Abstract
In the microbial reductive dechlorination of tetrachloroethene (PCE) and trichloroethene (TCE), dechlorinators usually produce cis-1,2-dichloroethene (cis-DCE) as the predominant product or an intermediate. This study shows that dechlorination of PCE and TCE can also lead to the generation of trans-1,2-dichloroethene (trans-DCE) by a co-culture MB. During its enrichment process, the ratio of trans- to cis-DCE increased from 1.4 (±0.1):1–3.7 (±0.4):1, whereas the TCE reductive dechlorination rate went up from ∼26.2 to ∼68.8 μmol l−1 day−1. PCR–denaturing gradient gel electrophoresis (PCR-DGGE) revealed that the increased ratio of trans- /cis-DCE was well correlated with the increased proportions of Dehalococcoides and the disappearance of Desulfuromonas during the enrichment process. As shown by PCR-DGGE, similar Dehalococcoides species were consistently present in another three sediment-free cultures with various trans- /cis-DCE ratios. The 16S rRNA gene sequence of this Dehalococcoides sp. in co-culture MB is 100% identical (over 1489 bp) to that of Dehalococcoides ethenogenes strain 195 (CP000027), which belongs to the Cornell subgroup of the Dehalococcoides cluster. The other bacterium in this co-culture MB was a Sedimentibacter species, which showed no PCE or TCE dechlorination activity. Results from this study show that microbial dechlorination of chloroethenes by this particular subgroup of Dehalococcoides could result in significant accumulation of trans-DCE in the environment if no trans-DCE dechlorinators coexist in the contaminated sites.
Similar content being viewed by others
Introduction
Large quantities of chlorinated organic solvents tetrachloroethene (PCE) and trichloroethene (TCE) have been released into the environment because of their widespread usage (for example, metal degreasing or dry cleaning) (Abelson, 1990). Under anaerobic conditions, PCE and TCE can be dechlorinated to dichloroethenes (DCEs) by a variety of microbes (for example, Dehalobacter, Desulfuromonas, Sulfurospirillum and Dehalococcoides) (Maymó-Gatell et al., 1997; He et al., 2003b, 2005; Duhamel et al., 2004; Smidt and de Vos, 2004; Sung et al., 2006). However, only Dehalococcoides species are involved in the complete reductive dechlorination of PCE/TCE beyond DCEs to vinyl chloride (VC) and ethene (Smidt and de Vos, 2004). The functional reductive dehalogenase (RDase) genes in Dehalococcoides sp. code for proteins that catalyze these dechlorination steps, such as pceA (for PCE-to-TCE), tceA (for TCE-to-VC), bvcA and vcrA (for DCEs-to-ethene) genes (Magnuson et al., 1998; Krajmalnik-Brown et al., 2004; Müller et al., 2004). In particular, Dehalococcoides sp. strains, BAV1 and FL2, are able to dechlorinate all DCE isomers to VC or ethene (He et al., 2003b, 2005). Multiple Dehalococcoides strains have also been found in mixed cultures that work together to dechlorinate PCE completely to ethene (Hölscher et al., 2004; Waller et al., 2005; Holmes et al., 2006). To date, the characterized PCE and TCE dechlorinators usually produce cis-1,2-dichloroethene (cis-DCE) predominantly and trans-1,2-dichloroethene (trans-DCE) negligibly (Smidt and de Vos, 2004). However, trans-DCE was found in at least 563 sites of the 1430 National Priority List Superfund sites identified by the US Environmental Protection Agency, whereas cis-DCE was detected only in 146 locations (ATSDR, 2007). Ratios of trans- /cis-DCE ranging from (1 to 7):1 have been documented in the TCE-contaminated subsurface of the Naval Air Force Station (Fort Worth, TX, USA) (http://pubs.usgs.gov/sir/2005/5176/) and in the TCE-contaminated wells of Key West, Florida (SWMU9, 2002). Significant amounts of trans-DCE were also detected at concentrations of up to 18 000 μg l–1 in groundwater samples taken from the upper tertiary aquifer in Bitterfeld (Germany) (Nijenhuis et al., 2007). The accumulated trans-DCE in the chloroethene-contaminated sites tends to persist and disperse in the subsurface (He et al., 2003a; Smidt and de Vos, 2004), hindering the complete removal of chloroethenes.
A few laboratory-scale studies reported that certain microorganisms in microcosms or mixed cultures could produce more trans- than cis-DCE during the reductive dechlorination of PCE/TCE (Löffler et al., 1997; Griffin et al., 2004; Miller et al., 2005; Kittelmann and Friedrich, 2008). For instance, a number of uncultured microbes of the Dehalococcoides sp. and DF-1 in the Chloroflexi cluster were capable of producing trans- /cis-DCE in various ratios ((1.3–3.5):1) when fed with PCE/TCE (Futamata et al., 2007). Although some trans-DCE may be produced through abiotic processes (Arnold and Roberts, 1998; ATSDR, 2007), a large fraction of accumulated trans-DCE at contaminated sites could be a result of microbial reductive dechlorination of PCE and TCE. Enrichment of such dechlorinating bacteria, in particular of the genus Dehalococcoides, has been proven to be laborious because of their long doubling time (>1 day). Therefore, limited information is available for dechlorinating microbes possessing specific RDase(s) involved in trans-DCE production and its further detoxification. So far, only Dehalococcoides sp. strain BAV1 could completely dechlorinate trans-DCE to the benign ethene (He et al., 2003b). Owing to the persistent nature of trans-DCE and limited information on trans-DCE-dechlorinating bacteria, it would be of interest to look further into trans-DCE detoxification by multiple dechlorinators when trans-DCE was formed predominantly at the contaminated sites.
