Abstract
The etiopathogenesis of cardiovascular diseases (CVDs) is multifactorial. Adverse blood pressure (BP) is a major independent risk factor for epidemic CVD affecting ~40% of the adult population worldwide and resulting in significant morbidity and mortality. Metabolic phenotyping of biological fluids has proven its application in characterizing low-molecular-weight metabolites providing novel insights into gene–environmental–gut microbiome interaction in relation to a disease state. In this review, we synthesize key results from the INTERnational study of MAcro/micronutrients and blood Pressure (INTERMAP) Study, a cross-sectional epidemiologic study of 4680 men and women aged 40–59 years from Japan, the People’s Republic of China, the United Kingdom and the United States. We describe the advancements we have made regarding the following: (1) analytical techniques for high-throughput metabolic phenotyping; (2) statistical analyses for biomarker identification; (3) discovery of unique food-specific biomarkers; and (4) application of metabolome-wide association studies to gain a better understanding into the molecular mechanisms of cross-cultural and regional BP differences.
Similar content being viewed by others
Introduction
Adverse blood pressure (BP) (prehypertension and hypertension) is a major independent risk factor for epidemic cardiovascular diseases (CVDs), affecting ~40% of the adult population worldwide.1, 2, 3, 4 Its causation is multifactorial, encompassing both environmental—diet and other aspects of lifestyle—and genetic factors. Public health policies aiming to improve the prevention of high BP and/or maintain optimal BP levels typically involve efforts to tackle known modifiable risk factors such as reduction of high salt intake, moderation of alcohol intake, maintenance of normal weight and increased physical activity. Although large-scale genome-wide association studies have identified common variants, novel loci and pathways associated with BP,5, 6, 7, 8, 9 the genetic contribution to BP variation in the population is modest.5, 7, 8 Despite the effectiveness of non-pharmacological approaches to BP control, many people with high BP are reliant on antihypertensive drugs, or their BP remains untreated or poorly controlled.10, 11
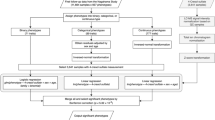
The INTERnational study of MAcro/micronutrients and blood Pressure (INTERMAP) is a cross-sectional epidemiological study of 4680 men and women aged 40–59 years from Japan, the People's Republic of China, the United Kingdom and the United States.12 The main aim of the INTERMAP Study is to investigate the etiology of adverse BP with an emphasis on studying diet–BP associations (Figure 1).12, 13 Participants were selected randomly from population lists, stratified by age/sex. Each participant attended four visits: visits 1 and 2 on consecutive days and visits 3 and 4 on consecutive days, on average 3 weeks later. At each of the four visits, BP was measured twice with a random zero sphygmomanometer, and dietary data were collected by a trained certified interviewer with use of the in-depth multipass 24-h recall method.14 Height and weight were measured twice on the first and third visits. At the first visit, questionnaire data were obtained on demographic, medical history, medication intake, and other possible confounders. Each participant provided two 24-h urine collections, with the beginning and end timed at the research center. The two timed 24-h urine collections per person on each of the 4680 INTERMAP participants are a pivotal feature of the study design that enabled the introduction of metabolome-wide association (MWA) study (see later section). The study received institutional ethics committee approval for each site, and all participants gave written informed consent.
In the past decade, the INTERMAP Study has added new knowledge to the limited data previously available on the effects of nutritional factors on BP.15 These advances include the inverse relationship between BP and intakes of vegetable protein,16 glutamic acid,17 total and insoluble fiber,18 raw and cooked vegetables,19 total polyunsaturated fatty acids and linoleic acid,20 oleic acid from vegetable sources,21 total omega-3 fatty acids and linolenic acid,22 phosphorus (P), calcium (Ca) and magnesium (Mg),23 non-heme iron (Fe) and total Fe,24 and starch25 and the direct associations of sugars (fructose, glucose and sucrose),26 cholesterol,27 glycine and alanine,28 raw fruits29 and oleic acid from animal sources.21 The INTERMAP Study showed that diet-induced metabolic acidosis was positively associated with BP (not significant after controlling for body mass index, BMI),30 whereas a cohort of over 61 000 persons reported a positive association between the metabolic acid load (such as serum bicarbonate, urine acidity) and cardiovascular mortality.31 While the INTERMAP Study reported a small nonsignificant inverse relationship between urinary Mg and BP,32 the World Health Organization-coordinated Cardiovascular Diseases and Alimentary Comparison (CARDIAC) Study showed that urinary Mg-to-creatinine ratio was inversely associated with CVD risk factors such as BMI and BP.33 Although important advances have been made by the INTERMAP Study in improving understanding of the etiology of high BP, together with other research into the physiology of BP regulation such as the control of kidney fluid and salt balance via the renin–angiotensin–aldosterone system,34, 35, 36 sympathetic nervous system activity37, 38 and the role of the structure and function of blood vessels,39 gaps remain in our knowledge of the causes and mechanisms of adverse BP levels. New approaches are needed to enhance the understanding of the multifactorial etiopathogenesis of BP.
