Abstract
Sponges harbour complex communities of diverse microorganisms, which have been postulated to form intimate symbiotic relationships with their host. Here we unravel some of these interactions by characterising the functional features of the microbial community of the sponge Cymbastela concentrica through a combined metagenomic and metaproteomic approach. We discover the expression of specific transport functions for typical sponge metabolites (for example, halogenated aromatics, dipeptides), which indicates metabolic interactions between the community and the host. We also uncover the simultaneous performance of aerobic nitrification and anaerobic denitrification, which would aid to remove ammonium secreted by the sponge. Our analysis also highlights the requirement for the microbial community to respond to variable environmental conditions and hence express an array of stress protection proteins. Molecular interactions between symbionts and their host might also be mediated by a set of expressed eukaryotic-like proteins and cell–cell mediators. Finally, some sponge-associated bacteria (for example, a Phyllobacteriaceae phylotype) appear to undergo an evolutionary adaptation process to the sponge environment as evidenced by active mobile genetic elements. Our data clearly show that a combined metaproteogenomic approach can provide novel information on the activities, physiology and interactions of sponge-associated microbial communities.
Similar content being viewed by others
Introduction
Marine sponges represent a significant component of the marine, benthic communities throughout the world. Sponges harbour diverse communities of microorganisms, which often form stable and specific associations with their symbiotic host (Taylor et al., 2007; Webster et al., 2008; Zhu et al., 2008). While much progress has been made over the past decade in defining the phylogenetic diversity and patterns of sponge-associated microbial communities (Taylor et al., 2007; Webster and Taylor, 2011), information on the function of individual symbionts or the microbial community as a whole is limited.
Examples where specific members have been assigned functional roles include cyanobacterial symbionts, which can provide photosynthetically fixed carbon to the sponge host (Wilkinson, 1983) and the bacterial production of biologically active metabolites that may play a role in host defence (Unson et al., 1994; Schmidt et al., 2000). The processes of nitrification/denitrification and anaerobic ammonium oxidation (anammox) have been well investigated in the sponges Geodia barretti (Hoffmann et al., 2009b), Dysidea avara and Chondrosia reniformis (Schlappy et al., 2010) using stable isotope experiments. In addition, by the identification of 16S rRNA gene sequences, the anammox process was putatively linked to planctomycetes in a reef sponge (Mohamed et al., 2010). These approaches require, however, an a priori knowledge of the processes performed in the sponge holobiont or the establishment of an irrevocable link between microbial phylogeny and function. Combining these limitations with the inherent difficulty of culturing (potentially obligate) symbionts has left the field of sponge microbiology with a rudimentary understanding of the ecological functions of sponge-associated microorganisms and the nature of host–symbiont interactions (Webster and Blackall, 2008).
The whole-community approaches of metagenomics, metatranscriptomics and metaproteomics provide a promising avenue to explore the function of uncultured organisms and for substantially advancing the field of sponge–microorganism symbiosis research. For example, our recent study using metagenomics on the microbial community associated with the sponge Cymbastela concentrica led to the recognition of many novel genomic markers that may provide specific mechanisms for bacteria to persist within and interact with their sponge host (Thomas et al., 2010). To further explore the genetic potential offered by metagenomic data sets, metaproteomics is being increasingly employed to describe the expressed protein profile of microbial communities. Low-diversity microbial systems, such as those of acid mine drainage (Ram et al., 2005) and lake water from Antarctica (Ng et al., 2010), and also more complex systems in waste water sludge (Wilmes et al., 2008), the human microbiome (Chen et al., 2008), the hindgut microbiome (Burnum et al., 2011) and open ocean (Sowell et al., 2009, 2011; Morris et al., 2010) have been studied in this way. For these systems, the combination of high-throughput protein mass spectrometry with extensive metagenomic data sets has provided novel and direct insights into functions expressed by microorganisms.
To further our understanding of microbial functions in sponges, we report here an integrated approach of using metagenome sequencing and metaproteomics on the microbial community associated with C. concentrica, an abundant marine sponge found in shallow, temperate waters of the Australian east coast. This sponge contains a stable and diverse microbial community, with predominantly uncultured phylotypes belonging to the Gammaproteobacteria, Phyllobacteriaceae, Sphingomondales, Piscirickettsiaceae and Deltaproteobacteria among others (Thomas et al., 2010). We show in the microbial community the expression of transport functions relevant to host-derived nutrients, aerobic and anaerobic metabolism, stress responses for the adaptation to variable conditions inside the sponge, as well as proteins that could facilitate a direct molecular interaction between the symbionts and the host. Our data also reveal specific protein expression by a Phyllobacteriaceae bacterium and a Nitrosopumilus-like crenarchaeon, thus linking particular functions to uncultured phylotypes.
Materials and methods
Sampling, cell separation and preparation
Triplicate samples of C. concentrica were collected by SCUBA diving from Botany Bay, near Bare Island, Sydney, Australia (S 33.59.461; E 151.13.946) at 1000 hours on 15 September 2009. Sponges were directly transferred upon surfacing into sterile calcium- and magnesium-free seawater with protease inhibitor and transported on ice back to the laboratory (15 min drive) for direct processing (see Supplementary Information for details). The microbial fraction was collected from each sponge sample through a series of centrifugation and filtration steps (see Supplementary Information for details). Three microbial cell pellets were collect separately and kept at −80 °C till further extraction.
DNA was extracted from the cell pellets according to the procedure previously described in Thomas et al. (2010). For protein preparation, each cell pellet from the triplicate samples was processed separately through a one-dimensional sodium dodecyl sulphate-polyacrylamide gel electrophoresis gel and the sample lane was sliced into 12 fractions. Protein fractions were reduced, alkylated and digested overnight with trypsin (see Supplementary Information for details).
