Abstract
Human dental pulp cells (hDPCs) possess the capacity to differentiate into odontoblast-like cells and generate reparative dentin in response to exogenous stimuli or injury. Ten–eleven translocation 1 (TET1) is a novel DNA methyldioxygenase that plays an important role in the promotion of DNA demethylation and transcriptional regulation in several cell lines. However, the role of TET1 in the biological functions of hDPCs is unknown. To investigate the effect of TET1 on the proliferation and odontogenic differentiation potential of hDPCs, a recombinant shRNA lentiviral vector was used to knock down TET1 expression in hDPCs. Following TET1 knockdown, TET1 was significantly downregulated at both the mRNA and protein levels. Proliferation of the hDPCs was suppressed in the TET1 knockdown groups. Alkaline phosphatase activity, the formation of mineralized nodules, and the expression levels of DSPP and DMP1 were all reduced in the TET1-knockdown hDPCs undergoing odontogenic differentiation. Based on these results, we concluded that TET1 knockdown can prevent the proliferation and odontogenic differentiation of hDPCs, which suggests that TET1 may play an important role in dental pulp repair and regeneration.
Similar content being viewed by others
Introduction
Human dental pulp cells (hDPCs) are mesenchymal cells derived from the neural crest that exhibit plasticity and multipotency; these cells can generate reparative dentin to resist and repair injury due to infection or trauma.1, 2 A great deal of attention has been focused on the mechanisms involved in the odontogenic differentiation process for reparative dentine formation and dental pulp regeneration.3, 4 Numerous studies have proven that signal pathways play critical roles in regulating gene expression of the core transcriptional network of hDPCs, such as Wnt, transforming growth factor (TGF)-β, and Notch1 signaling.5, 6, 7 Several growth factors, including bone morphogenic proteins (BMPs) and growth and differentiation factor-5 (GDF-5), have been identified as chemotactic signals to recruit progenitor cells and stimulate their proliferation and differentiation in hDPCs.8 There is increasing evidence that epigenetic regulatory mechanisms such as histone acetylation and microRNAs are involved in reparative dentinogenesis.9, 10, 11 DNA methylation is a vital epigenetic modification that serves as a critical switch for gene expression, genomic consistency, and other processes of epigenetic inheritance.12, 13, 14 However, the role of DNA methylation in the regulation of the biological properties of hDPCs remains elusive.
Our previous study indicated that the DNA methyltransferase inhibitor 5-Aza-2'-deoxycytidine (5-Aza-CdR) promotes the odontogenic differentiation capacity of hDPCs, suggesting that DNA demethylation may provide a new mechanism for the regulation of odontogenic differentiation.15 Recent studies have shown that ten–eleven translocation 1 (TET1), a recently discovered DNA dioxygenase, could catalyze the addition of covalent hydroxyl modifications to methylated DNA and promote DNA demethylation. This reaction influences gene transcription, presumably by converting 5-methylcytosine (5mC) to 5-hydroxymethylcytosine (5hmC) at specific genes.16, 17, 18, 19 Increasing evidence indicates that TET1 is involved in the epigenetic regulation of proliferation and differentiation in various cells such as embryonic stem cells (ESCs), adult neural progenitor cells, muscle progenitor cells, and cancer cells.20, 21, 22, 23, 24 Previously, we reported that TET1 was expressed in hDPCs and that its expression increased during early natural differentiation and odontogenic induction.25 However, the role of TET1 in the odontogenic differentiation of hDPCs remains unknown.
In the present study, shRNA was used to knock down TET1 expression in hDPCs. The effect of TET1 on the proliferation and odontogenic differentiation of hDPCs was then investigated. Our results demonstrate the role of TET1-dependent DNA demethylation in the regulation of the differentiation potential of hDPCs.
