Abstract
Sphingopyxis alaskensis is a marine member of the Alphaproteobacteria that is adapted to heterotrophic growth under nutrient-depleted (oligotrophic) conditions. S. alaskensis strain RB2256 is an ultramicrobacterium (cell volume <0.1 μm3), and has a genome size larger than that of the ultramicrobacterium ‘Candidatus Pelagibacter ubique’ HTCC1062 (SAR11 clade of Alphaproteobacteria): 3.35 versus 1.31 Mbp. In this study, we investigate the carbon and nitrogen metabolism of strain RB2256 using an integrated approach that combines growth and enzyme assays, proteomics and genome analysis. S. alaskensis is able to use specific amino acids and putrescine as a sole carbon and nitrogen source, and higher energy-yielding substrates such as glucose and trehalose as carbon sources. Alanine, in particular, emerges as a very important substrate in S. alaskensis metabolism. In an oligotrophic environment where competition for nutrients is intense, our data support a simplified metabolism for S. alaskensis in which the fate of certain substrates is constrained, especially at the intersections of central carbon and nitrogen metabolism, in order to ensure optimal disposition of scarce resources. This is the first investigation of central metabolism for an oligotrophic ultramicrobacterium that possesses a relatively large genome size. In contrast to the behavior so far observed for SAR11 oligotrophic bacteria, S. alaskensis shows a physiological capacity to exploit increases in ambient nutrient availability and thereby achieve high-population densities.
Similar content being viewed by others
Introduction
The Earth's oceans cover the majority of the surface of the planet, and have the highest cellular production rate of any ecosystem on the planet (Whitman et al., 1998). In the open ocean, the vast majority of this production is carried out by bacteria, which, despite the bulk nutrient-depleted (oligotrophic) nature of the open ocean, can achieve densities in the order of 0.5–5 × 105 cells ml−1 (Schut et al., 1997a; Whitman et al., 1998). The majority of these bacteria are free-living (planktonic) forms that include the smallest of all living cells, with constant cell volumes of not more than 0.1 μm3. These bacteria have been termed ‘ultramicrobacteria’, with cell volume (<0.1 μm3) as the defining criterion (Schut et al., 1997a; Cavicchioli and Ostrowski, 2003). This is particularly useful for studies of natural communities as a variety of cell shapes is often encountered, and volume provides a measurement of cell size that is independent of morphology (Schut et al., 1997a; Cavicchioli and Ostrowski, 2003). Ultramicrobacteria represent a major source of biomass and metabolic activity in oceanic ecosystems, and express higher metabolic activity per unit of volume of seawater than larger bacterial cells (Schut et al., 1997a). As such, in marine oligotrophic environments, these small microbial cells play an essential role in regulating the accumulation, remineralization and transformation of the earth's largest pool of organic carbon. However, the physiology of ultramicrobacteria has remained largely uncharacterized, in a large part because of their resistance to cultivation.
The successful isolation and axenic cultivation of the ultramicrobacterium Sphingopyxis alaskensis strain RB2256 (formerly Sphingomonas alaskensis) has provided opportunities for studying oligotrophic growth and metabolism (Schut et al., 1993, 1995, 1997a, 1997b; Eguchi et al., 1996; Fegatella et al., 1998; Fegatella and Cavicchioli, 2000; Ostrowski et al., 2001; Cavicchioli et al., 2003). S. alaskensis cells are exceptionally small (cell volume <0.1 μm3), and remain essentially constant in size between starvation and growth conditions (Schut et al., 1993, 1997a, 1997b; Cavicchioli et al., 2003). The reduced cell size of ultramicrobacteria provides high surface-to-volume ratios, which facilitates growth under oligotrophic conditions (Button, 1991), resistance to grazing by predatory zooplankton (González et al., 1990), and the ability to partition biomass among a greater number of progeny from a given substrate pool (Fegatella et al., 1998). S. alaskensis strain RB2256 was isolated from surface waters (10 m depth) of Resurrection Bay, Alaska, where it was isolated by an extinction dilution method as a numerically abundant bacterium (>105 cells ml−1) (Schut et al., 1993). S. alaskensis strain AF01 was also an abundant bacterial species sampled from another North Pacific site, in oligotrophic waters (350 m deep) off the coast of Japan (Eguchi et al., 2001), and similar isolates have been obtained from the North Sea (Schut et al., 1993). However, in contrast to ‘Candidatus Pelagibacter ubique’ from the SAR11 clade (Giovannoni et al., 2005), S. alaskensis does not seem to have been abundant in the samples taken for the Global Ocean Survey (which, however, does not include samples from the North Pacific) (Thomas et al., 2007). Although it is presently unclear how widespread this species is in ocean waters, its isolation over a period spanning about a decade and abundance at the time of sampling in North Pacific locations indicates that S. alaskensis has the capacity to proliferate in oligotrophic marine waters.
The ability to grow slowly on low concentrations (nanomolar) of substrates and maintain a relatively constant cell size in the shift between starvation and growth conditions is an oligotrophic trait (Eguchi et al., 1996; Schut et al., 1997a, 1997b; Fegatella and Cavicchioli, 2000; Cavicchioli et al., 2003). On the basis of the Michaelis–Menten constants for substrate transport (Kt), and the available concentrations of dissolved free amino acids (DFAAs) in the ocean, S. alaskensis is predicted to be able to grow by using DFAAs at an in situ doubling time of 12 h to 3 days (Schut et al., 1995), which compares favorably with measured doubling times for bacteria in oligotrophic waters of 5–15 days (Fuhrman et al., 1989). These traits distinguish S. alaskensis from typical copiotrophic bacteria, which exhibit a ‘feast-and-famine’ response that entails rapid growth under nutrient-enriched conditions, but respond to nutrient depletion (including starvation) by undergoing reductive cell division to form resting-stage cells (Srinivasan and Kjelleberg, 1998; Cavicchioli et al., 2003). Earlier studies on the physiology of S. alaskensis have shown a number of properties relevant to its oligotrophic ecology, including the ability to simultaneously take up mixed substrates, irrespective of concentration; a constitutive broad-specificity uptake system for amino acids; and inducible glucose uptake, with this substrate immediately converted to storage product, even during glucose-limiting growth (Schut et al., 1995, 1997a). It has also been shown earlier that the slow growth rate of S. alaskensis (<0.2 h−1) (for example, compared with a copiotroph), even under favorable growth conditions (30 °C, millimolar nutrient levels), is not the result of insufficient ribosome synthesis, despite possessing only a single rRNA copy per genome (Fegatella et al., 1998). Such experiments emphasize the physiological versatility of an ultramicrobacterium that is not confined to oligotrophic nutrient levels in order to grow, but also highlight gaps in our understanding of the critical aspects of S. alaskensis metabolism that may constrain growth, even under nutrient-sufficient conditions.
Earlier kinetic and metabolic data for S. alaskensis are consistent with a metabolism by which certain amino acids (such as alanine) represent important natural growth substrates, but glucose does not (Schut et al., 1995, 1997a). Although they are relatively poor sources of carbon and energy, amino acids are ubiquitous in the open ocean, with alanine, glutamate, glycine and serine being major components of the DFAA reservoir (Lee and Bada, 1975, 1977; Andersson et al., 1985; Ishida et al., 1986; Eguchi and Ishida, 1990). Around 90% of the nitrogen requirements of bacteria in oligotrophic ocean waters are believed to be served by free ammonia and DFAAs (Keil and Kirchman, 1991). However, key aspects of the S. alaskensis metabolism remain unknown, including the processes by which ammonia and amino acids are metabolized by the cell. The aim of this study was to link the analysis of the genome sequence of S. alaskensis RB2256 to proteomic, growth and biochemical analyses in order to elucidate the carbon and nitrogen metabolism of this marine bacterium. One outcome of this approach was the identification of characteristics that helped to clarify how S. alaskensis could compete in oceanic environments, and how this bacterium compares with model oligotrophs such as ‘Cand. P. ubique’ and specialist copiotrophs. This study is the first of this kind for a sphingomonad, a group known for its physiological and ecological versatility (White et al., 1996; Cavicchioli et al., 1999). Given the importance of ultramicrobacteria to carbon and nitrogen cycling in the open ocean, our study aims to provide important new insights into the evolution of metabolic strategies that have been selected in order for marine bacteria to proliferate in oceanic environments.