DNA sequencing, assembly and functional annotation
Extracted microbial DNA for each biological triplicate was sequenced separately using the Roche 454 Titanium platform at the J. Craig Venter Institute (Rockville, MD, USA) and used for the generation of metagenomic data sets to accompany the metaproteome analysis in this study. For functional annotation, protein sequences were searched against the Clusters of Orthologous Group (COG) (Tatusov et al., 2003), the Pfam database for protein families (Finn et al., 2010) and CDD for conserved protein domains (Marchler-Bauer et al., 2011). A subsystem-based approach to genome annotations as implemented in the SEED platform was also utilized to help reveal metabolic pathways (Overbeek et al., 2005). The details of sequencing assembly and functional annotation are outlined in the Supplementary Information. The shotgun sequencing is available through the Community Cyberinfrastructure for Advanced Marine Microbial Ecology Research and Analysis (CAMERA) website (http://camera.calit2.net/) under project accession ‘CAM_PROJ_BotanyBay’.
High-performance liquid chromatography, mass spectrometry and data analysis
The digested peptides of each biological replicate were analysed by online nano-liquid chromatography–tandem mass spectrometry on an LTQ FT Ultra (Thermo Electron, Karlsruhe, Germany). Each peptide sample was subjected to three separate liquid chromatography–tandem mass spectrometry analyses and this resulted in a total of 108 sample runs (3 sponge samples, 12 slices per sample and 3 runs per sliced sample) (see Supplementary Information for details). Peak lists of these samples were generated using Mascot Daemon/extract_msn (Matrix Science, Thermo, London, UK) using the default parameters (version 2.3; Matrix Science).
All 108 spectrometry spectra were analysed using Mascot (version 2.3; Matrix Science), Sequest (Thermo Fischer Scientific, San Jose, CA, USA; version 1.0.43.0) and X! Tandem (The GPM, thegpm.org; version 2007.01.01.1) with an ion mass tolerance of 0.40 Da and a parent ion tolerance of 4.0 p.p.m. against the metagenomic database (Supplementary Information for details). Sequest searches were performed with a fragment ion mass tolerance of 0.50 Da and a parent ion tolerance of 3.0 p.p.m.
Spectra for each sponge sample were pooled together (36 spectra per sample) and loaded into Scaffold (version Scaffold_2_00_05; Proteome Software Inc., Portland, OR, USA) as categorical samples for analysis. Peptide and protein identifications were performed as outlined in the Supplementary Information and resulted in a false discovery rate, as estimated by searches against a decoy database, of below 1%.
Mass spectrometry spectra for the metaproteomic analysis are available through the PRIDE database (www.ebi.ac.uk/pride; accession numbers: 20959–20969) and raw peptide identifications are given in Supplementary Table S3.
Phylogenetic analysis of 16S rRNA genes and a crenarchaeal ammonia monooxygenase subunit-α
The 16S rRNA gene sequence of the Phyllobacteriaceae phylotype was taken from the previously classified partial genome from C. concentrica (Thomas et al., 2010), which was also found in the current metagenomic data set. The 16S and α-subunit of the ammonia monooxygenase (AmoA) sequence for the Nitrosopumilus-like crenarchaeon was extracted from the current metagenomic contigs. Phylogenetic analysis of both genes is described in the Supplementary Information.
Localisation of the Phyllobacteriaceae phylotype
To localise the Phyllobacteriaceae bacterium in the sponge tissue, fresh C. concentrica samples were collected (as above) and rinsed with calcium- and magnesium-free seawater and immediately transferred to 15% sucrose solution. The specimens were then transferred back to the laboratory on ice within 1 h. Fluorescence in situ hybridization was performed as described in Liu et al. (2011) and in the Supplementary Information with a validated probe targeting the Phyllobacteriaceae phylotype.
Reconstruction of partial genomes and comparative genomics
Assembly and binning of Sanger-shotgun sequencing data from the metagenome of C. concentrica allowed for the reconstruction of five partial genomes from distinct bacterial lineages based on the binning of the tetranucleotide patterns (assigned to the Phyllobacteriaceae, Piscirickettsiaceae, Gammaproteobacteria, Sphingomondales and Deltaproteobacteria) (Thomas et al., 2010). These partial genomes were further expanded by binning of the current pyrosequencing data using PhymmBL (Brady and Salzberg, 2009). We searched all the proteins identified in the metaproteome against the five partial genomes using both protein similarity and contig classification (see Supplementary Information for details).
Statistical, pairwise comparison of the COG category profile and individual COGs between the proteome and partial genome data were performed by re-sampling (n=1000) of COG subsamples (n=60) as outlined in Lauro et al. (2009).
Results and discussion
Overview of metaproteogenomic data
Sequencing of DNA extracted from the microbial communities associated with three samples of C. concentrica resulted in a total of 2.8 million unique sequencing reads, which assembled into 988 317 contigs or singletons bigger than 100 nt. Of those, 687 588 passed a filtering procedure for eukaryotic contamination. For each of the three sponge samples, we identified 342 235, 550 559 and 555 480 proteins, respectively, of which an average 235 288, 208 852, 209 592 and 284 549 could be annotated to CDD, COG, PFAM and SEED, respectively (see Supplementary Table S2).
The three proteomic data sets had a total of 765 non-redundant proteins identified from 5275 peptide fragments. Taxonomic analysis indicated that 139 proteins were possibly from eukaryotic origin leaving 626 proteins for further functional characterization. Of these, 367, 364 and 395 proteins were found in each of the three individual C. concentrica samples and 186 proteins were common to all three samples (Figure 1) (see Supplementary Table S1). Protein sequences were annotated and clustered together into functional categories. This showed that the expressed proteins in the sponge contained a substantial number of proteins (34%) with no assignment, or of hypothetical nature, suggesting the presence of many unrecognized functions in the sponge's microbial community.
To further compare the expressed functional profile of the microbial communities with their underlying genetic potential, we analysed the metaproteomic and metagenomic data sets based on COG functional categories (Figure 2). Specific COGs that were relatively over-represented in the metaproteome included carbohydrate transport and metabolism, post-translational modification, protein turnover, chaperone functions and signal transduction. A relative under-representation was observed for functional groups associated with coenzyme transport and metabolism, transcription, translation, replication recombination and repair, when compared against the metagenome data set. Further analysis at the individual COG level indicated an abundance (both in terms of proteins and peptides detected) of chaperonin GroEL (HSP60 family) (COG0459), an array of specific transporters (COG0747, COG0834, COG4663, COG0683, COG1653), dehydrogenases with different specificities, FabG (COG1028) and outer membrane receptor proteins CirA and OmpA (COG1629, COG2885) (Figure 3). Analysis of these specific functions within each functional category allowed us to specify physiological properties and activities of the sponge's microbial community.
