Abstract
The soil microbiome is responsible for mediating key ecological processes; however, little is known about its sensitivity to climate change. Observed increases in global temperatures and alteration to rainfall patterns, due to anthropogenic release of greenhouse gases, will likely have a strong influence on soil microbial communities and ultimately the ecosystem services they provide. Therefore, it is vital to understand how soil microbial communities will respond to future climate change scenarios. To this end, we surveyed the abundance, diversity and structure of microbial communities over a 2-year period from a long-term in situ warming experiment that experienced a moderate natural drought. We found the warming treatment and soil water budgets strongly influence bacterial population size and diversity. In normal precipitation years, the warming treatment significantly increased microbial population size 40–150% but decreased diversity and significantly changed the composition of the community when compared with the unwarmed controls. However during drought conditions, the warming treatment significantly reduced soil moisture thereby creating unfavorable growth conditions that led to a 50–80% reduction in the microbial population size when compared with the control. Warmed plots also saw an increase in species richness, diversity and evenness; however, community composition was unaffected suggesting that few phylotypes may be active under these stressful conditions. Our results indicate that under warmed conditions, ecosystem water budget regulates the abundance and diversity of microbial populations and that rainfall timing is critical at the onset of drought for sustaining microbial populations.
Similar content being viewed by others
Introduction
The importance of the soil microbiome in cycling of key nutrients such as carbon and nitrogen is well established. However, because of the microbiome's complexity little is understood about how climate change (for example, warming and precipitation patterns) will affect the diversity, abundance and structure of the community. Climate change is thought to exert pressure on the soil environment and ultimately the soil microbiome through direct and indirect pathways (Shaver et al., 2000). Direct influence of climate change on bacteria will likely occur through increased seasonal temperature maxima, as temperature has long been known to be a determinative selector of microbial growth (Ratkowsky et al., 1982) and physiological activity of soil respiration (Lundegärdh, 1927). In grassland soils, experimental warming has been shown to increase bacterial biomass in winter and spring (Belay-Tedla et al., 2009); however, warming negatively affected bacterial biomass (Liu et al., 2009) and 16S rRNA gene abundance (Castro et al., 2010) in summer and early fall, respectively, suggesting that warming may have a seasonal affect on the soil community. In situ microbial activity as measured by soil CO2 efflux has been shown to be temperature sensitive (Luo et al., 2001), thermally adaptive (Bradford et al., 2008) and constrained by soil moisture (Garten et al., 2009; Liu et al., 2009), indicating that warming is likely also having an effect on the physiology of the microbial community. Soil respiration is a summation of total CO2 efflux that includes Eukarya in addition to Bacteria and Archaea. Hence, drawing specific conclusions regarding the response of the soil microbial community to climate change using these data may be arguable. Nonetheless, these studies show that climate warming can have a strong influence on the microbiome.
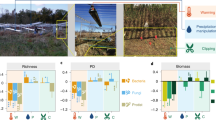
As a result of increased temperatures, a concomitant increase in the frequency and duration of drought events in mesic ecosystems is anticipated (Knapp et al., 2008). The expected exposure to water stress is likely to affect both microbial and plant communities by interrupting key nutrient cycles and plant–microbe feedbacks. Single-factor experiments focusing on precipitation manipulations show that rainfall additions had little effect on the composition of grassland microbial communities except under drought (Cruz-Martinez et al., 2009). In addition, rainfall frequency did not alter microbial community composition in laboratory incubations of grassland soil (Fierer et al., 2003). Multifactor warming experiments show that microbial community composition is altered under warming and precipitation additions in a constructed natural grass prairie (Castro et al., 2010). In a recent review, Bardgett et al. (2008) suggests that plant-mediated indirect effects of climate change (that is, alterations in plant community physiology or composition) are likely to have a significant role in how the soil microbiome responds to climate change. Experimental warming has been shown to influence the phenology (Parmesan, 2006) and extend the growing season of plant communities (Wan et al., 2005). As a result, warming causes increases in plant biomass (shoot and root) (Luo, 2007), however, a concomitant increase in water uptake from the soil is needed to account for the increased biomass. Such an anticipated effect from climate warming on the plant community has consequences on the total soil water budget and ultimately will affect the microbiome if frequency and duration of water stress periods increases (Porporato et al., 2004; Knapp et al., 2008). Drought is therefore likely to affect the microbiome due to the combined indirect effect on plants and the direct effects of increases in temperature and decreased soil moisture.