Materials and methods
Cell culture
This study was approved by the Ethical Review Board of the Guanghua School of Stomatology of Sun Yat-sen University. All of the patients enrolled in this study gave written informed consent. hDPCs were isolated and cultivated as previously described by Gronthos et al.26 Briefly, pulp tissues were minced into small pieces and digested in a solution containing 3 mg·mL−1 collagenase type I (Gibco, Carlsbad, CA, USA) for 20 min at 37 °C. The pulp tissue was then cultured in complete medium containing Dulbecco minimum essential medium (DMEM) supplemented with 10% fetal bovine serum (FBS), 100 u·mL−1 penicillin, and 100 mg·mL−1 streptomycin (Gibco, Carlsbad, CA, USA) at 37 °C in an atmosphere of 95% O2 and 5% CO2. The media were changed every 3 days. When the cells reached 80% confluence, they were harvested using trypsin/ethylene diamine tetraacetic acid (EDTA) (Gibco, Carlsbad, CA, USA) and subcultured at a ratio of 1:3.
Flow cytometry assay
The stem cell phenotypic markers of hDPCs were identified by flow cytometry; 105 cells were resuspended in 100 μL phosphate-buffered saline (PBS) and incubated with primary STRO-1 and CD146 antibodies at 4 °C for 1 h, using 1:100 dilutions. The labeled cells were suspended in 100 μL PBS with 1 μL anti-mouse IgG conjugated with fluorescein isothiocyanate (Chemicon, Temecula, CA, USA) at 4 °C for 1 h and then examined with a FACS Calibur apparatus (Becton Dickinson, La Jolla, CA, USA).
Knockdown of TET1 in hDPCs using shRNA
To determine the role of TET1 in the differentiation of hDPCs, TET1 expression was knocked down using short hairpin RNA (shRNA). The TET1-targeting shRNA sequences (TET1 shRNA1: 5'-CAGAAGATTTAGAATTGAT-3' and TET1 shRNA-2: 5'-AGCTAATGAAGGTCCAGAAC-3') were designed and cloned into the hU6-MCS-CMV-Puromycin vector. The recombinant construct or a non-specific shRNA construct (control group), as well as three helper vectors (pLP1, pLP2, and pLP/VSVG), were transfected into 293FT cells. The viruses were collected 72 h later and transfected into hDPCs.
Cell proliferation assay
The cell proliferation of hDPC was assessed using the Cell Counting Kit-8 (CCK8) assay (Dojindo, Kumamoto, Japan) according to the manufacturer’s protocol. After 1, 2, 3, 4, 5, and 6 days of incubation, the supernatant of each group was removed, and the hDPCs were incubated in DMEM containing CCK8 for another 2 h at 37 °C. The optical density (OD) of each well was read at 450 nm using an automated microplate reader (Sunrise, Tecan, Switzerland).
Odontogenic induction of hDPCs
The cells were seeded in six-well plates at a density of 1 × 105 cells per well and cultured in DMEM containing 10% FBS until they reached 80% confluence. The medium was then replaced with odontogenic differentiation media containing DMEM supplemented with 5% FBS, 50 μg·mL−1 ascorbic acid, 10 mmol·L−1 b-glycerophosphate, and 10−7 mol·L−1 dexamethasone (Sigma-Aldrich, St Louis, MO, USA). The medium was changed every 3 days.
Alkaline phosphatase activity analysis
After 7 days of incubation, the cells of each group were rinsed twice with ice-cold PBS (pH 7.4) and solubilized in 0.1% Triton X-100 for 15 h at 4 °C. An ALP assay kit (Nanjing Jiancheng Bioengineering Institute, Nanjing, China) was used to assess alkaline phosphatase (ALP) activity according to the manufacturer’s instructions. The absorbance was measured at 520 nm using an automated microplate reader (Sunrise, Tecan, Switzerland). Total protein content was quantified using a bicinchoninic acid protein assay (Beyotime, Haimen, China) and the ALP levels were normalized to the total protein.
Alizarin red S staining
The cells were seeded in six-well plates and mineralization was assessed via alizarin red staining after a culture period of 21 days. The mineralized nodules were observed and photographed using an inverted microscope (Zeiss, Jena, Germany). The amount of mineralization matrix was determined by dissolving alizarin red S in 100 g·L−1 cetylpyridinium chloride (CPC; Sigma-Aldrich, St Louis, MO, USA) and 10 mM sodium phosphate. The OD was read at 540 nm using an automated microplate reader (Sunrise, Tecan, Switzerland), and the quantitative measurements were made using the generated standard curve.
