Abstract
Estimates of the number of species of bacteria per gram of soil vary between 2000 and 8.3 million (Gans et al., 2005; Schloss and Handelsman, 2006). The highest estimate suggests that the number may be so large as to be impractical to test by amplification and sequencing of the highly conserved 16S rRNA gene from soil DNA (Gans et al., 2005). Here we present the use of high throughput DNA pyrosequencing and statistical inference to assess bacterial diversity in four soils across a large transect of the western hemisphere. The number of bacterial 16S rRNA sequences obtained from each site varied from 26 140 to 53 533. The most abundant bacterial groups in all four soils were the Bacteroidetes, Betaproteobacteria and Alphaproteobacteria. Using three estimators of diversity, the maximum number of unique sequences (operational taxonomic units roughly corresponding to the species level) never exceeded 52 000 in these soils at the lowest level of dissimilarity. Furthermore, the bacterial diversity of the forest soil was phylum rich compared to the agricultural soils, which are species rich but phylum poor. The forest site also showed far less diversity of the Archaea with only 0.009% of all sequences from that site being from this group as opposed to 4%–12% of the sequences from the three agricultural sites. This work is the most comprehensive examination to date of bacterial diversity in soil and suggests that agricultural management of soil may significantly influence the diversity of bacteria and archaea.
Similar content being viewed by others
Introduction
Recent estimates of the number of species in a gram of soil have garnered much attention. Such estimates are of particular interest as soil is considered to harbor the most diverse populations of bacteria of any environment on earth. Even those soils with large particle sizes provide an enormous surface area for many bacteria. The number of potential spatial microhabitats within a small soil sample is extraordinary. Temporal variation in the physicochemical factors of soils such as moisture, temperature and nutrients within each of those spatial microhabitats further increases the diversity of niches available to support microbial populations.
In the first culture-independent analysis of a bacterial species census in soil, DNA was isolated from extracted cells and the number of bacterial genomes present was estimated in a mixed sample using DNA:DNA hybridization (Torsvik et al., 1990). Taking the admittedly risky approach of extrapolating such data from a mixture containing a few species to a complex mixture of DNA from soil, the number of bacterial species in a gram of boreal forest soil was estimated to be approximately 10 000 (Torsvik et al., 1990). Recently, this approach was reevaluated and nearly 107 microbial species per gram of soil were predicted (Gans et al., 2005). This estimate was thought to be too large to be practically verified with current DNA sequencing technology (Gans et al., 2005).
Others have attempted to estimate the number of bacterial species in a gram of soil by extrapolation based on amplified, cloned and sequenced 16S rRNA genes. Defining a species as 3% dissimilarity in the comparison of 16S rRNA gene sequences (Stackebrandt and Goebel, 1994) and analyzing two sequenced clone libraries from different sites, a recent estimate for the number of operational taxonomic units (OTUs) estimated in a gram of soil was between 2000 and 5000 (Schloss and Handelsman, 2006). These two libraries were derived from Alaskan and Minnesota soils and contained 1033 and 600 16S rRNA clones, respectively. In a separate study, a nonparametric approach in the analysis of 556 16S rRNA clones was used to estimate that the number of species per gram of marine sediment was between 2000 and 3000 (Hong et al., 2006).
None of the estimates to date have been based on a sufficient number of sequences to provide reasonable extrapolations. Here we used pyrosequencing to obtain well over 25 000 16S rRNA gene fragment sequences from each of four soils. This number is more than an order of magnitude higher than obtained previously. Soil DNA was isolated and 454 Life Sciences (Branford, CT, USA) pyrosequencing was employed following the amplification of the hypervariable V9 region of the highly conserved 16S rRNA gene. Highly conserved primers were used to amplify this hypervariable region using a low number of PCR cycles to minimize species structure distortion due to amplification. A statistical transformation of the resulting rarefaction curves and species richness indicators permitted estimates of the number of OTUs, as defined by specific levels of sequence identity, for each soil.
Another objective of this project was to determine how this estimate of microbial diversity might vary between soils across a broad geographical scale (Table 1). Soils from four sites across the western hemisphere were chosen and included three agricultural soils from a maize field in the southernmost state of Brazil, Rio Grande do Sul, a sugarcane field in the Everglades Agricultural Area in Florida, and a soil from the Morrow Plots at the University of Illinois in Urbana. The fourth soil was collected from a boreal forest site in northwestern Ontario, Canada.
