Abstract
We used molecular techniques to analyze basalts of varying ages that were collected from the East Pacific Rise, 9° N, from the rift axis of the Juan de Fuca Ridge and from neighboring seamounts. Cluster analysis of 16S rDNA terminal restriction fragment polymorphism data revealed that basalt endoliths are distinct from seawater and that communities clustered, to some degree, based on the age of the host rock. This age-based clustering suggests that alteration processes may affect community structure. Cloning and sequencing of bacterial and archaeal 16S rRNA genes revealed 12 different phyla and subphyla associated with basalts. These include the Gemmatimonadetes, Nitrospirae, the candidate phylum SBR1093 in the bacteria, and in the Archaea Marine Benthic Group B, none of which have been previously reported in basalts. We delineated novel ocean crust clades in the γ-Proteobacteria, Planctomycetes and Actinobacteria that are composed entirely of basalt-associated microflora, and may represent basalt ecotypes. Finally, microarray analysis of functional genes in basalt revealed that genes coding for previously unreported processes such as carbon fixation, methane oxidation, methanogenesis and nitrogen fixation are present, suggesting that basalts harbor previously unrecognized metabolic diversity. These novel processes could exert a profound influence on ocean chemistry.
Similar content being viewed by others
Introduction
Oceanic basalts are one of the most abundant rock types on Earth, covering upward of 60% of the Earth's surface. These rocks typically have high permeabilities, which enables infiltration and circulation of large quantities of seawater (Fisher, 1998; Fisher and Becker, 2000). The rock–seawater interaction results in a significant flux of energy and solutes between basalt crust and the overlying seawater (Fisher, 1998). Recent quantitative analyses revealed that basalts harbor 6 × 105–4 × 106 and 3 × 106–1 × 109 cells per g rock (Einen et al., 2008; Santelli et al., 2008). In fact, Einen et al. (2008) suggested that the total number of microorganisms present in ocean crust exceeds the number present in seawater. These observations raise intriguing questions about the role that microorganisms play in biogeochemical cycling in basalts. Biological alteration of basalt by microorganisms has been the focus of numerous studies, with compelling evidence suggesting that they do play a part in this process (Thorseth et al., 1995; Giovannoni et al., 1996; Fisk et al., 1998, 2003; Torsvik et al., 1998; Furnes and Staudigel, 1999; Furnes et al., 2001, 2004; Banerjee and Muehlenbachs, 2003).
Alteration, whether abiotic or biotic, intrinsically changes the chemistry and mineralogy of the rock. For example, alteration of reactive primary minerals to secondary minerals, changing rock permeabilities, and changes in the oxidation state of the rocks alter the chemical milieu in which endolithic microorganisms reside. These changes may result in shifts in the microbial community. Analysis of prokaryotic communities associated with marine basalts revealed that several clades appear to be cosmopolitan in their distribution, as they are associated with globally distributed basalts (Mason et al., 2007; Santelli et al., 2008), regardless of rock age and degree of alteration. The ubiquity of certain clades, such as the α-Marine Group I ocean crust clade IX delineated by Mason et al. (2007), regardless of the age of the host rock, suggests that overall basalt microflora do not change on a temporal scale. However, Lysnes et al. (2004) reported that basalts of varying ages support different microbial phyla and subphyla. For example, the Actinobacteria were associated with older basalts, but were absent in recently erupted material. Therefore, certain clades may, in fact, respond to alteration processes, which could, for example, affect the available electron donors and acceptors.
Certain microbial taxa may be associated with rocks of varying ages, as suggested by Lysnes et al. (2004); however, it is unclear what factors contribute to this habitat specificity. Fresh basalts are ∼8% wt FeO and 2% wt Fe2O3. The increasing oxidation of reduced iron with time could lead to a shift in the microbial community from oxidizers to reducers. In fact, Edwards et al. (2003b) demonstrated that chemolithoautotrophic, iron-oxidizing α- and γ-Proteobacteria isolated from sulfides and metalliferous sediments are able to grow on basalt glass. These isolates are capable of using oxygen and nitrate as electron acceptors. The ability to use multiple electron acceptors would be a requisite as basalt alteration progresses and the in situ redox conditions change.
Alternately, the reduced iron available to iron-oxidizing prokaryotes may become hydrated during fluid–rock interactions. This reaction can evolve hydrogen (Janecky and Seyfried Jr, 1986; Berndt et al., 1996), which can serve as an electron donor for numerous microorganisms including methanogens and sulfate reducers. In fact, Bach and Edwards (2003) estimated that autotrophic sulfate reduction and methanogenesis in marine basalts could result in substantial prokaryotic biomass (9±7 × 1010 and 3±2 × 1010 g C per year, respectively).
Although the geological characteristics of basalts, such as the availability of FeO for microbial iron oxidation, discussed above, do provide some insight into the potential metabolic functions in this environment, examination of the in situ metabolic diversity of prokaryotes by cultivation efforts is limited to one study. Templeton et al. (2005) isolated Mn-oxidizing, heterotrophic bacteria from Loihi Seamount. Thus, there is a need to circumvent the lack of cultured microorganisms using a molecular approach to determine metabolic diversity in basalts. GeoChip is a molecular tool that does not rely on cultivation-based methods to assay for functional diversity. Specifically, it is a functional gene microarray that has 24 243 oligonucleotide probes covering >10 000 genes in >150 functional groups involved in nitrogen, carbon, sulfur and phosphorus cycling (He et al., 2007). GeoChip can provide significant insight into metabolic potential in a given environment, such as marine basalts.