This study describes dechlorination of PCE/TCE to various amounts of trans-DCE by Dehalococcoides-containing microcosms, enrichment and a co-culture MB. The microbial populations associated with various trans- /cis-DCE generation ratios were investigated in detail. We have also identified the first Dehalococcoides sp. belonging to the Cornell subgroup responsible for trans- and cis-DCE production in culture MB. In contrast to previous reports linking the Dehalococcoides sp. to VC and ethene generation, in this study, we concluded that the specific Dehalococcoides sp., which produced trans-DCE predominantly, could not dechlorinate DCEs further to VC and ethene. However, complete dechlorination of PCE to ethene through predominant intermediate trans-DCE was achieved by co-inoculating the enrichment culture MB and a trans-DCE-dechlorinating culture 11a that contained multiple Dehalococcoides sp. The study of trans-DCE-producing consortia will provide more information on the diversity of dechlorinators involved in the generation of various dechlorination products (such as trans-DCE, VC or ethene) on exposure to PCE/TCE. These findings could then be extrapolated to contaminated sites and potentially aid in the determination of the type of bioremediation strategy, such as bioaugmentation, with the appropriate dechlorinators.
Materials and methods
Chemicals
The chlorinated ethenes, ethene, and other chemicals were purchased from Sigma-Aldrich (Saint Louis, MO, USA) with a minimum purity of 99.5%. Hydrogen was produced from a hydrogen generator (NM-H250, Schmidlin-DBS AG, Neuheim, Switzerland) and used as indicated.
Microcosm preparation
Sediments or slurry used for setting up microcosms were collected from San Francisco Bay Area (CA, USA), Sungei Buloh Wetland Reserve (Singapore), Wuhan industrial districts (Hubei, China) and Guiyu landfill sites receiving electrical waste (Guangdong, China). Triplicate microcosm studies were conducted in 60-ml serum bottles containing ∼10 g of sediments or slurry, 25 ml autoclaved, bicarbonate-buffered mineral salts medium reduced with L-cysteine (0.2 mM), sodium sulfide (0.2 mM) and DL-dithiothreitol (0.5 mM) as previously described (Wolin et al., 1963; Cole et al., 1994; Löffler et al., 1997; He et al., 2002; He et al., 2003a). The bottles were sealed with black butyl rubber septa (Geo-Microbial Technologies, Inc, Ochelata, OK, USA) and secured with aluminum crimp caps. All microcosms in triplicates were amended with 10 mM of lactate and 55 mM of PCE per bottle. Abiotic controls were carried out by autoclaving another set of microcosm bottles that were set up in the same way as the sample bottles. All bottles were inverted and incubated quiescently in the dark at room temperature.
Culture and growth conditions
After screening the above microcosms, two microcosms MB (San Francisco, CA, USA) and 11a (Hubei, China) were selected for further enrichment as they showed rapid dechlorination rates. After transferring 20 times in the presence of PCE or TCE, serial dilutions (10%, v/v) were used to enrich the cultures, which were carried out in 20-ml vials filled with 10 ml of mineral salts medium. In addition, ampicillin (50–300 mg l–1) (Maymó-Gatell et al., 1997; He et al., 2003b) was added to the dilution series spiked with TCE (0.2 mM), acetate (10 mM) and hydrogen (500 000 ppmv). Initially, the cultures from the first three consecutive transfers received 200 mgl−1 of ampicillin. Subsequently, three consecutive dilution-to-extinction series and agar shakes were conducted. During these treatments, >50 colonies were picked up and re-inoculated back to liquid medium to test their dechlorination activity on exposure to TCE. Dechlorination time-course studies were conducted in 160-ml serum bottles containing 100 ml of mineral salts medium amended with TCE or PCE (∼50 μmoles, nominal concentration 0.5 mM), lactate (10 mM) or acetate (10 mM)/hydrogen (500 000 ppmv), a vitamin solution including 0.05 mg l–1 of vitamin B12 (He et al., 2007), and 2% of inocula. All time-course studies were conducted in triplicates along with an extra abiotic control. All bottles were inverted and incubated quiescently in the dark at 30 °C.
Analytical methods
Chloroethenes and ethene were measured with a gas chromatograph (GC-6890, Agilent, Wilmington, DE, USA) equipped with a flame ionizing detector and a capillary column (GS-GasPro, 30-m length, 0.32-mm i.d., J&W Sci., Folsom, CA, USA). The oven temperature was initially held at 50 °C for 2 min, increased at 30 °C min−1 to 220 °C, and held for 1 min. The supernatants carrying volatile fatty acids (VFAs) were obtained by centrifugation of 1 ml of sample at 14 000 g for 10 min at 4 °C. VFAs were determined on a high-performance liquid chromatograph (Agilent 1100 HPLC system, Palo Alto, CA, USA) equipped with a UV detector (set at 210 nm). Separation of VFAs was conducted on an organic acid analysis column Aminex HPX-87 H (300 × 7.8 mm, Bio-Rad, Hercules, CA, USA) at 40 °C, and 5 mM H2SO4 was used as the eluent at a flow rate of 0.5 ml min−1.
DNA extraction and PCR amplification
Cells for DNA extraction were collected periodically from 1 ml of culture samples by centrifugation (5 min at 20 000 g, 4 °C) in DNase/RNase-free microcentrifuge tubes. Cell pellets were stored at −20 °C until further processing. The genomic DNA was extracted with Qiagen DNeasy Tissue Kit (QIAGEN GmbH, Hilden, Germany) according to the manufacturer's instructions. PCR (Eppendorf, Hamburg, Germany) amplification was carried out by targeting the genomic DNA with universal Eubacterial primers 8F (Zhou et al., 1995) and 1541R (Lane et al., 1985) to obtain the 16S rRNA genes. Genomic DNA was also screened with the following species-specific primer pairs: Dehalococcoides, Desulfuromonas and Dehalobacter (Holliger et al., 1998; Löffler et al., 2000; Bunge et al., 2003) as previously described. The primer sequences used in this study were included in the Supplementary material Table S1.