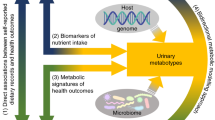
High-resolution proton nuclear magnetic resonance (NMR) spectroscopy has been successfully applied for the investigation of drug metabolism and toxicology, as well as disease development within a biological system, using biological fluids such as urine, plasma/serum, bile, cerebrospinal fluids and dialysates.40, 41 Recent studies have shown the importance of the gut microbiome in the etiology of a number of chronic diseases such as atherosclerosis, diabetes and the metabolic syndrome, obesity and elevated BP.42, 43, 44, 45, 46, 47 Metabolic phenotyping of biological fluids using spectroscopic methods enables the investigation of gene–environmental–gut microbiome interactions on disease risk and is thus an attractive approach for gaining new insights into BP mechanisms and its associated pathways.45, 48, 49, 50 The INTERMAP Study has capitalized on the evolving technologies in metabolic phenotyping to enhance its rich nutrient and anthropometric data by incorporating this top-down system approach to investigate the association of BP and urinary markers that are linked to environmental exposures, including diets and xenometabolomes.48, 49, 50, 51, 52 This review demonstrates the key progress made by the INTERMAP Study and the introduction of MWA studies on diet and BP.13 However, readers may wish to refer to other recent reviews on the principles of metabolic phenotyping.53, 54, 55
Analytical method development
Metabolic phenotyping involves the application of high-throughput advanced spectroscopic methods, such as proton nuclear magnetic resonance (1H NMR) spectroscopy and mass spectroscopy (MS), to biological samples. In INTERMAP, comprehensive metabolic phenotyping by 1H NMR spectroscopy of the two 24-h urine samples from each participant (N=4630) has been performed.13 The repeatability, accuracy and stability of the 1H NMR spectral profiles were evaluated, and the overall analytical reproducibility of the 1H NMR was found to be >98%.56 In INTERMAP, boric acid (borate, a preservative known to bind covalently to vicinal diols and some amino acids) was added to the urine collection jars in the field to prevent bacterial overgrowth. The effect of the boric acid on the 1H NMR urinary spectra was also assessed.37, 38 It was shown that the overall changes in the urinary 1H NMR spectral profile caused by borate addition were negligible compared with the physiological and metabolic differences between individuals.57 These studies have led to recommendations for improved sample preparation and handling of urine samples, as well as processing methods for large-scale epidemiologic research.58
In addition to metabolic phenotyping via 1H NMR spectroscopy, we obtained extensive data on 20 urinary amino acids using conventional cation-exchange chromatography followed by postcolumn derivatization (Biochrom 20 and Biochrom 30). These data provide a unique population-based data set on urinary amino-acid excretion levels in different populations. We then applied gas chromatography mass spectrometry and liquid chromatography-tandem mass spectrometry (LC-MS/MS) for high-throughput urinary amino-acid analysis and compared their sample preparation, run time, number of analytes amenable to quantification, cost, limit of quantification, reproducibility and validity with conventional amino-acid analysis.59 The amounts of urine needed for GS-MS and LC-MS/MS were 40–50 μl, much less than the 200 μl required for the amino-acid analyzer. Moreover, the run time for the amino-acid analyzer was ~5 to 6 times longer than that of gas chromatography mass spectrometry and LC-MS/MS. The Pearson's correlation coefficients of amino acids measured by gas chromatography mass spectrometry and the amino-acid analyzer ranged from 0.80 (tryptophan) to 0.98 (glycine); correlation coefficients comparing LC-MS/MS with the amino-acid analyzer ranged from 0.56 (arginine) to 0.95 (lysine). Our findings showed that gas chromatography mass spectrometry offered higher reproducibility and completely automated sample pre-treatment compared with LC-MS/MS and conventional amino-acid analysis, which covered more amino acids and related amines.
We also applied ultra-performance liquid chromatography-triple quadruple-tandem mass spectrometry for simultaneous detection (both positive and negative electrospray ionization modes) and quantification of three gut microbial cometabolites, phenylacetylglutamine, 4-cresyl sulfate and hippurate in the urinary specimens from 2000 US INTERMAP participants.60 This targeted high-throughput ultra-performance liquid chromatography-triple quadruple-tandem mass spectrometry method was developed specifically to measure these gut microbial cometabolites, which may be implicated in obesity61 and other chronic diseases such as cardiovascular and kidney diseases.62, 63 Following the US Food and Drug Administration guidelines, the imprecision (coefficient of variation) and inaccuracy (recovery) of the method were assessed using replicates of quality control urine samples at different concentration levels. The coefficient of variation and recovery were found to be within the acceptable limits of the Food and Drug Administration guidelines. The study demonstrated the applicability of metabolic phenotyping by ultra-performance liquid chromatography-triple quadruple-tandem mass spectrometry in a large-scale epidemiological study.
