Abstract
Symbiont choice has been proposed to play an important role in shaping many symbiotic relationships, including the fungus-growing ant–microbe mutualism. Over millions of years, fungus-growing ants have defended their fungus gardens from specialized parasites with antibiotics produced by an actinomycete bacterial mutualist (genus Pseudonocardia). Despite the potential of being infected by phylogenetically diverse strains of parasites, each ant colony maintains only a single Pseudonocardia symbiont strain, which is primarily vertically transmitted between colonies by the founding queens. In this study, we show that Acromyrmex leaf-cutter ants are able to differentiate between their native actinomycete strain and a variety of foreign strains isolated from sympatric and allopatric Acromyrmex species, in addition to strains originating from other fungus-growing ant genera. The recognition mechanism is sufficiently sensitive for the ants to discriminate between closely related symbiont strains. Our findings suggest that symbiont recognition may play a crucial role in the fungus-growing ant–bacterium mutualism, likely allowing the ants to retain ecological flexibility necessary for defending their garden from diverse parasites and, at the same time, resolve potential conflict that can arise from rearing competing symbiont strains.
Similar content being viewed by others
Introduction
Fungus-growing ants (tribe Attini) obligately depend on fungi they cultivate for food (Weber, 1972), and this mutualistic association is believed to have originated more than 50–65 million years ago in the Amazon Basin (Mueller et al., 2001). In addition to providing substrate for its growth, the ants weed and groom their fungus garden to remove competing microbes or pathogens (Quinlan and Cherrett, 1977; Currie and Stuart, 2001), especially Escovopsis – a specialized microparasite capable of rapidly devastating the entire fungus garden (Currie et al., 1999a; Currie and Stuart, 2001). Besides engaging in specific behaviors to defend their fungus gardens, the ants have a mutualism with bacteria that produce antibiotics with apparent specific activity against Escovopsis (Currie et al., 1999b, 2003a). These bacteria, in the genus Pseudonocardia (Cafaro and Currie, 2005), are typically housed on the ants’ cuticles in elaborate structures linked to exocrine glands (Currie et al., 2006). Molecular phylogenetic studies of the parasite Escovopsis (Currie et al., 2003b; Gerardo et al., 2004) and the distribution of specialized structures for culturing the filamentous bacteria (Currie et al., 2006) suggest that fungus-growing ants have employed Pseudonocardia symbionts to deal with the garden parasite from the earliest stages of the ant–fungus mutualism (Poulsen and Currie, 2006).
Like the fungal cultivar, the antibiotic-producing bacterial mutualist is vertically transmitted from parent to offspring nests by colony-founding queens (Currie et al., 1999a). As predicted by this mode of transmission, phylogenetic comparisons between Pseudonocardia and fungus-growing ants reveal some degree of broad-scale coevolution between the mutualists (Cafaro and Currie, 2005). However, strict cocladogenesis between the ants and the bacteria is disrupted, especially at the finer phylogenetic levels, by Pseudonocardia strains switching between attine ant species, both within and between genera (Poulsen et al., 2005; Cafaro et al., unpublished data). For example, Poulsen et al. (2005) found evidence for frequent Pseudonocardia switches between sympatric ant species within the genus Acromyrmex, indicating that horizontal symbiont transmission occurs and may be common.
In contrast to the primarily vertically transmitted cultivar and Pseudonocardia symbionts, the garden parasite Escovopsis is horizontally transmitted (Currie et al., 1999a). As predicted by this mode of transmission, the gardens of fungus-growing ants can be infected with phylogenetically diverse pathogen strains (Gerardo et al., 2006; Taerum et al., in press). Furthermore, bioassay and in vivo infection experiments indicate differences in the inhibitory capabilities of different ant-associated Pseudonocardia strains, as well as variation in Escovopsis strain susceptibility to the different antibiotics (Poulsen et al., unpublished data). Thus, to deal with the phylogenetically diverse strains of Escovopsis, fungus-growing ants would be expected to benefit from: (1) maintaining multiple strains of the Pseudonocardia mutualist and/or (2) having the flexibility to acquire novel strains of antibiotic-producing bacteria when their gardens are infected with strains of Escovopsis that are relatively resistant to the antibiotics produced by their resident Pseudonocardia strain. Only a single strain of Pseudonocardia has been isolated within a single ant colony (Poulsen et al., 2005), although additional symbiont strains have the potential to enter nests when ants from other colonies enter, through forage material added to the garden, if foraging workers come in contact with strains outside the nest, and/or via fungus-growing ant colony-associated invertebrates that could act as vectors (Currie et al., 1999a). Maintenance of a single symbiont strain fits current evolutionary theory, which predicts that mixing of Pseudonocardia strains may result in competition and conflict between strains, at a cost to the ants (Frank, 1996, 2003; Poulsen et al., submitted). On the other hand, a potential disadvantage of having a single Pseudonocardia strain is the limited ecological flexibility in defending the fungus garden against infections by diverse Escovopsis strains.
