 90s. His work follows a line of research established in the Netherlands by his colleague, the late Antoon Akkermans. One of the key foci of this Winogradksy Review in fact highlights the appropriately named gut microbe, Akkermansia muciniphila, due to its important role in mucin degradation in the intestine. Willem is also at the forefront in the development and application of molecular and functional ‘omics’ approaches for gaining a better understanding of the role of gut microbes in human health, as also outlined in this review.
Introduction
Early stages of vertebrate development typically occur in the protected confines of the chorion, an environment free of microorganisms. Upon birth, the gastrointestinal tract is colonized by a rapidly developing microbial ecosystem—our microbes inside. The environmental conditions within the intestinal tract vary considerably and hence result in evolutionarily adapted microbial communities that show specific, temporal and spatial organization. The primary function of the intestinal tract is to generate appropriately processed food components that can be sequestered by the host. This requires harsh food deconstruction conditions that are determined by the architecture, size and development phase of the intestinal tract. In humans, these conditions include steep gradients of hydrochloric and bile acids, hydrolytic enzymes and oxygen levels or redox potentials. The intestinal tract of a human adult is characterized by a progressive microbial colonization that is also related to the food retention time (Figure 1). The microbial residents that can cope with these specific conditions mainly consist of anaerobic Bacteria and to a lesser extent Archaea, Eukarya and their viruses (Rajilic-Stojanovic et al., 2007; Camp et al., 2009; Reyes et al., 2010). Changes in environmental conditions and microbial infections may have pervasive effects on the composition of the intestinal microbiota, but the microbiota in general has shown a marked resilience. Hence, it is assumed that maintenance and restoration of the indigenous microbiota is an important contribution to the health of the host.
From diversity to function. (a) Schematic outline of the human digestive tract with its characteristics and microbiota. The stomach is characterized by low pH conditions and a short retention time and colonization by bacteria is minimal. The small intestine (duodenum, jejunum and proximal ileum) has a short retention time (2–6 h) and high pH, the residing microflora is exposed to secreted bile salts and pancreatic juices at the proximal part. The lower digestive tract, comprising the terminal ileum and colon, is in contrast characterized by a longer retention time, neutral pH and is the most densely populated by microorganisms. (b) Overview of the major ‘omics’ approaches using DNA, RNA, protein or culture-based techniques.
Despite the substantial environmental differences between the major sections of the intestinal tract, the mucous lining that covers the epithelial cells forms a consistent layer along its internal surface (Johansson et al., 2011). Notably, this protective lubricant layer of highly glycosylated mucins serves as the initiation surface for many host–microbe interactions. Various bacteria have developed mechanisms that allow them to adhere to and to use mucus as a source of carbon and energy. In this way, these bacteria do not compete with the microbiota in the highly populated lumen and do not depend on nutrients deriving from host food consumption (Derrien et al., 2011). It has been suggested that mucus-colonizing microbes can protect the host against intestinal pathogens and contribute to restoration of the microbiota (Reid et al., 2011). A plausible mechanism for this effect is competitive exclusion. In order for indigenous bacteria to compete with transient pathogens, they must have attributes that strengthen their ability to colonize the host. For instance, human isolates of Bacteroides fragilis produce multiple capsular polysaccharides that are essential for colonization of the intestinal tract (Liu et al., 2008). These polysaccharides not only aid in persistence but also function in immune regulation, helping to exclude pathogens and restore homeostasis. Similarly, Lactobacillus rhamnosus GG that is globally marketed as a probiotic (Saxelin, 2008) carries micron-long pili that strongly bind to mucus and may give this strain a competitive advantage while outcompeting pathogens (Kankainen et al., 2009). Another example of a potentially beneficial microbe is Faecalibacterium prausnitzii. The presence of bacteria related to this species was strongly reduced in fecal and intestinal mucosal samples from patients with Crohn's disease, one of the prominent inflammatory bowel diseases (IBDs; Sokol et al., 2008, 2009; Willing et al., 2010). The type strain was found to trigger an anti-inflammatory response that decreased disease symptoms in a colitis mouse model (Sokol et al., 2008). Among the mucus-colonizing bacteria, the recently characterized Akkermansia muciniphila is highly specialized as it is capable of utilizing mucus as a sole carbon and nitrogen source (Derrien et al., 2004). Because of its abundance in healthy mucosa and the inverse correlation between its abundance and several intestinal disorders including IBD, both Crohn's disease and ulcerative colitis, and appendicitis, members of the genus Akkermansia have been suggested as biomarkers for a healthy intestine (Png et al., 2010; Swidsinski et al., 2011). Here, we will focus on the diversity and function of the intestinal microbiota with a special emphasis on the mucus-degrading Akkermansia spp. as they serve as a paradigm for establishing functions of an important and widely spread intestinal bacterium.
From diversity to function
The vertebrate intestinal microbiota is one of the most complex ecosystems on this planet. The number of microbial cells outnumber the cells in the human body and its metagenome harboring several millions of genes exceeds, by far, the number of genes from the host (Qin et al., 2010). The microbiota is also immensely diverse with over 1000 phylotypes and only includes several hundred cultured species (Sears, 2005; Rajilic-Stojanovic et al., 2007). Important aspects of diversity are the range of processes, complexity of interactions, and number of trophic levels. Moreover, it is of great importance to characterize the nature of specific host–microbe interactions and to link those to the health status of the host.