Materials and methods
Soil sampling, DNA extraction, PCR and pyrosequencing
Soil was collected from the top 10 cm of the surface. For the forest soil, sphagnum and duff layers were removed before collection. All soil was packed on ice upon collection and transported to labs for extraction. DNA was isolated from at least 1 g of mixed soil using the FastDNA Kit (Qbiogene Inc., CA, USA) and purified on agarose gels (Moreira, 1998). To amplify a 16S rRNA gene fragment of the appropriate size and sequence variability for 454 pyrosequencing, primers 787f and a modification of 1492r were chosen (Baker et al., 2003). The 1492r and 786f primers were modified slightly to create 1492-rm (5′-GNTACCTTGTTACGACTT-3′) and 787f (5′-ATTAGATACCCNGGTAG-3′) to broaden the taxonomic range of 16S rRNA genes amplified.
To obtain sequences from the equivalent of a gram of each of the soils, we calculated the theoretical amount of bacterial DNA present. Assuming 1 × 109 cells/g of soil, an average bacterial genome size of 3 Mb and an average 16S rRNA copy number of one per cell, we estimated 1 g of soil to contain 3.2 μg of genomic bacterial DNA, and decided to use a total of 10 μg total community DNA as template. Approximately 20 PCR cycles were required to obtain 3 μg of amplified 16S rRNA gene product with 96 PCR reactions each having 100 ng of template DNA. A total of 3 μg of amplified 16S rRNA gene product from each soil was required to construct the four libraries for 454 sequencing. The PCR conditions used were 94°C for 2 min, 20 cycles of 94°C, 45 s denaturation; 55°C, 45 s annealing and 72°C, 1 min extension; followed by 72°C, 6 min. After 20 rounds of amplification, another 3 rounds of amplification was done to add the A and B adapters required for 454 pyrosequencing to specific ends of the amplified 16S rRNA fragment for library construction (Margulies et al., 2005). For this purpose, two new primers were synthesized where the A and biotinylated B adapter sequences were immediately upstream of the 1492-rm and 787f primer sequences, respectively. The resulting sequences were 5′-(CCATCTCATCCCTGCGTGTCCCATCTGTTCCCTCCCTGTCTCAG)GITACCTTGTTACGACTT-3′ and 5′-(BioTEG/CCTATCCCCTGTGTGCCTTGCCTATCCCCTGTTGCGTGTCTCAG)ATTAGATACCCIGGTAG-3′ with the A and B adapter sequence in parentheses, respectively. The resulting PCR product was used in the emPCR process necessary to make the single strands on beads as required for 454 pyrosequencing (Margulies et al., 2005).
Phylogenetic assignment, alignment and clustering of 16S rRNA gene fragments
The 16S rRNA gene fragments were phylogenetically assigned according to their best matches to sequences in the Nearest Alignment Space Termination (NAST, http://greengenes.lbl.gov/NAST) database (DeSantis et al., 2006). Multiple sequence alignment was done using MUSCLE (with parameter -maxiters 1, -diags1 and -sv) (Edgar, 2004). Based on the alignment, a distance matrix was constructed using DNAdist from PHYLIP version 3.6 with default parameters (Felsenstein, 1989, 2005). These pairwise distances served as input to DOTUR (Schloss and Handelsman, 2005) for clustering the sequences into OTUs of defined sequence similarity that ranged from 0% to 20% dissimilarity. These clusters served as OTUs for generating rarefaction curves and for making calculations with the richness and diversity indexes, Ace and Chao1, in DOTUR. These programs were run on an HP DL585 Proliant Server with 64 GB of RAM and 2 dual core Opteron processors at 1.8 GHz, running Linux OS. Of the 139 918 bacterial 16S rRNA sequences used in these analyses, 562 were shorter than 50 bp in length. As Genbank does not provide accession numbers for sequences of this length, these 562 sequences are provided in the supplementary material (Table S1).