We set out to examine the effects of warming on soil microbial communities at a previously described long-term global warming experimental field site located at KFFL (Kessler Farm Field Laboratory) in central Oklahoma (Luo et al., 2001). However, during our sampling period the site experienced a moderate drought thereby allowing us to evaluate the interactive effects of warming and ecosystem water budgets on microbial communities. We tracked the abundance, diversity and structure of microbial populations before (August 2004), at the onset (April 2005), and during (August 2005 and April 2006) a moderate drought, using quantitative PCR (qPCR) and pyrosequencing. Results show that when moisture was abundant, the warming treatment significantly shifted the community structure and furthermore the indirect effect of global warming, specifically the decreases in soil moisture, significantly altered the abundance of the soil microbial community more than the direct effects of warming. Our results therefore suggest that rain fall periodicity and timing coupled to warming will have a significant impact on microbial communities and will have an important role in regulating microbial carbon and nitrogen cycling.
Materials and methods
Site description
The experimental warming site (Wan et al., 2002) is characterized as a tall grass prairie that receives ∼90.0 cm of rainfall annually with an average temperature of 16.3 °C. The site consists of six paired (control and warmed) 2 × 2 m plots. Warmed plots are heated to ∼2.0 °C above ambient temperature using infrared heaters suspended above the plot. All plots had been continuously heated for ∼4 years before the beginning of our sampling. Soil moisture was taken at regular intervals using a portable time domain reflectance probe (Wan et al., 2002). Soil samples were taken from unclipped subplots (to simulate a native grassland) over a 2-year period starting in August 2004 and ending in April 2006.
Soil sampling and DNA extraction
Soil cores were collected from the top 15 cm of soil from the unclipped subplots of the control and warmed plots using sterile stainless steel coring tools. Two cores were randomly taken from each plot (one core per subplot) combined and stored on ice for transport. The combined core material was then screened for roots, homogenized and stored at −20 °C. DNA was extracted in duplicate from soils using the MoBio Ultraclean Soil DNA Isolation Kit (MoBio Laboratories, Carlsbad, CA, USA). Extraction of DNA from 0.5-g soil was done according to the manufacturers methods except for the following modification. In lieu of shaking for 10 min on a flat bed vortex, samples were treated in a Mini-bead beater (Biospec Products Inc., Bartlesville, OK, USA) for 1 min at maximum speed and then placed on ice for 1 min. Isolated DNA was then diluted 1/10 in sterile, nuclease-free water and stored at −20 °C.
Quantitative PCR
16S rRNA gene copy numbers of targeted phylogenetic groups (Supplementary Table 3) were quantified over the course of two seasons (August 2004–2005 and April 2005–2006) in five pairs of plots. qPCR methods were adapted from Fierer et al. (2005), with alterations to DNA purification and adaptation to the MyIQ real-time PCR system (Bio-Rad, Hercules, CA, USA). To ensure qPCR was not inhibited, prior testing of DNA clean-up methods was done (see Supplementary Materials). We found that in our soil, a 1/10 dilution worked as well or outperformed popular cleaning methods for qPCR analysis. To quantify the 16S rRNA genes, two DNA extractions were done from each plot at each of the four time points. qPCR was then performed in triplicate on both DNAs. Each reaction mixture (25 μl total volume) consisted of 12.5 μl IQ SYBR Green Supermix (Bio-Rad), 9.5 μl of water, 1.0 μl (10 μM) of each primer and 2.0 μl of diluted DNA. qPCR generally followed a standard two-step protocol consisting of 5 min at 95 °C, followed by 40 cycles of 95 °C for 30 s, 54 °C for 30 s and 72 °C for 30 s. Specific qPCR annealing temperatures and times for each primer set are specified in Supplementary Table 3. To quantify the number of gene copies, standard curves were constructed using tenfold serial dilutions of pCR4-TOPO plasmid (Invitrogen, Carlsbad, CA, USA) containing a representative amplicon derived from the environment from each of the primer sets. Plasmids were extracted using a Qiagen Plasmid Miniprep kit (Qiagen, Valencia, CA, USA). Plasmids for each primer set were sequenced before quantification to ensure specificity of the target. Each plasmid standard was quantified using a Nanodrop spectrophotometer system (Thermo Scientific, Wilmington, DE, USA) then converted to copy number from plasmid molecular weight. PCR conditions were modified for each primer set to reach as closely as possible an amplification efficiency of −3.3. No primer dimers were observed in all primer sets as determined by melting curves of qPCR amplicons. The soil copy number was determined from the standard curve and subsequently standardized to copy numbers per gram of dry soil. Seasonal averages of qPCR data are given in Supplementary Table 2. To control for spatial variation in microbial population response at the sampling site, Multilevel Modeling (MLM) was chosen. MLM allowed data to be structured by individual bacterial groups, experimental plot and time. This permitted us to represent the interactive effect of warming on microbial populations, as plots and microbial populations may not respond similarly to the warming treatment. Log transformed qPCR values were the model's-dependent variable, which took into account all six populations plus total bacteria. Full methods and equations are included in the Supplemental Information.