The presence of chloroethene RDase genes (pceA, tceA, bvcA and vcrA) in the cultures of this study was examined as previously described (Magnuson et al., 1998; Krajmalnik-Brown et al., 2004; Müller et al., 2004). The PCR products were subsequently visualized on a Molecular Imager Gel Doc XR System (Bio-Rad) or on a Bioanalyzer using a DNA7500 Labchip Kit (Agilent Technologies Inc, Palo Alto, CA, USA).
The 16S rRNA genes copies of Dehalococcoides sp. in co-culture MB were determined by quantitative real-time PCR (qPCR) (ABI 7500 Fast Real-Time PCR system, Foster, CA, USA) as previously described (He et al., 2003a, 2003b; Sung et al., 2006). Standard curves spanned a range of 102–107 gene copies per microliter of template DNA with a R2 linear regression of 99.9%.
PCR–DGGE
PCR–denaturing gradient gel electrophoresis (PCR-DGGE) analyses were carried out for both universal bacterial 16S rRNA gene (with the primer pair 341F-GC and 534R) and Dehalococcoides-specific 16S rRNA gene fragments (1F-GC and 259R primer set) with a touch-down thermal program as previously described (Duhamel et al., 2004). PCR-amplified fragments were electrophoresed on an 8% polyacrylamide gel with a 30–60% urea–formamide gradient for 16 h at 120 V and 60 °C.
Clone library
A clone library of 16S rRNA genes of co-culture MB was established by using the TOPO-TA cloning kit (Invitrogen, Carlsbad, CA, USA) and all further clone-based experiments were carried out as previously described (Löffler et al., 2000; He et al., 2003a). The clones' purified plasmid DNA concentrations were measured by Nanodrop-1000 (NanoDrop Technologies Inc, Wilmington, DE, USA). The extracted plasmid DNA was subjected to restriction fragment length polymorphism analysis with the restriction endonucleases HhaI and MspI (NEB, Ipswich, MA, USA). The 16S rRNA gene inserts representing groups of distinct enzyme restriction pattern were subsequently sequenced with an ABI 3100 Sequencer (Applied Biosystems, Foster City, CA, USA) by using primers M13F-20, M13R-24, 533F, 529R and 907F (http://www.genomics.msu.edu). Sequences were aligned with BioEdit assembly software (http://www.mbio.ncsu.edu/BioEdit/bioedit.html) and were analyzed with BLASTN (http://www.ncbi.nlm.nih.gov/). The nearly complete Dehalococcoides-like 16S rRNA gene sequence (1489 bp) was deposited under the GenBank accession number EU073964. The other bacterium was characterized as being a Sedimentibacter species under GenBank accession number FJ593657.
Results
Dechlorination of PCE to predominant trans-DCE
From the 23 microcosms established with sediments collected from various locations, 3 (namely, MB, 11a and GY from CA, USA; Hubei, China; and Guangdong, China; respectively) generated more trans-DCE than cis-DCE in the reductive dechlorination of PCE, whereas SB (Singapore) generated relatively lower trans-/cis-DCE ratio (Table 1). No PCE dechlorination or lactate fermentation was observed in the abiotic control bottles (autoclaved microcosms). Microcosms, MB, SB and GY, generated trans-DCE and cis-DCE as the final dechlorination products, whereas 11a could dechlorinate PCE to ethene through intermediates trans- and cis-DCEs. Among these microcosms, MB exhibited the highest PCE dechlorination rate of 26.2 μmol l−1 day−1 with a trans- /cis-DCE generation ratio of (1.4±0.1):1 and was selected for further transfers and investigation.
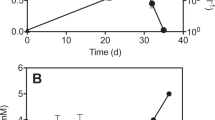
Enrichment culture MB was obtained by sequentially transferring the above PCE-to- trans/cis-DCE-dechlorinating microcosm. Methane was below detection limit after 10 transfers with PCE as substrate, though it was detected in the microcosms and the previous batches of sediment-free culture. The subsequent 10 transfers were fed with TCE and the time-course studies on the dechlorination of TCE were carried out thereafter. In defined mineral salts medium amended with lactate (10 mM) and TCE (0.2 mM), culture MB dechlorinated ∼50 μmoles of TCE completely to trans-/cis-DCEs with a ratio of (1.43±0.04):1 within 20 days, and a dechlorination rate of 30.6 μmol l−1 day−1 (Figure 1a). Dechlorination of TCE to trans- and cis-DCEs (in a ratio of 1.4:1) also occurred when culture MB grew in the same mineral salts medium amended with acetate (10 mM) only, or acetate (10 mM) and H2 (170 000 ppmv). The sole acetate-grown culture (no H2 supplied) showed the slowest dechlorination rate. Thereafter, the acetate-grown sub-culture was used as inocula for subsequent transfer to fresh acetate and H2 medium so as to facilitate the enrichment and isolation of the dechlorinators.
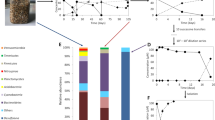
Reductive dechlorination of trichloroethene (TCE) to trans-1,2-dichloroethene (trans-DCE) and cis-1,2-dichloroethene (cis-DCE) by culture MB at different enrichment phases. (a) Enrichment culture MB after 20 transfers in lactate-amended medium. (b) Further enrichment culture MB grown with acetate (10 mM) and H2 (260 000 ppmv) after ampicillin treatment. (c) Highly enriched culture MB after three more transfers in acetate-/H2-amended medium. No vinyl chloride, ethene, ethane or methane was detected on extended incubation. Datum points were averaged from triplicate cultures. Error bars indicate s.d. and are not shown when they are smaller than the symbol.