What maintains single-clone rearing in the binary association between fungus-growing ants and their Pseudonocardia symbiont is unknown. Secondary metabolites produced by the resident symbiont strain are capable of inhibiting other actinomycete strains and hence may preclude the rearing of multiple strains within individual nests (Poulsen et al., submitted). In addition, given that fungus-growing ants have been shown to exert symbiont choice by behaviorally selecting certain cultivar strains over others (Bot et al., 2001; Mueller et al., 2004), it is also possible that fungus-growing ants can recognize and differentiate between native and nonnative Pseudonocardia strains, allowing active removal of potential competitors of the resident strain. Since the ants incur a significant metabolic cost to support the growth of Pseudonocardia symbiont (Poulsen et al., 2003b; Currie et al., 2006), and the energy consumed by the bacteria for the production of antibiotics against Escovopsis may be diverted to competition when strains mix, maintaining a single Pseudonocardia strain would likely reduce the energetic costs associated with competition, reducing the cost–benefit ratio of the association. At the same time, the ability to distinguish between different Pseudonocardia strains would allow ants to acquire novel strains when confronted with infections by a resistant strain of Escovopsis parasite.
In this study, we conduct behavioral experiments to test the hypothesis that Acromyrmex ants are able to differentiate between their native and a variety of foreign Pseudonocardia strains. This is done by presenting individual ants with a choice between two fragments of their native fungal cultivar, one containing their native Pseudonocardia strain and one containing a nonnative strain. In addition, by sequencing bacterial strains used in these choice experiments, we were able to determine if the ants’ ability to discriminate between strains depends on genetic differences between the bacterial symbionts.
Materials and methods
Isolation and phylogenetic placement of Pseudonocardia strains
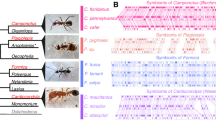
Before selecting the strains for use in the choice assays, we isolated Pseudonocardia symbionts from six colonies, representing three genera of fungus-growing ants (Atta and Acromyrmex leaf-cutting ants and Trachymyrmex higher attine non-leaf-cutting ants; Schultz and Meier, 1995). Pseudonocardia symbionts were isolated by crushing one ant in 500 μl of autoclaved milliQ water and plating the mixture on chitin agar (Cafaro and Currie, 2005; Poulsen et al., 2005). White colonies with morphologies consistent with Pseudonocardia were subcultured until pure colonies were obtained. DNA isolation was performed from pure cultures using a standard cetyltrimethylammonium bromide (CTAB) extraction protocol, and partial ribosomal 16S rDNA and nuclear EF-Tu sequences were obtained using standard PCR and sequencing techniques (Cafaro and Currie, 2005; Poulsen et al., 2005). Positive bands after PCR were sequenced at the University of Wisconsin-Madison Biotechnology Center (http://www.biotech.wisc.edu), corrected in Sequencher 4.6 for Windows (Gene Codes Corporation, Ann Arbor, MI, USA), and aligned using Clustal X 1.83 (Thompson et al., 1997) and MacClade 4.07 OS X (Maddison and Maddison, 2005). Sequences from three free-living actinomycetes (Pseudonocardia saturnae, P. thermophila and Streptomyces griseus) were generated and included in the phylogenetic analyses. The analyses were performed in PAUP* 4.0b 10 (Swofford, 2002) and bootstrap support was assessed with 1000 pseudoreplicates under maximum parsimony (MP), maximum likelihood (ML) and neighbor-joining (NJ) conditions. Before performing the ML analysis, the model of sequence evolution was determined in Modeltest 3.7 (Posada and Crandall, 1998) to be a general time-reversible (GTR) model with among-site substitution rate variation (R(A-C)=0.7393, R(A-G)=1.4226, R(A-T)=0.6941, R(C-G)=1.5957, R(C-T)=3.1495 and R(G-T)=1.0000), with the proportion of invariable sites (I) being 0.4913, and with a gamma distribution shape parameter G of 0.8059. These phylogenetic analyses allowed us to place actinomycetes from Acromyrmex colonies within either of the two clades known to associate with this ant genus (Poulsen et al., 2005; Clades I and II of Figure 1), in addition to providing information on placement of Pseudonocardia strains from the two other ant genera (Figure 1). All sequences have been submitted to GenBank (accession numbers EF588205–EF588262).