Pyrosequencing
PCR libraries were generated using modified 338 forward and 518 reverse primers listed in Supplementary Table 3. Modifications to the primer set included: adding the A-adapter and a unique 8 base barcode to the 5′ end of the 338 forward primer while the B-adapter was added to the 5′ end of the reverse primer (sequences for A and B adapters, taken from http://www.454.com, 454 Life Sciences, Branford, CT, USA). Duplicate DNA extractions from each plot (five plots per treatment) were amplified with two PCRs for a total of 20 reactions per treatment (control and warmed) at each of the four time points. The products of the 20 PCR reactions were combined before sequencing. DNA from control and warmed plots was amplified using a separate barcode per treatment and time point. DNA was PCR amplified in 50 μl reactions containing (final concentration) 2 μl of 1/10 diluted DNA, 1 × Flexi buffer (Promega, Madison, WI, USA), 2.5 mM MgCl2, 0.2 mM deoxynucleoside triphosphate mixture (Promega), 2.5 U GoTaq DNA polymerase (Promega) and 500 nM of the forward and reverse primer. PCR amplification protocol was as follows: 5 min at 95 °C, followed by 30 cycles of 95 °C for 1 min, 54 °C for 1 min and 72 °C for 1 min with a final extension of 15 min at 72 °C. PCR reactions were screened using gel electrophoresis and positive reactions were pooled (time and treatments kept separate) and concentrated using a speed vac concentrator (Savant DNA120, Thermo Electron Corporation, Waltham, MA, USA). Concentrated PCR products were then purified using the PureLink PCR purification kit (Invitrogen) and purity was confirmed using gel electrophoresis. PCR products were then sent to Engencore (http://engencore.sc.edu/) for pyrosequencing using FLX chemistry.
Data processing of pyrosequencing reads
Raw pyrosequencing data were processed for quality and barcode recovery using the software package Mothur (Schloss et al., 2009) version 1.7.0 (http://www.mothur.org). Clean sequences were then aligned to the Greengenes core sequence set using NAST algorithm (DeSantis et al., 2006a), distances were calculated, and operational taxonomic units (OTUs) generated all within Mothur. Representative OTUs0.03 were classified (Hugenholtz taxonomy) using the Greengenes database (DeSantis et al., 2006b). Diversity indices (Chao, abundance-based coverage estimator, Shannon and Simpson) and rarefaction curves were also generated using Mothur. Diversity ordering methods (Renyi generalized entropy and Hulbert family of diversity indices) were chosen because of their insensitivity to sample size and indices were calculated as previously described (Liu et al., 2007; Youssef and Elshahed, 2008). A warming treatment effect was scored as significant if two of the three diversity ordering indices were higher/lower than the control treatment and do not cross. Aligned OTUs0.03 for each treatment and time point were lane masked within Mothur and a phylogenetic tree was generated using FastTree (Price et al., 2009) (FastTree flagged conditions: -fastest, -gtr, -nt). The phylogenetic tree was uploaded to the Fast Unifrac (Hamady et al., 2009) web interface and an analysis of β diversity (community structure) was performed using the P-test and (un)weighted Unifrac metric. Heatmaps and clustering analyses were generated using the web interface (http://www.hiv.lanl.gov/content/sequence/HEATMAP/heatmap.html) that implements the heatmap tool (heatmap.2) in the gplots package within the statistical program R (http://www.r-project.org/). The relative abundances of the top 50 OTUs from each phylum were used to generate heatmaps using the Euclidean distances. Clustering of the sample times and individual OTUs0.03 was done using the complete method (Johnson, 1967).