Relative abundance of COG categories in sponge-associated microbial community based on metaproteome and metagenome data. COG counts were normalized and percentage of total counts in each COG categories is presented here. The error bars show calculated standard variation of triplicate samples and *indicate a statistical significant with a P-value <0.05 in a t-test.
Active transport systems involved in nutrient acquisition
The sponge-associated community showed an abundant expression of high-affinity and broad-specificity uptake systems, like ATP-binding cassette (ABC) transporters and tripartite ATP-independent periplasmic (TRAP) transporters. The most abundant transporter components detected in the metaproteome were periplasmic substrate-binding domains associated with ABC transporters, in particular for amino acids. DppA (COG0747) (Figure 3) is the substrate-binding component of the DppABCDEF dipeptide transport system, which has been demonstrated to transport proline-containing dipeptides (Olson et al., 1991). Proline-containing dipeptides have been previously isolated from marine sponges (Liu et al., 2009), however, the exact production source (sponge or symbiont) is not known. Nevertheless, DppA-type transporters were not noted to be abundant in the metaproteome of planktonic bacteria (Sowell et al., 2009, 2011), highlighting clear nutritional differences in the free-living and sponge-associated environment. Dipeptide transporters of the DppA type have also been found to be capable of transporting heme and the heme precursor (Letoffe et al., 2006), indicating the potential scavenging of these iron-containing compounds from the surrounding or the sponge host. Other abundant proteins detected were HisJ and LivK, which are the periplasmic components of the high-affinity histidine- and leucine-specific transport system, respectively.
The over-representation of the COG category carbohydrate transport and metabolism (Figure 2) is mainly due to a high number of proteins associated with the glycerol-3-P ABC-type transporter UgpB (COG1653) and the TRAP system DctP (COG1638). The UgpB is the periplasmic binding protein of the glycerol-3-phosphate uptake system (Brzoska et al., 1994), which transports glycerol-3-phosphate for use as a carbon source and/or phosphate source. However, glycerol-3-phosphate uptake through the Ugp system is not able to supply enough carbon for bacterial growth, but instead increases the internal phosphate concentration (Boos, 1998). Therefore, the Ugp system is ideally geared for scavenging phosphate-containing compounds (Boos, 1998). The DctP system is well characterized in Rhodobacter capsulatus (Forward et al., 1997) and has also been found in Wolinella succinogenes (Ullmann et al., 2000), where it is responsible for the transport of the C4-dicarboxylates fumarate and malate (Ullmann et al., 2000). Structurally, the known TRAP substrates are united by the presence of a carboxylate group and this means that many hundreds of organic acids could be potentially transported by the system (Mulligan et al., 2011). Large numbers of evolutionarily diverse TRAP transporters have been found in marine environments, especially in the SAR11 clade (Morris et al., 2002), and this suggests an important role in the transport of diverse substrates.
Another TRAP transporter expressed in the sponge-associated community belongs to the FcbT1 type, which has been shown in Comamonas sp. DJ-12 to transport halogenated, aromatic substrates, such as 4-chlorobenzoate (Chae et al., 2000). The genetic organization for this TRAP transporter is an operon-encoding enzyme responsible for hydrolytic dechlorination of 4-chlorobenzoate, which can be further metabolized to succinyl-CoA and acetyl-CoA (Nichols and Harwood, 1995). The fcbT genes can also be induced by benzoate derivatives, like 4-bromobenzoate, indicating a wider potential substrate range of aromatic compounds (Chae et al., 2000). Marine holobionts, including algae (Pedersen et al., 1974), jellyfish (White and Hager, 1977), polychaetes (Ashworth and Cormier, 1967) and sponges (Schmitz and Gopichand, 1978) have been recognized as a rich source of naturally occurring halogenated compounds, many of which have antibiotic or antagonistic activities. A number of sponge species, such as Psammopemma sp., Psammaplysilla purpurea, Aplysina aerophoba and Dysidea herbacea, produce brominated aromatic metabolites, including bromoindoles, bromophenol, polybrominated diphenyl ethers and dibromodibenzo-p-dioxins (Norte et al., 1988; Ebel et al., 1997; Gribble, 1999; Utkina et al., 2001). These observations are therefore consistent with the uptake of halogenated aromatic compounds by the sponge's symbionts. Alternatively, the bidirectional nature of TRAP transporters (Poolman and Konings, 1993) could facilitate export, making the symbionts the actual producers of the halogenated aromatics. Whichever way, our data show an intimate symbiotic relationship of the sponge with its symbionts through the transport of halogenated aromatic compounds.
TonB-dependent transporters were also expressed by the sponge-associated microbial community and this has also been observed for the microbial membrane metaproteome of samples from the South Atlantic (Morris et al., 2010). Specifically, we detected the outer membrane receptor proteins CirA and OmpA, which utilize a proton motive force to transport nutrients across the outer membrane of Gram-negative bacteria. Genome studies on bacterioplankton have revealed TonB-dependent transporters to be enriched among marine bacterial species (Giovannoni and Stingl, 2007). The transport activities of TonB-dependent transporters were thought to be restricted to iron complexes (siderophores) and vitamin B12 (cobalamin), but recent experimental and bioinformatic studies indicate that nickel, cobalt, copper, maltodextrins, sucrose, thiamin and chito-oligosaccharides are also suitable substrates (Schauer et al., 2008).
Overall, the sponge-associated community clearly expressed a large number of transporters for the acquisition of various substrates, and in this respect behaved similar to the planktonic, bacterial communities from oligotrophic open oceans and productive coastal ecosystems (Sowell et al., 2009, 2011; Morris et al., 2010). Despite these broad similarities, there were clear differences in transport (for example, dipeptides, halogenated aromatics) that reflect the nutrients specific for the microhabitats of the sponge.