When culture MB was treated with ampicillin (50 mg l–1) in the mineral salts medium amended with acetate and H2, dechlorination of TCE occurred at a slower rate as compared with dechlorination without ampicillin. The treatment of acetate/H2 cultures with 0–300 mg l–1 of ampicillin resulted in the variation of trans-DCE to cis-DCE ratio, ranging from (1.4 to 3.0):1. The highest ratio was achieved in the culture receiving 200 mg l–1 of ampicillin dosage. After growing in ampicillin-amended medium for two more transfers, the active cultures were then continuously transferred to medium amended with acetate and H2 only. Consequently, culture MB was capable of dechlorinating ∼50 μmoles of TCE to trans- and cis-DCEs with an increased ratio of 3.7:1 (Figure 1b) within 15 days. However, when culture MB was transferred back to the mineral salts medium amended with lactate, an extended lag phase (∼13 days) was observed and the dechlorination of TCE (∼50 μmoles) took another 17 days in all the triplicate bottles (data not shown). Acetate and propionate were detected as the fermentation products measured with high performance liquid chromatograph, indicating that certain fermentative microbes had yet to be diluted out of the culture.
After three more transfers in the absence of antibiotics, the ampicillin-treated culture MB with acetate and H2 achieved its highest TCE dechlorination rate (68.8 μmoles l−1 day−1) but the trans-/cis-DCE generation ratio remained unchanged (Figure 1c). Culture MB was continuously transferred to fresh medium (every 3–4 weeks) over a period of 2 years and showed consistent PCE/TCE dechlorination rates and trans-/cis-DCE generation ratio. At this stable dechlorination stage, culture MB was observed under light microscopy, which showed the prevalence of small disc-shaped bacteria and a lower abundance of rod-shaped bacteria. The size of the disc-shaped microbes coincides with that of the reported Dehalococcoides sp. (∼1.0 μm in diameter). Attempts had been made to isolate each of them by both serial dilutions and agar shakes amended with acetate and H2. However, only the rod-shaped bacteria were able to grow in pure culture successfully. Its 16S rRNA gene sequence was shown to be closely (99% identity over 1504 bp, 5 bp difference) related to Sedimentibacter sp. C7, which was reported to grow in co-culture with a Dehalobacter species (van Doesburg et al., 2005). This isolated Sedimentibacter sp. did not show dechlorination activity on PCE or TCE. Therefore, culture MB should mainly consist of the PCE-dechlorinating Dehalococcoides and the Sedimentibacter sp.
Identification of the trans-DCE-producing microbes
To identify the dechlorinating microbes in the sediment-free cultures, MB, 11a, GY and SB, genus-specific primers targeting the 16S rRNA genes of Dehalobacter, Desulfuromonas and Dehalococcoides were tested on genomic DNA of the above cultures by PCR. When targeted with Dehalococcoides-specific primers, the PCR amplicons yielded the expected DNA size of 620 bp for all the cultures (data not shown). When using Desulfuromonas-specific primers to amplify the genomic DNA of the above trans-DCE-producing cultures, only culture MB exhibited a very weak band of 815 bp. However, the Desulfuromonas-like species was no longer detected in MB when the ratio of trans- to cis-DCE reached ∼3.7:1. Amplification of the genomic DNA using Dehalobacter-specific primers yielded no amplicons for all the cultures. Therefore, the PCR analysis based on 16S rRNA genes suggests that the rod-shaped Desulfuromonas or Dehalobacter species may not contribute to trans-DCE formation, and further confirm the existence of Dehalococcoides-like species in the above cultures.
Dehalococcoides species diversity versus the ratio of trans- to cis-DCE
To understand the elevated ratio (1.4:1–3.7:1) of trans-/cis-DCE in culture MB, PCR-DGGE was used to track the community structures with universal bacterial primer pair and the diversity of Dehalococcoides sp. with genus-specific primer pair by targeting the genomic DNA of culture MB at different enrichment stages. Figure 2a shows the microbial community transitions from mixed culture (before ampicillin treatment), highly enriched culture (after one-time ampicillin treatment), to apparent co-culture (after ampicillin treatment), reflected by detecting multiple DGGE bands to two major bands. The band on the top was getting thicker, which turned out to be Dehalococcoides sp. This was verified by the positive control, plasmid DNA containing the 16S rRNA gene fragment of Dehalococcoides ethenogenes strain 195. The second major band in Figure 2a corresponded to the minor group in the co-culture, Sedimentibacter, which was confirmed by the positive control of this isolate. It is to be noted that the Desulfuromonas-like band disappeared in the post-amplicillin-treated MB cultures (Figure 2a). Only one band representing the Cornell subgroup of Dehalococcoides sp. (Figure 2b) was detected in the trans- and cis-DCE-producing culture MB. Therefore, the detected Desulfuromonas-like species before amplicillin treatment (Figure 1a) could have contributed to cis-DCE production in the mixed culture MB. In all, the DGGE profiles confirmed that culture MB consisted of the Cornell subgroup of Dehalococcoides and Sedimentibacter only, whereas the former was responsible for trans-DCE production from PCE/TCE.
PCR–denaturing gradient gel electrophoresis (PCR-DGGE) profiles of trans-1,2-dichloroethene (trans-DCE)-producing cultures. (a) DGGE profile of PCR amplified with a universal bacterial primer pair targeting genomic DNA of culture MB. (b) DGGE profile of PCR amplified with a Dehalococcoides-specific primer pair targeting genomic DNA of culture MB. The template DNA for lane 1, 2 and 3 was extracted from cultures as shown in the time-course studies of Figures 1a–c, respectively. Lane 1, culture MB before ampicillin treatment; lane 2, culture MB after one-time ampicillin treatment; lane 3: culture MB after ampicillin treatment; lane 4, positive control of D. ethenogenes 195 clone DNA; lane 5, a trichloroethene (TCE)-to-cis-1,2-dichloroethene (cis-DCE)-to-vinyl chloride culture, which shared 100% 16S rRNA gene as Dehalococcoides species strain FL2; lane 6, Desulfuromonas species strain BB1; lane 7, an isolate of Sedimentibacter. (c). PCR-DGGE profile of Dehalococcoides sp. present in different TCE-dechlorinating cultures. The number on the top of the lane indicates the ratio of trans-DCE to cis-DCE produced from various microcosms as described in Table 1. Dehalococcoides sp. strain FL2-like culture serves as a control. NA, no trans-DCE was accumulated during the dechlorination of TCE.