Real-time quantitative polymerase chain reaction
Total RNA was extracted from the hDPCs using TRIzol reagent according to the manufacturer’s instructions (Invitrogen, Carlsbad, CA, USA). Then, 2 μg of RNA was reverse-transcribed for cDNA synthesis using a RevertAid first strand cDNA synthesis kit (Fermentas, Ontario, Canada) and random primers. Real-time quantitative polymerase chain reaction (qRT-PCR) was performed using the LightCycler 480 SYBR Green I Master (Roche, Basel, Switzerland) with specific primers according to the manufacturer’s instructions. The expression data were normalized to the geometric mean of the housekeeping gene GAPDH. The following primers were synthesized by Invitrogen (Life Technologies, Carlsbad, CA, USA): TET1: 5'-CATCAGTCAAGACTTTAAGCCCT-3'(forward), 5'-CGGGTGGTTTAGGTTCTGTTT-3'(reverse); DSPP: 5'-GCCACTTTCAGTCTTCAAAGAGA-3' (forward), 5'-GCCCAAATGCAAAAATATGTAA-3'(reverse); DMP1: 5'-AAAATTCTTTGTGAACTACGGAGG-3'(forward), 5'-GAGCACAGGATAATCCCCAA-3'(reverse); and GAPDH: 5'-GGCATGGACTGTGGTCATGAG-3'(forward), 5'-TGCACCACCAACTGCTTAGC-3' (reverse). The mRNA levels were normalized to the mRNA level of GAPDH.
Western blotting analysis
The cells were harvested using RIPA lysis buffer (Beyotime, Haimen, China). In total, 40 μg of protein was subjected to 6% sodium dodecyl sulfate—polyacrylamide gel electrophoresis and transferred to polyvinylidene fluoride membranes (Millipore, Billerica, MA, USA) in transfer buffer containing 10% methanol. The membranes were blocked in TBST containing 5% skim milk at room temperature for 1 h and then incubated with an anti-TET1 (Genetex, Irvine, CA, USA), anti-DMP1 (Abcam, Cambridge, UK), anti-DSPP antibody (Abcam, Cambridge, UK), anti-β-ACTIN (Beyotime, Haimen, China) or an anti-GAPDH antibody (Beyotime, Haimen, China) overnight at 4 °C. After the cells were incubated with a secondary antibody (Abcam, Cambridge, UK) for 1 h at room temperature, the immunoreactive bands were developed using Amersham’s enhanced chemiluminescence reagents (Millipore ECL Western Blotting Detection System, MA, USA) and observed using an ImageQuant LAS 4000 Mini system (GE Healthcare Life Sciences, Piscataway, NJ, USA). The blots were quantified and normalized using ImageJ 1.47 software (National Institutes of Health, Bethesda, MD, USA).
Statistical analysis
Each experiment was performed in triplicate, and each set was repeated at least three times. The experimental groups were compared using one-way analysis of variance (ANOVA) or repeated-measures ANOVA with SPSS16.0 software (SPSS, Chicago, IL, USA). The values are expressed as the mean±standard deviation. All P-values are two-tailed, and P<0.05 was considered statistically significant.
Results
Identification of stem cell phenotypic markers in primary hDPCs
STRO-1+ and CD146+ have been shown to exhibit mesenchymal stem cell properties, and these markers have been used to identify dental pulp stem cells.27 STRO-1 and CD146 were identified using flow cytometry in primary hDPCs. The results revealed the expression of STRO-1 (23.68%) and CD146 (89.96%), indicating that hDPCs contain mesenchymal progenitors.
TET1 expression levels in TET1-shRNA hDPCs
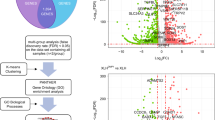
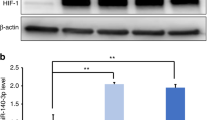
To investigate the role of TET1 in hDPCs, TET1 was knocked down using shRNA. As shown in Figure 1a, the TET1 mRNA expression level decreased by approximately 50% in the shRNA1 and shRNA2 groups compared with the control group, which was consistent with the reduction in protein expression determined by western blotting (Figure 1b).