In this study, we used terminal restriction fragment length polymorphism (T-RFLP), cloning and sequencing, and microarray analysis of functional genes to (1) assess successional changes in the microbial communities associated with basalts of varying ages and from different geographical locations, (2) examine species composition and distribution and (3) determine potential metabolic function in basalts by examining functional genes.
Our analyses revealed that rock age, or degree of alteration, may, to some degree, play a role in community succession. In addition, we report previously unrecognized phyla in basalts and several novel ocean crust clades of microorganisms that may represent basalt specialists. Finally, the examination of functional genes in basalt revealed the genetic potential for several novel metabolic processes. This analysis provides insight into biogeochemical cycling in this ocean crust environment.
Materials and methods
Sample collection
Glassy pillow basalts were collected from areas of low (or no) sediment accumulation using the DSV Alvin on two separate cruises to East Pacific Rise (9° N) and to the Co-axial segment of the Juan de Fuca Ridge (JdF) and neighboring seamounts (Table 1). Basalt samples were collected and placed inside a collection box, or ‘biobox’, which was designed to prevent sample exposure to ambient seawater during the ascent to the surface. Before the dive, the box was filled with either 0.2-μm filtered seawater or sterile Millipore water. During Alvin's descent, residual airspace was replaced with seawater that passed through 0.2 μm filters embedded in the lid. The biobox volume (16 l) allowed for several liters of ambient deep seawater to be collected with the basalts. Once on deck, the samples were removed from the biobox using sterile (flamed) tongs and placed into separate freezer bags. Samples were immediately frozen at −80 °C and remained frozen until shore-based analyses. To control for deep-sea planktonic organisms that may have found their way into fractures and pores in the basalt samples, the biobox water was filtered and the filters were frozen and analyzed along with the basalts (see below).
Nucleic acid extraction from the basalt samples
For molecular analyses, all rock sample-handling and all extraction steps were performed in a sterile laminar flow hood. Ceramic tumbling vessels, chisels, mortar and pestles were baked at 220 °C for at least 24 h. The outer rock surface was removed by tumbling the rock several times for 20 min, replacing with sterile grit each time. The glassy rind was pared away with a chisel and/or sterile rock splitter. Approximately 1 cm3 was powdered with a tungsten mortar and pestle, and 2 ml of the powder was used in each extraction. Two control DNA extractions, to which either 2 ml of the grit from the last tumbling step or no rock or grit material was added, were used to assess contamination introduced by the tumbling steps or by the DNA-extraction reagents, respectively.
The DNA-extraction protocol was optimized for extracting DNA from basalts. Specifically, each extraction tube contained 2 ml of rock powder and final concentrations of the following: 4.5 mM Tris·HCl, pH 7.4; 185 mM EDTA, pH 8.0; 4.5% Chelex-100 (w v−1); 0.7% sodium dodecyl sulfate (w v−1); and 2 mg proteinase K (Qiagen Inc., Valencia, CA, USA). Tubes were placed in a 37 °C rotator with gentle agitation (180 r.p.m.) overnight. The rock powder was separated from the supernatant by low-speed centrifugation and mixed with final concentrations of the following: 923 mM NaCl and 1.3% cetyltrimethylammonium bromide (w v−1). Samples were incubated at 65 °C for 30 min, extracted once with an equal volume of phenol/chloroform/isoamyl alcohol (25:24:1) (pH 7.4) and then extracted twice with an equal volume of chloroform/isoamyl alcohol (24:1). Nucleic acids were precipitated by adding one volume of isopropanol and incubating the samples overnight at 4 °C. Samples were spun at 24 000 g for 1 h at 4 °C in a TL-100 ultracentrifuge (Beckman Instruments Inc., Palo Alto, CA, USA) to pellet the precipitated nucleic acids. Pellets were washed with 70% ethanol (v v−1), dried in a laminar flow hood at room temperature and suspended in sterile water. Replicate extracts were combined (40 μl final volume) and stored at −80 °C.
DNA extraction from the filtered biobox water samples
At least 12 l of biobox water from each dive was filtered through a 142-mm 0.2-μm Supor filter (Pall Gelman Laboratory, Ann Arbor, MI, USA) in a polycarbonate filter holder (Geotech Environmental Equipment Inc., Denver, CO, USA) that was connected to a peristaltic pump. Filters were immediately preserved in 5 ml of sucrose lysis buffer (20 mM EDTA, 400 mM NaCl, 0.75 M sucrose, 50 mM Tris·HCl (pH 9.0)) and stored at −80 °C. Total community nucleic acids were extracted from the filters according to Giovannoni et al. (1990).
Terminal restriction fragment length polymorphism analysis
Terminal restriction fragment length polymorphism analysis was used to compare the archaeal and bacterial communities from several rocks and corresponding biobox seawater samples according to Moeseneder et al. (1999) with few modifications. The archaeal 16S rRNA genes were amplified using the primers Arch20F (DeLong et al., 1999) and Arch915R (Stahl and Amann, 1991), with the forward primer 5′-end-labeled with phosphoramidite fluorochrome 5-carboxyfluorescein (6-FAM) and the reverse primer labeled with 5-hexachlorofluorescein. Fifty PCR cycles were necessary to amplify archaeal DNA, whereas a semi-nested approach was required to amplify bacterial 16S rRNA genes from nearly all of the basalts, with primers 27F-B (5′-AGRGTTYGATYMTGGCTCAG) and 1492RY (5′-GGYTACCTTGTTACGACTT) modified from Lane (1991) used in the initial PCR (30 cycles) and primers 27F-B-[FAM] and 1391R (Lane, 1991) used in the second reaction (20 cycles). Only the forward strand of this PCR product was labeled for the T-RFLP analysis. For both archaeal and bacterial amplifications, three replicate PCRs (50 μl) for each DNA sample contained final concentrations of the following: 1μl of DNA extract; 1% (v v−1) PCR buffer (+ NH4SO4; MBI Fermentas, Hanover, MD, USA); 0.2 mM each deoxynucleotide triphosphate; 0.2 μM each primer; 2 mM MgCl2 (MBI Fermentas); 1.2 mg ml−1 bovine serum albumin (non-acetylated; Sigma, St Louis, MO, USA); 1% (w v−1) PVP (polyvinylpyrrolidone); and 2.5 U Taq polymerase (MBI Fermentas). PCR cycling consisted of denaturation at 94 °C for 1.5 min, annealing at 55 °C for 1.5 min and extension at 72 °C for 1.5 min. The filtered biobox seawater samples from each dive were also analyzed using the same cycling conditions as the basalts, except that the number of cycles was reduced to 30.