Phylogenetic placement of the strains used for the choice experiments. The figure shows a neighbor-joining (NJ) phylogeny of Pseudonocardia bacteria associated with fungus-growing ants based on partial sequences of ribosomal 16S rDNA and nuclear EF-Tu genes. Three free-living actinomycetes are included, two in the genus Pseudonocardia and one (outgroup) actinomycete from the genus Streptomyces. Ant-associated Pseudonocardia strains are labeled after the ants they were isolated from; ant colony codes, genus and species names are given; and arrows indicate the placement of the symbiont strains used in the choice essays. Bootstrap support values after 1000 pseudoreplicates under MP (maximum parsimony; top, left), ML (maximum likelihood; top, right), and NJ (bottom) conditions are given for the branches separating the major clades in the phylogeny.
Ant–actinomycete choice experiments
Based on the phylogenetic placement of the isolated Pseudonocardia symbionts, four Acromyrmex ant nests were selected for the behavioral experiments, with two from each of the clades of Pseudonocardia associated with this ant genus (A. octospinosus CC031210-22 and A. echinator CC031212-01 collected in Gamboa, Panama, Clade I; and A. laticeps UGM030330-04 and A. niger C030327-02 from Argentina, Clade II). All colonies were maintained at the University of Wisconsin-Madison in plastic chambers placed on islands surrounded by mineral oil to prevent transfer of microbes between nests by mites. Three times per week, colonies were provided with maple or oak leaves in an external plastic chamber connected to the main chamber. The colonies were watered once a week and dump chambers cleaned every 2 weeks.
In addition to the Acromyrmex nests, other Pseudonocardia symbionts included in the choice assays were isolated from an Atta colombica nest (CC031208-10, occurring within one of the clades of actinomycetes associated with Acromyrmex), and a Trachymyrmex zeteki nest (AL040410-06, in Clade III of Pseudonocardia associated with fungus-growing ants; see Figure 1). Both of these colonies were collected from the canal zone of Panama. Pure cultures of all Pseudonocardia were maintained on Yeast Malt Extract Agar (YMEA: 4 g yeast extract, 10 g malt extract, 4 g dextrose and 20 g agar in 1 l water) before the choice experiments were performed.
On the day of testing, fungus garden samples were collected from the parent nest and all the ants present in garden material were removed. Garden material was provided in all assays to mimic natural colony conditions, with both garden material and ants originating from the same parent colony. Approximately 100–150 mg of garden material was placed at two predetermined locations in a sterile 100 × 15 mm Petri dish (Figure 2). Pseudonocardia inoculum was prepared by scraping ∼100–150 mg of pure bacterial biomass from YMEA, the bacteria were suspended in 20 ml of autoclaved milliQ water in a 50 ml Falcon tube and the tube was shaken vigorously using a vortex to obtain a homogenous suspension of bacterial cells. Each of the two pieces of fungus garden material was then inoculated with 100 μl Pseudonocardia suspension, one garden with the native strain and one garden with a foreign strain. Shortly after application, before any potential effects on the fungus fragment of the inoculation of bacterial suspensions, an ant was randomly selected, regardless of caste, from the corresponding parent colony and added to the dish at equal distance from both garden fragments. The Petri dish was covered during the assay to prevent ant escape and was randomly turned between trials and replicates, so that any potential bias from its placement was avoided. Symbiont choice was scored when the ant groomed a cultivar-Pseudonocardia mixture with its mouthparts. This method of scoring was straightforward because it involved the ant moving to one fragment and subsequently spending a considerable amount of time (at least 1 min) grooming the garden piece. The trials were not performed blindly, however, scoring involved such a clear choice that it is very unlikely that it was affected by observer bias. For each choice combination, a total of 10 replicate Petri dishes were prepared and only 10 ants were tested per dish to reduce any possible effects of trail pheromones. Furthermore, should pheromone trails be important, we would have expected an effect also in control experiments (Experiment 1; Table 1), and since this was not observed, such effects appear unlikely. One hundred different ants, randomly picked from the nest, were used to ensure statistical independence between the replicates.