Sediment-free cultures obtained from microcosms shown in Table 1 also generated trans- and cis-DCEs in various ratios. The presence of multiple Dehalococcoides sp. bands (for example, GY and 11a) explained the various ratios of trans-/cis-DCE, as shown in this study (Figure 2c). Figure 2c also shows the stronger the MB-like bands, the higher trans-/cis-DCE generation ratios in the sediment-free cultures; conversely, the stronger the FL2-like bands, the lower the trans-/cis-DCE ratios in these cultures.
Clone library and sequence analysis of culture MB
To further determine the phylogeny of the PCE-to-trans/cis-DCEs dechlorinator(s), a clone library based on 16S rRNA genes was established with the genomic DNA extracted from the apparent co-culture MB fed with acetate and H2. The restriction fragment length polymorphism analysis of 57 clones revealed two different digestion patterns (56:1). Twenty-eight of the 56 clones, and the last single clone were sequenced for their 16S rRNA gene inserts in the plasmids. After aligning the sequences with the BioEdit assembly software, the contigs were analyzed using BLASTN and compared with known existing 16S rRNA gene databases. Results showed that the 56 clones possessed identical 16S rRNA gene sequence, which shares 100% identity of the 16S rRNA gene sequence of D. ethenogenes strain 195 (accession number CP000027) from its complete genome. The 16S rRNA gene sequence of the uncultured Dehalococcoides sp. strain MB (the Cornell subgroup of Dehalococcoides sp.) was deposited in GenBank under accession number of EU073964 (1489 bp). In addition to the above Dehalococcoides sp. found in the culture MB, the sequence of the other clone showed that it belonged to the genus of Sedimentibacter as isolated in pure culture.
The presence of the known RDase genes was also examined in the trans- and cis-DCE-producing cultures. The following genes, pceA, tceA, bvcA or vcrA, were not detected in culture MB when fed with PCE or TCE. The results suggested that a novel RDase gene, instead of pceA or tceA gene, might be responsible for the production of trans-DCE by the Dehalococcoides in culture MB. Therefore, on the basis of the above analysis (for example, constant trans- /cis-DCE ratio, light microscopy, DGGE, clone library, RDase genes), we were able to ascertain that the culture was further enriched to a co-culture MB consisting only of Dehalococcoides sp. of the Cornell subgroup and Sedimentibacter sp.
Growth of Dehalococcoides-like species in co-culture MB
Quantitative real-time PCR was carried out to further confirm that Dehalococcoides-like species was responsible for the generation of trans-DCE during the dechlorination of TCE. The Dehalococcoides cells were measured by targeting the genomic DNA extracted from co-culture MB. qPCR results showed the increase of Dehalococcoides 16S rRNA gene copies with the dechlorination of TCE (Figure 3). The total Dehalococcoides cell number reached 2.54 × 108 cells per ml from the initial 5.91 × 106 cells per ml (∼43 times increase) in the co-culture, verifying that the increase of the Dehalococcoides 16S rRNA gene copies was directly correlated with the dechlorination of TCE to the predominant trans-DCE.
Complete dechlorination of TCE to ethene through trans- and cis-DCEs by culture MB and 11a
Similar to culture MB, the sequential transfers of another microcosm (Hubei, China) generated enrichment culture 11a, which was capable of completely dechlorinating TCE to ethene through intermediates trans- and cis-DCEs (in a ratio of 1.7:1) in lactate-amended mineral salts medium (Figure 4a). After 40 days, the generated DCEs were completely dechlorinated to ethene. DGGE profile showed that culture 11a contained multiple Dehalococcoides sp. (Figure 2c), one of which was MB-like to produce trans-DCE, whereas another may function to further dechlorinate trans-DCE. With the trans-DCE-dechlorinating mixed culture 11a in hand, it was inoculated together with culture MB to the bottles spiked with TCE to examine the possibility of complete reductive dechlorination of the persistent trans-DCE generated biologically by culture MB. As shown in Figure 4b, complete dechlorination to ethene through trans-DCE occurred within 80 days. The maximum trans- to cis-DCE ratio reached ∼1.75:1 on day 11 when cis-DCE concentration started to decrease. During the reductive dechlorination process, trans-DCE was found to be the main intermediate, accumulating up to 18.6 μmoles on day 21, whereas small amounts of cis-DCE and VC (about 6.9 and 7.8 μmoles, respectively) appeared and the amount of 1,1-DCE produced was negligible. After day 21, trans-DCE started to be dechlorinated to ethene without the accumulation of VC.
Reductive dechlorination of trichloroethene (TCE) to ethene through trans-1,2-dichloroethene (trans-DCE) predominantly. (a) Culture 11a alone amended with lactate. (b) Culture MB and 11a amended with acetate (10 mM) and H2 (400 000 ppmv). (Datum points were averaged from triplicate cultures. Error bars indicate s.d. and are not shown when they are smaller than the symbol). Figure 4b shares the same symbols as Figure 4a.