PCR products (50 ng) were digested with 10 U of enzyme for 6 h at 37 °C with each of three separate restriction enzymes: AluI, BsuRI (HaeIII), and Hin6I (HhaI) (MBI Fermentas). Samples were run on an ABI 3100 (Applied Biosystems Inc. (ABI), Foster City, CA, USA). The fingerprint patterns for the rock and seawater communities were compared according to Moeseneder et al. (1999); however, only peaks longer than 70 bp in length were included in the analysis. Data were standardized by inclusion of peaks that represented >1% of the total peak height for each fingerprint and were then converted to binary matrices. Binary data were analyzed by the unweighted pair group with mathematical averages method in Phylogenetic Analysis Using Parsimony *(and other methods) version 4.0 b10 (Swofford, 1998) using the site distance-matrix method of Nei and Li (1979) according to Moeseneder et al. (1999).
PCR amplification and cloning of prokaryotic 16S rRNA genes
Data from the unweighted pair group with mathematical averages analysis were used to select three basalt samples that differed in age and community structure (D3718B, 9° N EPR, D3815F and D3823M JdF) for cloning and sequencing of archaeal and bacterial 16S rRNA genes. The archaeal communities were amplified according to the PCR conditions described above for T-RFLP analysis, except that the primers were not fluorescently labeled. To amplify archaeal 16S rDNA from D3815F, a semi-nested approach was employed using the primers Arch20F and 1492RY in the initial reaction and Arch20F and Arch915R in the semi-nested reaction. The bacterial community from D3718B was amplified using the semi-nested approach described above. Amplification of bacterial 16S rDNA from D3823M did not require a semi-nested approach. As with the T-RFLP analysis, the corresponding seawater samples from each dive were also cloned for comparison. PCRs (50 μl volume) were cloned into the pGEM-T Easy vector (Promega Corp., Madison, WI, USA). Clone libraries were constructed and screened according to the methods of Vergin et al. (2001). Briefly, clones were assigned to clone families based upon shared patterns for two separate restriction digests. Digested PCR products were resolved on a 3% agarose gel. One clone from each unique RFLP pattern was sequenced using an ABI 3730 capillary sequencer. Full-length sequences were obtained for clones representing each phylotype. Clones with restriction patterns that appeared only once in the library were designated ‘unique’. Percent coverage was calculated based upon the number of unique clones versus total clones according to the method of Good (1953). Chimeric sequences were identified with the CHECK_CHIMERA program (Maidak et al., 1997, 1999) and Mallard (Ashelford et al., 2006).
Phylogenetic analysis
The phylogenies of microorganisms from D3718B and D3823M were extensively reviewed in Mason et al. (2007). Clones from these libraries are presented here in phylogenetic dendrograms only if they are part of novel ocean crust clades delineated here or if they are highly similar to clones from D3815F. However, clones from all libraries were analyzed during phylogenetic reconstruction. Phylogenetic analyses and clade delineations were carried out according to Mason et al. (2007), using the Greengenes database (DeSantis et al., 2006). Briefly, neighbor-joining, maximum parsimony and maximum-likelihood trees of near-full-length sequences were generated in ARB (Ludwig et al., 2004). Maximum-likelihood trees were generated using Tree-Puzzle (Schmidt et al., 2002), with the Hasegawa–Kishino–Yano model (Hasegawa et al., 1985). Quartet-puzzling (QP) reliability values are not shown at bifurcations if they are below 50%. In determining clades, QP values from 90% to 100% are strongly supported; however, QP values less than 70% can also be trusted (Schmidt et al., 2002). Clades delineated here with QP values lower than 70% were analyzed relative to QP support values of the other branches in the tree (Schmidt et al., 2002).
Nucleotide sequence accession numbers
The 16S rRNA gene sequences for the archaeal and bacterial clones were submitted to the GenBank database and have been assigned the following accession numbers: DQ070750–DQ070835 (D3718B and D3823M) and FJ024305–FJ024341 (D3815F).