Materials and methods
Genome analysis
The complete and auto-annotated genome of S. alaskensis RB2256 was searched for genes of potential relevance to central carbon and nitrogen metabolism. Coding regions, automated annotation and manual curation of the S. alaskensis RB2256 genome were carried out as described for other JGI genomes (for example, Klotz et al., 2006; Ivanova et al., 2007). Functional assignments for genes were manually evaluated against experimental data from the literature and the confidence of each gene's predicted function was assigned an evidence rating (ER) value based on a system of manual annotation developed for Methanococcoides burtonii (Allen et al., 2009): ER1, S. alaskensis protein had been experimentally characterized; ER2, the most closely related functionally characterized ortholog share ⩾35% sequence identity along the entire length of the protein; ER3, the most closely related functionally characterized homolog shares <35% sequence identity along the length of the protein, but all required motifs/domains for function are present; ER4, an experimentally characterized full-length homolog is not available but conserved protein motifs or domains can be identified; ER5 (hypothetical protein), no functionally characterized homolog can be found, and no characterized protein domains above the Pfam and InterProScan cut-off thresholds can be identified. The genome sequences of ‘Cand. P. ubique’ HTCC1062 (Rappé et al., 2002; Giovannoni et al., 2005), Silicibacter pomeroyi DSS-3 (González et al., 1999; Moran et al., 2004), Photobacterium angustum S14 (Humphrey et al., 1983; Kjelleberg et al., 1993) and Pseudoalteromonas haloplanktis TAC125 (Médigue et al., 2005; Stocker et al., 2008) were used for comparative analysis as representatives of marine bacteria with diverse strategies for nutrient acquisition. ‘Cand. P. ubique’ HTCC1062 is a non-motile oligotroph isolated from the coast of Oregon (Rappé et al., 2002). It is the first cultured member of the ubiquitous SAR11 clade and has an exceptionally small genome (1.31 Mb) for a free-living bacterium, and represents an example of extensive genomic streamlining in a marine bacterium (Giovannoni et al., 2005). S. pomeroyi DSS-3 (Roseobacter group, Alphaproteobacteria) was isolated off the coast of Georgia (González et al., 1999) and is thought to have a trophic strategy involving association (including direct attachment) with algal blooms (Moran et al., 2004). Its genome has an exceptionally high number of ABC transporter genes, but no identifiable genes associated with chemotaxis (Moran et al., 2004). P. haloplanktis TAC125 (Gammaproteobacteria) was isolated in Antarctic coastal waters (Médigue et al., 2005), and is a marine bacterium that specializes in exploiting ephemeral nutrient pulses and plumes through chemotaxis (Jackson, 1989; Stocker et al., 2008). Photobacterium (formerly Vibrio) angustum S14 (Gammaproteobacteria) was isolated from coastal waters in southeastern Australia, and is a model copiotrophic bacterium that undergoes reductive cell division and mounts a strong starvation-induced stress protection response (Humphrey et al., 1983; Kjelleberg et al., 1993).
Growth conditions and preparation of cell extract
S. alaskensis RB2256 was grown in artificial seawater (ASW) medium (Eguchi et al., 1996) at 30 °C in 250 ml side-armed conical flasks on a rotary shaker at 150 r.p.m. Growth was monitored at 433 nm on various carbon and nitrogen compounds (stipulated in Results). The concentrations of substrates used in the study were at levels higher than typically found in seawater; this was for two reasons. First, conditions that promote strong S. alaskensis growth were used to gauge the activities of enzymes that were potentially important to carbon and nitrogen metabolism. This was important in order to ascertain which metabolic pathways associated with ammonia assimilation and amino acid catabolism were represented in S. alaskensis, especially considering that genes for certain metabolic enzymes seem to be absent from the S. alaskensis genome. Second, the ability of S. alaskensis to use substrates for growth is dependent upon appropriate transport mechanisms; using millimolar levels ensured that intracellular fluxes of these substrates were sufficient, even if transport mechanisms were inefficient in certain cases.
To prepare crude cell extract for enzyme assays, S. alaskensis was grown in ASW medium with the addition of various carbon and nitrogen sources (stipulated in Results). Cultures were grown at 30 °C and harvested at mid-logarithmic phase and centrifuged at 10 000 g for 20 min at 4 °C. The cell pellet was washed once in 50 mM Tris-HCl buffer (pH 7.6) containing 1 mM ethylenediaminetetraacetic acid (EDTA) and stored at −20 °C until required. The cell pellet was resuspended in 50 mM Tris-HCl buffer (pH 7.6) containing 1 mM EDTA and 1 mM phenylmethanesulfonylfluoride (PMSF). The cells were disrupted on ice by sonication for 3 min or by a passage through a French pressure cell (prechilled, at 18 000 lb in–2) and centrifuged at 21 000 × g for 30 min at 4 °C. The resulting supernatant solution was used to assay enzyme activity.
Enzyme assays
Activities of glutamate dehydrogenase (GDH; EC 1.4.1.2 (NAD)/EC 1.4.1.4 (NADP)), alanine dehydrogenase (AlaDH; EC 1.4.1.1), and glutamate synthase (GOGAT) (EC 1.4.1.13) were measured spectrophotometrically from the rate of NAD(P)H oxidation or NAD(P)+ reduction at 340 nm at room temperature (ɛ340=6.22 mM−1 cm−1). Values were corrected for endogenous rates measured in the absence of substrates. The low NADH oxidase activity present in the crude extract was determined with appropriate reagent blanks. One unit (U) of enzyme activity is defined as the amount of enzyme that catalyzes the oxidation of 1 μmol of NADPH or NADH or reduction of 1 μmol of NAD(P)+ per min. The amount of protein in the crude extract was estimated at 595 nm using the method of Bradford (1976) with bovine serum albumin as the standard. All results are averages of values derived from at least two independent experiments.
NAD(P)H-GDH (aminating) reaction mixture contained 100 mM Tris-HCl buffer, pH 8.0 (pH 7.5 for NADH-GDH assay), 50 mM NH4Cl, 15 mM 2-oxoglutarate, 0.2 mM NADH or NADPH, and crude extract. The NAD(P)+-GDH (deaminating) activities were measured in the reaction mixture containing 100 mM Tris-HCl buffer, pH 9.0, 100 mM glutamate, 1 mM NAD(P)+ and crude extract. NADH-AlaDH (aminating) assay mixture contained 100 mM Tris-HCl buffer, pH 8.8, 50 mM NH4Cl, 20 mM pyruvate, 0.2 mM NADH and crude extract. The NAD+-AlaDH (deaminating) assay mixture contained 100 mM 3-(cyclohexylamino)-1-propanesulfonic acid (CAPS) pH 10.0, 40 mM alanine, 1 mM NADH and crude extract. Binding order of substrates in AlaDH activity was determined by adding reagents sequentially to the reaction mixture containing reaction buffer and crude extract. NADPH-GOGAT reaction mixture contained 100 mM Tris-HCl buffer, pH 7.6, or 100 mM K-buffer, pH 7.6, and 10 mM glutamine, 2 mM 2-oxoglutarate (neutralized with KOH), 0.2 mM NADPH and crude extract.
Glutamine synthetase (GS; EC 6.3.1.2) activity in S. alaskensis crude extracts was measured by its transferase and biosynthetic hydroxamate activities, based on Ertan (1992a). One unit of GS activity is defined as the amount of enzyme that catalyzes the formation of 1 μmol γ-glutamyl hydroxamate per min. In this study, GS levels were measured by transferase assay because the state of adenylation has little effect on the γ-glutamyl transferase activity (Shapiro and Stadtman, 1970). The effect of the glutamate analog L-methionine-S-sulfoximine (MSO) (2 mM), an irreversible inhibitor of GS (Ronzio et al., 1969), was also tested for its effect on S. alaskensis growth (Ertan, 1992b).
Glutaminase (EC 3.5.1.2) catalyzes the reversible hydrolysis of glutamine to glutamate, yielding ammonium. Glutaminase activity was measured in both forward and reverse directions, by coupled reaction of bovine liver GDH (Sigma, St Louis, MO, USA, G-2501) and γ-glutamyl hydroxamate assays, respectively (Prusiner et al., 1972, 1976).
Alanine aminotransferase (AlaAT; EC 2.6.1.2) activity was assayed spectrophotometrically by monitoring NADH oxidation at 340 nm for 3 min. Activity in the pyruvate-to-alanine direction was determined by coupling the reaction to NADH oxidation by GDH (Bergmeyer, 1983). The reaction was assayed in a final 2.4-ml mixture containing 5 U of GDH (Sigma G2501), 100 mM Tris-HCl pH 8.0, 50 mM NH4Cl, 0.18 mm NADH, 10 mM pyruvate, 0.025 mM pyridoxal phosphate, 15 mM glutamate and crude extract. One unit of AlaAT was defined as the amount catalyzing the formation of 1.0 μmol of product per min at 30 °C.
α,α-Trehalose phosphorylase (EC 2.4.1.64) activity was measured in the direction of trehalose phosphorolysis as described earlier (Aisaka and Masuda, 1995), with the exception that the trehalose concentration was 200 mM. One unit of enzyme activity was defined as the amount of the enzyme that liberates 1 μmol of glucose per min.