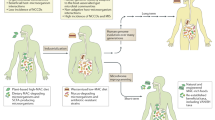
A number of ‘omics’-based approaches have been recently developed to probe the culturable and non-cultured microbial diversity and function (summarized in Figure 1b). Most of the studies addressing the microbial community structure are based on comparative sequence analyses of highly conserved genes such as the small subunit rRNA genes. Current public sequence databases contain over a million full-length 16S rRNA sequences spanning a broad phylogenetic range (Cole et al., 2009). It is important to realize that all of these omics approaches suffer from the fact that the information obtained on the macromolecules (DNA, RNA and protein) is collected at the ecosystem level. By contrast, all physiological, or other information collected from isolated strains relates to the pure culture grown in an in vitro or gnotobiotic system. What needs to be done is to integrate these two worlds of chemical and microbiological information by linking specific molecules to cells (Figure 2). This integration will be addressed here with the complexity of the human intestinal microbiota in mind and focusing on the intestinal inhabitant, Akkermansia.
Integrating omics and culturing approaches. All metagenomics and other omics approaches suffer from the fact that information on the macromolecules (DNA, RNA or protein) is collected at the ecosystem level. What needs to be done is to integrate these two worlds of predominantly chemistry and biology by linking the molecules to the cells. Red arrows indicate the experimental approaches related to this integration and the other arrows describe other activities and approaches. See text for further explanations.
Phylogenetic diversity in health and disease
Recent application of high throughput methods based on phylogenetic microarrays or parallel sequencing of barcoded small subunit rRNA amplicons, in combination with improved bioinformatics tools, have allowed the large-scale description of microbial communities in the human gut (Sogin et al., 2006; Zoetendal et al., 2008; Schloss et al., 2009). In recent years, next generation sequencing has been heavily implemented in the studies of the vertebrate intestinal microbial diversity (Dethlefsen et al., 2008; Claesson et al., 2009; Turnbaugh et al., 2009; Caporaso et al., 2010). In particular, sequencing of barcoded amplicons has extended the depth of biodiversity analyses. These studies have rapidly expanded our knowledge about the diversity of the intestinal microbiota and only a decade after their introduction, the number of detected phylotypes has outnumbered that of cultivated species by orders of magnitude. Most importantly, these high throughput sequencing methods also have provided insights into microbiota structure and population dynamics in relation to intestinal disorders such as IBD and a panoply of other diseases that include the rapidly developing obesity, diabetes and metabolic syndromes (Turnbaugh et al., 2009; Larsen et al., 2010; Willing et al., 2010). However, while these diversity-based approaches are promising in suggesting links between microbes and a certain physiological status, they do not provide causal relationships. Moreover, measures of microbial diversity and composition have to be coupled to measures of functional genes and activity measurements to gain an insight in the structural and functional diversity in the intestine. While the amount of sequence data produced by high throughput techniques is accumulating, perhaps the greatest challenge facing microbiologists today is the problem of linking the phylogeny information to function (Figure 2). Approaches such as RNA sequencing, metaproteomics, microarray and stable isotope probing allow to link metabolic conversions to phylogeny, as will be discussed below.
Function of the intestinal microbiota
Current developments in metaomics approaches provide a portal into the functional potential and activity of the intestinal microbiota. These approaches have made it possible to get a molecular snapshot at a certain time and location (Figure 1b). Notably, the genomic toolbox is rapidly expanding and has been instrumental in the generation of draft genome sequences of over 1000 human-associated microorganisms (Nelson et al., 2010) as well as an astonishing 3.3 million unique microbial genes derived from the intestinal tract of over 100 European adults (Qin et al., 2010). The human intestinal microbial metagenome further revealed unique functions carried out in the intestinal environment and elucidated novel mechanisms for signaling, vitamin production and glycan, amino-acid and xenobiotic biosynthesis.
A detailed analysis of the metagenomic sequences from the human microbiota allowed the discovery of a limited number of networks, so-called enterotypes, which are robustly found in global populations and driven by certain groups of intestinal taxa (Arumugam et al., 2011). These metagenomics developments obtained in the European MetaHIT project (Qin et al., 2010; Arumugam et al., 2011) provided new insight into the functions of the intestinal microbiota and also generated a wealth of genomic baseline information that now can be used in functional genomics.
A major limitation of DNA-based approaches is that they predict potential functions, but it is not known whether the predicted genes are transcribed and expressed or if so, under which conditions and to what extent. In addition, it is not possible to determine whether the DNA is from cells that are active and viable, dormant or even dead. These limitations can be overcome by directly probing messenger RNA or proteins.