Experimental setup for the choice experiments. (a) The picture shows a newly excavated 1-year-old colony of A. echinator in which a major worker (center of the picture) is completely covered with the mutualistic actinomycete bacterium (photo, courtesy of David R Nash). (b) The experimental setup of the Pseudonocardia symbiont choice assay, where native cultivar fragments from the colony of the ant tested were placed at both ends of the Petri dish setup and aqueous suspensions of Pseudonocardia strains were subsequently added to the cultivar: one of the cultivar fragments (top of picture) received a native while the other received a foreign strain (bottom of picture). The ant tested was placed anywhere along the dotted line, at equal distance to either of the fungus fragments.
Table 1 lists the different choice combinations for each experiment. All four Acromyrmex nests were tested in each experiment to determine the ability to discriminate between symbiont strains. Experiment 1 represented the controls for the Pseudonocardia choice assays: the ants were given the choice between two cultivar samples both containing their native Pseudonocardia symbiont. Experiment 2 evaluated the ability of the ants to differentiate between native and foreign Pseudonocardia symbiont, by providing the native symbiont and a symbiont from the T. zeteki nest (AL040410-06), which is relatively distantly related to all of the symbionts reared by the four Acromyrmex colonies (Clade III of Figure 1). We expected this combination to elicit a strong behavioral response from the ants if recognition was dependent on phylogenetic placement of the actinomycete strain. Experiments 3 and 4 examined how genetic distance between the Pseudonocardia strains affected the ants’ ability to choose between closely related bacterial strains; thus, providing information on the sensitivity of symbiont recognition in Acromyrmex. In experiment 3, the foreign symbiont used was from the A. colombica colony (CC031208-10), which was genetically more similar to Acromyrmex symbionts than the Trachymyrmex symbionts (Clade II of Figure 1). Experiment 4 investigated the ants’ ability to differentiate between Acromyrmex symbionts originating from the same phylogenetic clade as symbionts reared by the ants themselves (that is, both from within Clade I or both from within Clade II of Figure 1). In this experiment, ants from all four Acromyrmex colonies were presented with a choice between their native symbiont and one from the same phylogenetic clade (Table 1; Figure 1).
Statistical analyses
Ant preference between native and foreign Pseudonocardia symbionts was analyzed using two-sided paired t-tests. Within each of the 10 replicates, the difference was defined as the number of ants choosing the native symbiont over the number choosing the foreign symbiont. In these tests, the null hypotheses were that there was no difference between the numbers of ants choosing the native vs the foreign symbiont. For experiments 2–4 (choice experiments between native and nonnative strains), analysis of variance (ANOVA) tests were performed to determine if results between nests were significantly different. Pooling of the samples for second paired t-test analyses were justified since no significance was observed in the ANOVA comparisons.
Results
Table 1 summarizes the results of the symbiont choice assays. Experiment 1, with ants being presented with two samples of their fungus garden that were both inoculated with their native Pseudonocardia symbiont, revealed no clear preference for either sample (P-values of 0.8225, 0.5569, 1.000 and 0.7049 for the four colonies, respectively; see Table 1). In experiment 2, which tested the ability of the ants to differentiate between their native Pseudonocardia symbiont and the isolate from the T. zeteki nest, we found significantly more ants from two nests chose their native symbiont over the T. zeteki symbiont (A. laticeps UGM030330-04, P=0.0438; A. octospinosus CC031210-22, P=0.0205). Even though marginally significant effects were detected for A. niger CC0303027-02 (P=0.0571) and A. echinator CC031212-01 (P=0.0510), pooling the results from all four colonies was justified because the ANOVA did not reveal any significant nest-dependent effect (P=0.3089). The pooled data uncovered a significant preference for the native symbiont by all four Acromyrmex ant colonies (P=1.40E−05; Table 1).