Results
Rainfall patterns
During the course of our sampling, the experimental site experienced a moderate drought receiving ∼47% less rainfall in 2005 (51.5 cm) than 2004 (96.5 cm) (Supplementary Figure 1), which had received above average precipitation (Luo et al., 2009). The drought extended through the winter of 2005 and up to the time of sampling in April 2006. During the months leading up to the start of the drought in April 2005, monthly rainfall totals steadily declined to ∼74% of those of the previous spring (February–April cumulative rainfall totals were 19.7 cm (2004) and 10.4 cm (2005)), resulting in a net deficit in soil moisture reserves at the beginning of the growing season (Figure 1).
Average soil moisture taken at 10 cm over the 2-year sampling period. Solid lines represent warm (plus sign) and control (circles) treatments while grey bands represent the standard deviation associated with each time point. Dashed lines represent the total microbial community modeled abundance from the Multi-Level Modeling (MLM). Arrows represent dates at which samples were taken.
Assessment of bacterial population size in control and warmed plots using qPCR
Quantification of dominant bacterial and archaeal phyla (Total, Actinobacteria, Acidobacteria, α-Proteobacteria, Planctomycetes, Verrucomicrobia and Crenarchaeota) revealed that both warming and warming coupled to drought had a significant influence on soil microbial population size. During the period of above normal precipitation (August 2004), warming increased numbers of all microbial groups tested by 40–150% (Figures 2a–g). At the onset of the drought (April 2005), the majority of control-plot bacterial and archaeal population sizes were elevated above those in the warmed plots. Exceptions were members of the Acidobacteria and Verrucomicrobia (Figures 2c and g). During the drought period (August 2005), the warming treatment caused a significant decrease (50–80%) in numbers (Figures 2a–g) of all groups tested. The following spring (April 2006) most of the phyla tested had recovered from the drought and returned back to population sizes comparable to those in April 2005, however, most were still below or similar to control plots. Interestingly, the warming treatment increased numbers of Acidobacteria and Crenarchaeota (Figures 2c and e) populations in April 2006 samples while Actinobacteria and Verrucomicrobia (Figures 2b and g) never recovered from the August drought. Bayesian MLM results revealed that the warming treatment significantly stimulated microbial populations in August 2004 when soil moisture was abundant. However, as the experimental plots transitioned into drought, soil moisture became a limiting factor and when coupled to the warming treatment caused significant decreases in microbial populations (Figure 2h).
Residual plots of qPCR generated 16S rRNA gene abundance data over time for warmed (plus sign) and control (circles) plots. Residual plots of total bacteria (a) and individual phyla (b–g) were generated using the baseline average for each plot across all four seasons. The shaded bands depict the standard deviation. (h) Bayesian MLM takes into account all microbial groups assayed and grey bands represent the bootstrapped 95% confidence intervals. Raw values are in Supplementary Table 2.
Assessment of the diversity and community structure of control and warmed plots using pyrosequencing
A total of 260 499 sequences were generated resulting in ∼10 000–45 000 sequences for each treatment and time point (four control and four warmed libraries combined, Supplementary Table 1). In all control and warmed data sets, phylum level distributions for each of the four time points revealed an overall pattern similar to other soil studies (Elshahed et al., 2008; Fulthorpe et al., 2008) whereby Actinobacteria, Proteobacteria and Acidobacteria were the most abundant phyla represented (Figure 3). When rainfall was abundant (August 2004), the warming treatment increased the relative abundance of sequences of the Proteobacteria (11%), Acidobacteria (50%), Bacteroidetes (35%), Verrucomicrobia (3%) and SPAM (68%) (Figure 3). However, during drought (that is, August 2005) the warming treatment had little effect on the relative abundance of the dominant phyla with the exception of the Planctomycetes, which saw an increase of 48% over the control (Figure 3).
Relative abundance of dominant bacterial phyla in pyrosequencing generated 16S rRNA genes sequence libraries at the KFFL warming site. Each bar is color-coded and represents a single sampling time. Abundance was normalized by treatment and time. Abundance of Proteobacteria is included in the subset. F(C/W)4=August 2004, S(C/W)=April 2005, F(C/W)5=August 2005, S(C/W)6=April 2006; C=control, W=warmed.