While metatranscriptomic studies have been present applied to environmental samples, such as water and soil, only a few reports focus on the intestinal tract (Goodman et al., 2011). Two reports study cDNA microarray analyses of single bacterial species in the human intestine. These studies reveal that the transcriptome response of these bacteria depends on each other's presence and nutrient availability (Klaassens et al., 2009). Metatranscriptome analyses of the total bacterial community from human fecal material using RNAseq or cDNA-AFLP have also been reported (Booijink et al., 2010; Turnbaugh et al., 2010; Gosalbes et al., 2011). From both cDNA-AFLP and mRNA data, around 50% of the sequences were assigned to a COG category. In a recent study, RNAseq was applied on intestinal samples derived from 10 healthy individuals (Gosalbes et al., 2011). Notably, Firmicutes and Bacteroidetes transcripts turned out to be most abundant and involved in nearly all functional categories. These results further indicate that the main functional roles of the gut microbiota involved carbohydrate metabolism, energy production and synthesis of cellular components, while housekeeping activities such as amino-acid and lipid metabolism were underrepresented in the metatranscriptome. In contrast with metagenomic data, the phylogenetic composition of the active microbiota was uniform among the 10 subjects. This homogeneity even increased when clusters of genes with the same function were compared. However, as it has been shown that on average half of the cells isolated from fecal samples are dead or damaged (Ben-Amor et al., 2005), the question arises whether this material is the best source to study gene expression in the human intestine.
Metaproteomics is the study of all the proteins recovered directly from complex environmental ecosystems like the human intestine. Thus far, only few reports describe this technique to study the gut metaproteome (Klaassens et al., 2007; Verberkmoes et al., 2009; Rooijers et al., 2011). Results in these studies are in line with the predictions from the metatranscriptome and metagenome data. Most bacterial proteins were matched to Bacteriodes, Bifidobacterium or Firmicutes species, emphasizing the dominance of these groups and their functional significance in the human distal intestine (Dicksved et al., 2008; Verberkmoes et al., 2009; Rooijers et al., 2011). The main discrepancy with the metagenome data has been the skewed distribution of proteins related to translation, energy production and carbohydrate metabolism that appeared more abundant in the metaproteome data (Verberkmoes et al., 2009).
While the intestinal metaproteome and transcriptome data have been very informative, the technique's biggest challenge is to link proteins and RNA derived from a complex multiorganism environment to a non-matched metagenome data set, and hence these approaches promise to capitalize strongly on the intestinal metagenome databases that have made tremendous developments in the recent years (Qin et al., 2010). Moreover, the availability of genome sequences of intestinal isolates (Nelson et al., 2010) as well as new bioinformatic tools (Rooijers et al., 2011) will contribute to further expanding these approaches (see Figure 2).
Stable isotope probing is another technique to link metabolic conversions to phylogeny (Boschker et al., 1998; Radajewski et al., 2000; Kovatcheva-Datchary et al., 2009). While this has been successfully applied to the intestinal tract microbiota and to a limited extend used in human health (Kovatcheva-Datchary, 2010), it requires specifically labeled substrates and can only be used to address very specific questions.
Finally, metabolomic studies have been instrumental in describing microbe–host mutualism as it allows detecting and tracking diverse microbial metabolites from different non-digestible food ingredients. Moreover, metabolomics has been used for discriminating between phenotypes with different microbiota and for potentially diagnosing infection and gastrointestinal diseases, as reviewed in Holmes et al. (2011).
Culturing the unculturable
The enormous amounts of data from sequenced genomes and metagenomes are difficult to interpret as annotation of genomic sequences is often poorly supported by experimental data. The number of new microbial isolates is still increasing and various high throughput approaches have been proposed varying from culturing in germ-free animals to single cell characterization on million well plates (Ingham et al., 2007; Goodman et al., 2011). However, until recently few isolates have been sufficiently characterized at the genomic and physiologic level to directly link metabolic traits to specific microbial lineages. A big advantage of characterized isolates is that when these are detected in a certain environment, predictions can be made as to their function and behavior. We will discuss this here using A. muciniphila as an example. Remarkably, A. muciniphila is a common inhabitant of the intestine of a broad range of animals that has previously been overlooked because of its inconspicuous cell morphology, small size and specific carbon source requirements (Derrien et al., 2004, 2008; Collado et al., 2007). Hence, this case illustrates how development of alternative cultivation methods for microorganisms that appear to be inherently resistant to artificial culture remains most important.
The case of Akkermansia
A. muciniphila is a member of the Verrucomicrobia phylum and was isolated in 2004 in a quest to identify new mucus-degrading bacteria from human feces. It was named after the late Dr Antoon Akkermans, a dutch microbiologist recognized for his many contributions to microbial ecology (Akkermans et al., 1996). This bacterium appears to be a true symbiont of humans, detectable in the majority of tested subjects (Collado et al., 2007). It is one of the driving forces in two of the three recently presented human gut microbiome enterotypes (Arumugam et al., 2011) and abundantly present in the human intestinal tract, making up to 1–4% of the bacterial population in the colon (Collado et al., 2007; Derrien et al., 2008). Metagenome data suggest that at least eight different species of the Akkermansia genus colonize the intestines of humans apart from A. muciniphila, and even simultaneous colonization by different species can occur (van Passel et al., 2011). This finding is further strengthened by the fact that Akkermansia affiliated 16S rRNA sequences derived from mammalian intestinal samples form five distinct clades, four of them containing sequences associated with human gut samples. The sequence similarity between the type strain A. muciniphila and other sequences within these four clades ranges from 80% to 100%, making human colonization with different Akkermansia strains and genera plausible (Figures 3a and b; Supplementary Figure S1).