The level of TET1 expression after TET1 knockdown in hDPCs. (a) TET1 mRNA expression was assessed using qRT-PCR. GAPDH was used as an internal control. (b) TET1 protein expression was assessed using western blotting and densitometric evaluation. β-ACTIN was used as an internal control. The expression level of TET1 decreased by approximately 50% in the shRNA1 and shRNA2 groups compared with the control group. All of the results represent the mean±standard deviation of three independent experiments (n=3). *Significant difference compared with the control (P<0.05). hDPC, human dental pulp cell; qRT-PCR, real-time quantitative polymerase chain reaction; shRNA, short hairpin RNA; TET1, ten–eleven translocation 1.
Effect of TET1 knockdown on the proliferation of hDPCs
To analyze the effect of TET1 knockdown on the proliferation of the hDPCs, growth rates were measured using the CCK8 assay. As shown in Figure 2, the growth rates of the shRNA1 and shRNA2 groups decreased after 2, 3, 4, 5, and 6 days compared with the control group.
Effect of TET1 knockdown on the proliferation of hDPCs. Cell growth of each group was measured using the CCK8 assay. The growth curves showed that the growth rates of the hDPC/shTET1 cells were significantly decreased from the second day onward. All of the results represent the mean±standard deviation of three independent experiments (n=3). *Significant difference compared with the control (P<0.05). D, day; hDPC, human dental pulp cell; TET 1, ten–eleven translocation 1.
Effects of TET1 knockdown on the mineralization potential of hDPCs
To determine the effect of TET1 knockdown on the mineralization potential of hDPCs, ALP activity and the formation of mineralized nodules were assessed. The ALP activity of all of the groups did not significantly differ under normal culture conditions. However, in the odontogenic induction medium, ALP activity decreased by 54.92% in the shRNA1 group and by 79.52% in the shRNA2 group compared with the control group (Figure 3a). Similarly, alizarin red S staining indicated that the formation of mineralized nodules was inhibited in the TET1-knockdown hDPCs undergoing odontogenic induction (Figure 3b).
Effect of TET1 knockdown on the mineralization potential of hDPCs. Cells were cultured in GM and DM. (a) ALP activity was measured at day 7. The ALP activity of all of the groups did not significantly differ under normal culture conditions. However, the ALP activity decreased by 54.92% in the shRNA1 group and by 79.52% in the shRNA2 group compared with the control group in the odontogenic differentiation medium. (b) Mineralization was analyzed using alizarin red S staining at day 21. The formation of mineralized nodules was inhibited in the TET1-knockdown hDPCs undergoing odontogenic induction. All of the results represent the mean±standard deviation of three independent experiments (n=3). *Significant difference compared with the control (P<0.05). ALP, alkaline phosphatase; DM, differentiation medium; GM, growth medium; hDPC, human dental pulp cell; shRNA, short hairpin RNA; TET1, ten–eleven translocation 1.
Effects of TET1 knockdown on odontogenic differentiation markers in hDPCs
To further identify the role of TET1 in the regulation of the odontogenic differentiation potential of hDPCs, expression of the odontogenic markers DSPP and DMP1 was detected using qRT-PCR and western blotting. TET1-shRNA led to a decrease in the mRNA expression levels of DMP1 and DSPP with or without odontogenic induction (Figure 4a). The protein expression level of DSPP decreased in culture media, whereas the protein expression of DMP1 did not significantly differ in normal culture conditions but decreased in the TET1-shRNA groups after odontogenic induction (Figure 4b).
Effect of TET1 knockdown on DSPP and DMP1 in hDPCs. (a) The mRNA expression levels of DSPP and DMP1 were determined using qRT-PCR. GAPDH was used as an internal control. The mRNA expression levels of DMP1 and DSPP decreased in the TET1-shRNA groups with or without odontogenic induction. (b) Protein expression levels of DMP1 and DSPP were assessed by western blot. GAPDH was used as an internal control. The protein expression of DSPP decreased in both culture media, whereas the protein expression of DMP1 did not significantly differ under normal culture conditions but decreased in the TET1-knockdown hDPCs undergoing odontogenic induction. All of the results represent the mean±standard deviation of three independent experiments (n=3). *Significant difference compared with the control (P<0.05). DM, differentiation medium; GM, growth medium; hDPC, human dental pulp cell; qRT-PCR, real-time quantitative polymerase chain reaction; shRNA, short hairpin RNA; TET1, ten–eleven translocation 1.