Statistical analysis of OTU richness: rarefaction, Chao1 and Ace
At each dissimilarity level, a Michaelis–Menten (MM) equation was fit to the rarefaction curves generated by DOTUR. For example, for the Brazil data, the DOTUR output was obtained for sample sizes of 5000, 10 000, 15 000, 20 000, 25 000 and 26 140, sequences and 1%, 3%, 5%, 10% and 20% dissimilarities. The MM equation has parameters V and K with the equation y=Vx/(K+x), and the estimate of V is our estimate of the asymptote or maximum number of OTUs at a given % dissimilarity. The MM equation was not a good fit to the rarefaction curves, as two parameters were not able to sufficiently model the curvature. The resulting asymptote estimates were actually below the observed data by a substantial amount. The problem was that the curvature at the beginning of the rarefaction curve was different from that at the end. To solve this problem we used a model of two MM equations, one for the beginning and one for the end of the curve, that were joined continuously at a point that was optimized by the fit. The curves were fit by the method of maximum likelihood, using the transform-both-sides methods (Rupert et al., 1989). This approach better predicts the error terms and provides improved estimators over standard approaches such as Lineweaver–Burk. The result of this analysis was an estimate of the rarefaction asymptote for each dissimilarity level. These rarefaction asymptote estimates, and the ACE and Chao1 estimates, still varied with sample size. For each region, we have an estimate for each sample size. For example, for the Brazil data, we obtained separate rarefaction, ACE and Chao1 estimates for sample sizes of 500, 10 000, 15 000, 20 000, 25 000 and 26 140 sequences. In these estimate-sample size curves, MM models were again used to estimate the three overall asymptotic richness values. The bootstrap estimated the standard deviations for both V and K for each curve. Standard deviations were also determined for estimating proportions of V. For example, the necessary sample for getting pV species is sample size (p/(1–p))K with standard error (p/(1–p))Std(K). At low % dissimilarities, the estimates are quite variable, but similar while at high % dissimilarities, the estimates become remarkably similar.
Results
The numbers of species detected in a sample, or of the numbers of organisms discerned at any given phylogenetic level, are strongly affected by the number of sequences analyzed (Schloss and Handelsman, 2005). Estimates of OTUs increase with number of sequences and a plot of OTUs versus the 1 number of sequences yields a rarefaction curve that approaches a maximum (Figure 1). To estimate the true maximum value at any phylogenetic level it is necessary to model and extrapolate from the rarefaction curve, or to use nonparametric methods to estimate the true OTU richness by taking into account the population structure. The nonparametric methods provide estimates that also vary with sample size, so their approach to a richness maximum must also be modeled and extrapolated. The results obtained from three methods are presented and they converge.
To estimate the ability of the richness estimators used here to predict the number of species and genera in a sample, a collection of 2702 known 16S rRNA genes from 2410 species and 685 genera of bacteria from the Ribosomal Database Project II was aligned using the same section of the gene sequenced from the soil samples (Table 2). At 0% dissimilarity, the nonparametric ACE and Chao1 estimators overestimate the number of species while the rarefaction estimator underestimates number of species. At 5% dissimilarity, the rarefaction estimator accurately predicted the number of genera.
Small subunit rRNA gene fragments of 800 bp were amplified from an amount of DNA equivalent to that found in a gram of soil. This size is optimal for 454 pyrosequencing library construction. These fragments were amplified from DNA isolated from each of the soils. Over 149 000 sequences of an average of 103 nucleotides in length were obtained. The number of OTUs present in each sample was determined after defining an OTU at five levels of phylogenetic resolution or sequence similarity (Figures 2 and 3). At the highest level of resolution (0% dissimilarity), the maximum number of OTUs in any one soil with any of the three estimators used was just under 52 000 (Figure 3). Data for 0%, 3%, 5%, 10% and 20% are presented so that the reader can choose a level of discrimination of interest (Table S2).
The total number of OTUs obtained, and the maximum number estimated by three different methods are shown for five different phylogenetic levels (Tables S2 and S3). A supplementary table includes the standard deviations about these estimates. One of these estimates is parametric and based on rarefaction curves while two of these are based on nonparametric estimates, Chao1 and ACE (Schloss and Handelsman, 2006). The estimates improve as the definition of an OTU declines in resolution from species, through genera, to something approaching the level of phyla. These data suggest that the original DNA reassociation estimates of 2000–10 000 species per gram were underestimates (Schloss and Handelsman, 2006; Torsvik et al., 1990), but these estimates do not approach the higher maximum proposed recently (Gans et al., 2005).
Using these OTU richness estimates, the number of sequences required to reach 90% and 95% of the maximum number of OTUs at each % dissimilarity are shown (Table S3). As a result, we disagree with the statement that ‘the survey size required for accurate analysis of soil communities is impractically large’ (Gans et al., 2005). In fact, with the latest improvement in throughput in 454 pyrosequencing, this survey can be completed in less than 1 day of operation of the Roche Genome Sequencer FLX system. The maximum number of sequences required to identify 90% of the 52 000 OTUs is less than 713 000. Thus, using the methods here, 95% of all OTUs can be identified with over 10-fold fewer sequences than the number of species suggested recently (Gans et al., 2005). Further, we present a number of new observations, beyond the estimates of OTU richness.