Functional genes
Basalt sample D3815F was selected for functional gene analysis because it had several clades that have not been previously reported from this environment, particularly the Marine Benthic Group B. We hypothesized that this diversity of species would be mirrored in the diversity of functional genes. Second, thin sections of this sample showed textures that suggest bioalteration; therefore, analysis of functional genes in this sample would provide insight into the biological processes that may result in these textural features. Functional genes were assayed for using the GeoChip 2.0 (He et al., 2007) microarray following previously described methods (Wu et al., 2006; He et al., 2007). Briefly, DNA from D3815F was amplified in triplicate using a Templiphi 500 amplification kit (Amersham Biosciences, Piscataway, NJ, USA) following the manufacturer's protocol. To facilitate amplification, spermidine (0.1 μg μl−1) and single-strand binding protein (0.04 mM) were added to the reaction mixture. Amplified DNA was fluorescently labeled with Cy5. Hybridizations were performed using a HS4800Pro Hybridization Station (TECAN US, Durham, NC, USA) overnight at 42 °C. Microarrays were scanned using a ProScanArray (PerkinElmer, Waltham, MA, USA). Images were then analyzed using ImaGene 6.0 (BioDiscovery, El Segundo, CA, USA) to designate the identity of each spot and to determine spot quality. Data were processed as described by Wu et al. (2006). Briefly, raw data from Imagene were analyzed using a GeoChip data analysis pipeline. A signal-to-noise ratio of ⩾3 was considered a positive signal. A positive signal in at least one-third of the probes for a particular gene (minimum of two probes) was required for a gene to be considered positive. Each gene had one, two or three probes per array based on the number of probes available, meeting the criteria described by He et al. (2007).
Results and discussion
T-RFLP
Unweighted pair group with mathematical averages cluster analysis of T-RFLP data revealed that the archaeal and bacterial communities were distinct from deep seawater communities (Figure 1). Further, there was striking congruency in the unweighted pair group with mathematical averages clustering patterns for the four oldest JdF samples. These old samples, ranging from a few thousand to about 3 Myr in age, clustered together, whereas the younger basalts from 9° N (from an eruption in 1991) clustered with one JdF sample of a similar age (D3826U, from the 1993 lava flow). This clustering is evidence that there are differences in microbial communities present in recently erupted basalts compared with older, more weathered rocks. The observed clustering is supported, to some degree, by phylogeny. For example, the Planctomycetes ocean crust clade XIV members (see below) are from recently erupted to medium-aged basalts. Overall, however, there is a distinct overlap in the microbial communities regardless of rock age. For example, the basalt-specific ocean crust clade presented here, such as the γ-Proteobacteria ocean crust clade XII, is composed of microorganisms from young, fresh basalts to 3.3-Myr-old basalts. This pattern suggests that basalt microflora are largely associated with rocks of varying ages, but that a minority may reside in, for example, younger, less-altered rocks to the exclusion of older, more-weathered rocks. This finding is consistent with that of Lysnes et al. (2004), who reported that specific bacterial species are found only in rocks of a certain age.
UPGMA analysis of T-RFLP fingerprint patterns for the bacterial (left) and archaeal (right) communities recovered from basalts (above the dashed line) and background seawater from 9° N and JdF (below the dashed line). Older basalts (>20 years) from JdF are underlined. All 9° N samples are less than 20 years. Sample numbers indicate Alvin dive number and location: 9N is 9° N on the East Pacific Rise and JdF is Juan de Fuca Ridge and Cobb Seamount. The scale bar represents similarity. T-RFLP, terminal restriction fragment length polymorphism; UPGMA, unweighted pair group with mathematical averages.
Phylogenetic analysis
A total of 547 bacterial and archaeal 16S rDNA clones were analyzed and 173 unique clones were sequenced (Table 2). This analysis revealed that Gemmatimonadetes, Nitrospirae, SBR1093 and in the Archaea, the Marine Benthic Group B (Figures 2, 3 and 4), were present in basalt samples. None of these clades have been previously reported in marine basalts. In addition, microorganisms in the α-, δ- and γ-Proteobacteria, Acidobacteria, Actinobacteria, Bacteroidetes and Planctomycetes in the bacterial domain are reported (Figures 2 and 3). The most prevalent microorganisms were Proteobacteria (56%), the majority of which were γ- (25%), α- (15%) and δ- (13%), followed by Bacteroidetes (10%), Actinobacteria (9%), Planctomycetes (7%), Acidobacteria (6%) and Gemmatimonadetes (3%). The remaining clades were observed in a single rock sample.
Maximum-likelihood phylogenetic tree of proteobacterial 16S rRNA gene sequences from basalt samples. The Proteobacteria tree was constructed with 25 000 puzzling steps. A general Bacteria filter was used. The 16S rDNA sequence of Aquifex pyrophilus (M83548) was used as the outgroup (not shown). The α-Proteobacteria ocean crust clade II, designated by an (*), was delineated by Mason et al. (2007) and is included here because new basalt sequences extend this clade. The scale bar indicates 0.1 nucleotide substitutions per site.
Maximum-likelihood phylogenetic tree of Actinobacteria, Cyanobacteria, Bacteroidetes, Planctomycetes, Gemmatimonadetes, Acidobacteria, Nitrospirae and SBR1093 16S rRNA gene sequences from basalt samples. The bacterial tree was constructed with 25 000 puzzling steps. A general Bacteria filter was used. The 16S rDNA sequence of Aquifex pyrophilus (M83548) was used as the outgroup (not shown). The scale bar indicates 0.1 nucleotide substitutions per site.
Maximum-likelihood phylogenetic tree of archaeal 16S rRNA gene sequences from basalt samples. The archaeal tree was constructed with 25 000 puzzling steps. A general Archaea filter was used. The 16S rDNA sequence of Aquifex pyrophilus (M83548) was used as the outgroup (not shown). The α-MGI ocean crust clade VIII, designated by an (*), was delineated by Mason et al. (2007) and is included here because new basalt sequences extend this clade. The scale bar indicates 0.1 nucleotide substitutions per site. MGI, Marine Group I Crenarchaeota.