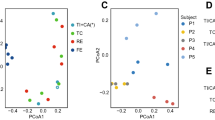
Akkermansia muciniphila is universally distributed in intestinal tracts all over the animal kingdom. (a) Phylogenetic tree indicating the position of A. muciniphila among selected full-length 16S rRNA clones from mammalian gut samples. Red colored samples derive from human sources. Thermotoga thermarum is used as an outgroup. The tree was generated using the neighbor joining method. Full details and high-resolution information are provided in Supplementary Figure S1. (b) Schematic representation of the tree in (a) with the five different clades their position and similarity to A. muciniphila. (c) Taxonomic tree of mammals generated using iTol webtool from tree of life project using all available sequences from NCBI (Letunic and Bork). Animal silhouettes indicate single species as a representative of that order. When an animal species from the mammalian orders was positive for Akkermansia-like sequences the animal logo belonging to that order is colored red, when it was negative the animal logo is colored gray. No Akkermansia sequences have been reported yet in any of the animals belonging to the mammalian orders depicted in black.
Akkermansia-like organisms are universally distributed in the intestines of the animal kingdom ranging from humans and other mammals (Turnbaugh et al., 2006; Stecher et al., 2007; Ley et al., 2008; Tsukinowa et al., 2008; Sonoyama et al., 2009, 2010; Presley et al., 2010) to non-mammals such as zebrafish (Roeselers et al., 2011), chickens (Dr O Perez, Wageningen University, personal communication) and Burmese pythons (Costello et al., 2010). Mammalian gut-derived Verrucomicrobia sequences form distinct clades within the verrucomicrobial tree and all those sequences show 80% or more similarity to A. muciniphila (Figures 3a and b). Notably, most of the other 9–14 bacterial phyla described in the mammalian gut (Eckburg et al., 2005; Rajilic-Stojanovic et al., 2007) are highly diverse in their numbers of families, genera and species with numerous different functions, niches and physiological properties.
Within the Akkermansia phylogenetic tree from mammalian-derived samples (Figure 3), clade number 1 has the lowest sequence similarity to the human-derived type strain A. muciniphila and contains no other human-associated sequences. The non-human derived sequences within this clade are highly diverse and span the complete mammalian tree (Figure 3b, clade number 1). Clade number 2 includes A. muciniphila and contains sequences with a high similarity (97–100%) to A. muciniphila. Sequences within this clade are all derived from primates (human, lemur and gorilla) and mice. Clades 3, 4 and 5 contain samples from various species of mammalian taxa, including humans. As Akkermansia affiliated 16S rRNA gene sequences have been detected in both domesticated and wild animals, this genus might be considered as an indigenous member of the microbiota within a broad range of animals. Akkermansia is not detected in all taxa within the order of the Mammalia, but the majority of these animals have yet to be sampled. It should be noted that in some animal gut microbiota studies Akkermansia sequences might have been overlooked because sequence depth was limited. Apart from mammals, Akkermansia-like sequences are also found in other vertebrates. The intestinal content of the Burmese python, zebrafish and deep-sea grenadier fish (Coryphaenoides yaquinae) retrieved from the depth of 5800 m in the northwestern Pacific Ocean, for example, is positive for Akkermansia 16S rRNA gene sequences (Costello et al., 2010; Roeselers et al., 2011) (Genbank AB591745-592640; A Nakayama and R Saito, unpublished). Clearly, the evolutionary distances and environmental differences between Akkermansia-positive animals are tremendous. The abundance and distribution of Akkermansia along the guts of animals suggests co-evolution of these bacteria with their host and their potential functionality in the intestinal tract.
The mucus-degrading abilities of A. muciniphila and its localization within the mucus layer reveal its specific niche and function within the gut (Swidsinski et al., 2009, 2011; Png et al., 2010). The A. muciniphila genome analyses predict a large secretome with over 61 (11%) of proteins to be involved in the degradation of mucin. Metaomic data sets can easily be assessed for A. muciniphila as it is the single intestinal representatives of the deeply rooted Verrucomicrobia. Proteome analyses from human fecal samples indicate that a high proportion of A. muciniphila mucus-degrading proteins are also expressed in vivo (Rooijers et al., 2011). Furthermore, in-vitro experiments have shown A. muciniphila is able to degrade both human (muc2) and porcine gastric mucus (muc5ac) (Swidsinski et al., 2009; Png et al., 2010). Clearly, the case of Akkermansia exemplifies the importance of integrating ‘omics’ approaches and functional data from isolates as summarized in Figure 2, to elucidate connections between complex ecosystems and individual constituents.
The specialization of mucin-degrading bacteria is a competitive advantage during nutrient deprivation, such as during fasting, malnutrition or total parenteral nutrition. An example of the advantages of Akkermansia adaptation to the gut mucosa can be estimated from the fact that the organism benefits from conditions of nutrient deprivation in the gut. Increased populations of A. muciniphila in the cecal contents of fasted active hamsters suggest that a lack of food-derived enteral nutrients encourages the growth of A. muciniphila (Sonoyama et al., 2009). In addition, more Verrucomicrobia exclusively of the genera Akkermansia were found in fasting Burmese phytons, notably this host is distantly related to mammals and a strict carnivore (Costello et al., 2010). Moreover, Akkermansia colonization of the colon may also be promoted by the administration of prebiotic substrates as shown in animal models (Everard et al., 2011; van den Abbeele et al., 2011). Humanized rats that were fed either inulin or arabinoxylans had increased colonal mucus levels accompanied by increased levels of Akkermansia (van den Abbeele et al., 2011). Genetic-obese mice with diet-induced leptin resistance showed an 80-fold increase in A. muciniphila levels after prebiotic, Fructooligosaccharides, administration, accompanied with increased colon length, size and enteroendocrine L cells (Everard et al., 2011).