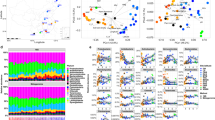
With over 40% of the total bacterial sequences, the Proteobacteria represented the dominant phylum in each soil (Figure 4). The Betaproteobacteria were the dominant class among the Proteobacteria in all soils except Brazil where the Gammaproteobacteria were dominant. With 15%–25% of the bacterial sequences in each sample, the second most abundant phylum in all four soils was the Bacteriodetes. Other prominent phyla were the Acidobacteria, the Actinobacteria and the Firmicutes. Among the other phyla, the Gemmatimonadetes represented 3.5% of the sequences in the Canadian sample while in Florida and Brazil, the Nitrospira represented 3% and 2% of total sequences, respectively. In Illinois and Canada, the Verrucomicrobia were represented by more than 2% of the sequences. In Illinois, nearly 4% of the sequences were in the TM7 phylum. Approximately 6%–12% of the sequences from each sample remained unclassified. Many unusual bacterial sequences were found that were at least 10% dissimilar to all previously described groups (Figure S1).
Relative abundance of phyla and proteobacterial classes for each soil library, in which 16S sequences were classified according to the nearest neighbor in the Greengenes database (http://greengenes.lbl.gov).
The primers used in this work also amplify the 16S rRNA gene from the Archaea (Table 1, Figure S2). In the agricultural soils, the Crenarchaeota represented a fairly significant proportion, 4.6%–12.5%, of the total sequences. Of the Crenarchaeota observed from each of the agricultural soils, about one-third of the sequences are closely related to the ammonia oxidizing archaea. While 16S rRNA gene sequence alone cannot describe the physiology of an organism, it is not unreasonable to expect a high number of ammonia oxidizing archaea in these soils given recent work (Treusch et al., 2005; Leininger et al., 2006). The surprising result was the relative dearth of crenarchaeotal sequences in the Canadian sample with just five of 53 251 sequences being in this group. These five sequences were all related to the ammonia oxidizing Archaea and the low number of these sequences may be attributable to the low pH of this soil. However, the Brazilian soil had an even lower pH and yet over 4% of the total sequences from Brazil were crenarchaeotal. Further work is needed to confirm this observation of a low proportion of crenarchaeotal sequences in forest soils. The cause of this may be that nitrogen is more limiting in forest soils compared to fertilized agricultural soils. The low number of crenarchaeotal sequences in Canada is not substituted by ammonia oxidizing bacterial sequences. Very few sequences from the genera responsible for ammonia oxidation in bacteria were found in each soil (Table 3).
The patterns of the rarefaction curves for the Canadian forest sample are very different from those of the agricultural soils (Figures 2 and 3). In contrast to the situation with archaeal diversity, bacterial diversity is much higher in the forest sample compared to the others. In the Canadian sample, OTU abundance is particularly high at high levels of dissimilarity while in the other samples the diversity is low at high levels of dissimilarity. Conversely at low levels of dissimilarity the OTU abundance of the Canadian sample is lower that that of the other samples. This result is interpreted as the boreal forest sample being very phylum rich while the agricultural samples are phylum poor.
Discussion
In this paper, three significant findings are presented. First, the number of OTUs in each soil is estimated with the largest 16S rRNA libraries sequenced to date. Although biases are likely present in this work at the steps of DNA extraction and PCR amplification, these biases are unlikely to have missed so many taxa that our estimate of the number of OTUs could be incorrect by over 100-fold. That is, if the number of species in a gram of soil is 8.3 million, our methods would have to have missed over 99.5% of the OTUs in soil. Given the expected biases and limitations of our methods, that is unlikely.