Our observations are consistent with those reported by Santelli et al. (2008), who analyzed basalts from the East Pacific Rise and from Hawaii and found 68%/66% (EPR%/Hawaii%) Proteobacteria, 8%/5% Planctomycetes, 7%/8% Actinobacteria, 4%/1% Bacteroidetes and 3%/4% Acidobacteria. The similarity in bacterial communities associated with basalts from a broad geographic distribution suggests cosmopolitan distributions of these clades, which is in agreement with the findings presented by Mason et al. (2007) and Santelli et al. (2008).
Phylogenetic reconstruction revealed three novel ocean crust clades composed entirely of microorganisms associated with basalt. These new clades are the γ-Proteobacteria ocean crust clade XII (Figure 2), Actinobacteria ocean crust clade XIII (Figure 3) and Planctomycetes ocean crust clade XIV (Figure 3). These clades are comprised of bacteria sampled from JdF (this study), East Pacific Rise, 9° N (this study and Santelli et al., 2008), and Hawaiian (Santelli et al., 2008) basalts. These cosmopolitan basalt clades may represent ecotypes of bacteria that are specifically adapted to this environment.
Cloning and sequencing of archaeal 16S rDNA revealed that Marine Benthic Group B were present in basalts (Figure 4). This is the first report of this clade in this environment, as previous studies that examined the archaeal communities in basalts revealed only Marine Group I Crenarchaeota (Thorseth et al., 2001; Fisk et al., 2003; Lysnes et al., 2004; Mason et al., 2007). Recently, quantitative analyses of the microbial communities in basalts revealed that Archaea comprise 4–12% and 0.02% or less of the prokaryotic communities (Santelli et al., 2008; Einen et al., 2008), respectively. Although these estimates are disparate, they do reveal that Archaea are a minor component in the overall microbial communities that reside in basalt. Although Archaea are less prevalent, they are ubiquitous in basalts and have been reported in all studies that assayed for their presence (Thorseth et al., 2001; Fisk et al., 2003; Lysnes et al., 2004; Mason et al., 2007; Einen et al., 2008; Santelli et al., 2008). Further, as discussed previously, a clade of Marine Group I Archaea appears to be endemic to basalt (Mason et al., 2007). This habitat specificity and global distribution indicate that some Archaea, although less abundant than bacteria, are particularly adapted to life in basalt and likely play a role in biogeochemical cycling.
Functional genes
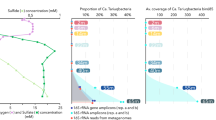
GeoChip (He et al., 2007) microarray analysis of functional genes in basalt sample D3815F revealed the presence of genes coding for metabolic processes previously unrecognized in this environment. In this analysis, a total of 604 probes of the 24 243 total probes present on GeoChip were positive. Specifically, we found genes coding for carbon fixation, methane production and oxidation, nitrogen fixation, ammonium oxidation, nitrate and nitrite reduction, dissimilatory sulfate reduction and iron reduction (see Supplementary Table 1 for a complete list).
Here we report genes coding for carbon fixation. Basalts lacking a sediment layer are considered to be an oligotrophic, low-carbon environment (Edwards et al., 2003a); thus carbon cycling in this habitat is particularly significant. The oligotrophic nature of this environment suggests that carbon fixation would be paramount in this habitat. In fact, chemolithoautotrophic processes in marine subsurface ridge flank hydrothermal environments have been theoretically shown to provide energy that could result in significant microbial biomass (∼1 × 1012 g C per year) (Bach and Edwards, 2003). Therefore, chemolithoautotrophic processes occurring in situ could serve to underpin a basalt-hosted biosphere. One such process is methanogenesis, where hydrogen can serve as the electron donor to reduce carbon dioxide, evolving methane. During fluid–rock interactions when the basalt minerals olivine and pyroxene react with water, hydrogen may be evolved (Janecky and Seyfried Jr, 1986; Berndt et al., 1996). Thus, the requisite electron donor may be present as a result of this abiotic reaction.
Here, we report that genes coding for methanogenesis are present in basalt. Methanogens have not been reported in molecular analyses of Archaea in basalts conducted to date (Thorseth et al., 2001; Fisk et al., 2003; Lysnes et al., 2004; Mason et al., 2007). However, Lysnes et al. (2004) reported that methane was evolved in enrichment cultures inoculated with marine basalts. Although the Marine Benthic Group B clade currently lacks a cultured representative (Knittel et al., 2005), they are frequently associated with environments dominated by methane, methanogens and methanotrophs (Knittel et al., 2005; Kendall and Boone, 2006; Kendall et al., 2007). The role of this clade in the environment is unknown, but it is plausible that they are involved in methane biogeochemical cycling. Although no known methanogens were observed in our study, the diversity of mcr genes (Supplementary Table 1) in conjunction with a clade typically observed in methane-rich environments suggests that this metabolic process may be occurring in basalts. Methane resulting from biological processes could serve as a carbon and energy source for heterotrophic processes. In fact, we found genes coding for methane oxidation. Methane cycling in marine basalts would have a direct impact on the overlying hydrosphere.
As discussed above, basalts are not carbon replete. Similarly, they are composed of only a small amount of nitrogen, averaging approximately 2 p.p.m. (Marty et al., 1995). Therefore, the detection of genes coding for nitrogen fixation is intriguing. Cowen et al. (2003), and more recently Huber et al. (2006), investigated ocean crust fluids and reported elevated levels of ammonium compared with seawater. Cowen et al. (2003) suggested that nitrogen fixation may serve as the source of this excess ammonium. Mehta et al. (2005) attributed nitrogen fixation in crustal fluids and in deep seawater to non-methanogenic Archaea, which are the only known archaeal nitrogen fixers. In that study, nifH genes were detected in crustal fluids. Nitrogen fixation may also be taking place in the host rocks themselves, given the presence of nifH genes in our basalt sample.