Proteomics
In separate proteomic studies focusing on the response of S. alaskensis to growth at different temperatures (L Ting et al., not shown), 2135 proteins (66% proteome coverage) were identified from cells harvested at late-logarithmic phase (OD433 of 0.3) grown in ASW medium containing 3 mM glucose and 9.4 mM ammonia. Briefly, cell lysates were separated using gel-based fractionation, followed by in-gel tryptic digestion, and the resulting peptides were analyzed using nanoLC and data-dependent tandem mass spectrometry (MS/MS) against a S. alaskensis genome database. In this study, the proteomics analysis was limited to proteins that were potentially relevant to carbon and nitrogen metabolism in S. alaskensis.
Results
Nutrient uptake
The S. alaskensis genome contains seven putative ATP-binding cassette (ABC) transport systems for nutrient uptake: two for sugar transport, and one each for general amino acids (Aap-type), polyamines (putrescine/spermidine), phosphate, Fe3+-siderophore and molybdate. Despite high proteomics coverage, only a single complete ABC transporter system (phosphate uptake) was detected during growth of S. alaskensis on glucose and ammonia (Table 1), and no uptake system responsible for glucose uptake could be identified. The high concentrations (millimolar) of glucose and ammonia used in the growth media may have suppressed expression of energetically expensive ABC transport systems responsible for importing other substrates (such as amino acids). In addition to primary (ABC-mediated) uptake, glucose has been shown to be imported in S. alaskensis by a second mechanism that is independent of group translocation (Schut et al., 1995). Both TRAP (tripartite ATP-independent periplasmic) transporter genes and those for PTS (phosphotransferase system) transport are absent from the S. alaskensis genome. Symporter and permease genes for the uptake of sugars, carboxylic acids, amino acids and oligopeptides are present in the genome, although substrate specificity for these systems was often difficult to infer (Table 1). Such transporters are presumed to include the mechanisms for the uptake of acetate, glutamate and other growth substrates not imported by primary transport.
S. alaskensis grows poorly or does not grow at all with particular carbon or nitrogen compounds (Table 2, and described below). This may be because of inefficient or unavailable transport mechanisms, such as for fructose (consistent with lack of PTS transport), pyruvate, tricarboxylic acid (TCA) intermediates and glutamate. However, in other cases, this could also reflect an absence of appropriate enzymes (for example, gluconokinase/gluconate).
Central carbon metabolism
S. alaskensis is an obligate heterotroph, which is consistent with the absence of any genes associated with known autotrophic pathways (for example, Calvin–Benson–Bassham cycle; reductive TCA cycle). S. alaskensis has a complete set of genes for a functional TCA cycle (Figure 1). The ability to grow on acetate as the sole carbon source, combined with predicted genes for isocitrate lyase and malate synthase, points to the operation of the glyoxylate bypass in S. alaskensis (Tables 1 and 2). S. alaskensis has a pathway for polyhydroxyalkanoic acid (PHA) synthesis and degradation, and several PHA pathway enzymes were identified in the proteomic data (Table 1). PHA synthesis in this species (26% cellular dry weight; Godoy et al., 2003) is indicative of S. alaskensis diverting surplus carbon and energy to storage material. Further, operation of the glyoxylate bypass in S. alaskensis would allow acetyl-coenzyme A (acetyl-CoA) derived from PHA degradation to be fed into the TCA cycle without subsequent loss of carbon.
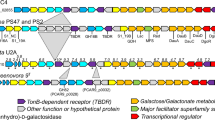
Simplified pathway of central carbon and nitrogen metabolism in Sphingopyxis alaskensis RB2256, based on genomic, proteomic and biochemical data. Pathways that exist in S. alaskensis are shown as solid arrows; pathways that are absent are shown with stippled arrows. Enzymes involved directly in ammonia assimilation are boxed: GS (glutamine synthetase), GOGAT (glutamate synthase), AlaDH (alanine dehydrogenase) and GDH (glutamate dehydrogenase). The major ammonia assimilatory enzymes in S. alaskensis are shaded in gray. Also shown is a unique putative bifunctional enzyme involved in trehalose cleavage, which is inferred to have both trehalose phosphorylase and β-phosphoglucomutase activity.
Genome analysis indicates that, as in other sphingomonads, S. alaskensis uses the Entner–Doudoroff pathway for glucose catabolism (Vartak et al., 1995; Seo et al., 2004; Table 1). The only missing gene is 6-phosphogluconolactonase; neither the pgl or ybhE forms (Zimenkov et al., 2005) are present, indicating that hydrolysis of the ester linkage may occur spontaneously (non-enzymatically) (Kupor and Fraenkel, 1972). For the gluconeogenic direction, it is to be noted that the conversion of fructose-1,6-bisphosphate to fructose-6-phosphate seems to be catalyzed by a GlpX-type (type II) fructose-1,6-bisphosphatase (FBPase II) (Donahue et al., 2000). The auto-annotated genome sequence has two candidates for 6-phosphogluconate dehydrogenase. The protein Sala_1843, but not Sala_0114, was identified by proteomics to be expressed in S. alaskensis during growth on glucose (Table 1). This suggests that Sala_1843 is a 6-phosphogluconate dehydrogenase involved in the pentose phosphate cycle. Proteomic data also suggest that operation of the Entner–Doudoroff pathway is accompanied by expression of ‘gluconeogenic’ enzymes (Table 1). Such enzymes may be involved in the generation of fructose-6-phosphate through isomerization of glucose-6-phosphate, or through the two-step conversion from glyceraldehyde-3-phosphate. The latter could also be converted to dehydroxyacetone phosphate for synthesis of glycerol for lipid backbones.
Genes for pyruvate kinase and pyruvate dehydrogenase complex are present to facilitate the entry of acetyl-CoA into the TCA cycle. Anaplerotic replenishment of oxaloacetate from phosphoenolpyruvate (PEP) is presumably carried out by PEP carboxylase. PEP can be supplied from the TCA cycle intermediates malate and oxaloacetate using either PEP carboxykinase, or a combination of malic enzyme and pyruvate phosphate dikinase.
Trehalose utilization
Trehalose can be used as a carbon source as readily as glucose (Tables 2 and 3), but the only identifiable gene in the S. alaskensis genome for the cleavage of trehalose encodes a fusion protein (Sala_0304) with a trehalose phosphorylase or trehalase amino domain and a β-phosphoglucomutase carboxy domain. Trehalose phosphorylase activity was 125 mU mg−1 for S. alaskensis cells grown in 2 mM trehalose+ 9 mM NH4Cl, compared with 11 and 8 mU mg−1 for cells grown in 3 mM glucose+5 mM glutamate and 3 mM glucose+10 mM glutamine, respectively. The enzyme assays show that minimal glucose was liberated in the absence of phosphate, indicating that S. alaskensis does not possess trehalase activity. This putative bifunctional enzyme suggests that two steps are catalyzed by a single protein: trehalose cleavage into glucose and glucose 1-phosphate, and subsequent glucose 1-phosphate conversion into glucose 6-phosphate. Higher trehalose phosphorylase activity occured in trehalose-grown cells, with low but reproducibly detectable levels present in the absence of trehalose indicating that this enzyme is constitutively expressed; the latter is supported by identification of trehalose phosphorylase from proteomics data of glucose-grown cells (Table 1).
Ammonia and nitrate assimilation
S. alaskensis can use both NH4Cl and KNO3 as the sole source of nitrogen in defined medium in combination with different carbon sources (Table 2). The ability to assimilate nitrate is concordant with the presence of a gene cluster that encodes proteins responsible for nitrate transport and nitrate reduction to ammonia. These proteins were not detected during growth on media containing ammonia (9.4 mM), suggesting that nitrate utilization machinery is not expressed when ammonia (rather than nitrate) is the nitrogen source (Table 1). A predicted ammonia transporter gene (amtB) is present in the genome, and the protein was detected during growth on ammonia (Table 1).
The S. alaskensis genome sequence has a complete set of Ntr regulatory genes, most of which were detected by proteomics analysis (Table 1). Enzyme activity data indicate that the GS in S. alaskensis is not repressed by high concentrations of exogenous ammonia (Table 3). Nitrogen metabolism is governed by a phosphorylation cascade involving PtsP that phosphorylates PtsO and the regulator PtsN, and controls expression of all σ54-dependent genes (Reizer et al., 1992; Commichau et al., 2006). The detection of PtsNOP and Ntr proteins indicates that nitrogen metabolism is tightly regulated in S. alaskensis.
S. alaskensis has a number of genes for enzymes implicated in ammonia assimilation, including GS (type 1), GOGAT (α and β subunits), GDH (NADP-dependent) and AlaDH. The auto-annotated S. alaskensis genome originally included three genes annotated as GS, but we re-annotated two (Sala_1118, Sala_1121) as putative γ-glutamylputrescine synthetase genes, involved in putrescine degradation (Table 1). There are two predicted GDH genes in S. alaskensis: NADP-dependent GDH (Sala_2771) and NAD-dependent GDH (Sala_2258). No glutaminase activity was detected in S. alaskensis, which is consistent with the absence of a glutaminase gene.