Next to the advantages that Akkermansia spp. might have due to its ability to colonize mucus, the host probably also benefits from its presence and activity. A. muciniphila has the ability to produce acetate, and propionate as a result of mucus degradation (Derrien et al., 2004), these short chain fatty acids (SCFAs) are being produced within the mucus layer, closely to the epithelial cells (50 μm), and will therefore be easily available to the host (Derrien et al., 2011; Figure 4). The propionate produced by Akkermansia-like bacteria can signal to the host via the Gpr43 receptor and other SCFAs may also do the same via Gpr41 (Le Poul et al., 2003; Maslowski et al., 2009). This may trigger a cascade of responses in the host expression machinery and together with other signaling pathways has shown to result in immune stimulation and metabolic signaling in monoassociated germ-free mice (Derrien et al., 2011). Furthermore, A. muciniphila was the most abundantly identified mucolytic mucosa-associated bacterium in healthy controls when compared with patients with IBD (Png et al., 2010), and the amount of Akkermansia spp. was found to be inversely related to the severity of appendicitis (Swidsinski et al., 2011; Table 1), obesity (Zhang et al., 2009; Santacruz et al., 2010) and children with autism (Wang et al., 2011). The authors reporting these data suggest that A. muciniphila could be associated with a protective or anti-inflammatory role, which may be lost in IBD (Png et al., 2010). In terms of host responses, it was shown that in the epithelial transcriptomes from gnotobiotic mice colonized with A. muciniphila; cecal colonization by A. muciniphila resulted in upregulation of genes involved in antigen presentation of leukocytes. In the colon, A. muciniphila induced multiple immune response-related pathways, involved in chemotaxis and complement cascade, parts of the innate immune response, but also in cell adhesion and the maturation of B and T cells. Finally, ileal colonization by A. muciniphila led to differential expression of genes involved in metabolic and signaling pathways, mainly via modulation of PPARα-dependent processes (Derrien et al., 2011). In line with its potential function as a beneficial microbe, germ-free mice colonized by high numbers of A. muciniphila did not develop microscopically visible inflammation, nor did they show any sign of discomfort. This suggests that the transcriptional profiles obtained are involved in the regulation of immune tolerance toward A. muciniphila (Derrien et al., 2011).
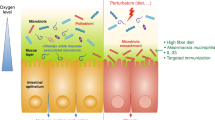
Akkermansia muciniphila activity and interactions in the intestine. Schematic overview of the metabolic activities of A. muciniphila in the gut and the microbiota and host response as a result of A. muciniphila colonization. As a result of mucus degradation, A. muciniphila produces oligosaccharides and SCFAs. These products can stimulate microbiota interactions and host response. Oligosaccharides and acetate stimulate growth and metabolic activity of bacteria that colonize close to the mucus layer. This may provide colonization resistance to pathogenic bacteria that have to cross the mucus layer to reach the intestinal cells. The propionate produced by Akkermansia-like bacteria can signal to the host via the Gpr43 receptor and other SCFA may also do the same via Gpr41 (Le Poul et al., 2003; Maslowski et al., 2009). This may trigger a cascade of responses in the host expression machinery and together with other signaling pathways has shown to result in immune stimulation and metabolic signaling in monoassociated germ-free mice (Derrien et al., 2011).
Akkermansia spp. can also be considered to contribute to a healthy mucus-associated microbiota composition. The microbiota serves as a buffer and its composition can reestablish after turbulences caused by environmental disturbances such as infectious agents. Akkermansia spp. are expected to colonize the mucus layer and initiate mucus degradation. As a result of mucus degradation, A. muciniphila produces oligosaccharides and SCFAs (Derrien et al., 2004). These products can stimulate microbiota interactions and host response. Oligosaccharides and acetate stimulate growth and metabolic activity of bacteria that colonize close to the mucus layer. This may provide colonization resistance to pathogenic bacteria that have to cross the mucus layer to reach the intestinal cells (Figure 4). It is likely that during infection or inflammation the mucus layer gets damaged and the growth of Akkermansia spp. is automatically inhibited, causing an inhibition of microbes that coexist with Akkermansia spp.
In conclusion, our present knowledge suggests human Akkermansia spp. to be important for a healthy mucus layer in the human gut with respect to mucus production and thickness. The mucus layer is continuously reshaped and refreshed and in this way a healthy environment is created for the epithelial cells that lay underneath. Apart from degradation of the luminal flow of food particles, bacteria also have an important role in the degradation of the upper loose layer of the mucus. The communication between host and bacteria creates a positive feedback loop; the production of new mucus stimulates bacterial growth and degradation stimulates mucus production. In this way, the activity of Akkermansia spp. at the mucosal surface can help to keep the mucus layer in shape.
Conclusion
The integration of the omics and culturing approaches as shown here are expected to be instrumental in advancing insight into microbial structure and function in the intestine as well as its dynamics in other ecosystems. This is described here for the intestinal tract ecosystem and is exemplified with A. muciniphila, a human intestinal isolate that is widely spread among the animal kingdom and capable of degrading mucin, a host-generated substrate.