Discussion
In this study, a highly enriched PCE-to-trans/cis-DCE-dechlorinating co-culture MB was characterized and the growth of the newly identified trans-DCE-producing Dehalococcoides sp. was verified by qPCR. A number of identified trans-DCE-producing Dehalococcoides sp., including the Cornell (in this study) and Pinellas (Griffin et al., 2004) subgroups, showed a common inability to dechlorinate PCE/TCE beyond DCEs to VC and ethene. The dechlorination pathway of MB is different from the previously discovered Dehalococcoides sp. linking to VC or ethene production from polychloroethenes, including those that produced predominant 1,1-DCE isomers during the reductive dechlorination of TCE (Zhang et al., 2006). Complete detoxification of PCE/TCE through intermediates trans- and cis-DCEs also occurred when the required dechlorinating microbes were present in the community, as shown in the mixed culture of 11a and MB. The results from this study indicate that accumulation of trans-DCE through microbial reductive dechlorination in contaminated sites could be significant when the trans-DCE dechlorinator(s) are absent or are operating at a lower rate compared with that of cis-DCE and VC dechlorinators. The chloroethene distribution in contaminated sites is significantly affected by different types of indigenous dechlorinators (Smidt and de Vos, 2004). The trans-DCE-generating microbe discovered in this study would enhance the current understanding of diverse dechlorinating capabilities of the Dehalococcoides genus.
During the enrichment process from sediment-free culture to co-culture MB, the ampicillin-treatment step significantly increased the culture's TCE dechlorination rates (from 26.2 to 68.8 μmoles l−1 day−1) and the ratio of trans-DCE to cis-DCE (from (1.4±0.1):1 to (3.7±0.4):1). Previous studies showed that sediment-free cultures generated trans-DCE to cis-DCE in ratios of (3±0.5):1 (Griffin et al., 2004) and (1.3±0.2):1 (Miller et al., 2005), accompanying with an average TCE dechlorination rate of 4.8 μmoles l−1 day−1 (Griffin et al., 2004) and 20 μmoles l−1 day−1 (Miller et al., 2005), respectively. The increased dechlorination rates and trans- to cis-DCE ratios for culture MB could be explained by that (i) the competitors for electron donor H2 (for example, methanogens) were diluted out; (ii) culture MB contained higher number of trans-DCE-producing Dehalococcoides cells, but less or negligible amount of the cis-DCE-producing Desulfuromonas-like cells (revealed by DGGE profiles), compared with the previous cultures. This is also supported by the fact that the majority of the clones (56 out of 57) belong to Dehalococcoides and the Desulfuromonas band disappeared in the co-culture MB. However, culture MB was not a pure culture yet as lactate fermenters (for example, Sedimentibacter) were still present. This is reflected by their recovered dechlorination activity after an extended incubation time when fed from acetate/H2- back to lactate-amended medium.
By sequencing of the representative clones (established with PCR 16S rRNA gene inserts), Dehalococcoides sp. was identified to be responsible for PCE/TCE dechlorination to trans- and cis-DCEs. On the basis of the 16S rRNA gene sequence similarity, Dehalococcoides sp. MB belongs to the Cornell subgroup of the Dehalococcoides cluster (Hendrickson et al., 2002), which is different from the trans- and cis-DCE-producing Dehalococcoides populations in the Pinellas group (Griffin et al., 2004) (refer to Supplementary Table S2). Supplementary Table S2 also shows that bacterium DF-1 and TFCC groups (T-RF 513 bp and T-RF 143 bp), which are distantly related to Dehalococcoides sp., contributed to the production of trans-DCE. We could not exclude the possibility of other bacteria having a role in trans-DCE formation in the environment. However, this study provides conclusive proof that the Cornell subgroup of Dehalococcoides sp. in co-culture MB grew with the dechlorination of TCE to predominant trans-DCE by both qualitative (for example, DGGE and clone library) and quantitative (real-time PCR) molecular tools. On the other hand, owing to the high similarity of 16S rRNA gene sequences for the genus Dehalococcoides, it is difficult to rule out the presence of other Dehalococcoides sp. Nevertheless, the Cornell subgroup of Dehalococcoides has never been shown to have this dechlorination ability of producing trans-DCE predominantly. The stable trans-/cis-DCE ratio, two morphologies by light microscopy and the culture-independent approaches (DGGE, clone sequencing, RDase gene analysis) strongly suggest that co-culture MB contained only one Dehalococcoides sp. The interactions between Dehalococcoides and Sedimentibacter are not clear but may be nutritional in nature. Isolation of this particular strain of Dehalococcoides will lead to an improved understanding of the nutritional requirements and physiological characterization of this novel Dehalococcoides sp. Therefore, this study and previous reports (Griffin et al., 2004; Miller et al., 2005; Kittelmann and Friedrich, 2008) suggest that these diverse trans-DCE-producing dechlorinators belong to the green non-sulfur bacteria, including Dehalococcoides, the Tidal Flat Chloroflexi cluster and bacterium DF-1.
Despite the high conservation of the 16S rRNA gene among the Dehalococcoides sp., RDase genes (pceA, tceA, bvcA and vcrA) have been identified to differentiate reductive dechlorination of chloroethenes (Magnuson et al., 1998; Krajmalnik-Brown et al., 2004; Müller et al., 2004). In the enrichment process of culture MB, no pceA or tceA gene was detected in its genomic DNA. This observation suggests that the tceA gene in D. ethenogenes 195 is different from the functional trans-DCE-producing gene(s) in the Dehalococcoides sp. of culture MB. Similarly, bacterium DF-1 was reported to dechlorinate PCE/TCE to significant amounts of cis- and trans-DCEs, whereas no tceA gene was reported in its genome (Miller et al., 2005). Therefore, the trans-DCE-producing gene could be quite different from the currently identified RDase genes, which may also be potentially acquired through horizontal gene transfer as suggested for the known RDases in Dehalococcoides sp. (Regeard et al., 2005; Seshadri et al., 2005). PCR-DGGE identified a common band, whose presence may indicate trans-DCE production from PCE or TCE. However, the detection of this band in DGGE would not guarantee the generation of trans-DCE in the Dehalococcoides sp., for example, D. ethenogenes strain 195.