Nitrogen fixation could augment the low nitrogen concentrations in basalts and may ultimately support ammonium-oxidizing microorganisms. This hypothesis is supported by the presence of genes that code for ammonium oxidation in our basalt sample. As reported by Mason et al. (2007) (see Figures 3 and 4), basalt sequences similar to Nitrosococcus oceani (89% similar) and Nitrosospira multiformis (96% similar), both of which are known ammonium-oxidizing microorganisms (Watson, 1965; Watson et al., 1971), were derived from basalts from JdF and 9° N (this study), and Mohns Ridge (Einen et al., 2006). Thus, phylogenetic and functional gene analyses both suggest that ammonium oxidation may be occurring in basalts.
Further, basalt clones closely related to the nitrite-oxidizing Nitrospina gracilis (91–94% similar) and Nitrospira marina (95–96% similar) (Watson and Waterbury, 1971; Tal et al., 2003) were found (Figures 2 and 3), suggesting that nitrite oxidation, the second step in nitrification, may be occurring in basalts. This observation could not be confirmed using GeoChip, however, because genes coding for nitrite oxidation are not present on the gene chip.
Nitrification could provide the substrate for both denitrification and anaerobic ammonium oxidation (anammox), both of which lead to loss of nitrogen (Lam et al., 2007). In fact, we found numerous genes coding for nitrate and nitrite reduction; therefore, the genetic potential for denitrification is present in this environment. Recently, Edwards et al. (2003b) demonstrated that chemolithoautotrophic iron-oxidizing bacteria are able to grow on basalt glass using nitrate as the electron acceptor. Whether anaerobic ammonium oxidation is occurring in basalts remains unclear. Although Planctomycetes have been reported in basalts, including the novel ocean crust clade Planctomycetes XIV delineated here, microorganisms closely related to known anammox bacteria, such as Kuenenia stuttgartiensis (77% similar to basalt-associated microorganisms), have not been detected. Therefore, it is unclear if this process is important in considering nitrogen loss from the basalt layer. Our data do suggest, however, that nitrogen could be lost from marine crust by denitrification processes.
In addition to genes coding for denitrification processes, we also detected genes coding for iron reduction and dissimilatory sulfate reduction in basalt. Together, these genes suggest that anaerobic respiration may be occurring in basalt. The presence of genes that code for aerobic respiration (for example, ammonium oxidation) in the same sample indicates that aerobic and anaerobic processes may occur simultaneously on a small spatial scale, suggesting, perhaps that microniches are occupied by prokaryotes in basalt. Consistent with our findings, it was reported that in upper basaltic crust, redox conditions are such that aerobic and anaerobic processes are likely supported (Bach and Edwards, 2003).
Conclusion
Basalts from JdF, neighboring seamounts and 9° N, EPR harbor cosmopolitan microorganisms that are distinct from seawater prokaryotes. Several novel ocean crust clades, composed only of microorganisms from basalts, suggest that some bacteria are specifically adapted to this ocean crust environment. Our analysis of geochemically important functional genes revealed the potential for several metabolic processes not known to be occurring in basalts, particularly carbon fixation, methanogenesis, methane oxidation, nitrogen fixation and denitrification. Our data suggest that basalts not only harbor a diversity of broadly distributed microbial species, but also unexpected metabolic diversity. Future studies should utilize culture-dependent and culture-independent methods to analyze biogeochemical cycling in basalts to better understand the biological processes in this vast subsurface environment and how these processes ultimately affect ocean chemistry.
Accession codes
References
Ashelford KE, Chuzhanova NA, Fry JC, Jones AJ, Weightman AJ . (2006). New screening software shows that most recent large 16S rRNA gene clone libraries contain chimeras. Appl Environ Microbiol 72: 5734–5741.
Bach W, Edwards KJ . (2003). Iron and sulfide oxidation within the basaltic ocean crust: implications for chemolithoautotrophic microbial biomass production. Geochim Cosmochim Acta 67: 3871–3887.
Banerjee NR, Muehlenbachs K . (2003). Tuff life: Bioalteration in volcaniclastic rocks from the Ontong Java Plateau. Geochem Geophys Geosyst 4 (doi:10.1029/2002GC000470).
Berndt ME, Allen DE, Seyfried WE . (1996). Reduction of CO2 during serpentinization of olivine at 300 °C and 500 bar. Geology 24: 351–354.
Cowen JP, Giovannoni SJ, Kenig F, Johnson HP, Butterfield D, Rappe MS et al. (2003). Fluids from aging ocean crust that support microbial life. Science 299: 120–123.
DeLong EF, Taylor LT, Marsh TL, Preston CM . (1999). Visualization and enumeration of marine planktonic archaea and bacteria by using polyribonucleotide probes and fluorescent in situ hybridization. Appl Environ Microbiol 65: 5554–5563.
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K et al. (2006). Greengenes, a Chimera-Checked 16S rRNA Gene Database and Workbench Compatible with ARB. Appl Environ Microbiol 72: 5069–5072.
Desonie DL, Duncan RA . (1990). The Cobb-Eickelberg seamount chain: Hotspot volcanism with mid-ocean ridge basalt affinity. J Geophys Res 95: 12697–12711.
Edwards KJ, Bach W, Rogers DR . (2003a). Geomicrobiology of the ocean crust: a role for chemoautotrophic Fe-bacteria. Biol Bull 204: 180–185.
Edwards KJ, Rogers DR, Wirsen CO, McCollom TM . (2003b). Isolation and characterization of novel psychrophilic, neutrophilic, fe-oxidizing, chemolithoautotrophic alpha- and gamma-proteobacteria from the deep sea. Appl Environ Microbiol 69: 2906–2913.