References
Akkermans ADL, van Elsas JD, de Bruijn FJ . (1996). Molecular Microbial Ecology Manual. Kluwer Academic Publishers: Dordrecht, The Netherlands.
Arumugam M, Raes J, Pelletier E, Le Paslier D, Yamada T, Mende DR et al. (2011). Enterotypes of the human gut microbiome. Nature 473: 174–180.
Ben-Amor K, Heilig H, Smidt H, Vaughan EE, Abee T, de Vos WM . (2005). Genetic diversity of viable, injured, and dead fecal bacteria assessed by fluorescence-activated cell sorting and 16S rRNA gene analysis. Appl Environ Microbiol 71: 4679–4689.
Booijink CC, El-Aidy S, Rajilic-Stojanovic M, Heilig HG, Troost FJ, Smidt H et al. (2010). High temporal and inter-individual variation detected in the human ileal microbiota. Environ Microbiol 12: 3213–3227.
Boschker HTS, Nold SC, Wellsbury P, Bos D, de Graaf W, Pel R et al. (1998). CappenbergDirect linking of microbial populations to specific biogeochemical processes by 13C-labelling of biomarkers. Nature 392: 801–805.
Camp JG, Kanther M, Semova I, Rawls JF . (2009). Patterns and scales in gastrointestinal microbial ecology. Gastroenterology 136: 1989–2002.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7: 335–336.
Claesson MJ, O’Sullivan O, Wang Q, Nikkilä J, Marchesi JR, Smidt H et al. (2009). Comparative analysis of pyrosequencing and a phylogenetic microarray for exploring microbial community structures in the human distal intestine. PLoS One 4: e6669.
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ et al. (2009). The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37: D141–D145.
Collado MC, Derrien M, Isolauri E, de Vos WM, Salminen S . (2007). Intestinal integrity and Akkermansia muciniphila, a mucin-degrading member of the intestinal microbiota present in infants, adults, and the elderly. Appl Environ Microbiol 73: 7767–7770.
Costello EK, Gordon JI, Secor SM, Knight R . (2010). Postprandial remodeling of the gut microbiota in Burmese pythons. ISME J 4: 1375–1385.
Derrien M, Collado MC, Ben-Amor K, Salminen S, de Vos WM . (2008). The Mucin degrader Akkermansia muciniphila is an abundant resident of the human intestinal tract. Appl Environ Microbiol 74: 1646–1648.
Derrien M, van Baarlen P, Hooiveld G, Norin E, Muller M, de Vos WM . (2011). Modulation of mucosal immune response, tolerance and proliferation in mice colonized by the mucin-degrader Akkermansia muciniphila. Front Cell Infect Microbiol 2: 166.
Derrien M, Vaughan EE, Plugge CM, de Vos WM . (2004). Akkermansia muciniphila gen. nov., sp. nov., a human intestinal mucin-degrading bacterium. Int J Syst Evol Microbiol 54: 1469–1476.
Dethlefsen L, Huse S, Sogin ML, Relman DA . (2008). The pervasive effects of an antibiotic on the human gut microbiota, as revealed by deep 16S rRNA sequencing. PLoS Biol 6: e280.
Dicksved J, Halfvarson J, Rosenquist M, Jarnerot G, Tysk C, Apajalahti J et al. (2008). Molecular analysis of the gut microbiota of identical twins with Crohn's disease. ISME J 2: 716–727.
Eckburg PB, Bik EM, Bernstein CN, Purdom E, Dethlefsen L, Sargent M et al. (2005). Diversity of the human intestinal microbial flora. Science 308: 1635–1638.
Everard A, Lazarevic V, Derrien M, Girard M, Muccioli GM, Neyrinck AM et al. (2011). Responses of gut microbiota and glucose and lipid metabolism to prebiotics in genetic obese and diet-induced leptin-resistant mice. Diabetes 60: 2775–2786.
Goodman AL, Kallstrom G, Faith JJ, Reyes A, Moore A, Dantas G et al. (2011). Extensive personal human gut microbiota culture collections characterized and manipulated in gnotobiotic mice. Proc Natl Acad Sci USA 108: 6252–6257.
Gosalbes MJ, Durban A, Pignatelli M, Abellan JJ, Jimenez-Hernandez N, Perez-Cobas AE et al. (2011). Metatranscriptomic approach to analyze the functional human gut microbiota. PLoS One 6: e17447.
Holmes E, Li JV, Athanasiou T, Ashrafian H, Nicholson JK . (2011). Understanding the role of gut microbiome-host metabolic signal disruption in health and disease. Trends Microbiol 19: 349–359.
Ingham CJ, Sprenkels A, Bomer J, Molenaar D, van den Berg A, van Hylckama Vlieg JE et al. (2007). The micro-Petri dish, a million-well growth chip for the culture and high-throughput screening of microorganisms. Proc Natl Acad Sci USA 104: 18217–18222.
Johansson ME, Ambort D, Pelaseyed T, Schutte A, Gustafsson JK, Ermund A et al. (2011). Composition and functional role of the mucus layers in the intestine. Cell Mol Life Sci 68: 3635–3641.