In experiment 3, ants from all four nests tested were able to distinguish between their native Pseudonocardia symbiont and that of an A. colombica nest (P-values of 0.0138, 0.0138, 0.0399 and 0.0330), including even in the two cases where the resident strain and the Atta strain were closely related (Table 1; Figure 1). After eliminating the possibility of significant differences between the nests (ANOVA; P=0.7467), analyzing the pooled results with paired t-test yielded the same overall result (P=1.62E−06; Table 1).
In experiment 4, ants from three colonies were found to differentiate between their native bacterial mutualist and that of a closely related Pseudonocardia strain isolated from another colony of Acromyrmex and occurring within the same phylogenetic clade as their own symbiont (P=0.0368, 0.0004, 0.0051 for the three colonies, respectively; Table 1). The one exception involved the colony of A. echinator CC031212-01, which did not have a statistically significant preference (P=0.0947; Figure 1; Table 1). Pooling the samples (ANOVA; P=0.9803) nevertheless supported the overall finding that colonies of Acromyrmex can distinguish between closely related Pseudonocardia symbionts originating from the same phylogenetic clade as their own (P=1.65E−07; Figure 1; Table 1).
Discussion
When provided with a choice between fungus garden material inoculated with Pseudonocardia bacteria from either their own or another colony, Acromyrmex ants from all four colonies preferentially chose their native strain. Since one ant colony for each of four Acromyrmex species was tested, recognition appears to be consistent across this ant genus, and does not appear to depend on the relatedness of the symbiont strains. The only exception involved the colony A. echinator CC031212-01, where ants did not make a clear choice when presented with their native and a symbiont from A. octospinosus CC031210-22 (Table 1; P=0.0947). Although this could potentially be explained by the high degree of genetic similarity found between the two Pseudonocardia isolates tested (Figure 1), the reverse combination revealed a significant preferential choice (P=0.0051). While this finding indicates that A. echinator workers may be less choosy than other Acromyrmex species, more data is needed to determine if this is the case. Should it prove valid, it would indicate a general characteristic of A. echinator ant-symbiont specificity, because ants of this species also are less choosy of their fungus cultivar than sympatric A. octospinosus colonies (Bot et al., 2001).
Acromyrmex ant colonies have complex social structures with workers engaging in task partitioning, which is believed to increase the efficiency of the worker force and improve nest functioning. Previous studies have revealed that the distribution and abundance of Pseudonocardia on individual Acromyrmex workers is integrated into the colony social structure, depending both on worker size and age caste (Poulsen et al., 2002; Currie et al., 2003a; Poulsen et al., 2003a). Pseudonocardia is most abundant on youngest garden-tending workers as well as on major workers predominantly found in the bottom portions of gardens where Escovopsis is most prevalent (Currie et al., 2003a; Poulsen et al., 2002). Furthermore, actinomycete abundance can be upregulated in response to Escovopsis infection (Currie et al., 2003a). The ability of ants to recognize their Pseudonocardia strain likely influences these functional aspects of the association. Similarly, recognition likely plays an important role in the maintenance of the ant–bacterium association through the transmission of bacterial inocula from adults to newly emerged workers within colonies (Poulsen et al., 2003b) and by prospective queens between ant generations (Currie et al., 1999b).
The mechanism of Pseudonocardia recognition likely involves specific chemical profiles of the bacteria that allow for precise recognition even between very closely related strains (Figure 1; Table 1). This would mirror the ant–fungus mutualism, where ants can differentiate between different cultivar strains (Bot et al., 2001; Viana et al., 2001), presumably by recognizing clone-specific chemical signatures (Richard et al., 2007). Furthermore, both the forage that the ants use to manure their cultivar (Richard et al., 2004), and the chemical signatures of the fungal cultivar (Richard et al., 2007), affect Acromyrmex workers so that ants from different colonies adapted to the same forage or cultivar clone exhibit reduced aggression when introduced. While it is unknown whether Pseudonocardia affects the ant nest-mate recognition, the location of Pseudonocardia on the ant cuticle (Currie et al., 2006), and the ability for very precise ant–actinomycete recognition, make it conceivable that the bacterial symbiont also plays a role in this aspect of the symbiosis.