Einen J, Kruber C, Øvreas L, Thorseth IH, Torsvik T . (2006). Microbial colonization and alteration of basaltic glass. Biogeosci Discuss 3: 273–307.
Einen J, Thorseth IH, Ovreas L . (2008). Enumeration of Archaea and Bacteria in seafloor basalt using real-time quantitative PCR and fluorescence microscopy. FEMS Microbiol Lett 282: 182–187.
Embley RW, Chadwick WW, Perfit MR, Smith MC, Delaney JR . (2000). Recent eruptions on the CoAxial segment of the Juan de fuca ridge: Implications for mid-ocean ridge accretion processes. J Geophys Res 105: 16501–16525.
Fisher AT . (1998). Permeability within basaltic ocean crust. Rev Geophys 36: 143–182.
Fisher AT, Becker K . (2000). Channelized fluid flow in oceanic crust reconciles heat-flow and permeability data. Nature 403: 71–74.
Fisk MR, Giovannoni SJ, Thorseth IH . (1998). Alteration of oceanic volcanic glass: textural evidence of microbial activity. Science 281: 978–980.
Fisk MR, Storrie-Lombardi MC, Douglas S, Popa R, McDonald G, Di Meo-Savoie CA . (2003). Evidence of biological activity in Hawaiian subsurface basalts. Geochem Geophys Geosyst 4 (doi:10.1029/2002GC000387).
Fornari DJ, Embley RW . (1995). Tectonic and volcanic controls on hydrothermal processes at the mid-ocean ridge: an overview based on near-bottom and submersible studies. In: Humphris SE, Zierenberg RA, Mullineaux LS, Thompson RE (eds). Seafloor Hydrothermal Systems: Physical Chemical, Biological, and Geological Interactions. Washington, DC, American Geophysical Union Monograph, v. 91. pp 1–46.
Furnes H, Banerjee NR, Muehlenbachs K, Staudigel H, de Wit M . (2004). Early life recorded in Archean Pillow Lavas. Science 304: 578–581.
Furnes H, Staudigel H . (1999). Biological mediation in ocean crust alteration: how deep is the deep biosphere? Earth Planet Sci Lett 166: 97–103.
Furnes H, Staudigel H, Thorseth IH, Torsvik T, Muehlenbachs K, Tumyr O . (2001). Bioalteration of basaltic glass in the oceanic crust. Geochem Geophys Geosyst 2 (doi:10.1029/2000GC000150).
Giovannoni SJ, DeLong EF, Schmidt TM, Pace NR . (1990). Tangential flow filtration and preliminary phylogenetic analysis of marine picoplankton. Appl Environ Microbiol 56: 2572–2575.
Giovannoni SJ, Fisk MR, Mullins TD, Furnes H . (1996). Genetic evidence for endolithic microbial life colonizing basaltic glass-seawater interfaces. Proc ODP, Sci Results 148: 207–214.
Good IJ . (1953). The population frequencies of species and the estimation of population parameters. Biometrika 40: 237–264.
Hasegawa M, Kishino H, Yano T . (1985). Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol 22: 160–174.
Haymon RM, Fornari DJ, Von Damm KL, Lilley MD, Perfit MR, Edmond JM et al. (1993). Volcanic eruption of the mid-ocean ridge along the East Pacific Rise crest at 9°45–52′N: direct submersible observations of seafloor phenomena associated with an eruption event in April, 1991. Earth Planet Sci Lett 119: 85–101.
He Z, Gentry TJ, Schadt CW, Wu L, Liebich J, Chong SC et al. (2007). GeoChip: a comprehensive microarray for investigating biogeochemical, ecological and environmental processes. ISME J 1: 67–77.
Huber JA, Johnson HP, Butterfield DA, Baross JA . (2006). Microbial life in ridge flank crustal fluids. Environ Microbiol 8: 88–99.
Janecky DR, Seyfried Jr WE . (1986). Hydrothermal serpentinization of peridotite within the oceanic crust: experimental investigations of mineralogy and major element chemistry. Geochim Cosmochim Acta 50: 1357–1378.
Kendall MM, Boone DR . (2006). Cultivation of methanogens from shallow marine sediments at Hydrate Ridge, Oregon. Archaea 2: 31–38.
Kendall MM, Wardlaw GD, Tang CF, Bonin AS, Liu Y, Valentine DL . (2007). Diversity of Archaea in marine sediments from Skan Bay, Alaska, including cultivated methanogens, and description of Methanogenium boonei sp. nov. Appl Environ Microbiol 73: 407–414.
Knittel K, Losekann T, Boetius A, Kort R, Amann R . (2005). Diversity and distribution of methanotrophic archaea at Cold Seeps. Appl Environ Microbiol 71: 467–479.
Lam P, Jensen MM, Lavik G, McGinnis DF, Müller B, Schubert CJ et al. (2007). Linking crenarchaeal and bacterial nitrification to anammox in the Black Sea. Proc Natl Acad Sci USA 104: 7104–7109.
Lane DJ . (1991). 16S/23S rRNA sequencing. In: Stackenbrandt E, Goodfellow M (eds). Nucleic Acid Techniques in Bacterial Systematics. John Wiley: New York. pp 115–148.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al. (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: 1363–1371.
Lysnes K, Thorseth IH, Steinsbu BO, Øvreas L, Torsvik T, Pedersen RB . (2004). Microbial community diversity in seafloor basalt from the Arctic spreading ridges. FEMS Microbiol Ecol 50: 213–230.