Findings in this and previous studies suggest that diverse trans-DCE-producing Dehalococcoides sp. are present at chloroethene-contaminated sites and these Dehalococcoides sp. are capable of producing high levels of trans-DCE (Löffler et al., 1997; Griffin et al., 2004; Miller et al., 2005; Futamata et al., 2007; Kittelmann and Friedrich, 2008). For the first time, we reported that the trans-DCE-producing Dehalococcoides sp. was unable to generate VC and ethene during the dechlorination process through characterization of the co-culture MB (∼98% of the bacteria belonging to the Cornell subgroup of Dehalococcoides sp. and ∼2% affiliated with Sedimentibacter), whereas previous Dehalococcoides cultures were linked to VC and ethene generation. Hence, the large amounts of trans-DCE detected in the TCE-contaminated sites might be a result of the presence of trans-DCE-producing microbes and the lack of trans-DCE dechlorinators. Fortunately, culture 11a in this study proves to be a promising candidate in the complete dechlorination of PCE/ TCE to harmless ethene through the cooperation of multiple Dehalococcoides sp. Thus great care must be taken when bioremediation techniques are being applied to such trans-DCE-formation sites, as the trans-DCE-producing culture (for example, MB) is also capable of using the same electron donor as the other Dehalococcoides strains, such as BAV1. The identification and administration of the microbe(s) responsible for rapid and complete dechlorination at any particular chloroethene-contaminated site may provide a better bioremediation strategy.
References
Abelson PH . (1990). Inefficient remediation of ground-water pollution. Science 250: 73.
Arnold WA, Roberts AL . (1998). Pathways of chlorinated ethylene and chlorinated acetylene reaction with Zn(0). Environ Sci Technol 32: 3017–3025.
ATSDR (2007). Agency for Toxic Substances and Disease Registry. US Public Health Service, U S Department of Health and Human Services: Atlanta, GA.
Bunge M, Adrian L, Kraus A, Opel M, Lorenz WG, Andreesen JR et al. (2003). Reductive dehalogenation of chlorinated dioxins by an anaerobic bacterium. Nature 421: 357–360.
Cole JR, Cascarelli AL, Mohn WW, Tiedje JM . (1994). Isolation and characterization of a novel bacterium growing via reductive dehalogenation of 2-chlorophenol. Appl Environ Microbiol 60: 3536–3542.
Duhamel M, Mo K, Edwards EA . (2004). Characterization of a highly enriched Dehalococcoides-containing culture that grows on vinyl chloride and trichloroethene. Appl Environ Microbiol 70: 5538–5545.
Futamata H, Yoshida N, Kurogi T, Kaiya S, Hiraishi A . (2007). Reductive dechlorination of chloroethenes by Dehalococcoides-containing cultures enriched from a polychlorinated-dioxin-contaminated microcosm. ISME J 1: 471–479.
Griffin BM, Tiedje JM, Löffler FE . (2004). Anaerobic microbial reductive dechlorination of tetrachloroethene to predominately trans-1,2-Dichloroethene. Environ Sci Technol 38: 4300–4303.
He J, Holmes VF, Lee PKH, Alvarez-Cohen L . (2007). Influence of vitamin B12 and cocultures on the growth of Dehalococcoides isolates in defined medium. Appl Environ Microbiol 73: 2847–2853.
He J, Ritalahti KM, Aiello MR, Löffler FE . (2003a). Complete detoxification of vinyl chloride by an anaerobic enrichment culture and identification of the reductively dechlorinating population as a Dehalococcoides species. Appl Environ Microbiol 69: 996–1003.
He J, Ritalahti KM, Yang K-L, Koenigsberg SS, Löffler FE . (2003b). Detoxification of vinyl chloride to ethene coupled to growth of an anaerobic bacterium. Nature 424: 62–65.
He J, Sung Y, Dollhopf ME, Fathepure BZ, Tiedje JM, Löffler FE . (2002). Acetate versus hydrogen as direct electron donors to stimulate the microbial reductive dechlorinaton process at chloroethene-contaminated sites. Environ Sci Technol 36: 3945–3952.
He J, Sung Y, Krajmalnik-Brown R, Ritalahti KM, Löffler FE . (2005). Isolation and characterization of Dehalococcoides sp. strain FL2, a trichloroethene (TCE)- and 1,2-dichloroethene-respiring anaerobe. Environ Microbiol 7: 1442–1450.
Hendrickson ER, Payne JA, Young RM, Starr MG, Perry MP, Fahnestock S et al. (2002). Molecular analysis of Dehalococcoides 16S ribosomal DNA from chloroethene-contaminated sites throughout North America and Europe. Appl Environ Microbiol 68: 485–495.
Holliger C, Hahn D, Harmsen H, Ludwig W, Schumacher W, Tindall B et al. (1998). Dehalobacter restrictus gen. nov. and sp. nov., a strictly anaerobic bacterium that reductively dechlorinates tetra- and trichloroethene in an anaerobic respiration. Arch Microbiol 169: 313–321.
Holmes VF, He J, Lee PKH, Alvarez-Cohen L . (2006). Discrimination of multiple Dehalococcoides strains in a trichloroethene enrichment by quantification of their reductive dehalogenase genes. Appl Environ Microbiol 72: 5877–5883.
Hölscher T, Krajmalnik-Brown R, Ritalahti KM, von Wintzingerode F, Görisch H, Löffler FE et al. (2004). Multiple nonidentical reductive-dehalogenase-homologous genes are common in Dehalococcoides. Appl Environ Microbiol 70: 5290–5297.
Kittelmann S, Friedrich MW . (2008). Novel uncultured Chloroflexi dechlorinate perchloroethene to trans-dichloroethene in tidal flat sediments. Environ Microbiol 10: 1557–1570.
Krajmalnik-Brown R, Hölscher T, Thomson IN, Saunders FM, Ritalahti KM, Löffler FE . (2004). Genetic identification of a putative vinyl chloride reductase in Dehalococcoides sp. strain BAV1. Appl Environ Microbiol 70: 6347–6351.