Kankainen M, Paulin L, Tynkkynen S, von Ossowski I, Reunanen J, Partanen P et al. (2009). Comparative genomic analysis of Lactobacillus rhamnosus GG reveals pili containing a human- mucus binding protein. Proc Natl Acad Sci USA 106: 17193–17198.
Klaassens ES, Boesten RJ, Haarman M, Knol J, Schuren FH, Vaughan EE et al. (2009). Mixed-species genomic microarray analysis of fecal samples reveals differential transcriptional responses of bifidobacteria in breast- and formula-fed infants. Appl Environ Microbiol 75: 2668–2676.
Klaassens ES, de Vos WM, Vaughan EE . (2007). Metaproteomics approach to study the functionality of the microbiota in the human infant gastrointestinal tract. Appl Environ Microbiol 73: 1388–1392.
Kovatcheva-Datchary P . (2010). Linking phylogenetic identities of bacteria to starch fermentation in an in vitro model of the large intestine by RNA-based stable isotope probing. PhD thesis, Microbiology Department, Wageningen University.
Kovatcheva-Datchary P, Egert M, Maathuis A, Rajilic-Stojanovic M, de Graaf AA, Smidt H et al. (2009). Linking phylogenetic identities of bacteria to starch fermentation in an in vitro model of the large intestine by RNA-based stable isotope probing. Environ Microbiol 11: 914–926.
Larsen N, Vogensen FK, van den Berg FW, Nielsen DS, Andreasen AS, Pedersen BK et al. (2010). Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS One 5: e9085.
Le Poul E, Loison C, Struyf S, Springael JY, Lannoy V, Decobecq ME et al. (2003). Functional characterization of human receptors for short chain fatty acids and their role in polymorphonuclear cell activation. J Biol Chem 278: 25481–25489.
Ley RE, Hamady M, Lozupone C, Turnbaugh PJ, Ramey RR, Bircher JS et al. (2008). Evolution of mammals and their gut microbes. Science 320: 1647–1651.
Liu CH, Lee SM, Vanlare JM, Kasper DL, Mazmanian SK . (2008). Regulation of surface architecture by symbiotic bacteria mediates host colonization. Proc Natl Acad Sci USA 105: 3951–3956.
Maslowski KM, Vieira AT, Ng A, Kranich J, Sierro F, Yu D et al. (2009). Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature 461: 1282–1286.
Nelson KE, Weinstock GM, Highlander SK, Worley KC, Creasy HH, Wortman JR et al. (2010). A catalog of reference genomes from the human microbiome. Science 328: 994–999.
Png CW, Linden SK, Gilshenan KS, Zoetendal EG, McSweeney CS, Sly LI et al. (2010). Mucolytic bacteria with increased prevalence in IBD mucosa augment in vitro utilization of mucin by other bacteria. Am J Gastroenterol 105: 2420–2428.
Presley LL, Wei B, Braun J, Borneman J . (2010). Bacteria associated with immunoregulatory cells in mice. Appl Environ Microbiol 76: 936–941.
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C et al. (2010). A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464: 59–65.
Radajewski S, Ineson P, Parekh NR, Murrell JC . (2000). Stable-isotope probing as a tool in microbial ecology. Nature 403: 646–649.
Rajilic-Stojanovic M, Smidt H, de Vos WM . (2007). Diversity of the human gastrointestinal tract microbiota revisited. Environ Microbiol 9: 2125–2136.
Reid G, Younes JA, Van der Mei HC, Gloor GB, Knight R, Busscher HJ . (2011). Microbiota restoration: natural and supplemented recovery of human microbial communities. Nat Rev Microbiol 9: 27–38.
Reyes A, Haynes M, Hanson N, Angly FE, Heath AC, Rohwer F et al. (2010). Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature 466: 334–338.
Roeselers G, Mittge EK, Stephens WZ, Parichy DM, Cavanaugh CM, Guillemin K et al. (2011). Evidence for a core gut microbiota in the zebrafish. ISME J 5: 1595–1608.
Rooijers K, Kolmeder C, Juste C, Dore J, de Been M, Boeren S et al. (2011). An iterative workflow for mining the human intestinal metaproteome. BMC Genomics 12: 6.
Santacruz A, Collado MC, Garcia-Valdes L, Segura MT, Martin-Lagos JA, Anjos T et al. (2010). Gut microbiota composition is associated with body weight, weight gain and biochemical parameters in pregnant women. Br J Nutr 104: 83–92.
Saxelin M . (2008). Probiotic formulations and applications, the current probiotics market, and changes in the marketplace: a European perspective. Clin Infect Dis 46 (Suppl 2): S76–S79; discussion S144–S151.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB et al. (2009). Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75: 7537–7541.
Sears CL . (2005). A dynamic partnership: celebrating our gut flora. Anaerobe 11: 247–251.
Sogin ML, Morrison HG, Huber JA, Mark Welch D, Huse SM, Neal PR et al. (2006). Microbial diversity in the deep sea and the underexplored ″rare biosphere″. Proc Natl Acad Sci USA 103: 12115–12120.
Sokol H, Pigneur B, Watterlot L, Lakhdari O, Bermudez-Humaran LG, Gratadoux JJ et al. (2008). Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc Natl Acad Sci USA 105: 16731–16736.