Attempts to measure GOGAT in S. alaskensis crude extract were unsuccessful; activities were below detectable levels, despite testing different disruption techniques and buffers. We attribute the lack of activity to the instability of the enzyme under experimental conditions, as encountered by other studies (for example, Brown and Dilworth, 1975; Boland and Benny, 1977; Chen et al., 1987). The absence of glutaminase activity rules out the possibility of a competing two-step reaction occurring in place of GOGAT reaction that would involve the liberation of ammonia from glutamine through glutaminase and subsequent assimilation of ammonia through GDH to produce glutamate.
GS activity was detected across all growth conditions, and showed a trend of increasing activity with increasing exogenous ammonia concentration (Table 3). The transferase assay for GS biosynthetic activity was successful, but the hydroxamate assay did not yield activity. The transferase assay facilitates an examination of the effect of different carbon and nitrogen sources on repression/derepression of GS levels, and provides a good assessment of GS levels under all growth conditions (the biosynthetic hydroxamate assay only measures unadenylated GS). pH optimum of GS transferase activity was 6.8 in imidazole-HCl buffer. GS activity was low when alanine was used as the sole carbon and nitrogen source, but high when alanine was combined with glucose in the medium. Thus, the presence of alanine in the growth medium elicited a strong response from both GS (when alanine was combined with glucose) and AlaDH (when used as the sole source of carbon and nitrogen). The glutamate analog MSO inhibited S. alaskensis growth (determined by maximum OD433) by approximately 50% compared with untreated cultures. However, adding glutamine to the MSO-containing medium restored growth. The data indicate that, although GS-GOGAT is the major pathway for ammonia assimilation in S. alaskensis, ammonia assimilation can proceed when GS is inhibited.
Levels of AlaDH activity generally rose with increasing ammonia concentration (Table 3), and the data strongly support a predominantly assimilatory role for AlaDH in S. alaskensis. Proteomics data show that AlaDH is one of the major proteins in S. alaskensis grown on alanine-free media containing glucose and ammonia (Table 1). The enzyme activity data show that S. alaskensis AlaDH is NAD-dependent in both oxidative and reductive directions (there is no NADP-dependent activity). The Km values of S. alaskensis AlaDH for L-alanine and ammonia were 5.5 and 4.9 mM, respectively. The Km value for ammonia is among the lowest recorded for bacterial AlaDH for which a role in ammonia assimilation has been shown (cf. rhizobial bacteroids: 5–9 mM (Smith and Emerich, 1993; Allaway et al., 2000); Anabaena cylindrica: <8 mM (Rowell and Stewart, 1975); Rhodobacter capsulatus: 16 mM (Caballero et al., 1989a); Streptomyces clavuligerus: 20 mM (Aharonowitz and Friedrich, 1980)). In the reductive amination direction, substrates were bound to S. alaskensis AlaDH in the order NH4+, NADH, pyruvate. In the reverse direction, there was no order of substrate binding. Thus, ammonia can bind the active site without the requirement for binding of other substrates, in contrast to other AlaDH enzymes, in which NH4+ binding is dependent upon the binding of NADH or NADH/pyruvate complex (for example, Grimshaw and Cleland, 1981; Smith and Emerich, 1993). For S. alaskensis, reductive aminating activity was significantly higher than oxidative deamination under most growth conditions (Table 3), and the optimum pH was closer to physiological pH: 8.8–9.0 for amination, compared with 10.0 for deamination. In all, 20 mM alanine inhibited NADH-AlaDH activity by around 50%, and 50 mM NH4Cl inhibited NAD+-AlaDH activity by around 90%. These data indicate that product inhibition by reaction products is one of the regulatory mechanisms for S. alaskensis AlaDH activity, in both directions.
AlaDH activities were influenced by the nature and concentration of the carbon and nitrogen sources. Activities rose with increasing ammonia concentration (consistent with an assimilatory role), and increasing glucose concentrations (likely because of increased availability of pyruvate) up to 12 mM, when AlaDH activity was low. The low activity at high concentrations is consistent with earlier findings that showed that high glucose represses alanine oxidation (Schut et al., 1995). The low AlaDH activities in response to nitrate and glutamate indicate that, in contrast to GS and NADP-GDH, AlaDH seems to have a limited role in assimilating endogenously generated ammonia, such as resulting from nitrate reduction or glutamate catabolism. Higher AlaDH activity in response to glutamine is not unexpected, given that glutamine typically represses GS activity.
Both NADP- and NAD-dependent GDH activities were detected, although those for NAD-GDH were low (Table 3). The optimum pH values for reductive and oxidative activities of NADP-GDH were 8.0 and 8.5, respectively. For NAD-GDH, the reductive amination of 2-oxoglutarate had a pH optimum of 9, and the oxidative deamination of glutamate had a broad optimum spanning pH 8.5–10.0. The Km value of NADP-GDH for glutamate was 7.7 mM. GDH clearly has a limited role in exogenous ammonia assimilation in S. alaskensis. NADP-GDH was not responsive to increasing ammonia concentration in growth medium, which would be otherwise expected for an ammonia assimilatory enzyme. The better growth of S. alaskensis on glutamine compared with ammonia as the sole nitrogen source is also consistent with low assimilatory GDH activity (Rossi et al., 1989; Patriarca et al., 2002). However, GDH seems to play a role in the incorporation of endogenously generated ammonia. The highest NADP-GDH values were found with alanine as the sole carbon and nitrogen source, or when nitrate was the nitrogen source (Table 3). This is interpreted as a response to the high levels of intracellular ammonia liberated from alanine by AlaDH, or by the reduction of nitrate. Nevertheless, GDH catabolic function is presumably sufficient for S. alaskensis to grow on glutamate and glutamine, with the latter catabolized by the combined action of GOGAT and GDH.
In summary, GS and AlaDH were found to be the principal assimilatory enzymes under all conditions tested in our enzyme assays. Proteomic data showed GS, GOGAT and AlaDH to be among the most frequently detected proteins during growth on high-ammonia media. These data underscore the important role for AlaDH in ammonia assimilation in S. alaskensis and, conversely, the relatively minor role of GDH. GS is a major ammonia assimilatory enzyme, but there is no evidence that GS activity is repressed by high ammonia levels in the medium, even up to 18 mM. Nevertheless, some regulation of nitrogen metabolism is apparent, given that GS and AlaDH show contrasting responses to individual carbon and nitrogen sources. The combination of a rich carbon source (glucose) and a poor nitrogen source (nitrate, glutamate, alanine) resulted in high GS activity, presumably a result of derepression of GS. This response contrasts with AlaDH activity, which showed an opposite response to high glucose and/or poor nitrogen sources (nitrate, glutamate and glutamine, but not alanine). The high GS activities in response to nitrate and glutamate presumably result from low intracellular ammonia concentrations, which limits glutamine production, and thereby derepresses GS.
Amino acid metabolism
Growth data show that S. alaskensis is evidently capable of utilizing alanine, glutamate and glutamine as the sole source of carbon and nitrogen (Table 2). In addition, alanine, glutamate, glutamine, glycine and serine can each be used as a nitrogen source when glucose is present in the medium. A gene for D-amino acid dehydrogenase (DadA) could not be identified in the genome, indicating that AlaDH is responsible for alanine catabolism, as well as being an assimilatory enzyme (see the section ‘Ammonia and nitrate assimilation’). AlaDH has been observed to substitute for the Dad system when overexpressed in a rhizobial strain that lacks the Dad system, which is normally the principal mechanism for alanine catabolism (Allaway et al., 2000; Lodwig et al., 2004). A catabolic role for AlaDH in S. alaskensis is consistent with the observation that elevated AlaDH values were attained for cells grown on alanine. The presence of nitrate, glutamate and glutamine in the media resulted in low AlaDH activities, as did high glucose (12 mM).
The NAD-GDH belongs to the ‘large GDH’ subfamily, which has been ascribed a role in glutamate catabolism (Miñambres et al., 2000). Owing to the apparent absence of a glutamate decarboxylase gene in the S. alaskensis genome glutamate cannot be catabolized by the γ-aminobutyrate (GABA) shunt. The putative NAD-GDH was among the most frequently detected proteins during growth on high-ammonia media (Table 1), with endogenous glutamate generated from GS-GOGAT able to serve as a substrate for catabolic GDH. Glutamate supplied in the media could be used as a sole carbon and nitrogen source for S. alaskensis growth when present at 10 mM, but not at 5 mM (Table 2). Glutamate could also serve as a nitrogen source at 5 mM when accompanied by glucose or trehalose as a carbon source (Table 2). Earlier, it was found from a mixture of 10 amino acids (1 mM each; including alanine, glutamate, glutamine and serine) that glutamate was the only amino acid not to be utilized by S. alaskensis (Schut et al., 1995). These data indicate that the utilization of glutamate as a growth substrate is dependent upon glutamate concentration and/or the identity of the (other) carbon source. Given that the single amino acid ABC transport system in S. alaskensis is not receptive to glutamate, this amino acid must be imported by other mechanisms that require high solute concentrations, or an energy-rich carbon source.