Stress response
The abundance of expressed proteins associated with post-translational modification, protein turnover and chaperone functions (Figure 2) reflected a high number of chaperone proteins GroEL (HSP60, COG0459), membrane proteases HflC (COG0330) and DnaK (COG0443). These chaperones and proteases are essential for the elimination of denatured or damaged proteins, which could result from stress conditions, such as temperature shifts, osmotic pressure, presence of reactive oxygen species and toxic compounds. In addition, the peptide methionine sulphoxide reductase MsrA, which repairs proteins that have been inactivated by oxidation (Ezraty et al., 2005), is expressed. We also detected a number of proteins that are annotated to be heme-dependent peroxidases as well as the superoxide dismutase SodA, which eliminates harmful oxidation products like hydrogen peroxide (Perry et al., 2010). In addition, peroxiredoxin (COG0450) and glutathione-S-transferases (COG0625) were expressed and these proteins might be important to control cytoplasmic redox balance (Vuilleumier, 1997; Hofmann et al., 2002). The choline dehydrogenase BetA was also detected, which catalyzes the oxidation of choline to glycine betaine (Landfald and Strom, 1986). Betaines are potent and frequently used osmolytes that ensure osmotic balance in the cytoplasm and their production are often induced by osmotic stress.
Stress-related functions have previously been noted to be abundant in the genomes of bacteria associated with C. concentrica, when compared with planktonic bacteria of the surrounding water (Thomas et al., 2010). The cycling pumping activity of sponges (Vogel, 1977) and steep local gradients (Hoffmann et al., 2009a) would expose bacteria to variable environmental conditions, in terms of availability of nutrients and electron acceptors (for example, oxygen), and here we show that symbionts of C. concentrica are indeed dealing with such fluctuations in an evolutionary (abundance of genes) and physiological way (expression of those genes).
Metabolism
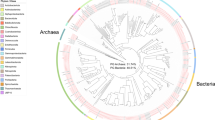
The nitrogen metabolism of bacterial and archaeal symbionts is closely linked to the sponge host, which secretes and accumulates ammonium (Taylor et al., 2007; Bayer et al., 2008; Hoffmann et al., 2009b; Schlappy et al., 2010). It is therefore not surprising that we detected the expression of the ammonia monooxygenase membrane-bound subunits β and γ (AmoB and C) and an ammonia transporter (AmtB) in the microbial community of C. concentrica (see Supplementary Table S1). Both sequences were most closely related (BlastN identity: 92%) to those of the marine crenarchaeon Nitrosopumilus maritimus (Walker et al., 2010). The genes encoding AmoB and C were adjacent and orientated in opposite transcriptional directions on a contig of the C. concentrica metagenome (Supplementary Figure S1a). The contig also contains a gene for the α-subunit (amoA) and has overall striking synteny with a genomic region of N. maritimus (Supplementary Figure S1a, b). Phylogenetic analysis of AmoA further confirmed the close relationship with N. maritimus (see Supplementary Figure 2). Interestingly, putative nitric oxide reductase subunits (NorQ and NorD) are also encoded in the genomic region (Supplementary Figure S1a, b) and might have a particular role in determining tolerance to nitric oxides under limiting oxygen concentrations or to allow for the use of nitrous oxide as an alternative electron acceptor (Schmidt and Bock, 1997).
N. maritimus belongs in the C1a-α subgroup of the crenarchaeal Marine Group I, which contains many sequences obtained from sponges and plankton (Holmes and Blanch, 2007). To further define the phylogeny for the Nitrosopumilus-like crenarchaeon in C. concentrica, we identified a 16S rRNA gene sequence in a metagenomic contig (Supplementary Figure S1c) and constructed its phylogeny with other selected Group C1a-α members. This C. concentrica-derived crenarchaeal sequence clustered with a mix of 16S rRNA sequences from sponge symbionts and free-living archaea (see Supplementary Figure 3). Taken together, our data show that aerobic nitrification and transport of ammonia are active in C. concentrica and that those functions are being carried out by a Nitrosopumilus-like crenarchaeon, which is a representative of common symbiotic and planktonic archaea. These results also indicate that members of this nitrifying, archaeal clade might exist in either a host-associated or a free-living form, which could explain why crenarchaeal sequences were not detected in a previous metagenomic analysis of C. concentrica (Thomas et al., 2010).
Given that both aerobic and anaerobic conditions might exist within microhabitats of the sponge tissue, we queried our metaproteomic data set for functions related to anaerobiosis. We found the expression of proteins annotated to COG0076 (glutamate decarboxylase and related pyridoxal 5-phosphate-dependent proteins) (Supplementary Table S1), which is part of the glutamate-dependent acid resistance systems (AR2). The AR2 system protects cells during anaerobic phosphate starvation, when glutamate is available, by preventing damage from weak acids produced from carbohydrate fermentation. Although no proteins directly associated with carbohydrate fermentation were detected, acetoacetate decarboxylase was expressed (Supplementary Table S1), which is involved in the solventogenesis of the typical fermentation products butyric and acetic acid into acetone and butanol (Schaffer et al., 2002). Anaerobic degradation of amines and polyamines may also occur as we detected the expression of crotonobetainyl-CoA hydratase (CaiD) (COG1024) (Supplementary Table S1), which is part of the carnitine degradation pathway (Elssner et al., 2001). In Escherichia coli, this pathway, which includes the dehydration and reduction of L-carnitine to γ-butyrobetaine, is induced during anaerobic growth. The carnitine pathway has also been found to generate the osmoprotectant betaine during anaerobic respiration (Kleber, 1997; Preusser et al., 1999; Elssner et al., 2001). Recent analysis of the anaerobic carnitine reduction in E. coli has also shown that the electron transfer of flavoproteins FixA and FixB are necessary for the transfer of electrons to crotonobetaine reductase (CaiA) (Walt and Kahn, 2002) and we found FixA expressed in the sponge community (Supplementary Table S1). Taken together, our data show the existence of both aerobic and anaerobic metabolism in the sponges and future work could target the exact localisation of these processes.