The second important finding is the discovery that the forest soil is very phylum rich compared to the three agricultural soils. The reasons for this are unknown but are worthy of continued study, particularly to determine whether this result is generally observed in forest soils or otherwise undisturbed soils. Perhaps the average temperature of the soil is low enough to prevent rapid cell division and slows the process of speciation although recent work showed no correlation between mean average soil temperature and bacterial diversity (Fierer and Jackson, 2006). However, other work supports such a trend (Neufeld and Mohn, 2005). Undisturbed sites, relative to the agricultural areas, may have a higher diversity of flora and fauna and that positively impacts microbial diversity although recent work suggests that the inverse may be true (Fierer and Jackson, 2006). The relatively low pH of the Canadian soil would suggest that this soil may have a lower microbial diversity relative to soils with higher pH (Fierer and Jackson, 2006) but the Brazilian soil has an even lower pH and its diversity is much lower at the phylum level. However, the trend observed between phylotype richness and pH by Fierer and Jackson (2006) cannot be applied to predict the diversity of individual soils but does appear to be a trend when many soils are sampled.
However, comparing this work to previous analyses is complicated since previous studies have used methods with a very different level of resolution such as terminal restriction fragment length polymorphism (Fierer and Jackson, 2006) or have sequenced far fewer 16S rRNA fragments (Neufeld and Mohn, 2005). Also, each study assesses diversity at different sites. To date, there has been no systematic analysis of the effect of disturbance on soil microbial diversity by examining many thousands of 16S rRNA fragments. The three agricultural sites are physically, chemically and geographically very different from each other. They vary greatly in soil texture, pH, soil type and the agricultural system applied to the soil. Nevertheless, their phylum richness is very low at each site compared to the forest soil. More such comparisons are needed at more sites to determine whether forest soils are generally more diverse than agricultural soils.
Worldwide, the total population of bacteria in soil seems to be limited to 1 billion cells per gram, which suggests a limit on the niche space that bacteria can occupy. Without major population losses and new habitats to move into, speciation might be slow, as it is for macrospecies in continental systems where niches are filled. In agricultural systems, with low vegetative diversity and high inputs of xenobiotics, overall species diversity may be reduced to a bottleneck, from which species diversification is possible, albeit from a limited number of phyla. The boreal forest sample may have higher phylum diversity because forest soils have archived a high diversity of ancient populations both through the low diversity of organic inputs and the long periods of inactivity during cold periods. More analyses are required of disturbed and undisturbed sites to begin to resolve the causes of the soil diversity differences observed here.
The third finding of this work is the astonishing low diversity and presumably abundance of archaea at the forest site. As with the differences in phylum richness, the cause of the much higher archaeal diversity in agricultural soils is unknown. Any of the factors discussed above may be relevant but the importance of nitrogen cycling in agricultural soils may play a key role. The discovery of the abundance and activity of the ammonia oxidizing archaea in agricultural soils may be important here (Treusch et al., 2005; Leininger et al., 2006). Agricultural soils are commonly fertilized with N sources such as anhydrous ammonia or urea. These N sources are often quickly oxidized to nitrate in soil. In an environment where fertilization is common, the number and diversity of archaea may be expected to be very high. The site that has been in use for agriculture for the longest time may be the Morrow Plots at the University of Illinois where the experimental plots began in 1876 with agriculture initiated on the site perhaps 40 years earlier (Aref and Wander, 1998). Agricultural experimentation at the Florida site in the Everglades began in 1923 (Phillips, 1985). The agricultural plots in Brazil were initiated in 1962. The longer a site has been cultivated, the higher the proportion of sequences that are archaea. More experiments are needed to assess archaeal abundance in relation to agriculture and N fertilization but the data here are intriguing and encourage further work.
The age of inexpensive, high throughput pyrosequencing allows the assessment of the full taxonomic diversity of bacteria in soil for the first time. As a result, much better estimates of OTU richness can be obtained from any sample. A significant debate exists in the literature regarding the species concept for bacteria. No judgment is made on this here as no attempt is made to define a bacterial species. However, it is clear that regardless of the 16S rRNA percent dissimilarity chosen by the reader to define a particular level of taxonomic discrimination, the number of unique sequences in a gram of soil by any definition cannot be as high as 8.3 million per gram (Gans et al., 2005). But beyond estimating the number of OTUs in a soil, large-scale sequencing of 16S rRNA in multiple soils can lead to intriguing observations that encourage future analyses.
Accession codes
Accessions
GenBank/EMBL/DDBJ
References
Aref S, Wander MM . (1998). Long-term trends of corn yield and soil organic matter in different crop sequences and soil fertility treatments on the Morrow Plots. Adv Agron 62: 153–202.
Baker GC, Smith JJ, Cowan DA . (2003). Review and re-analysis of domain-specific 16S primers. J Microbiol Methods 55: 541–555.