Lane DJ, Pace B, Olsen GJ, Stahl DA, Sogin ML, Pace NR . (1985). Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci USA 82: 6955–6959.
Löffler FE, Champine JE, Ritalahti KM, Sprague SJ, Tiedje JM . (1997). Complete reductive dechlorination of 1,2-dichloropropane by anaerobic bacteria. Appl Environ Microbiol 63: 2870–2875.
Löffler FE, Sun Q, Li J, Tiedje JM . (2000). 16S rRNA gene-based detection of tetrachloroethene-dechlorinating Desulfuromonas and Dehalococcoides species. Appl Environ Microbiol 66: 1369–1374.
Magnuson JK, Stern RV, Gossett JM, Zinder SH, Burris DR . (1998). Reductive dechlorination of tetrachloroethene to ethene by a two-component enzyme pathway. Appl Environ Microbiol 64: 1270–1275.
Maymó-Gatell X, Chien Y-T, Gossett JM, Zinder SH . (1997). Isolation of a bacterium that reductively dechlorinates tetrachloroethene to ethene. Science 276: 1568–1571.
Miller GS, Milliken CE, Sowers KR, May HD . (2005). Reductive dechlorination of tetrachloroethene to trans-dichloroethene and cis-dichloroethene by PCB-dechlorinating bacterium DF-1. Environ Sci Technol 39: 2631–2635.
Müller JA, Rosner BM, von Abendroth G, Meshulam-Simon G, McCarty PL, Spormann AM . (2004). Molecular identification of the catabolic vinyl chloride reductase from Dehalococcoides sp. strain VS and its environmental distribution. Appl Environ Microbiol 70: 4880–4888.
Nijenhuis I, Nikolausz M, Köth A, Felföldi T, Weiss H, Drangmeister J et al. (2007). Assessment of the natural attenuation of chlorinated ethenes in an anaerobic contaminated aquifer in the Bitterfeld/Wolfen area using stable isotope techniques, microcosm studies, and molecular biomarkers. Chemosphere 67: 300–311.
Regeard C, Maillard J, Dufraigne C, Deschavanne P, Holliger C . (2005). Indications for acquisition of reductive dehalogenase genes through horizontal gene transfer by Dehalococcoides ethenogenes strain 195. Appl Environ Microbiol 71: 2955–2961.
Seshadri R, Adrian L, Fouts DE, Eisen JA, Phillippy AM, Methe BA et al. (2005). Genome sequence of the PCE-dechlorinating bacterium Dehalococcoides ethenogenes. Science 307: 105–108.
Smidt H, de Vos WM . (2004). Anaerobic microbial dehalogenation. Annu Rev Microbiol 58: 43–73.
Sung Y, Ritalahti KM, Apkarian RP, Löffler FE . (2006). Quantitative PCR confirms purity of strain GT, a novel trichloroethene-to-ethene respiring Dehalococcoides isolate. Appl Environ Microbiol 72: 1980–1987.
SWMU9 (2002). Treatability study annual monitoring report. Naval air facility: Key West, Florida. Contract number N62407-94-D-0888, contract task order 0188. Tetra Tech NUS, Inc for Southern Division Naval Facilities Engineering Command.
van Doesburg W, van Eekert MHA, Middeldorp PJM, Balk M, Schraa G, Stams AJM . (2005). Reductive dechlorination of β-hexachlorocyclohexane (β-HCH) by a Dehalobacter species in coculture with a Sedimentibacter sp. FEMS Microbiol Ecol 54: 87–95.
Waller AS, Krajmalnik-Brown R, Löffler FE, Edwards EA . (2005). Multiple reductive-dehalogenase-homologous genes are simultaneously transcribed during dechlorination by Dehalococcoides-containing cultures. Appl Environ Microbiol 71: 8257–8264.
Wolin EA, Wolin MJ, Wolfe RS . (1963). Formation of methane by bacterial extracts. J Biol Chem 238: 2882–2886.
Zhang J, Joslyn AP, Chiu PC . (2006). 1,1-Dichloroethene as a predominant intermediate of microbial trichloroethene reduction. Environ Sci Technol 40: 1830–1836.
Zhou J, Fries MR, Chee-Sanford JC, Tiedje JM . (1995). Phylogenetic analysis of a new group of denitrifiers capable of anaerobic growth on toluene and description of Azoarcus tolulyticus sp. nov. Int J Syst Bacteriol 45: 500–506.
Acknowledgements
This research was supported by the Academic Research Fund from the Singapore Ministry of Education under Project No.: R-288-000-024-112/133 and R-288-000-041-112. The sequences of primers used in this study and the comparison of the 16S rRNA gene sequences for selected chloroethene-dechlorinating isolates and uncultured clones are listed in the supporting information.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Supplementary information
Rights and permissions
About this article
Cite this article
Cheng, D., Chow, W. & He, J. A Dehalococcoides-containing co-culture that dechlorinates tetrachloroethene to trans-1,2-dichloroethene. ISME J 4, 88–97 (2010). https://doi.org/10.1038/ismej.2009.90
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2009.90
Keywords
This article is cited by
-
Interspecies metabolite transfer and aggregate formation in a co-culture of Dehalococcoides and Sulfurospirillum dehalogenating tetrachloroethene to ethene
The ISME Journal (2021)
-
Growth of Dehalococcoides mccartyi species in an autotrophic consortium producing limited acetate
Biodegradation (2018)
-
Isolation, identification and characterization of a novel Ralstonia sp. FD-1, capable of degrading 4-fluoroaniline
Biodegradation (2014)
-
Site-specific pre-evaluation of bioremediation technologies for chloroethene degradation
International Journal of Environmental Science and Technology (2014)
-
Dechlorination and organohalide-respiring bacteria dynamics in sediment samples of the Yangtze Three Gorges Reservoir
Environmental Science and Pollution Research (2013)