Molecular symbiont–host interactions
A substantial over-representation of ankyrin repeat (AR) and tetratricopeptide proteins was observed in our recent metagenomic study of the bacterial communities associated with C. concentrica (Thomas et al., 2010) and the genes for these eukaryotic-like proteins also clustered in the genome of an uncultured, deltaproteobacterial symbiont of the same sponge (Liu et al., 2011). Genes encoding AR proteins have been reported to be abundant in the genomes of obligate and facultative symbionts, such as Wolbachia pipientis (Iturbe-Ormaetxe et al., 2005), Ehrlichia canis (Mavromatis et al., 2006), Legionella pneumophila (Habyarimana et al., 2008) and Coxiella butnetii (Voth et al., 2009), and might play a role in mediating host–bacteria associations. This has been highlighted recently by a mutant study of L. pneumophila, which showed that certain AR proteins are controlling its intracellular replication within the amoebal host (Habyarimana et al., 2008). An AR protein (COG0666) and tetratricopeptide protein (PFAM00515) were found in the metaproteomic data set (Supplementary Table S1), showing that sponge-associated microorganisms indeed express those proteins, potentially for mediating interactions with the sponge host, as was proposed previously (Thomas et al., 2010). Other proteins with roles in bacteria–eukaryote interactions were also expressed, including proteins with Hep/Hag domains (Supplementary Table S1). This seven-residue repeat domain of Hep/Hag is contained in the majority of the sequences of bacterial hemagglutinins and invasins. The adhesin YadA was also detected in the metaproteome and this protein has been shown to be responsible for phagocytosis resistance (Nummelin et al., 2004). Sponges are filter feeders, and the presence of such eukaryotic-like proteins and bacteria–eukaryote mediators suggests that sponge symbionts might use these proteins to escape phagocytosis and/ or control their symbiotic relationship with their hosts.
Linking phylotype to function–expression profiling of an uncultured Phyllobacteriaceae
Binning of our previous sequencing data for the metagenome of C. concentrica (Thomas et al., 2010) and our current pyrosequencing data set allowed for the reconstruction of partial genomes of five uncultured sponge symbiont, that is, phylotypes belonging to the Phyllobacteriaceae, Piscirickettsiaceae, Gammaproteobacteria, Sphingomondales and Deltaproteobacteria (Thomas et al., 2010) (see also Materials and methods section). We could confidently assign 65 proteins to the Phyllobacteriaceae genome, 17 to Piscirickettsiaceae, 15 to Gammaproteobacteria, one to Sphingomondales and one to Deltaproteobacteria. As the numbers of hits against the Phyllobacteriaceae genome was the highest, we further investigated its expression profile and genomic features.
Detailed phylogenetic analysis (see Materials and methods section) revealed that the 16S rRNA gene sequence within the Phyllobacteriaceae partial genome forms, together with two sequences previously amplified from C. concentrica (AY942778 and AY942764), a distinct clade that is deeply branched within the Phyllobacteriaceae family (see Supplementary Figure S4). A large majority of members of the Phyllobacteriaceae are plant associated and have been well studied for their potential to promote plant growth (Mantelin et al., 2006). They also occupy diverse habitats, such as soil (Jurado et al., 2005), water (Mergaert et al., 2001) and unicellular organisms (Alavi et al., 2001), suggesting a remarkable adaptive capacity to the environment. The sponge sequences of this study are also related to nitrogen-fixing Mesorhizobium species and denitrifying Nitratireductor species (Labbe et al., 2004), which could imply that the Phyllobacteriaceae phylotype in C. concentrica is potentially involved in nitrogen metabolism.
We compared the proteome assigned to the Phyllobacteriaceae phylotype with its partial genome and found over- and under-representation of functional COG categories (Figure 4a) similar to the overall metaproteome and metagenome comparison (Figure 2). Again COG categories of amino-acid transport and metabolism, post-translational modification, protein turnover, chaperone functions and signal transduction were over-represented in the expressed proteome data (Figure 4a). At the individual COG level, specific transport functions were over-represented (Figure 4b), with both ABC- and TRAP-type transporters being expressed. Of particular interest was the ABC-type nitrate/sulphonate/bicarbonate transport systems TauA, which can import nitrate across the cell membrane. A nitrate reductase gene cluster (narG, narH, narI and narY) was present in the partial genome of the Phyllobacteriaceae phylotype highlighting its potential for denitrification, however, the expressed proteins nitrate reductase NarG and NarY in our metaproteome data set (see Supplementary Table S1) could not be unambiguously assigned to this organism. Nevertheless, the proteomic data suggest that denitrification is actively expressed in the microbial community of C. concentrica and most likely take place in the anaerobic, inner part of the sponge (Hoffmann et al., 2008). We therefore investigated the physical location of the Phyllobacteriaceae phylotype by fluorescence in situ hybridization. No fluorescence in situ hybridization signal was observed near the outer surface of the sponge and the Phyllobacteriaceae phylotype was found to reside within the mesohyl, mainly near the chambers, where feeding takes place in sponges (see Supplementary Figure S5).
Comparison between expressed proteome and partial genome of the Phyllobacteriaceae phylotype on the level of COG categories (a) and COG level (b). Black and grey bars represent over- and under-representation, respectively, of identified proteins. The x axis displays the median value with significant cutoff value of −1 and 1.
We also assigned an expressed transposase gene to this Phyllobacteriaceae phylotype and its chromosomal location was flanked by two genes associated with lipid metabolism (cyclopropane-fatty-acyl-phospholipid synthase and acyl-CoA dehydrogenase) (data not shown). Transposons and transposase were found to be abundant in the metagenome of C. concentrica and speculated to be an important part of the genomic adaptation process towards a symbiotic relationship between bacteria and host (Thomas et al., 2010). Our observation shows that the transposase function is still active and that the transposase genes are not all just remnants of past events of intra- or intercellular horizontal gene transfer.
Conclusion
Our analysis has provided new insights into the activities of sponge-associated microbial communities, and for the Phyllobacteriaceae phylotype has offered a clear link between uncultivated phylotypes and functions. For the first time, specific transport functions for typical sponge metabolites (for example, halogenated aromatics, dipeptides) could be identified. While the coexistence of aerobic and anaerobic phases of the nitrogen cycles has been previously observed for another sponge system (Hoffmann et al., 2009b), here we link those functions to specific, expressed proteins and phylotypes in C. concentrica. Our analysis also indicated the requirement for the microbial community to respond to variable environmental conditions and hence express an array of stress protection proteins. Finally, molecular interactions between symbionts and their host might also be mediated by a set of expressed eukaryotic-like proteins and cell–cell mediators, and some sponge-associated bacteria (for example, the Phyllobacteriaceae phylotype) may be undergoing evolutionary adaptation process to the sponge environment, as evidenced by active mobile genetic elements.