Maidak BL, Cole JR, Parker Jr CT, Garrity GM, Larsen N, Li B et al. (1999). A new version of the RDP (ribosomal database project). Nucleic Acids Res 27: 171–173.
Maidak BL, Olsen GJ, Larsen N, Overbeek R, McCaughey MJ, Woese CR . (1997). The RDP (ribosomal database project). Nucleic Acids Res 25: 109–111.
Marty B, Lenoble M, Vassard N . (1995). Nitrogen, helium and argon in basalt: a static mass spectrometry study. Chem Geol 120: 183–195.
Mason OU, Stingl U, Wilhelm LJ, Moeseneder MM, Di Meo-Savoie CA, Fisk MR, Giovannoni SJ . (2007). The phylogeny of endolithic microbes associated with marine basalts. Environ Microbiol 9: 2539–2550.
Mehta MP, Huber JA, Baross JA . (2005). Incidence of novel and potentially archaeal nitrogenase genes in the deep Northeast Pacific Ocean. Environ Microbiol 7: 1525–1534.
Moeseneder MM, Arrieta JM, Muyzer G, Winter C, Herndl GJ . (1999). Optimization of terminal-restriction fragment length polymorphism analysis for complex marine bacterioplankton communities and comparison with denaturing gradient gel electrophoresis. Appl Environ Microbiol 65: 3518–3525.
Nei M, Li W-H . (1979). Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76: 5269–5273.
Santelli CM, Orcutt BN, Banning E, Bach W, Moyer CL, Sogin ML et al. (2008). Abundance and diversity of microbial life in ocean crust. Nature 453: 653–656.
Schmidt HA, Strimmer K, Vingron M, von Haeseler A . (2002). TREE-PUZZLE: maximum likelihood phylogenetic analysis using quartets and parallel computing. Bioinformatics 18: 502–504.
Stahl DA, Amann R . (1991). Development and application of nucleic acid probes. In: Stackebrandt E, Goodfellow M (eds). Nucleic Acid Techniques in Bacterial Systematics. John Wiley and Sons, Inc, New York, NY. pp 205–248.
Swofford DL . (1998). PAUP*. Phylogenetic analysis using parsimony (*and other methods). Version 4, Sinauer Associates, Sunderland, Massachusetts.
Tal Y, Watts JEM, Schreier SB, Sowers KR, Schreier HJ . (2003). Characterization of the microbial community and nitrogen transformation processes associated with moving bed bioreactors in a closed recirculated mariculture system. Aquaculture 215: 187–202.
Templeton A, Staudigel H, Tebo B . (2005). Diverse Mn(II)-oxidizing bacteria isolated from submarine basalts at Loihi Seamount. Geomicrobiol J 22: 127–139.
Thorseth IH, Torsvik T, Furnes H, Muehlenbachs K . (1995). Microbes play an important role in the alteration of oceanic crust. Chem Geol 126: 137–146.
Thorseth IH, Torsvik T, Torsvik V, Daae FL, Pedersen RB, Party K-S . (2001). Diversity of life in ocean floor basalt. Earth Planet Sci Lett 194: 31–37.
Torsvik T, Furnes H, Muehlenbachs K, Thorseth IH, Tumyr O . (1998). Evidence for microbial activity at the glass-alteration interface in oceanic basalts. Earth Planet Sci Lett 162: 165–176.
Vergin KL, Rappé MS, Giovannoni SJ . (2001). Streamlined method to analyze 16S rRNA gene clone libraries. BioTechniques 30: 938–943.
Watson SW . (1965). Characteristics of a marine nitrifying bacterium, Nitrosocystis oceanus sp. n. Limnol Oceanogr 10 (Suppl.): R274–R289.
Watson SW, Graham LB, Remsen CC, Valois FW . (1971). A lobular, ammonia-oxidizing bacterium, Nitrosolobus multiformis Nov. Gen. Nov. sp. Arch Microbiol 76: 183–203.
Watson SW, Waterbury JB . (1971). Characteristics of two marine nitrite oxidizing bacteria, Nitrospina gracilis nov. gen. nov. sp. and Nitrococcus mobilis nov. gen. nov. sp. Arch Microbiol 77: 203–230.
Wu L, Liu X, Schadt CW, Zhou J . (2006). Microarray-based analysis of subnanogram quantities of microbial community DNAs by using whole-community genome amplification. Appl Environ Microbiol 72: 4931–4941.
Acknowledgements
We gratefully acknowledge the captain and crew of the R/V Atlantis and the DSV Alvin group for their generous assistance in sample collection. We also thank Craig Cary (chief scientist, Extreme 2001 cruise) for inviting us to participate on the 9° N cruise. This research was supported by a Marine Microbiology Initiative Investigator Award from the Gordon and Betty Moore Foundation to SJ Giovannoni, the National Science Foundation with a LExEn grant awarded to SJ Giovannoni and MR Fisk (OCE-0085436), an NSF IGERT Subsurface Biosphere Fellowship to OU Mason and a RIDGE postdoctoral fellowship awarded to CA Di Meo-Savoie (OCE-0002411). The GeoChip microarray analysis was supported in part by the United States Department of Energy under the Genomics:GTL Program to JZ Zhou through the Virtual Institute of Microbial Stress and Survival (VIMSS; http://vimss.lbl.gov) of the Office of Biological and Environmental Research, Office of Science.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Supplementary information
Rights and permissions
About this article
Cite this article
Mason, O., Di Meo-Savoie, C., Van Nostrand, J. et al. Prokaryotic diversity, distribution, and insights into their role in biogeochemical cycling in marine basalts. ISME J 3, 231–242 (2009). https://doi.org/10.1038/ismej.2008.92
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2008.92