GDH activity in S. alaskensis was not induced by the presence of glutamate in the media (even in the presence of NAD+ or NADP+). For S. alaskensis, weak catabolic GDH that is not induced by glutamate may correlate with the absence of an ABC transport system capable of importing glutamate (see the section ‘Nutrient uptake’). Overall, this response to glutamate is in marked contrast to the response of S. alakensis to alanine, which is imported by active ABC transport, and results in elevated AlaDH levels. Nevertheless, GDH catabolic function is presumably sufficient for S. alaskensis to grow on glutamate.
No AlaAT activity was observed for S. alaskensis, which is consistent with the absence of a gene responsible for this transamination enzyme. Several aminotransferase genes are present, but only two (apartate aminotransferase, branched-chain-amino acid aminotransferase) seem to encode enzymes capable of mediating transaminations among the 20 common amino acids. The inability of S. alaskensis to grow on glycine as a carbon source, despite growth on alanine, is consistent with the absence of an alanine-glyoxylate aminotransferase gene. There is no aspartate decarboxylase gene for the synthesis of aspartate through the carboxylation of alanine. Thus, there is no available mechanism for the direct interconversion of alanine and glutamate, which can therefore only occur indirectly though central carbon metabolism in order for either to serve as both a carbon and nitrogen source for growth.
Glycine and serine can each serve as a nitrogen source for the growth of S. alaskensis, but not as a carbon source. Genes for the components of the glycine cleavage enzyme complex are present in the S. alaskensis genome. This complex liberates ammonia as a nitrogen source, but the carbon skeleton is first decarboxylated, then directed to C1 metabolism, precluding its use as a carbon source for growth. No deaminating glycine dehydrogenase gene is present. Genes are present for the interconversion of glycine and serine or threonine, the synthesis of serine from 3-phosphoglycerate and the deamination of serine; all the corresponding proteins were identified by proteomics analysis (Table 1). Thus, the inability to use glycine or serine as a carbon source is interesting in light of the genomic and proteomic data, which indicate that S. alaskensis is metabolically equipped to degrade these amino acids to pyruvate and ammonia. L-serine deaminase, which was expressed in S. alaskensis under conditions (high glucose) for which serine deamination should not be necessary, may therefore have a metabolic role that is not directly related to growth on amino acids (Su et al., 1989).
Putrescine catabolism
Strong growth was observed on putrescine when it was supplied either as the sole carbon and nitrogen source, or as a nitrogen source in combination with glucose (Table 2). The genome has a gene cluster for the uptake (ABC transport) and utilization of putrescine. This pathway catabolizes putrescine by way of glutamylated intermediates, and generates ammonia and succinate, which enters the TCA cycle (Kurihara et al., 2005). Many of the enzymes of this pathway were detected by proteomics from cells growing on glucose (Table 1), suggesting that this pathway may be constitutively expressed in S. alaskensis.
Discussion
Substrate preference and utilization
S. alaskensis is versatile in its ability to utilize carbon and nitrogen compounds as substrates for growth. Ammonia and nitrate can be used as nitrogen sources, despite the high energetic cost associated with nitrate utilization. This is in contrast to a number of other marine bacteria that lack the ability to assimilate nitrate, including ‘Cand. P. ubique’ (for example, Dufresne et al., 2003; García-Fernández et al., 2004; Giovannoni et al., 2005). S. alaskensis was isolated as an abundant bacterium from the oceanic waters of the North Pacific (Button et al., 1993; Schut et al., 1993; Eguchi et al., 2001), which indicates that it contributed to the assimilation of inorganic nitrogen in these waters at the time of sampling. Our data particularly highlight the important role of alanine in this process, as it is a major product of ammonia and nitrate assimilation by S. alaskensis.
Certain amino acids (alanine, glutamate, glutamine) and putrescine can each be used by S. alaskensis as sole carbon and nitrogen sources, and these substrates are ubiquitous at nanomolar concentrations in seawater (Lee and Bada, 1975, 1977; Eguchi and Ishida, 1990; Nishibori et al., 2001, 2003). Extracytoplasmic solute capture is extremely important under oligotrophic conditions (Schut et al., 1995, 1997a; Giovannoni et al., 2005; Sowell et al., 2009). S. alaskensis has only a single amino acid ABC transport system (Table 4) that has earlier been found to be constitutive for alanine, in contrast to the inducible uptake of glucose (Schut et al., 1995). The uptake of alanine was also found to be competitively inhibited by several amino acids (including glycine, although weakly), but not glutamate, glutamine or serine (Schut et al., 1995). Our data indicate that the earlier observation that 1 mM glutamate is not used by S. alaskensis (Schut et al., 1995) is likely a consequence of insufficient transport into the cytoplasm, rather than an absence of glutamate catabolic activity. The available data therefore indicate that alanine is a very important natural growth substrate for S. alaskensis, but that glutamate and serine are not. Furthermore, AlaDH activities were elevated in response to alanine, in contrast to low and non-inducible GDH activities in response to glutamate (Table 3). Thus, alanine is singled out by S. alaskensis as a preferred growth substrate, which is a further refinement of the preference for amino acids earlier documented for alphaproteobacteria (Cottrell and Kirchman, 2003; Malmstrom et al., 2004). This may represent a key aspect of the oligotrophic ecology of S. alaskensis, as an example of simplification at the level of metabolic processing. Pyruvate is an important intermediate positioned at the nexus of several metabolic pathways (TCA cycle, gluconeogenesis, fatty acid synthesis, PHA synthesis), and therefore potentially a more versatile intermediate than 2-oxoglutarate, the deamination product of glutamate. Also, sufficient glutamate may be generated from ammonia assimilation via GS-GOGAT to make uptake of glutamate unnecessary under oligotrophic conditions. Serine, which like alanine has pyruvate as a deamination product, was also not favored as a growth substrate, which might reflect a consolidation of the supply of pyruvate through the uptake and catabolism of alanine, with serine directed toward other fates, such as C1 metabolism or glycine and threonine synthesis.
AlaDH and ammonia assimilation
In addition to being an important catabolic substrate, alanine is also the product of one of two major ammonia assimilation pathways in S. alaskensis. We present several lines of evidence that AlaDH is a major ammonia assimilatory enzyme in S. alaskensis. In most bacteria, the sequential action of GS and GOGAT serves as either the sole pathway for ammonia assimilation, or together with GDH (Kondorosi et al., 1977; Westby et al., 1987; Brown and Herbert, 1977a, 1977b; Reitzer, 2003; Muro-Pastor et al., 2005; Li and Lu, 2007). However, AlaDH plays a role in ammonia assimilation in some bacteria that have low or absent GDH activities. In these cases, GS-GOGAT remains the principal mechanism for ammonia assimilation, and AlaDH is used as an alternative or auxiliary ammonia enzyme under certain conditions, such as excess ammonia, when GS activity is repressed (Aharonowitz and Friedrich, 1980); inactivation of GS by MSO (Herbert et al., 1978; Madigan and Cox, 1982; Caballero et al., 1989a); or high pyruvate levels (Johansson and Gest, 1976; Moreno-Vivián et al., 1983; Caballero et al., 1989a). By contrast, in S. alaskensis, AlaDH activity was constitutive, and AlaDH seems to play a role in ammonia assimilation during growth across a range of ammonia concentrations and in the presence of various carbon sources (Table 3). AlaDH requires a different carbon skeleton to GDH and GS-GOGAT, and (like GDH) does not require ATP. Therefore, employing AlaDH in preference to GDH as an alternative assimilatory enzyme allows ammonia assimilation to proceed in the absence of a ready supply of 2-oxoglutarate. Thus, the selection of GS-GOGAT and AlaDH as the two major mechanisms for ammonia assimilation might be important under conditions of carbon limitation if the intracellular replenishment of 2-oxoglutarate is limited, such as during growth on substrates that are processed by the glyoxylate bypass (for example, PHA, acetate).
The fate of alanine in S. alaskensis seems to differ from that of other bacteria in which AlaDH is used to assimilate ammonia taken up by the cell. Phototrophic bacteria that use AlaDH as an assimilatory enzyme can convert alanine to glutamate, either directly using AlaAT (Johansson and Gest, 1976; Caballero et al., 1989b) or through a series of transamination reactions (Herbert et al., 1978; Cárdenas et al., 1987). However, neither route seems to be available to S. alaskensis. Some of the accumulated alanine in S. alaskensis could be drawn off for biosynthetic purposes (for example, peptidoglycan, CoA, biotin, alanyl-tRNA), but the remainder cannot be directly converted to glutamate for further nitrogen metabolism. In the absence of suitable transaminases (for example, AlaAT) and a dedicated alanine deamination mechanism (Dad system), only AlaDH is available to catabolize intracellular alanine that is imported or generated endogenously. This again may constitute an example of metabolic simplification, by directing alanine away from nitrogen metabolism by way of glutamate or glutamine (Figure 1).