To assess the bacterial diversity of control and warmed plots at various sampling times we chose to employ both traditional species richness estimators and less used diversity ordering methods. In the grassland soil, species richness (Chao1), evenness (abundance-based coverage estimator), diversity (Shannon Index) and diversity ordering followed a similar pattern. During the non-drought period (August 2004) the warming treatment decreased richness, evenness and diversity; while during the drought (August 2005), warming increased richness, evenness and diversity of the community when compared with the controls (Table 1). Diversity ordering of the most dominant phyla in Kessler Farm soil reveals a similar pattern to that of the total data set with control plots more diverse in August 2004 and warmed plots more diverse in August 2005 (Supplementary Table 2). Spring time points (April 2005 and 2006) show that species richness, evenness and diversity patterns showed no clear treatment level effects. Diversity ordering patterns of individual phyla also show spring samples to be variable with most phyla not increasing with the warming treatment (Supplementary Table 2).
To assess how community structure is affected by the warming treatment we chose to use the β diversity measurements P-test (Martin, 2002) and (un)weighted Unifrac (Lozupone et al., 2006) metrics implemented in Fast Unifrac (Hamady et al., 2009) and traditional community similarity measures Bray–Curtis and Jaccard dissimilarity indices. Pairwise comparisons between warmed and control plots show warming had no significant effect (unweighted Unifrac P>0.05) on the overall structure of microbial communities at each time point, however, warming did significantly alter lineage-specific patterns in two out of the four seasons (P-test: August 2004, P<0.001; weighted Unifrac: August 2004, P=0.032 and April 2005, P=0.016). Transitioning into drought (August 2004 to April 2005) from normal moisture conditions had a significant effect on control and warmed plot community structures (unweighted Unifrac: control plots P=0.009; Warmed plots P=0.007). Bray–Curtis and Jaccard similarity indices corroborate Unifrac results such that when moisture was abundant (August 2004 and April 2005) the warmed and control plots had low similarity, but as the drought intensified, similarity between the two communities increased (Supplementary Figure 2). Cluster analysis of dominant genus level OTUs from each phylum revealed that the warming treatment continually altered the community profile such that the control and warmed plots never clustered with one another at any given time point (Supplementary Figure 3). Heatmaps in conjunction with the cluster analysis allow for visualization of the dominant genera distribution patterns captured at the time of sampling. Interestingly, these patterns were either constrained (that is, few OTUs0.03 dominant across all time points sampled) as in the Betaproteobacteria or highly variable (that is, many OTUs0.03 share the dominance but are sporadically distributed across time) like the Deltaproteobacteria (Supplementary Figure 3).
Discussion
The microbiome is an integral component of soil and is important to maintaining the function of all ecosystems. However, our understanding of how these microbial communities will respond to disturbances such as climate change is limited. Therefore, we sought to understand how bacterial and archaeal communities would respond to elevated temperatures associated with future, predicted global warming temperatures. Microbial communities were monitored during normal precipitation and drought periods using pyrosequencing and qPCR. Sampling during the transition from adequate precipitation to drought provides a glimpse of how microbial communities respond to the environmental stressors of warming alone and warming under drought.
Effects of warming under normal precipitation
Temperature has long been known to be a determinant for the growth and physiology of microorganisms and may be a determining factor for niche space competition between physiologically similar organisms (Bennett and Lenski, 1993). However, due to the dearth of uncultivated organisms in soil little is known of the temperature optima of the majority of microorganisms. We observed that under normal precipitation conditions (August 2004) the warming treatment had a significant effect on microbial populations and community structure when compared with the control. Under warming conditions microbial population abundance was significantly higher than the controls, however, diversity of the warmed plots was markedly lower than the control. Furthermore, shifts in lineage-specific community structure (unweighted vs weighted Unifrac) and community similarity (Bray–Curtis and Jaccard indices) affirm that the warming treatment is selecting for a subset of the total warming plot diversity. Assessing a physiological significance to the shift in community structure is difficult because many of the dominant phylotypes within our data set are uncharacterized. Under warming, the reduction of Actinobacteria suggests that increases in respiration associated with higher temperatures (Luo, 2007) may be negatively affecting this group as they are known to be sensitive to increased CO2 (Goodfellow and Williams, 1983). Furthermore, increases in the relative abundance of Acidobacteria may be due to the reduction in Actinobacteria, as these groups likely share similar niches. Increased abundance of the Crenarchaea, known archaeal ammonia oxidizers (Konneke et al., 2005), suggests that this group may be an important component of the nitrogen cycle in a warmed climate, especially under nitrogen limiting conditions. Recently, Crenarchaea have been shown to be active at ammonia concentrations far below those that are used by their bacterial counterparts (Martens-Habbena et al., 2009).