Sokol H, Seksik P, Furet JP, Firmesse O, Nion-Larmurier I, Beaugerie L et al. (2009). Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm Bowel Dis 15: 1183–1189.
Sonoyama K, Fujiwara R, Takemura N, Ogasawara T, Watanabe J, Ito H et al. (2009). Response of gut microbiota to fasting and hibernation in Syrian hamsters. Appl Environ Microbiol 75: 6451–6456.
Sonoyama K, Ogasawara T, Goto H, Yoshida T, Takemura N, Fujiwara R et al. (2010). Comparison of gut microbiota and allergic reactions in BALB/c mice fed different cultivars of rice. Br J Nutr 103: 218–226.
Stecher B, Robbiani R, Walker AW, Westendorf AM, Barthel M, Kremer M et al. (2007). Salmonella enterica serovar typhimurium exploits inflammation to compete with the intestinal microbiota. PLoS Biol 5: 2177–2189.
Swidsinski A, Dorffel Y, Loening-Baucke V, Theissig F, Ruckert JC, Ismail M et al. (2011). Acute appendicitis is characterised by local invasion with Fusobacterium nucleatum/necrophorum. Gut 60: 34–40.
Swidsinski A, Loening-Baucke V, Herber A . (2009). Mucosal flora in Crohn's disease and ulcerative colitis - an overview. J Physiol Pharmacol 60 (Suppl 6): 61–71.
Tsukinowa E, Karita S, Asano S, Wakai Y, Oka Y, Furuta M et al. (2008). Fecal microbiota of a dugong (Dugong dugong) in captivity at Toba Aquarium. J Gen Appl Microbiol 54: 25–38.
Turnbaugh PJ, Hamady M, Yatsunenko T, Cantarel BL, Duncan A, Ley RE et al. (2009). A core gut microbiome in obese and lean twins. Nature 457: 480–484.
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI . (2006). An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444: 1027–1031.
Turnbaugh PJ, Quince C, Faith JJ, McHardy AC, Yatsunenko T, Niazi F et al. (2010). Organismal, genetic, and transcriptional variation in the deeply sequenced gut microbiomes of identical twins. Proc Natl Acad Sci USA 107: 7503–7508.
Van den Abbeele P, Gerard P, Rabot S, Bruneau A, El Aidy S, Derrien M et al. (2011). Arabinoxylans and inulin differentially modulate the mucosal and luminal gut microbiota and mucin-degradation in humanized rats. Environ Microbiol 13: 2667–2680.
van Passel MW, Kant R, Zoetendal EG, Plugge CM, Derrien M, Malfatti SA et al. (2011). The genome of Akkermansia muciniphila, a dedicated intestinal mucin degrader, and its use in exploring intestinal metagenomes. PLoS One 6: e16876.
Verberkmoes NC, Russell AL, Shah M, Godzik A, Rosenquist M, Halfvarson J et al. (2009). Shotgun metaproteomics of the human distal gut microbiota. ISME J 3: 179–189.
Wang L, Christophersen CT, Sorich MJ, Gerber JP, Angley MT, Conlon MA . (2011). Low relative abundances of the mucolytic bacterium Akkermansia muciniphila and Bifidobacterium spp. in feces of children with autism. Appl Environ Microbiol 77: 6718–6721.
Willing BP, Dicksved J, Halfvarson J, Andersson AF, Lucio M, Zheng Z et al. (2010). A pyrosequencing study in twins shows that gastrointestinal microbial profiles vary with inflammatory bowel disease phenotypes. Gastroenterology 139: 1844–1854 e1.
Zhang H, DiBaise JK, Zuccolo A, Kudrna D, Braidotti M, Yu Y et al. (2009). Human gut microbiota in obesity and after gastric bypass. Proc Natl Acad Sci USA 106: 2365–2370.
Zoetendal EG, Rajilic-Stojanovic M, de Vos WM . (2008). High-throughput diversity and functionality analysis of the gastrointestinal tract microbiota. Gut 57: 1605–1615.
Acknowledgements
We are grateful to our colleagues and collaborators for stimulating discussions. This work was partly funded by grant 250172-MicrobesInside of the European Research Council.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Belzer, C., de Vos, W. Microbes inside—from diversity to function: the case of Akkermansia. ISME J 6, 1449–1458 (2012). https://doi.org/10.1038/ismej.2012.6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2012.6
Keywords
This article is cited by
-
Akkermansia muciniphila and its membrane protein ameliorates intestinal inflammatory stress and promotes epithelial wound healing via CREBH and miR-143/145
Journal of Biomedical Science (2023)
-
Akkermansia muciniphila, which is enriched in the gut microbiota by metformin, improves cognitive function in aged mice by reducing the proinflammatory cytokine interleukin-6
Microbiome (2023)
-
Metagenome and metabolome insights into the energy compensation and exogenous toxin degradation of gut microbiota in high-altitude rhesus macaques (Macaca mulatta)
npj Biofilms and Microbiomes (2023)
-
Colonization with ubiquitous protist Blastocystis ST1 ameliorates DSS-induced colitis and promotes beneficial microbiota and immune outcomes
npj Biofilms and Microbiomes (2023)
-
Relationship between obesity-related colorectal tumors and the intestinal microbiome: an animal-based trial
Journal of Cancer Research and Clinical Oncology (2023)