Our data have clearly shown that a combined metaproteogenomic approach can provide novel information on the activities and physiology of sponge-associated, microbial communities. We believe that this approach will not only be useful to further our understanding of the enormous microbial diversity found in sponges around the world, but also to investigate the functional behaviour of symbiont communities in response to environmental change or host physiology (Webster et al., 2008). These observations will be crucial to understand the dynamic and complex interactions between sponges and their associated microbial symbionts.
Accession codes
References
Alavi M, Miller T, Erlandson K, Schneider R, Belas R . (2001). Bacterial community associated with Pfiesteria-like dinoflagellate cultures. Environ Microbiol 3: 380–396.
Ashworth RB, Cormier MJ . (1967). Isolation of 2,6-dibromophenol from marine Hemichordate Balanoglossus biminiensis. Science 155: 1558–1559.
Bayer K, Schmitt S, Hentschel U . (2008). Physiology, phylogeny and in situ evidence for bacterial and archaeal nitrifiers in the marine sponge Aplysina aerophoba. Environ Microbiol 10: 2942–2955.
Boos W . (1998). Binding protein-dependent ABC transport system for glycerol 3-phosphate of Escherichia coli. Methods Enzymol 292: 40–51.
Brady A, Salzberg SL . (2009). Phymm and PhymmBL: metagenomic phylogenetic classification with interpolated Markov models. Nat Methods 6: 673–676.
Brzoska P, Rimmele M, Brzostek K, Boos W . (1994). The pho regulon-dependent Ugp uptake system for glycerol-3-phosphate in Escherichia coli is trans inhibited by Pi. J Bacteriol 176: 15–20.
Burnum KE, Callister SJ, Nicora CD, Purvine SO, Hugenholtz P, Warnecke F et al. (2011). Proteome insights into the symbiotic relationship between a captive colony of Nasutitermes corniger and its hindgut microbiome. ISME J 5: 161–164.
Chae JC, Kim Y, Kim YC, Zylstra GJ, Kim CK . (2000). Genetic structure and functional implication of the fcb gene cluster for hydrolytic dechlorination of 4-chlorobenzoate from Pseudomonas sp. DJ-12. Gene 258: 109–116.
Chen J, Ryu S, Gharib SA, Goodlett DR, Schnapp LM . (2008). Exploration of the normal human bronchoalveolar lavage fluid proteome. Proteomics Clin Appl 2: 585–595.
Ebel R, Brenzinger M, Kunze A, Gross HJ, Proksch P . (1997). Wound activation of protoxins in marine sponge Aplysina aerophoba. J Chem Ecol 23: 1451–1462.
Elssner T, Engemann C, Baumgart K, Kleber HP . (2001). Involvement of coenzyme A esters and two new enzymes, an enoyl-CoA hydratase and a CoA-transferase, in the hydration of crotonobetaine to L-carnitine by Escherichia coli. Biochemistry 40: 11140–11148.
Ezraty B, Aussel L, Barras F . (2005). Methionine sulfoxide reductases in prokaryotes. Biochim Biophys Acta 1703: 221–229.
Finn RD, Mistry J, Tate J, Coggill P, Heger A, Pollington JE et al. (2010). The Pfam protein families database. Nucleic Acids Res 38: D211–D222.
Forward JA, Behrendt MC, Wyborn NR, Cross R, Kelly DJ . (1997). TRAP transporters: a new family of periplasmic solute transport systems encoded by the dctPQM genes of Rhodobacter capsulatus and by homologs in diverse Gram-negative bacteria. J Bacteriol 179: 5482–5493.
Giovannoni S, Stingl U . (2007). The importance of culturing bacterioplankton in the ‘omics’ age. Nat Rev Microbiol 5: 820–826.
Gribble GW . (1999). The diversity of naturally occurring organobromine compounds. Chem Soc Rev 28: 335–346.
Habyarimana F, Al-Khodor S, Kalia A, Graham JE, Price CT, Garcia MT et al. (2008). Role for the ankyrin eukaryotic-like genes of Legionella pneumophila in parasitism of protozoan hosts and human macrophages. Environ Microbiol 10: 1460–1474.
Hofmann B, Hecht HJ, Flohe L . (2002). Peroxiredoxins. Biol Chem 383: 347–364.
Hoffmann F, Radax R, Woebken D, Holtappels M, Lavik G, Rapp HT et al. (2009a). Complex nitrogen cycling in the sponge Geodia barretti. Environ Microbiol 11: 2228–2243.
Hoffmann F, Radax R, Woebken D, Holtappels M, Lavik G, Rapp HT et al. (2009b). Complex nitrogen cycling in the sponge Geodia barretti. Environ Microbiol 11: 2228–2243.
Hoffmann F, Røy H, Bayer K, Hentschel U, Pfannkuchen M, Brümmer F et al. (2008). Oxygen dynamics and transport in the Mediterranean sponge Aplysina aerophoba. Mar Biol 153: 1257–1264.
Holmes B, Blanch H . (2007). Genus-specific associations of marine sponges with group I crenarchaeotes. Mar Biol 150: 759–772.
Iturbe-Ormaetxe I, Burke GR, Riegler M, O’Neill SL . (2005). Distribution, expression, and motif variability of ankyrin domain genes in Wolbachia pipientis. J Bacteriol 187: 5136–5145.
Jurado V, Laiz L, Gonzalez JM, Hernandez-Marine M, Valens M, Saiz-Jimenez C . (2005). Phyllobacterium catacumbae sp. nov., a member of the order ‘Rhizobiales’ isolated from Roman catacombs. Int J System Evoluti Microbiol 55: 1487–1490.
Kleber HP . (1997). Bacterial carnitine metabolism. FEMS Microbiol Lett 147: 1–9.