Discussion
Elucidating the molecular mechanisms that regulate the odontogenic differentiation of hDPCs would contribute greatly to advancing treatment strategies for regenerative endodontics. Our recent studies indicated that DNA demethylation may provide a way to creatively illustrate this problem.15, 25 TET1 is a key protein in the DNA demethylation pathway and has a profound impact on cell self-renewal and on the determination of lineage.28, 29 However, the mechanism by which TET1 is involved in the differentiation program of hDPCs remains unclear.
TET1 is known to catalyze the conversion of 5mC to 5hmC.16 5hmC can be further oxidized to 5-formylcytosine and 5-carboxylcytosine. These derivatives, the production of which is mediated by TET1, can be actively removed from the genome by thymine-DNA glycosylase through a base excision repair pathway and can then convert back to 5mC.30, 31 This process is called active DNA demethylation, which is associated with the activation of genes.32 Recent studies have demonstrated that TET1 is involved in the control of active and passive demethylation via different mechanisms and thus plays a vital role in the modulation of primordial germ cell formation, embryonic development, stem cell pluripotency, and nerve and brain development.20, 33, 34, 35 Our previous study demonstrated that TET1 is present in the hDPCs and that its expression increases in a time-dependent manner during odontogenic induction.25 The present study further investigated the role of TET1 in the proliferation and odontogenic differentiation of hDPCs.
Zhang et al.21 observed that the loss of Tet1 downregulated a cohort of genes involved in the proliferation of adult neural progenitor cells and impaired hippocampal neurogenesis in mice, which was accompanied by poor learning and memory. Tet1 depletion inhibits the growth of NIH3T3 cells by blocking cyclin D1 accumulation in G1 phase, inhibiting Rb phosphorylation and consequently delaying entrance to G1/S phase.36 The reduction of TET1 significantly downregulated proliferating cell nuclear antigen, thus inhibiting cell proliferation in human uterine leiomyoma.37 In the present study, TET1 knockdown suppressed cell growth from the second day onward, suggesting that the loss of TET1 inhibited the proliferation of the hDPCs. The inhibitory effect of TET1-shRNA on the proliferation of hDPCs was in accordance with these previous reports.
TET1 can influence the balance between 5hmC and 5mC in the genome, which is inextricably associated with lineage commitment.13, 19 Wu et al.28 found that Tet1 has a dual function in transcriptional regulation in mouse ESCs. It binds to and affects both repressed and actively transcribed genes. TET1-mediated hypomethylation of promoter regions leads to the upregulation of genes related to pluripotency and ES cell maintenance. Meanwhile, Tet1 also contributes to gene silencing by facilitating the recruitment of polycomb repressive complex 2 or SIN3A to genes that function in development and differentiation.38 Furthermore, TET1 could maintain a hypomethylated state at the regions to which it was bound, but its depletion did not lead to downregulation of all the TET1 targets.28 With regard to somatic cells, Fujiki et al.39 found that TET1 catalyzes the conversion of 5mC to 5hmC by binding to the PPARgamma co-activator complex, thereby inducing region-specific demethylation and activating the adipocyte differentiation process. Jin et al.40 recently reported that TET1 does not purposely decrease methylation levels; instead, it specifically prevents the spread of aberrant methylation into CpG islands in differentiated cells (HEK293T cells). These new findings suggest that TET1-dependent demethylation plays an important role in the lineage determination, but the underlying mechanism is complex and remains controversial. In the present study, TET1 knockdown decreased ALP activity and the formation of mineralized nodules, and downregulated the expression levels of DSPP and DMP1, which indicates that TET1 is positively involved in the regulation of the odontogenic differentiation of hDPCs. The exact mechanisms by which the TET1 enzyme regulates the differentiation properties of hDPCs are currently being investigated in our laboratory.
In conclusion, this study demonstrated that TET1 knockdown can suppress cell growth and prevent the odontogenic differentiation of hDPCs by inhibiting ALP activity, mineralized nodule formation, and the expression of DSPP and DMP1. These findings indicate that TET1 may promote the proliferation and odontogenic differentiation of hDPCs. More studies are necessary to further elucidate the epigenetic mechanisms by which TET1-dependent demethylation regulates the biological characteristics of hDPCs.