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Keller K, Huber T et al. (2006). Greengenes, a chimera-checked 16S rRNA database and workbench compatible with ARB. Appl Environ Microbiol 72: 5069–5072.
Edgar RC . (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32: 1792–1797.
Felsenstein J . (1989). PHYLIP—Phylogeny Inference Package (Version 3.2). Cladistics 5: 164–166.
Felsenstein J . (2005). PHYLIP (Phylogeny Inference Package) Version 3.6. Distributed by the author. Department of Genome Sciences, University of Washington: Seattle.
Fierer N, Jackson RB . (2006). The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci (USA) 103: 626–631.
Gans J, Woilinsky M, Dunbar J . (2005). Computational improvements reveal great bacterial diversity and high metal toxicity in soil. Science 309: 1387–1390.
Hong S-H, Bunge J, Jeon S-O, Epstein SS . (2006). Predicting microbial species richness. Proc Natl Acad Sci USA 103: 117–122.
Leininger S, Urich T, Schloter M, Schwark L, Qi J, Nicol GW et al. (2006). Archaea predominate among ammonia-oxidizing prokaryotes in soils. Nature 442: 806–809.
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA et al. (2005). Genome sequencing in microfabricated high-density picolitre reactors. Nature 437: 376–380.
Moreira D . (1998). Efficient removal of PCR inhibitors using agarose-embedded DNA preparations. Nucleic Acids Res 26: 3309–3310.
Neufeld JD, Mohn WW . (2005). Unexpectedly high bacterial diversity in arctic tundra relative to boreal forest soils, revealed by serial analysis of ribosomal sequence tags. Appl Environ Microbiol 71: 5710–5718.
Phillips N . (1985). The Everglades experience: sixty years of progress. EREC Research Report EV-1985-5. Belle Glade.
Rupert D, Cressie N, Carroll RJ . (1989). A transformation/weighting model for estimating Michaelis–Menton parameters. Biometrics 45: 637–656.
Schloss PD, Handelsman J . (2005). Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Appl Environ Microbiol 71: 1501–1506.
Schloss PD, Handelsman J . (2006). Toward a census of bacteria in soil. PLOS Computational Biology 2: 786–793.
Stackebrandt E, Goebel BM . (1994). Taxonomic note: a lace for DNA–DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Internat J Syst Bacteriol 44: 846–849.
Torsvik V, Goksoyr J, Daae FL . (1990). High diversity in DNA of soil bacteria. Appl Environ Microbiol 56: 782–787.
Treusch AH et al. (2005). Novel genes for nitrite reductase and Amo-related proteins indicate a role of uncultivated mesophilic crenarchaeota in nitrogen cycling. Environ Microbiol 7: 1985–1995.
Acknowledgements
EWT acknowledges support from the Florida Agricultural Experiment Station, the National Science Foundation (MCB-0454030) and the United States Department of Agriculture (National Research Initiative Competitive Grant 2005-35319-16300). GC was supported by the National Science Foundation (DEB-0540745). RRF received support from National Science and Engineering Research Council Discovery Grant. LFWR and FAOC were supported by the Brazilian Ministry of Education (CAPES) for a scholarship concession, the Estate Research Foundation (FAPERGS) and the Brazilian National Research Council (CNPq). We thank the people of the North Caribou Lake First Nation for guidance and permission to sample soil in their traditional territory in Ontario, Canada. We also thank Jennifer Drew for suggestions on the manuscript before submission.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej).
Rights and permissions
About this article
Cite this article
Roesch, L., Fulthorpe, R., Riva, A. et al. Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1, 283–290 (2007). https://doi.org/10.1038/ismej.2007.53
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2007.53
Keywords
This article is cited by
-
Different microbial communities in paddy soils under organic and nonorganic farming
Brazilian Journal of Microbiology (2024)
-
Tree and shrub richness modifies subtropical tree productivity by regulating the diversity and community composition of soil bacteria and archaea
Microbiome (2023)
-
Exploring soil bacterial diversity in different micro-vegetational habitats of Dachigam National Park in North-western Himalaya
Scientific Reports (2023)
-
Bacterial type VI secretion system (T6SS): an evolved molecular weapon with diverse functionality
Biotechnology Letters (2023)
-
A Matter of Metals: Copper but Not Cadmium Affects the Microbial Alpha-Diversity of Soils and Sediments — a Meta-analysis
Microbial Ecology (2023)