Oligotrophic lifestyle and trophic strategy of S. alaskensis
The relatively slow doubling time for S. alaskensis compared with specialist copiotrophs may be attributable to the loss of certain non-essential metabolic functions (for example, glutamate/alanine transamination; Dad system; 6-phosphogluconolactonase) (Figure 1); it has been shown earlier that the relatively slow growth rate (<0.2 h−1) is not the result of insufficient translational capacity (Fegatella et al., 1998). A buffering of maximum growth rate might also relate to a metabolic emphasis on the synthesis of storage material (PHA, intracellular polysaccharide) to maintain a steady rate of cell division throughout periods of relative nutrient surfeit.
In addition to the ability of S. alaskensis to take up mixed substrates simultaneously (Schut et al., 1995, 1997a), aspects of the carbon and nitrogen metabolism outlined above seem to be specifically geared toward an oligotrophic ecology, such as simplification of amino acid catabolism and alternate routes for ammonia assimilation that require different precursors, with the latter facilitating the efficient scavenging of ammonia under carbon-limited conditions (Figure 1). Overall, our data indicate an adaptive ecological strategy that is distinctive among marine bacteria, and adds to the repertoire of strategies so far ascribed to bacteria that survive and grow in the oceanic environment (for example, Moran et al., 2004; Médigue et al., 2005; Giovannoni et al., 2005; Stocker et al., 2008). The metabolic simplification of S. alaskensis is interpreted as an adaptation to conditions of nutrient depletion, especially carbon limitation, by focusing the uptake machinery on a narrow subset of bioavailable substrates, and ensuring that all aspects of metabolism are furnished by these substrates. This obviates the need for extensive gene regulation governing central carbon and nitrogen metabolism, especially regarding the fate of individual amino acids.
Nevertheless, our genomic analysis of S. alaskensis genome shows an array of regulatory genes, absent from SAR11. For S. alaskensis, this may reflect a capacity to alter metabolic strategies in response to ambient changes in the identity and concentration of both carbon and nitrogen substrates, such as in response to elevated nutrient levels (Table 4). Gene regulation is integral to an opportunistic strategy of efficiently exploiting competing substrates, whereas minimization of regulatory networks is one of the hallmarks of passive oligotrophs with streamlined genomes (Dufresne et al., 2003; Giovannoni et al., 2005). The small genome of ‘Cand. P. ubique’ lacks genes associated with nutrient detection, motility, and most regulatory mechanisms (Table 4; Giovannoni et al., 2005). The regulatory capacity of S. alaskensis is well illustrated by the expression of specific regulatory proteins associated with the Ntr and PtsNOP regulatory networks under nutrient-enriched conditions (Table 1), and the different responses of enzymes involved in ammonia assimilation to individual carbon and nitrogen sources (Table 3). The fact that S. alaskensis is flagellated (and many of the flagella and regulatory proteins associated with motility were detected by proteomics; Table 1) suggests that chemotactic ability provides a selective advantage in bulk low-nutrient waters. However, the possession of only a limited number of methyl-accepting chemotaxis proteins (MCPs) suggests that S. alaskensis has less scope to respond to diverse nutrient signals than copiotrophic bacteria.
The potential to detect, associate with and efficiently exploit elevated levels of substrates through a chemotactic response would distinguish S. alaskensis from passive oligotrophs (Polz et al., 2006; Stocker et al., 2008). The latter include SAR11 bacteria, for which the data indicate an oligotrophic strategy that entails obligate low-nutrient growth, low population densities, and a greater dependence on exogenous substrates to fulfill its metabolic requirements (Joint, 2008; Tripp et al., 2008). The ubiquity of SAR11 bacteria suggests that this bacterial group may be specialized for extreme low-nutrient environments that are exposed to minimal nutrient fluxes (such as the Sargasso Sea) without compromising their ability to survive in large numbers in nutrient-enriched environments (Giovannoni et al., 1990, 2005; Morris et al., 2002). In contrast to SAR11 bacteria, S. alaskensis is metabolically poised to exploit high concentrations of substrates (amino acids, polyamines, carbohydrates). In support of this, it has been shown earlier for S. alaskensis that after nutrient upshift, starved or nutrient-limited chemostat cultures (from a range of carbon sources), resume maximum growth rates without a detectable lag (Fegatella and Cavicchioli, 2000; Cavicchioli et al., 2003). However, S. alaskensis is not specialized for a strategy that entails rapid detection and exploitation of nutrient plumes or pulses, thereby setting it apart from specialist ‘feast-and-famine’ copiotrophs. Our genomic and experimental data for S. alaskensis are consistent with an ecophysiological trade-off between intense genomic streamlining and the ability to efficiently exploit fluxes in the quality and quantity of available nutrients.
References
Aharonowitz Y, Friedrich CG . (1980). Alanine dehydrogenase of the β-lactam antibiotic producer Streptomyces clavuligerus. Arch Microbiol 125: 137–142.
Aisaka K, Masuda T . (1995). Production of trehalose phosphorylase by Catellatospora ferruginea. FEMS Microbiol Lett 131: 47–51.
Allaway D, Lodwig EM, Crompton LA, Wood M, Parsons R, Wheeler TR et al. (2000). Identification of alanine dehydrogenase and its role in mixed secretion of ammonium and alanine by pea bacteroids. Mol Microbiol 36: 508–515.
Allen M, Lauro FM, Williams TJ, Burg D, Siddiqui KS, De Francisci D et al. (2009). The genome sequence of the psychrophilic archaeon, Methanococcoides burtonii: the role of genome evolution in cold-adaptation. ISME J (e-pub ahead of print, 30 April 2009; doi:10.1038/ismej.2009.45).
Andersson A, Lee C, Azam F, Hagström Å . (1985). Release of amino acids and inorganic nutrients by heterotrophic marine microflagellates. Mar Ecol Prog Ser 23: 99–106.
Bergmeyer H . (1983). Methods of Enzymatic Analysis, vol. 2. Verlag Chemie: Veinheim, p 277.
Boland MJ, Benny AG . (1977). Enzymes of nitrogen metabolism in legume nodules: purification and properties of NADH-dependent glutamate synthase from lupin nodules. Eur J Biochem 79: 355–362.
Bradford MM . (1976). A rapid and sensitive method for quantitation of microgram quantities of protein utilizing the principle of protein-dye-binding. Anal Biochem 72: 248–254.
Brown CM, Dilworth MJ . (1975). Ammonia assimilation by rhizobium cultures and bacteroids. J Gen Microbiol 86: 39–48.
Brown CM, Herbert RA . (1977a). Ammonia assimilation in purple and green sulphur bacteria. FEMS Lett 1: 39–42.
Brown CM, Herbert RA . (1977b). Ammonia assimilation in members of the Rhodospirillaceae. FEMS Lett 1: 43–46.
Button DK . (1991). Biochemical basis for whole-cell uptake kinetics: specific affinity, oligotrophic capacity, and the meaning of the Michaelis constant. Appl Environ Microbiol 57: 2033–2038.
Button DK, Schut F, Quang P, Martin R, Robertson BR . (1993). Viability and isolation of marine bacteria by dilution culture: theory, procedures, and initial results. Appl Environ Microbiol 59: 881–891.
Caballero F J, Cárdenas J, Castillo F . (1989a). Purification and properties of L-alanine dehydrogenase of the phototrophic bacterium Rhodobacter capsulatus E1F1. J Bacteriol 171: 3205–3210.
Caballero FJ, Igeño I, Cárdenas J, Castillo F . (1989b). Regulation of reduced nitrogen assimilation in Rhodobacter capsulatus E1F1. Arch Microbiol 152: 508–511.
Cárdenas J, Caballero FJ, Moreno-Vivián C, Castillo F . (1987). Enzymology of ammonia assimilation in purple nonsulfur bacteria. In: Ullrich WR, Aparicio PJ, Syrett PJ, Castillo F (eds). Inorganic Nitrogen Metabolism. Springer-Verlag: Berlin, pp 148–153.
Cavicchioli R, Fegatella F, Ostrowski M, Eguchi M, Gottschal J . (1999). Sphingomonads from marine environments. J Ind Microbiol Biotechnol 23: 268–272.
Cavicchioli R, Ostrowski M . (2003). Ultramicrobacteria. Encyclopedia of Life Sciences. Nature Publishing Group: London.
Cavicchioli R, Ostrowski M, Fegatella F, Goodchild A, Guixa-Boixereu N . (2003). Life under nutrient limitation in oligotrophic marine environments: an eco/physiological perspective of Sphingopyxis alaskensis (formerly Sphingomonas alaskensis). Microb Ecol 45: 203–217.
Chen CH, Van Baalen C, Tabita FR. . (1987). DL-7-azatryptophan and citrulline metabolism in the cyanobacterium Anabaena sp. strain 1F. J Bacteriol 169: 1114–1119.
Commichau FM, Forchhammer K, Stülke J . (2006). Regulatory links between carbon and nitrogen metabolism. Curr Opin Microbiol 9: 167–172.