While the presence of multiple strains of antibiotic-producing bacterial strains could potentially increase the diversity of compounds produced to defend colonies against Escovopsis (Poulsen et al., unpublished data), antagonistic reactions between genetically different strains (Poulsen et al., submitted) are predicted to decrease the benefit of the ant–bacterium association (cf. Frank, 1996, 2003). Current evidence indicates that a single Pseudonocardia strain is reared within individual ant colonies (Poulsen et al., 2005), mirroring the ant–cultivar association where ant removal of competing cultivar strains facilitates single-clone rearing within individual colonies (Bot et al., 2001; Poulsen and Boomsma, 2005). Discrimination between native and foreign Pseudonocardia symbionts thus allows the ants to monitor and manipulate symbiont diversity within colonies and avoid competition between symbiont strains. This would further enable ants to distinguish beneficial from cheating Pseudonocardia strains as well as parasites and competitors that may enter ant colonies (for example, Escovopsis: Currie and Stuart, 2001; entomopathogenic fungi: for example, Hughes et al., 2004).
Although single-strain rearing (Poulsen et al., 2005) and vertical Pseudonocardia transmission (Currie et al., 1999b) may impair counter-adaptation to the continuously evolving and horizontally transmitted Escovopsis parasites, the presence of Pseudonocardia symbionts in all attine ant genera studied (Currie et al., 2006) suggests that the bacterial symbionts, nevertheless, are crucial in the defense against Escovopsis (cf. Poulsen and Currie, 2006). However, unlike the ant–fungus–Escovopsis association, which is characterized by broad-scale coevolution with cocladogenesis between four main groups of each of the three associates (Currie et al., 2003b), the ant–bacterium association displays far more frequent events of horizontal transfers both within and between these broad groupings (Cafaro et al., unpublished data; Figure 1). Actinomycete recognition may be a necessary prerequisite for such acquisitions, which likely allow the ants to obtain a new symbiont strain after loss of their residing strain and/or to obtain strains that produce novel antibiotics against Escovopsis, potentially facilitating rapid changes in the defense against parasitism. Our finding that 30–41% of workers presented with native and nonnative Pseudonocardia strains chose the nonnative strain suggests that the ants may have some flexibility to select nonnative strain of Pseudonocardia strains. Such flexibility may help explain the observed frequency of switches occurring over evolutionary time (Figure 1; Poulsen et al., 2005). If switching was very rare, we would expect precise Pseudonocardia recognition and much higher selection of native bacteria symbiont strains. Our findings, in combination with the fact that Pseudonocardia strains vary in their inhibitory capabilities against Escovopsis (Poulsen et al., unpublished data), suggests that ant colonies infected with Escovopsis should choose the most potent Pseudonocardia strain available.
Symbiont choice has repeatedly been proposed to be important in mutualistic associations, and may function as either a choice of the most cooperative over cheating strains (cf. Sachs et al., 2004) or as a preferential choice of one symbiont over another irrespective of symbiont degree of cooperation (for example, Bot et al., 2001; Mueller et al., 2004). Partner choice can take place during the initiation of an association, for example, the colonization of the bobtail squid light organ by Vibrio fischeri symbionts (McFall-Ngai and Ruby, 1991; Visick et al., 2000; Nishiguchi, 2002), or as a means of avoiding cheating symbiont strains (for example, in the yucca–yucca moth symbiosis: Pellmyr and Huth, 1994; Huth and Pellmyr, 2000; in the squid–Vibrio symbiosis: Visick et al., 2000; and in the legume–Rhizobium symbiosis: Simms and Taylor, 2002; West et al., 2002). Our experiments do not evaluate if ant symbiont choice is a response to the degree of cooperation of the Pseudonocardia strains. While future studies exploring whether symbiont choice of more cooperative strains over cheating or inefficient strains help shape the ant–actinomycete association are warranted, the mere presence of symbiont recognition presented here indicates that a mechanism is present that allows the ants to actively regulate the functional aspects of the relationship, facilitate the prevention of conflict between competing Pseudonocardia strains within colonies, and potentially provide a prerequisite for the acquisition of suitable strains for more efficient antibiotic production.
Accession codes
References
Bot ANM, Rehner SA, Boomsma JJ . (2001). Partial incompatibility between ants and symbiotic fungi in two sympatric species of Acromyrmex leaf-cutting ants. Evolution 55: 1980–1991.
Cafaro M, Currie CR . (2005). Phylogenetic analysis of mutualistic filamentous bacteria associated with fungus-growing ants. Can J Microbiol 51: 441–446.