References
Téclès O, Laurent P, Zygouritsas S et al. Activation of human dental pulp progenitor/stem cells in response to odontoblast injury. Arch Oral Biol 2005; 50 (2): 103–108.
Cooper PR, Holder MJ, Smith AJ . Inflammation and regeneration in the dentin-pulp complex: a double-edged sword. J Endod 2014; 40 (4 Suppl): S46–S51.
Tamaoki N, Takahashi K, Tanaka T et al. Dental pulp cells for induced pluripotent stem cell banking. J Dent Res 2010; 89 (8): 773–778.
Rodríguez-Lozano FJ, Insausti CL, Iniesta F et al. Mesenchymal dental stem cells in regenerative dentistry. Med Oral Patol Oral Cir Bucal 2012; 17 (6): e1062–e1067.
Bae CH, Lee JY, Kim TH et al. Excessive Wnt/β-catenin signaling disturbs tooth-root formation. J Periodont Res 2013; 48 (4): 405–410.
Zhou C, Yang G, Chen M et al. Lhx8 mediated Wnt and TGFβ pathways in tooth development and regeneration. Biomaterials 2015; 63: 35–46.
He F, Yang Z, Tan Y et al. Effects of Notch ligand Delta1 on the proliferation and differentiation of human dental pulp stem cells in vitro. Arch Oral Biol 2009; 54 (3): 216–222.
Al-Sharabi N, Xue Y, Fujio M et al. Bone marrow stromal cell paracrine factors direct osteo/odontogenic differentiation of dental pulp cells. Tissue Eng Part A 2014; 20 (21/22): 3063–3072.
Liu HJ, Wang T, Li QM et al. Knock-down of p300 decreases the proliferation and odontogenic differentiation potentiality of HDPCs. Int Endod J 2015; 48 (10): 976–985.
Jang YE, Go SH, Lee BN et al. Changes in SIRT gene expression during odontoblastic differentiation of human dental pulp cells. Restor Dent Endod 2015; 40 (3): 223–228.
Gong Q, Wang R, Jiang H et al. Alteration of microRNA expression of human dental pulp cells during odontogenic differentiation. J Endod 2012; 38 (10): 1348–1354.
Bird A . DNA methylation patterns and epigenetic memory. Genes Dev 2002; 16 (1): 6–21.
Cimmino L, Abdel-Wahab O, Levine RL et al. TET family proteins and their role in stem cell differentiation and transformation. Cell Stem Cell 2011; 9 (3): 193–204.
Bhutani N, Burns DM, Blau HM . DNA demethylation dynamics. Cell 2011; 146 (6): 866–872.
Zhang D, Li Q, Rao L et al. Effect of 5-Aza-2'-deoxycytidine on odontogenic differentiation of human dental pulp cells. J Endod 2015; 41 (5): 640–645.
Tahiliani M, Koh KP, Shen Y et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 2009; 324 (5929): 930–935.
Ito S, Shen L, Dai Q et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 2011; 333 (6047): 1300–1303.
Koh KP, Yabuuchi A, Rao S et al. Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell 2011; 8 (2): 200–213.
Ficz G, Branco MR, Seisenberger S et al. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature 2011; 473 (7347): 398–402.
Ito S, D'Alessio AC, Taranova OV et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 2010; 466 (7310): 1129–1133.
Zhang RR, Cui QY, Murai K et al. Tet1 regulates adult hippocampal neurogenesis and cognition. Cell Stem Cell 2013; 13 (2): 237–245.
Tsumagari K, Baribault C, Terragni J et al. Early de novo DNA methylation and prolonged demethylation in the muscle lineage. Epigenetics 2013; 8 (3): 317–332.
Sun M, Song CX, Huang H et al. HMGA2/TET1/HOXA9 signaling pathway regulates breast cancer growth and metastasis. Proc Natl Acad Sci USA 2013; 110 (24): 9920–9925.
Feng J, Wang Q, Li G et al. TET1-mediated different transcriptional regulation in prostate cancer. Int J Clin Exp Med 2015; 8 (1): 203–211.