Cottrell MT, Kirchman DL . (2003). Contribution of major bacterial groups to bacterial biomass production (thymidine and leucine incorporation) in the Delaware estuary. Limnol Oceanogr 48: 168–178.
Donahue JL, Bownas JL, Niehaus WG, Larson TJ . (2000). Purification and characterization of glpX-encoded fructose 1, 6-bisphosphatase, a new enzyme of the glycerol 3-phosphate regulon of Escherichia coli. J Bacteriol 182: 5624–6427.
Dufresne A, Salanoubat M, Partensky F, Artiguenave F, Axmann IM, Barbe V et al. (2003). Genome sequence of the cyanobacterium Prochlorococcus marinus SS120, a nearly minimal oxyphototrophic genome. Proc Natl Acad Sci USA 100: 10020–10025.
Eguchi M, Ishida Y . (1990). Oligotrophic properties of heterotrophic bacteria and in situ heterotrophy activity in pelagic seawaters. FEMS Microbiol Ecol 73: 23–30.
Eguchi M, Nishikawa T, Macdonald K, Cavicchioli R, Gottschal JC, Kjelleberg S . (1996). Responses to stress and nutrient availability by the marine ultramicrobacterium Sphingomonas sp. strain RB2256. Appl Environ Microbiol 62: 1287–1294.
Eguchi M, Ostrowski M, Fegatella F, Bowman J, Nichols D, Nishino T et al. (2001). Sphingomonas alaskensis strain AFO1, an abundant oligotrophic ultramicrobacterium from the North Pacific. Appl Environ Microbiol 67: 4945–4954.
Ertan H . (1992a). Some properties of glutamate dehydrogenase, glutamine synthetase and glutamate synthase from Corynebacterium callunae. Arch Microbiol 158: 35–41.
Ertan H . (1992b). The effect of various culture conditions on the levels of ammonia assimilatory enzymes of Corynebacterium callunae. Arch Microbiol 158: 42–47.
Fegatella F, Cavicchioli R . (2000). Physiological responses to starvation in the marine oligotrophic ultramicrobacterium Sphingomonas sp. strain RB2256. Appl Environ Microbiol 66: 2037–2044.
Fegatella F, Lim J, Kjelleberg S, Cavicchioli R . (1998). Implications of rRNA operon copy number and ribosome content in the marine oligotrophic ultramicrobacterium Sphingomonas sp. strain RB2256. Appl Environ Microbiol 64: 4433–4438.
Fuhrman JA, Sleeter TD, Carlson CA, Proctor LM . (1989). Dominance of bacterial biomass in the Sargasso Sea and its ecological implications. Mar Ecol Prog Ser 57: 207–217.
García-Fernández JM, de Marsac NT, Diez J . (2004). Streamlined regulation and gene loss as adaptive mechanisms in Prochlorococcus for optimized nitrogen utilization in oligotrophic environments. Microbiol Mol Biol Rev 68: 630–638.
Giovannoni SJ, Britschgi TB, Moyer CL, Field KG . (1990). Genetic diversity in Sargasso Sea bacterioplankton. Nature 345: 60–63.
Giovannoni SJ, Tripp HJ, Givan S, Podar M, Vergin KL, Baptista D et al. (2005). Genome streamlining in a cosmopolitan oceanic bacterium. Science 309: 1242–1245.
Godoy F, Vancanneyt M, Martínez M, Steinbüchel A, Swings J, Rehm BHA . (2003). Sphingopyxis chilensis sp. nov.a chlorophenol-degrading bacterium that accumulates polyhydroxyalkanoate, and transfer of Sphingomonas alaskensis to Sphingopyxis alaskensis comb. nov. Int. J Syst Evol Microbiol 53: 473–477.
González JM, Kiene RP, Moran MA . (1999). Transformation of sulfur compounds by an abundant lineage of marine bacteria in the α-subclass of the class Proteobacteria. Appl Environ Microbiol 65: 3810–3819.
González JM, Sherr EB, Sherr BF . (1990). Size-selective on bacteria by natural assemblages of estuarine flagellates and ciliates. Appl Environ Microbiol 56: 583–589.
Grimshaw CE, Cleland WW . (1981). Kinetic mechanism of Bacillus subtilis L-alanine dehydrogenase. Biochemistry 20: 5650–5655.
Herbert RA, Siefert E, Pfennig N . (1978). Nitrogen assimilation in Rhodopseudomonas acidophila. Arch Microbiol 119: 1–5.
Humphrey B, Kjelleberg S, Marshall KC . (1983). Responses of marine bacteria under starvation conditions at a solid-water interface. Appl Environ Microbiol 45: 43–47.
Ishida Y, Eguchi M, Kadota H . (1986). Existence of obligately oligotrophic bacteria as a dominant population in the South China Sea and the West Pacific Ocean. Mar Ecol Prog Ser 30: 197–203.
Ivanova NN, Mavromatis K, Chen I-MA, Markowitz VM, Kyrpides NC . (2007). Standard operating procedure for the annotations of genomes and metagenomes submitted to the integrated microbial genomes expert review (IMG-ER) system. Available at http://imgweb.jgi-psf.org/w/doc/img_er_ann.pdf.
Jackson GA . (1989). Simulation of bacterial attraction and adhesion to falling particles in an aquatic environment. Limnol Oceanogr 34: 514–530.
Joint I . (2008). Unravelling the enigma of SAR11. ISME J 2: 455–456.
Johansson BC, Gest H . (1976). Inorganic nitrogen assimilation by the photosynthetic bacterium Rhodopseudomonas capsulata. J Bacteriol 128: 683–688.
Keil RG, Kirchman DL. (1991). Contribution of dissolved free amino acids and ammonium to the nitrogen requirements of heterotrophic bacterioplankton. Mar Ecol Prog Ser 73: 1–10.
Kjelleberg S, Flardh K, Nystrom T, Moriarty DJW . (1993). Growth limitation and starvation of bacteria. In: Ford T (ed). Aquatic Microbiology: An Ecological Approach. Blackwell Scientific Publications: Boston, pp 289–320.
Klotz MG, Arp DJ, Chain PSG, El-Sheikh AF, Hauser LJ, Hommes NG et al. (2006). Complete genome sequence of the marine, chemolithoautotrophic, ammonia-oxidizing bacterium Nitrosococcus oceani ATCC 19707. Appl Environ Microbiol 72: 6299–6315.
Kondorosi A, Svfib Z, Kiss GB, Dixon RA . (1977). Ammonia assimilation and nitrogen fixation in Rhizobium meliloti. Mol Gen Genet 151: 221–226.
Kupor SR, Fraenkel DG . (1972). Glucose metabolism in 6-phosphogluconolactonase mutants of Escherichia coli. J Biol Chem 247: 1904–1910.
Kurihara S, Oda S, Kato K, Kim HG, Koyanagi T, Kumagai H et al. (2005). A novel putrescine utilization pathway involves γ-glutamylated intermediates of Escherichia coli K-12. J Biol Chem 280: 4602–4608.
Lee C, Bada JF . (1975). Amino acids in equatorial Pacific Ocean water. Earth Planet Sci Lett 26: 61–68.
Lee C, Bada JF . (1977). Dissolved amino acids in the equatorial Pacific, the Sargasso Sea, and Biscayne Bay. Limnol Oceanogr 22: 502–510.
Li W, Lu CD . (2007). Regulation of carbon and nitrogen utilization by CbrAB and NtrBC two-component systems in Pseudomonas aeruginosa. J Bacteriol 189: 5413–5420.
Lodwig E, Kumar S, Allaway D, Bourdes A, Prell J, Priefer U et al. (2004). Regulation of L-alanine dehydrogenase in Rhizobium leguminosarum bv viciae and its role in pea nodules. J Bacteriol 186: 842–849.
Madigan MT, Cox SS . (1982). Nitrogen metabolism in Rhodopseudomonas globiformis. Arch Microbiol 133: 6–10.
Malmstrom RR, Kiene RP, Cottrell MT, Kirchman DL . (2004). Contribution of SAR11 bacteria to dissolved dimethylsulfoniopropionate and amino acid uptake in the North Atlantic Ocean. Appl Environ Microbiol 70: 4129–4135.
Médigue C, Krin E, Pascal G, Barbe V, Bernsel A, Bertin PN et al. (2005). Coping with cold: the genome of the versatile marine Antarctica bacterium Pseudoalteromonas haloplanktis TAC125. Genome Res 15: 1325–1335.
Miñambres B, Olivera ER, Jensen RA, Luengo JM . (2000). Biochemical and genetic characterization of the first member, the AMP-requiring NAD-specific GDH of Streptomyces clavuligerus: a new class of glutamate dehydrogenases (GDH). J Biol Chem 275: 39529–39542.
Moran MA, Buchan A, González JM, Heidelberg JF, Whitman WB, Kiene RP et al. (2004). Genome sequence of Silicibacter pomeroyi reveals adaptations to the marine environment. Nature 432: 910–913.