Labbe N, Parent S, Villemur R . (2004). Nitratireductor aquibiodomus gen. nov., sp. nov., a novel alpha-proteobacterium from the marine denitrification system of the Montreal Biodome (Canada). Int J Syst Evol Microbiol 54: 269–273.
Landfald B, Strom AR . (1986). Choline–glycine betaine pathway confers a high level of osmotic tolerance in Escherichia coli. J Bacteriol 165: 849–855.
Lauro FM, McDougald D, Thomas T, Williams TJ, Egan S, Rice S et al. (2009). The genomic basis of trophic strategy in marine bacteria. Proc Natl Acad Sci USA 106: 15527–15533.
Letoffe S, Delepelaire P, Wandersman C . (2006). The housekeeping dipeptide permease is the Escherichia coli heme transporter and functions with two optional peptide binding proteins. Proc Natl Acad Sci USA 103: 12891–12896.
Liu MY, Kjelleberg S, Thomas T . (2011). Functional genomic analysis of an uncultured delta-proteobacterium in the sponge Cymbastela concentrica. ISME J 5: 427–435.
Liu YH, Yang B, Dong JD, Zhou XF, Yang XW, Lee KJ et al. (2009). Proline-containing dipeptides from a marine sponge of a Callyspongia species. Helv Chim Acta 92: 1112–1117.
Mantelin S, Saux MF, Zakhia F, Bena G, Bonneau S, Jeder H et al. (2006). Emended description of the genus Phyllobacterium and description of four novel species associated with plant roots: Phyllobacterium bourgognense sp. nov., Phyllobacterium ifriqiyense sp. nov., Phyllobacterium leguminum sp. nov. and Phyllobacterium brassicacearum sp. nov. Int J Syst Evolut Microbiol 56: 827–839.
Marchler-Bauer A, Lu S, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C et al. (2011). CDD: a Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res 39: D225–D229.
Mavromatis K, Doyle CK, Lykidis A, Ivanova N, Francino MP, Chain P et al. (2006). The genome of the obligately intracellular bacterium Ehrlichia canis reveals themes of complex membrane structure and immune evasion strategies. J Bacteriol 188: 4015–4023.
Mergaert J, Boley A, Cnockaert MC, Muller WR, Swings J . (2001). Identity and potential functions of heterotrophic bacterial isolates from a continuous-upflow fixed-bed reactor for denitrification of drinking water with bacterial polyester as source of carbon and electron donor. System Aappl Microbiol 24: 303–310.
Mohamed NM, Saito K, Tal Y, Hill RT . (2010). Diversity of aerobic and anaerobic ammonia-oxidizing bacteria in marine sponges. ISME J 4: 38–48.
Morris RM, Nunn BL, Frazar C, Goodlett DR, Ting YS, Rocap G . (2010). Comparative metaproteomics reveals ocean-scale shifts in microbial nutrient utilization and energy transduction. ISME J 4: 673–685.
Morris RM, Rappe MS, Connon SA, Vergin KL, Siebold WA, Carlson CA et al. (2002). SAR11 clade dominates ocean surface bacterioplankton communities. Nature 420: 806–810.
Mulligan C, Fischer M, Thomas GH . (2011). Tripartite ATP-independent periplasmic (TRAP) transporters in bacteria and archaea. FEMS Microbiol Rev 35: 68–86.
Ng C, DeMaere MZ, Williams TJ, Lauro FM, Raftery M, Gibson JA et al. (2010). Metaproteogenomic analysis of a dominant green sulfur bacterium from Ace Lake, Antarctica. ISME J 4: 1002–1019.
Nichols NN, Harwood CS . (1995). Repression of 4-hydroxybenzoate transport and degradation by benzoate: a new layer of regulatory control in the Pseudomonas putida beta-ketoadipate pathway. J Bacteriol 177: 7033–7040.
Norte M, Rodriguez ML, Fernandez JJ, Eguren L, Estrada DM . (1988). Aplysinadiene and (R,R) 5 [3,5-dibromo-4-[(2-oxo-5-oxazolidinyl)] methoxyphenyl]-2-oxazolidinone, 2 novel metabolites from Aplysina–Aerophoba—synthesis of Aplysinadiene. Tetrahedron 44: 4973–4980.
Nummelin H, Merckel MC, Leo JC, Lankinen H, Skurnik M, Goldman A . (2004). The Yersinia adhesin YadA collagen-binding domain structure is a novel left-handed parallel beta-roll. EMBO J 23: 701–711.
Olson ER, Dunyak DS, Jurss LM, Poorman RA . (1991). Identification and characterization of dppA, an Escherichia coli gene encoding a periplasmic dipeptide transport protein. J Bacteriol 173: 234–244.
Overbeek R, Begley T, Butler RM, Choudhuri JV, Chuang HY, Cohoon M et al. (2005). The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes. Nucleic Acids Res 33: 5691–5702.
Pedersen M, Saenger P, Fries L . (1974). Simple brominated phenols in red algae. Phytochemistry 13: 2273–2279.
Perry JJ, Shin DS, Getzoff ED, Tainer JA . (2010). The structural biochemistry of the superoxide dismutases. Biochim Biophys Acta 1804: 245–262.
Poolman B, Konings WN . (1993). Secondary solute transport in bacteria. Biochim Biophys Acta 1183: 5–39.
Preusser A, Wagner U, Elssner T, Kleber HP . (1999). Crotonobetaine reductase from Escherichia coli consists of two proteins. Biochim Biophys Acta 1431: 166–178.
Ram RJ, Verberkmoes NC, Thelen MP, Tyson GW, Baker BJ, Blake II RC et al. (2005). Community proteomics of a natural microbial biofilm. Science 308: 1915–1920.
Schaffer S, Isci N, Zickner B, Durre P . (2002). Changes in protein synthesis and identification of proteins specifically induced during solventogenesis in Clostridium acetobutylicum. Electrophoresis 23: 110–121.
Schauer K, Rodionov DA, de Reuse H . (2008). New substrates for TonB-dependent transport: do we only see the ‘tip of the iceberg’? Trends Biochem Sci 33: 330–338.