Li Q, Rao L, Zhang D et al. Expression features of DNA methylcytosine dioxygenase ten-eleven translocation 1 in human dental pulp cells. J Endod 2014; 40 (11): 1791–1795.
Gronthos S, Mankani M, Brahim J et al. Postnatal human dental pulp stem cells (DPSCs) in vitro and in vivo. Proc Natl Acad Sci USA 2000; 97 (25): 13625–13630.
Huang GT, Gronthos S, Shi S . Mesenchymal stem cells derived from dental tissues vs those from other sources: their biology and role in regenerative medicine. J Dent Res 2009; 88 (9): 792–806.
Wu H, D'Alessio AC, Ito S et al. Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells. Nature 2011; 473 (7347): 389–393.
Dawlaty MM, Breiling A, Le T et al. Loss of Tet enzymes compromises proper differentiation of embryonic stem cells. Dev Cell 2014; 29 (1): 102–111.
He YF, Li BZ, Li Z et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 2011; 333 (6047): 1303–1307.
Maiti A, Drohat AC . Thymine DNA glycosylase can rapidly excise 5-formylcytosine and 5-carboxylcytosine: potential implications for active demethylation of CpG sites. J Biol Chem 2011; 286 (41): 35334–35338.
Shen L, Zhang Y . Enzymatic analysis of Tet proteins: key enzymes in the metabolism of DNA methylation. Meth Enzymol 2012; 512: 93–105.
Hackett JA, Sengupta R, Zylicz JJ et al. Germline DNA demethylation dynamics and imprint erasure through 5-hydroxymethylcytosine. Science 2013; 339 (6118): 448–452.
Gao Y, Chen J, Li K et al. Replacement of Oct4 by Tet1 during iPSC induction reveals an important role of DNA methylation and hydroxymethylation in reprogramming. Cell Stem Cell 2013; 12 (4): 453–469.
Guo JU, Ma DK, Mo H et al. Neuronal activity modifies the DNA methylation landscape in the adult brain. Nat Neurosci 2011; 14 (10): 1345–1351.
Huang S, Zhu Z, Wang Y et al. Tet1 is required for Rb phosphorylation during G1/S phase transition. Biochem Biophys Res Commun 2013; 434 (2): 241–244.
Navarro A, Yin P, Ono M et al. 5-Hydroxymethylcytosine promotes proliferation of human uterine leiomyoma: a biological link to a new epigenetic modification in benign tumors. J Clin Endocrinol Metab 2014; 99 (11): E2437–E2445.
Neri F, Incarnato D, Krepelova et al. Genome-wide analysis identifies a functional association of Tet1 and polycomb repressive complex 2 in mouse embryonic stem cells. Genome Biol 2013; 14 (8): R91.
Fujiki K, Shinoda A, Kano F et al. PPARγ-induced PARylation promotes local DNA demethylation by production of 5-hydroxymethylcytosine. Nat Commun 2013; 4: 2262.
Jin C, Lu Y, Jelinek J et al. TET1 is a maintenance DNA demethylase that prevents methylation spreading in differentiated cells. Nucleic Acids Res 2014; 42 (11): 6956–6971.
Acknowledgements
This work was supported by the National Nature Science Foundation of China (grant no. 81570971).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Rao, LJ., Yi, BC., Li, QM. et al. TET1 knockdown inhibits the odontogenic differentiation potential of human dental pulp cells. Int J Oral Sci 8, 110–116 (2016). https://doi.org/10.1038/ijos.2016.4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ijos.2016.4
Keywords
This article is cited by
-
DNA Methylation and Histone Modification in Dental-derived Mesenchymal Stem Cells
Stem Cell Reviews and Reports (2022)
-
Temporal Modulation of DNA Methylation and Gene Expression in Monolayer and 3D Spheroids of Dental Pulp Stem Cells during Osteogenic Differentiation: A Comparative Study
Tissue Engineering and Regenerative Medicine (2022)
-
Role of connexin 43 in odontoblastic differentiation and structural maintenance in pulp damage repair
International Journal of Oral Science (2021)
-
Specific functions of TET1 and TET2 in regulating mesenchymal cell lineage determination
Epigenetics & Chromatin (2019)
-
Decoding the role of TET family dioxygenases in lineage specification
Epigenetics & Chromatin (2018)