Effects of warming at onset of drought
As the plots transitioned from a normal precipitation year into drought (Spring 2005 to August 2005), precipitation patterns at the study site suggest that lack of rainfall before the 2005 growing season was more important for long-term survival of microbial communities than summer rainfall. Under decreasing rainfall conditions, the biome must survive on soil moisture reserves (often described as the bucket model; Porporato et al., 2004) until stocks are replenished through rainfall (Knapp et al., 2008). However under warmed conditions, plant communities respond by developing earlier (Parmesan, 2006), which is evident at our study site by elevated plant biomass in warmed plots relative to controls (Luo et al., 2009). The tax on soil moisture reserves due to early onset of plant growth is evident as plant evapotranspiration and increased evaporation rates contributed to the critically low soil moisture levels (Figure 2; March–April 2005) in warmed plots. As a result of the moisture deficit, microbial communities responded to warming differently than when moisture was abundant. Microorganisms in warmed plots were generally less abundant and community structure was significantly different when compared with the control. The significant shift in lineage-specific microbial community structure would again suggest the warming treatment is selecting for a subset of the total community. However because plant communities are advanced ahead of control plots, the shift in microbial community structure may also be due to increased labile carbon pools (Belay-Tedla et al., 2009) and the reduction of moisture. Furthermore, significant shifts in community structure from August 2004 to April 2005 in both control and warmed plots would suggest that seasonality is a large contributing factor to the structure of these communities and could correspond to life stage of the plant community.
Effects of warming under drought
Under drought conditions, the warming treatment had the most pronounced effect on microbial population abundance. All warmed plot populations tested in August 2005 decreased in size from the April 2005 samples and when compared with the control plots the decline is a stark contrast (Figure 2). MLM results indicate the decline in abundance is attributed to soil moisture at the time of sampling. Furthermore, soil conditions in warmed plots created by the early onset of the plant community in conjunction with the lack of rainfall are likely the underpinnings for the decline in microbial populations. Plant biomass data at the site also suggest that because of the lack of moisture, NPP (net primary production) was decreased relative to controls (Luo et al., 2009). The potential reduction in labile carbon and nitrogen entering the rhizosphere could also be a partial cause for the decline of microbial phyla such as Verrucomicrobia who are known to be heterotrophs (Hedlund et al., 1997) and are sensitive to nitrogen ratios (Keiblinger et al., 2010). An analysis of the effect of warming on community structure showed no significant difference despite the effect on abundance. Bray–Curtis similarity indices indicate that the drought constrained the abundance patterns between warmed and control communities as the plots transition into drought, whereby control and warmed communities became more similar as the drought intensified thus corroborating the community structure results.
Effects of warming transitioning out of drought
The spring 2006 samples present an interesting look at the community because recharging of soil moisture reserves, a process that normally occurs over winter months at the site, was absent. Instead large differences between control and warmed plot soil moisture was maintained throughout the fall of 2005 and just before the time of sampling in 2006. Abundance data show that some groups are much more resilient to drought and quickly return to numbers similar to (Planctomycetes) or above (Crenarchaea and Acidobacteria) control plot values, while others are slower to recover (Actinobacteria and Verrucomicrobia). The differential response of each phylum suggests that restructuring of communities may take long periods of time after extreme stress events such as drought and warming. Furthermore, the ability of microorganisms to lay dormant during periods of stress (Jones and Lennon, 2010) may be vital to the resiliency of prairie soil microbial communities (Fierer et al., 2003) and may explain why community similarity increased under drought.
Summary
In Oklahoma prairie soil, increases in temperature and the combination of drought and temperature had dramatic negative effects on the abundance, diversity and structure of soil bacterial communities. Our data suggest that shifts in rainfall patterns, specifically timing and amounts per event, can evoke long-term changes to the microbial community especially under a warming climate. These alterations to the community are likely to have overarching effects on the physiology of the microbiome, as portions of the community lay dormant in response to stress. Furthermore, recovery of microbial communities after perturbations, such as drought, may not occur immediately and thus may disrupt soil carbon and nitrogen cycling. We feel that further rainfall manipulation and warming experiments are needed to help elucidate the complex response of the microbiome, as the likelihood of experiencing water-stress events will increase with a warming climate.