Currie CR, Bot ANM, Boomsma JJ . (2003a). Experimental evidence of a tripartite mutualism: bacteria protect ant fungal gardens from specialized parasites. Oikos 101: 91–102.
Currie CR, Mueller UG, Malloch D . (1999a). The agricultural pathology of ant fungus gardens. Proc Natl Acad Sci USA 96: 7998–8002.
Currie CR, Poulsen M, Mendenhall J, Boomsma JJ, Billen J . (2006). Coevolved crypts and exocrine glands support mutualistic bacteria in fungus-growing ants. Science 311: 81–83.
Currie CR, Scott JA, Summerbell RC, Malloch D . (1999b). Fungus-growing ants use antibiotic-producing bacteria to control garden parasites. Nature 398: 701–704.
Currie CR, Stuart AE . (2001). Weeding and grooming of pathogens in agriculture by ants. Proc R Soc London B 268: 1033–1039.
Currie CR, Wong B, Stuart AE, Schultz TR, Rehner SA, Mueller UG et al. (2003b). Ancient tripartite coevolution in the attine ant-microbe symbiosis. Science 299: 386–388.
Frank SA . (1996). Host-symbiont conflict over the mixing of symbiotic lineages. Proc R Soc London B 263: 339–344.
Frank SA . (2003). Perspective: Repression of competition and the evolution of cooperation. Evolution 57: 693–705.
Gerardo NM, Jacobs SR, Currie CR, Mueller UG . (2006). Ancient host–pathogen associations maintained by specificity of chemotaxis and antibiosis. PLoS Biol 4: 1358–1363.
Gerardo NM, Mueller UG, Price SL, Currie CR . (2004). Exploiting a mutualism: parasite specialization on cultivars within the fungus-growing ant symbiosis. Proc R Soc London B 271: 1791–1798.
Hughes WOH, Thomsen L, Eilenberg J, Boomsma JJ . (2004). Diversity of entomopathogenic fungi near leaf-cutting ant nests in a neotropical forest, with particular reference to Metarhizium anisopliae var. anisopliae. J Inv Pathol 85: 46–53.
Huth CJ, Pellmyr O . (2000). Pollen-mediated selective abortion promotes evolutionary stability of mutualism between yuccas and yucca moths. Ecology 81: 1100–1107.
Maddison DR, Maddison WP . (2005). MacClade 4.07 for OS X. Sinauer Associates Inc.: Sunderland, MA.
McFall-Ngai MJ, Ruby EG . (1991). Symbiont recognition and subsequent morphogenesis as early events in an animal–bacterial mutualism. Science 254: 1491–1494.
Mueller UG, Poulin J, Adams RMM . (2004). Symbiont choice in a fungus-growing ant (Attini, Formicidae). Behav Ecol 15: 357–364.
Mueller UG, Schultz TR, Currie CR, Adams RMM, Malloch D . (2001). The origin of the attine ant-fungus mutualism. Q Rev Biol 76: 169–197.
Nishiguchi K . (2002). Host–symbiont recognition in the environmentally transmitted sepiolid squid–Vibrio mutualism. Microbial Ecol 44: 10–18.
Pellmyr O, Huth CJ. (1994). Evolutionary stability of mutualism between yuccas and yucca moths. Nature 372: 257–260.
Posada D, Crandall KA . (1998). Model test: testing the model of DNA substitution. Bioinformatics 14: 817–818.
Poulsen M, Boomsma JJ . (2005). Mutualistic fungi control crop-diversity in fungus-growing ants. Science 307: 741–744.
Poulsen M, Bot ANM, Boomsma JJ . (2003a). The effect of metapleural gland secretion on the growth of a mutualistic bacterium on the cuticle of leaf-cutting ants. Nature 90: 406–409.
Poulsen M, Bot ANM, Currie CR, Boomsma JJ . (2002). Mutualistic bacteria and a possible trade-off between alternative defence mechanisms in Acromyrmex leaf-cutting ants. Insectes Soc 49: 15–19.
Poulsen M, Bot ANM, Currie CR, Nielsen MG, Boomsma JJ . (2003b). Within colony transmission and the cost of a mutualistic bacterium in the leaf-cutting ant Acromyrmex octospinosis. Funct Ecol 17: 260–269.