Schlappy ML, Schottner SI, Lavik G, Kuypers MMM, de Beer D, Hoffmann F . (2010). Evidence of nitrification and denitrification in high and low microbial abundance sponges. Mar Biol 157: 593–602.
Schmidt EW, Obraztsova AY, Davidson SK, Faulkner DJ, Haygood MG . (2000). Identification of the antifungal peptide-containing symbiont of the marine sponge Theonella swinhoei as a novel delta-proteobacterium, ‘Candidatus Entotheonella palauensis’. Mar Biol 136: 969–977.
Schmidt II, Bock E . (1997). Anaerobic ammonia oxidation with nitrogen dioxide by Nitrosomonas eutropha. Arch Microbiol 167: 106–111.
Schmitz FJ, Gopichand Y . (1978). (7e, 13-Xi, 15z)-14,16-dibromo-7,13,15-hexadecatrien-5-ynoic acid—novel dibromo acetylenic acid from marine sponge Xestospongia muta. Tetrahedron Lett 19: 3637–3640.
Sowell SM, Abraham PE, Shah M, Verberkmoes NC, Smith DP, Barofsky DF et al. (2011). Environmental proteomics of microbial plankton in a highly productive coastal upwelling system. ISME J 5: 856–865.
Sowell SM, Wilhelm LJ, Norbeck AD, Lipton MS, Nicora CD, Barofsky DF et al. (2009). Transport functions dominate the SAR11 metaproteome at low-nutrient extremes in the Sargasso Sea. ISME J 3: 93–105.
Tatusov RL, Fedorova ND, Jackson JD, Jacobs AR, Kiryutin B, Koonin EV et al. (2003). The COG database: an updated version includes eukaryotes. BMC Bioinform 4: 41.
Taylor MW, Radax R, Steger D, Wagner M . (2007). Sponge-associated microorganisms: evolution, ecology, and biotechnological potential. Microbiol Mol Biol Rev 71: 295–347.
Thomas T, Rusch D, DeMaere MZ, Yung PY, Lewis M, Halpern A et al. (2010). Functional genomic signatures of sponge bacteria reveal unique and shared features of symbiosis. ISME J 4: 1557–1567.
Ullmann R, Gross R, Simon J, Unden G, Kroger A . (2000). Transport of C(4)-dicarboxylates in Wolinella succinogenes. J Bacteriol 182: 5757–5764.
Unson MD, Holland ND, Faulkner DJ . (1994). A brominated secondary metabolite synthesized by the cyanobacterial symbiont of a marine sponge and accumulation of the crystalline metabolite in the sponge tissue. Mar Biol 119: 1–11.
Utkina NK, Denisenko VA, Scholokova OV, Virovaya MV, Gerasimenko AV, Popov DY et al. (2001). Spongiadioxins A and B, two new polybrominated dibenzo-p-dioxins from an Australian marine sponge Dysidea dendyi. J Nat Prod 64: 151–153.
Vogel S . (1977). Current-induced flow through living sponges in nature. Proc Natl Acad Sci USA 74: 2069–2071.
Voth DE, Howe D, Beare PA, Vogel JP, Unsworth N, Samuel JE et al. (2009). The Coxiella burnetii ankyrin repeat domain-containing protein family is heterogeneous, with C-terminal truncations that influence Dot/Icm-mediated secretion. J Bacteriol 191: 4232–4242.
Vuilleumier S . (1997). Bacterial glutathione S-transferases: what are they good for? J Bacteriol 179: 1431–1441.
Walker CB, de la Torre JR, Klotz MG, Urakawa H, Pinel N, Arp DJ et al. (2010). Nitrosopumilus maritimus genome reveals unique mechanisms for nitrification and autotrophy in globally distributed marine crenarchaea. Proc Natl Acad Sci USA 107: 8818–8823.
Walt A, Kahn ML . (2002). The fixA and fixB genes are necessary for anaerobic carnitine reduction in Escherichia coli. J Bacteriol 184: 4044–4047.
Webster NS, Blackall LL . (2008). What do we really know about sponge–microbial symbioses? ISME J 3: 1–3.
Webster NS, Cobb RE, Negri AP . (2008). Temperature thresholds for bacterial symbiosis with a sponge. ISME J 2: 830–842.
Webster NS, Taylor MW . (2011). Marine sponges and their microbial symbionts: love and other relationships. Environ Microbiol; epub ahead of print 28 March 2011; doi: 10.1111/j.1462-2920.2011.02460.x.
White RH, Hager LP . (1977). Occurrence of fatty-acid chlorohydrins in jellyfish lipids. Biochemistry 16: 4944–4948.
Wilkinson CR . (1983). Net primary productivity in coral reef sponges. Science 219: 410–412.
Wilmes P, Wexler M, Bond PL . (2008). Metaproteomics provides functional insight into activated sludge wastewater treatment. PLoS One 3: e1778.
Zhu P, Li Q, Wang G . (2008). Unique microbial signatures of the alien Hawaiian marine sponge Suberites zeteki. Microb Ecol 55: 406–414.
Acknowledgements
We acknowledge the J. Craig Venter Institute (JCVI) Joint Technology Center under the leadership of Yu-Hui Rogers and the assistance of Matt Lewis for producing the sequencing data. This work was supported by funding from the Gordon and Betty Moore Foundation and the Australian Research Council (LP0668235).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Liu, M., Fan, L., Zhong, L. et al. Metaproteogenomic analysis of a community of sponge symbionts. ISME J 6, 1515–1525 (2012). https://doi.org/10.1038/ismej.2012.1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2012.1
Keywords
This article is cited by
-
Exploring Prokaryotic Communities in the Guts and Mucus of Nudibranchs, and Their Similarity to Sediment and Seawater Microbiomes
Current Microbiology (2023)
-
Ammonia-oxidizing archaea in biological interactions
Journal of Microbiology (2021)
-
Integrating novel tools to elucidate the metabolic basis of microbial symbiosis in reef holobionts
Marine Biology (2021)
-
Microbiome-mediated mechanisms contributing to the environmental tolerance of reef invertebrate species
Marine Biology (2021)
-
Characterization of a sponge microbiome using an integrative genome-centric approach
The ISME Journal (2020)