Accession codes
References
Bardgett RD, Freeman C, Ostle NJ . (2008). Microbial contributions to climate change through carbon cycle feedbacks. ISME J 2: 805–814.
Belay-Tedla A, Zhou X, Su B, Wan S, Luo Y . (2009). Labile, recalcitrant, and microbial carbon and nitrogen pools of a tallgrass prairie soil in the US Great Plains subjected to experimental warming and clipping. Soil Biol Biochem 41: 110–116.
Bennett AF, Lenski RE . (1993). Evolutionary adaptation to temperature II. Thermal niches of experimental lines of Escherichia coli. Evolution 47: 1–12.
Bradford MA, Davies CA, Frey SD, Maddox TR, Melillo JM, Mohan JE et al. (2008). Thermal adaptation of soil microbial respiration to elevated temperature. Ecol Lett 11: 1316–1327.
Castro HF, Classen AT, Austin EE, Norby RJ, Schadt CW . (2010). Soil microbial community responses to multiple experimental climate change drivers. Appl Environ Microbiol 76: 999–1007.
Cruz-Martinez K, Suttle KB, Brodie EL, Power ME, Andersen GL, Banfield JF . (2009). Despite strong seasonal responses, soil microbial consortia are more resilient to long-term changes in rainfall than overlying grassland. ISME J 3: 738–744.
DeSantis TZ, Hugenholtz P, Keller K, Brodie EL, Larsen N, Piceno YM . (2006a). NAST: a multiple sequence alignment server for comparative analysis of 16S rRNA genes. Nucleic Acids Res 34: W394–W399.
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K . (2006b). Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72: 5069–5072.
Elshahed MS, Youssef NH, Spain AM, Sheik C, Najar FZ, Sukharnikov LO . (2008). Novelty and uniqueness patterns of rare members of the soil biosphere. Appl Environ Microbiol 74: 5422–5428.
Fierer N, Jackson JA, Vilgalys R, Jackson RB . (2005). Assessment of soil microbial community structure by use of taxon-specific quantitative PCR assays. Appl Environ Microbiol 71: 4117–4120.
Fierer N, Schimel JP, Holden PA . (2003). Influence of drying–rewetting frequency on soil bacterial community structure. Microb Ecol 45: 63–71.
Fulthorpe RR, Roesch LF, Riva A, Triplett EW . (2008). Distantly sampled soils carry few species in common. ISME J 2: 901–910.
Garten C, Classen A, Norby R . (2009). Soil moisture surpasses elevated CO2 and temperature as a control on soil carbon dynamics in a multi-factor climate change experiment. Plant Soil 319: 85–94.
Goodfellow M, Williams ST . (1983). Ecology of Actinomycetes. Annu Rev Microbiol 37: 189–216.
Hamady M, Lozupone C, Knight R . (2009). Fast UniFrac: facilitating high-throughput phylogenetic analyses of microbial communities including analysis of pyrosequencing and PhyloChip data. ISME J 4: 17–27.
Hedlund BP, Gosink JJ, Staley JT . (1997). Verrucomicrobia div. nov., a new division of the Bacteria containing three new species of Prosthecobacter. Antonie van Leeuwenhoek 72: 29–38.
Johnson SC . (1967). Hierarchical clustering schemes. Psychometrika 32: 241–254.
Jones SE, Lennon JT . (2010). Dormancy contributes to the maintenance of microbial diversity. Proc Natl Acad Sci USA 107: 5881–5886.
Keiblinger KM, Hall EK, Wanek W, Szukics U, Hämmerle I, Ellersdorfer G et al. (2010). The effect of resource quantity and resource stoichiometry on microbial carbon-use-efficiency. FEMS Microbiol Ecol 73: 430–440.
Knapp AK, Beier C, Briske DD, Classen AeT, Luo Y, Reichstein M et al. (2008). Consequences of more extreme precipitation regimes for terrestrial ecosystems. BioScience 58: 811–821.
Konneke M, Bernhard AE, de la Torre JR, Walker CB, Waterbury JB, Stahl DA . (2005). Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature 437: 543–546.