Moreno-Vivián C, Cejudo FJ, Cárdenas J, Castillo F . (1983). Ammonia assimilation pathways in Rhodopseudomonas capsulata ElFl. Arch Microbiol 136: 147–151.
Morris RM, Rappé MS, Connon SA, Vergin KL, Siebold WA, Carlson CA et al. (2002). SAR11 clade dominates ocean surface bacterioplankton communities. Nature 420: 806–810.
Muro-Pastor MI, Reyes JC, Florencio FJ . (2005). Ammonium assimilation in cyanobacteria. Photosynthesis Res 83: 135–150.
Nishibori N, Matuyama Y, Uchida T, Moriyama T, Ogita Y, Oda M et al. (2003). Spatial and temporal variations in free polyamine distributions in Uranouchi Inlet, Japan. Mar Chem 82: 307–314.
Nishibori N, Yuasa A, Sakai M, Fujihara S, Nishio S . (2001). Free polyamines in coastal seawater during phytoplankton bloom. Fisheries Sci 67: 79–83.
Ostrowski M, Cavicchioli R, Blaauw M, Gottschal JC . (2001). Specific growth rate plays a critical role in hydrogen peroxide resistance of the marine oligotrophic ultramicrobacterium Sphingomonas alaskensis strain RB2256. Appl Environ Microbiol 67: 1292–1299.
Patriarca EJ, Taté R, Iaccarino M . (2002). Key role of bacterial NH4+ metabolism in rhizobium-plant symbiosis. Microbiol Mol Biol Reviews 66: 203–222.
Polz MF, Hunt DE, Preheim SP, Weinreich DM . (2006). Patterns and mechanisms of genetic and phenotypic differentiation in marine microbes. Phil Trans R Soc B 361: 2009–2021.
Prusiner S, Miller RE, Valentine RC . (1972). Adenosine 3′:5′-cyclic monophosphate control of the enzymes of glutamine metabolism in Escherichia coli. Proc Natl Acad Sci USA 69: 2922–2926.
Prusiner S, Davis JN, Stadtman ER . (1976). Regulation of glutaminase B in Escherichia coli. J Biol Chem 251: 3447–3456.
Rappé MS, Connon SA, Vergin KL, Giovannoni SJ . (2002). Cultivation of the ubiquitous SAR11 marine bacterioplankton clade. Nature 418: 630–633.
Reitzer L . (2003). Nitrogen assimilation and global regulation in Escherichia coli. Annu Rev Microbiol 57: 155–176.
Reizer J, Reizer A, Saier Jr MH, Jacobson GR . (1992). A proposed link between nitrogen and carbon metabolism involving protein phosphorylation in bacteria. Protein Sci 1: 722–726.
Ronzio R, Rowe W, Meister A . (1969). Studies on the mechanism of inhibition of glutamine synthetase by methionine sulfoximine. Biochemistry 8: 1066–1075.
Rossi M, Defez R, Chiurazzi M, Lamberti A, Fuggi A, Iaccarino M . (1989). Regulation of glutamine synthetase isoenzymes in Rhizobium leguminosarum biovar viciae. J Gen Microbiol 135: 629–637.
Rowell P, Stewart WD . (1975). Alanine dehydrogenase of the N2-fixing blue-green alga, Anabaena cylindrica. Arch Microbiol 107: 115–124.
Schut F, de Vries EJ, Gottschal JC, Robertson BR, Harder W, Prins RA et al. (1993). Isolation of typical marine bacteria by dilution culture: growth, maintenance, and characteristics of isolates under laboratory conditions. Appl Environ Microbiol 59: 2150–2160.
Schut F, Jansen M, Gomes TM, Gottschal JC, Harder W, Prins RA . (1995). Substrate uptake and utilization by a marine ultramicrobacterium. Microbiology 141: 351–361.
Schut F, Gottschal JC, Prins RA . (1997b). Isolation and characterization of the marine ultramicrobacterium Sphingomonas sp. strain RB2256. FEMS Microbiol Rev 20: 363–369.
Schut F, Prins RA, Gottschal JC . (1997a). Oligotrophy and pelagic marine bacteria: facts and fiction. Aquat Microb Ecol 12: 177–202.
Seo JS, Chong H, Park HS, Yoon KO, Jung C, Kim JJ et al. (2004). The genome sequence of the ethanologenic bacterium Zymomonas mobilis ZM4. Nat Biotechnol 23: 63–68.
Shapiro BM, Stadtman ER . (1970). Glutamine synthetase (E. coli). Methods Enzymol 17A: 910–922.
Smith MT, Emerich DW . (1993). Alanine dehydrogenase from soybean nodule bacteroids—kinetic mechanism and pH studies. J Biol Chem 268: 10746–10753.
Sowell SM, Wilhelm LJ, Norbeck AD, Lipton MS, Nicora CD, Barofsky DF et al. (2009). Transport functions dominate the SAR11 metaproteome at low-nutrient extremes in the Sargasso Sea. ISME J 3: 93–105.
Srinivasan S, Kjelleberg S . (1998). Cycles of famine and feast: the starvation and outgrowth strategies of a marine Vibrio. J Biosci 23: 501–511.
Stocker R, Seymour JR, Samadani A, Hunt DE, Polz MF . (2008). Rapid chemotactic response enables marine bacteria to exploit ephemeral microscale nutrient patches. Proc Natl Acad Sci USA 105: 4209–4214.
Su HS, Lang BF, Newman EB . (1989). L-serine degradation in Escherichia coli K-12: cloning and sequencing of the sdaA gene. J Bacteriol 171: 5095–5102.
Thomas T, Egan S, Burg D, Ng C, Ting L, Cavicchioli R . (2007). The integration of genomics and proteomics into marine, microbial ecology. Theme series on ‘Genomics, proteomics and metabolomics in marine ecology’. Mar Ecol Prog Ser 332: 291–299.
Tripp HJ, Kitner JB, Schwalbach MS, Dacey JW, Wilhelm LJ, Giovannoni SJ . (2008). SAR11 marine bacteria require exogenous reduced sulphur for growth. Nature 452: 741–744.
Vartak NB, Lin CC, Cleary JM, Fagan MJ, Saler MH . (1995). Glucose metabolism in Sphingomonas elodea: pathway engineering via construction of a glucose-6-phosphate dehydrogenase insertion mutant. Microbiology 141: 2339–2350.
Westby CA, Enderlin CS, Steinberg NA, Joseph CM, Meeks JC . (1987). Assimilation of 13NH4+ by Azospirillum brasilense grown under nitrogen limitation and excess. J Bacteriol 169: 4211–4214.
White DC, Sutton SD, Ringelberg DB . (1996). The genus Sphingomonas: physiology and ecology. Curr Opin Biotechnol 7: 301–306.
Whitman WB, Coleman DC, Weibe WJ . (1998). Prokaryotes—the unseen majority. Proc Natl Acad Sci USA 95: 6578–6583.
Zimenkov D, Gulevich A, Skorokhodova A, Biriukova I, Kozlov Y, Mashko S . (2005). Escherichia coli ORF ybhE is pgl gene encoding 6-phosphogluconolactonase (EC 3.1 1 31 ) that has no homology with known 6PGLs from other organisms. FEMS Microbiol Lett 244: 275–280.
Acknowledgements
This work was supported by the Australian Research Council. Mass spectrometric analysis for the work was carried out at the Bioanalytical Mass Spectrometry Facility, UNSW, with assistance from M Raftery, and was supported in part by grants from the Australian Government Systemic Infrastructure Initiative and Major National Research Facilities Program (UNSW node of the Australian Proteome Analysis Facility) and by the UNSW Capital Grants Scheme. HE acknowledges the support of the RCs group and UNSW for sponsoring him as a Visiting Professorial Scholar. This paper has benefited from an in-depth review process, and we wish to thank the Editor and all the reviewers.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Williams, T., Ertan, H., Ting, L. et al. Carbon and nitrogen substrate utilization in the marine bacterium Sphingopyxis alaskensis strain RB2256. ISME J 3, 1036–1052 (2009). https://doi.org/10.1038/ismej.2009.52
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2009.52
Keywords
This article is cited by
-
Molecular hydrogen in seawater supports growth of diverse marine bacteria
Nature Microbiology (2023)
-
The Epidermal Microbiome Within an Aggregation of Leopard Sharks (Triakis semifasciata) Has Taxonomic Flexibility with Gene Functional Stability Across Three Time-points
Microbial Ecology (2023)
-
Diagnosing bioremediation of crude oil-contaminated soil and related geochemical processes at the field scale through microbial community and functional genes
Annals of Microbiology (2020)
-
Microbial communities of aquatic environments on Heard Island characterized by pyrotag sequencing and environmental data
Scientific Reports (2017)
-
Genomic analysis of the nitrate-respiring Sphingopyxis granuli (formerly Sphingomonas macrogoltabida) strain TFA
BMC Genomics (2016)