Poulsen M, Cafaro M, Boomsma JJ, Currie CR . (2005). Specificity of the mutualistic association between actinomycete bacteria and two sympatric species of Acromyrmex leaf-cutting ants. Mol Ecol 14: 3597–3604.
Poulsen M, Currie CR . (2006). Complexity of insect-fungal associations: Exploring the influence of microorganisms on the attine ant-fungus symbiosis. In: Bourtzis K, Miller T (eds). Insect Symbiosis, Vol. II. CRC Press: Boca Raton, FL.
Quinlan RJ, Cherrett JM . (1977). The role of substrate preparation in the symbiosis between the leaf-cutting ant Acromyrmex octospinosus and its food fungus. Ecol Entomol 2: 161–170.
Richard FJ, Hefetz A, Christides JP, Errard C . (2004). Food influence on colonial recognition and chemical signature between nestmates in the fungus-growing ant Acromyrmex subterraneus. Chemoecology 14: 9–16.
Richard FJ, Poulsen M, Hefetz A, Errard C, Nash DR, Boomsma JJ . (2007). The origin of chemical profiles of fungal symbionts: their significance for nest-mate recognition in Acromyrmex leaf-cutting ants. Behav Ecol Sociobiol (doi:10.1007/s00265-007-0395-1).
Sachs JL, Mueller UG, Wilcox TP, Bull JJ . (2004). The evolution of cooperation. Q Rev Biol 79: 135–160.
Schultz TR, Meier R . (1995). A phylogenetic analysis of the fungus-growing ants (Hymenoptera: Formicidae: Attini) based on morphological characters of the larvae. Syst Entomol 20: 337–370.
Simms EL, Taylor DL . (2002). Partner choice in nitrogen-fixation mutualisms of legumes and rhizobia. Int Comp Biol 42: 369–380.
Swofford DL . (2002). PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods). Version 4. Sinauer Associates: Sunderland, MA.
Taerum SJ, Cafaro MJ, Little AEF, Schultz TR, Currie CR . (in press). Low host–pathogen specificity in the leaf-cutting ant–microbe symbiosis. Proc R Soc London B.
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG . (1997). The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24: 4876–4882.
Viana AMM, Frezard A, Malosse C, Della Lucia TMC, Errand C, Lenoir A . (2001). Colonial recognition of fungus in the fungus-growing ant Acromyrmex subterraneus subterraneus (Hymenoptera: Formicidae). Chemoecology 11: 29–36.
Visick KL, Foster J, Doino J, McFall-Ngai MJ, Ruby EG . (2000). Vibrio fischeri lux genes play an important role in colonization and development of the host light organ. J Bacteriol 182: 4578–4586.
Weber NA . (1972). Gardening Ants: The Attines. American Philosophical Society: Philadelphia.
West SA, Kiers ET, Simms EL, Denison RF . (2002). Sanctions and mutualism stability: why do rhizobia fix nitrogen? Proc R Soc London B 269: 685–694.
Acknowledgements
We are grateful to N Keuler (CALS Statistical Consulting) for statistics advice and to Eric Caldera, Adrian Pinto and Jarrod Scott for comments on a previous draft of this manuscript. This work was supported by Lundbeckfonden to MP and NSF (Integrative Research Challenges in Environmental Biology DEB-0110073) to CRC.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, M., Poulsen, M. & Currie, C. Symbiont recognition of mutualistic bacteria by Acromyrmex leaf-cutting ants. ISME J 1, 313–320 (2007). https://doi.org/10.1038/ismej.2007.41
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2007.41
Keywords
This article is cited by
-
Surface sterilization methods impact measures of internal microbial diversity in ticks
Parasites & Vectors (2019)
-
Enforcement is central to the evolution of cooperation
Nature Ecology & Evolution (2019)
-
Microbial Diversity and Chemical Multiplicity of Culturable, Taxonomically Similar Bacterial Symbionts of the Leaf-Cutting Ant Acromyrmex coronatus
Microbial Ecology (2019)
-
Detection and cell sorting of Pseudonocardia species by fluorescence in situ hybridization and flow cytometry using 16S rRNA-targeted oligonucleotide probes
Applied Microbiology and Biotechnology (2018)
-
Fungus-Farming Termites Selectively Bury Weedy Fungi that Smell Different from Crop Fungi
Journal of Chemical Ecology (2017)