Liu C, Whittaker RJ, Ma K, Malcolm JR . (2007). Unifying and distinguishing diversity ordering methods for comparing communities. Popul Ecol 49: 89–100.
Liu W, Zhang Z, Wan S . (2009). Predominant role of water in regulating soil and microbial respiration and their responses to climate change in a semiarid grassland. Glob Change Biol 15: 184–195.
Lozupone C, Hamady M, Knight R . (2006). UniFrac—an online tool for comparing microbial community diversity in a phylogenetic context. BMC Bioinformatics 7: 371.
Lundegärdh H . (1927). Carbon dioxide evolution of soil and crop growth. Soil Sci 23: 417–453.
Luo Y . (2007). Terrestrial carbon cycle feedback to climate warming. Annu Rev Ecol Evol Syst 38: 683–712.
Luo Y, Sherry RA, Zhou X, Wan S . (2009). Terrestrial carbon-cycle feedback to climate warming: experimental evidence on plant regulation and impacts of biofuel feedstock harvest. Glob Change Biol Bioenergy 1: 62–74.
Luo Y, Wan S, Hui D . (2001). Acclimatization of soil respiration to warming in tall grass prairie. Nature 413: 622–625.
Martens-Habbena W, Berube PM, Urakawa H, de la Torre JR, Stahl DA . (2009). Ammonia oxidation kinetics determine niche separation of nitrifying Archaea and Bacteria. Nature 461: 976–979.
Martin AP . (2002). Phylogenetic approaches for describing and comparing the diversity of microbial communities. Appl Environ Microbiol 68: 3673–3682.
Parmesan C . (2006). Ecological and evolutionary responses to recent climate change. Annu Rev Ecol Evol Syst 37: 637.
Porporato A, Daly E, Rodriguez-Iturbe I . (2004). Soil water balance and ecosystem response to climate change. Am Nat 164: 625–632.
Price MN, Dehal PS, Arkin AP . (2009). FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol Biol Evol 26: 1641–1650.
Ratkowsky DA, Olley J, McMeekin TA, Ball A . (1982). Relationship between temperature and growth rate of bacterial cultures. J Bacteriol 149: 1–5.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB et al. (2009). Introducing Mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75: 7537–7541.
Shaver GR, Canadell J, Chapin III FS, Gurevitch J, Henry G . (2000). Global warming and terrestrial ecosystems, a conceptual framework for analysis. BioScience 50: 871.
Wan S, Hui D, Wallace LL, Luo Y . (2005). Direct and indirect effects of experimental warming on ecosystem carbon processes in a tallgrass prairie. Glob Biogeochem Cycles 19: GB2014.
Wan S, Luo Y, Wallace LL . (2002). Changes in microclimate induced by experimental warming and clipping in tallgrass prairie. Glob Change Biol 8: 754–768.
Youssef NH, Elshahed MS . (2008). Diversity rankings among bacterial lineages in soil. ISME J 3: 305–313.
Acknowledgements
We thank Dr Ed Kessler and the University of Oklahoma for access to the Kessler Farm Field Laboratory, Jessica Sieber for editorial assistance and Dr Boris Wawrik and Dr Liz Karr for use of lab equipment. This work was supported by the National Science Foundation Microbial Observatories Program (Grant EF0801858) and the National Academy of Sciences (Grant PGA P280292). Sequence Data: Pyrosequencing data can be accessed through Genbank Sequence Read Archive accession number SRA11007.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Sheik, C., Beasley, W., Elshahed, M. et al. Effect of warming and drought on grassland microbial communities. ISME J 5, 1692–1700 (2011). https://doi.org/10.1038/ismej.2011.32
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2011.32
Keywords
This article is cited by
-
Nitrogen deposition mediates more stochastic processes in structuring plant community than soil microbial community in the Eurasian steppe
Science China Life Sciences (2024)
-
Climate change and cropland management compromise soil integrity and multifunctionality
Communications Earth & Environment (2023)
-
Seasonal precipitation and soil microbial community influence plant growth response to warming and N addition in a desert steppe
Plant and Soil (2023)
-
Unboxing the black box—one step forward to understand the soil microbiome: A systematic review
Microbial Ecology (2023)
-
Microbial diversity and ecological interactions of microorganisms in the mangrove ecosystem: Threats, vulnerability, and adaptations
Environmental Science and Pollution Research (2022)