Statistical method development
Metabolomic data pose special challenges for statistical analysis, including high dimensionality, strong colinearity, nonlinear and highly complex spectral profiles, and the presence of structured and unstructured noise (due to within- and between-individual variability).64, 65, 66, 67, 68 Spectroscopic data are first preprocessed, including spectral alignment and normalization of the full-resolution spectral data. Multiple statistical strategies are then applied, including the use of both unsupervised and supervised multivariate data analysis techniques to extract information from the data.69 Principal component analysis is routinely used to identify the main sources of variation in the data and detect outlying values with the goal of providing the most compact representation of the data.64, 67, 68, 69 Hierarchical cluster analysis is another method that is widely used in exploratory data analysis to provide an overview of the data by grouping phenotypes according to their similarity, without assuming any prior knowledge of the data.64, 68, 69 Other techniques include partial least-squares (PLS), orthogonal partial least-squares (OPLS) and orthogonal partial least-squares discriminant analysis (OPLS-DA)70, 71, 72 aim to extract discriminatory metabolic signals from the data sets. They are often applied after the initial exploratory analysis, whereas statistical spectroscopy methods, accommodating the complex spectral data structure and correlations, such as statistical total correlation spectroscopy,73, 74 iterative-statistical total correlation spectroscopy,75 cluster analysis statistical spectroscopy76 and subset optimization by reference matching,77 are used to aid in the biomarker identification process. Statistical heterospectroscopy is a statistical strategy for the analysis of multiple spectroscopic data sets, for example, 1H NMR and ultra-performance liquid chromatography-mass spectrometry on the same samples. This method has enabled the characterization of drug metabolites (xenometabolome) in an epidemiological study.78 Other recent statistical spectroscopic methods include statistical homogeneous cluster spectroscopy79 and automatic spectroscopic data categorization by clustering analysis.80 These methods aim to remove the influence of irrelevant interferences within the data set to enhance the biomarker selection process79 and to differentiate reliably potential discriminatory markers from non-discriminatory markers in a biological data set,80 respectively. Figure 2 shows the steps of statistical analysis of INTERMAP NMR metabolic phenotyping data.13, 81, 82
In MWA studies, hundreds to thousands of biomarkers are assayed leading to data that are highly multivariate and colinear. To detect statistically significant relationships between molecular variables and phenotype, we defined the metabolome-wide significance level, a threshold required to control the family-wise error rate through a permutation approach.83 Using the spectra of the INTERMAP Chinese participants (N=836) as the reference population, we investigated the influence of spectral resolution and the number of variables in the NMR spectra, and we examined population heterogeneity by repeating the analysis in the INTERMAP US population samples (N=2164). The results showed that metabolome-wide significance level of 2 × 10−5 and 4 × 10−6 for a family-wise error rate of 0.05 and 0.01, respectively, could be used as a benchmark for NMR-based MWA studies of human urine. For the subsequent INTERMAP MWA studies, metabolome-wide significance level of 4.0 × 10−6 is used to identify significant metabolic features as a conservative estimate taking into account the high degree of colinearity in urinary NMR spectral data.81, 82 A similar approach may be used for MS-based MWA studies.
Novel biomarker discovery
Identification of unknown discriminatory metabolites is a key bottleneck in MWA studies as the human metabolome is still largely unknown. Elucidating the chemical structure of unknown metabolites is often labor-intensive and requires multiple analyses using a series of analytical experiments. Within the INTERMAP, we have been successful in the identification of a number of metabolites, including metabolites derived from dietary and drug intake.84, 85, 86, 87
Ethyl glucoside
The INTERMAP MWA study has discovered novel metabolites related to the intake of alcohol among the Chinese and Japanese population samples. From the 1H NMR urinary spectra, a doublet at δ4.93 corresponding to ethyl glucoside was observed, and it was derived following the ingestion of rice wine and sake.84 This doublet was not observed among the western population samples in the INTERMAP Study.
Proline betaine
We used metabolic phenotyping by 1H NMR spectroscopy to detect an increased excretion of proline betaine, tartaric acid and hippurate after fruit consumption compared with baseline diet in a dietary intervention study.85 We then measured the concentrations of proline betaine in selected fruit and commercially available fruit juices by 1H NMR spectroscopy optimized for quantification of this compound. All citrus fruit tested contained proline betaine; concentrations varied from 75 mg l−1 in orange squash to 1316 mg l−1 in orange juice from concentrate. After the consumption of 250 ml orange juice, we found a singlet peak at δ3.11, representing the CH3 moiety of proline betaine. Most proline betaine excretion occurred in the first 14 h after consumption, peaked at the 2-h postintervention urine collection and declined to almost baseline levels after 24 h.
The 24-h dietary recalls of INTERMAP UK participants were then assessed to validate the use of proline betaine excretion as a biomarker of citrus fruit consumption and as a surrogate marker of healthier eating patterns.85 Proline betaine excretion differed significantly between individuals with no recorded citrus consumption in their 24-h dietary recall data and individuals with recorded citrus consumption (P<0.0001). Those who reported citrus consumption and had higher levels of proline betaine excretion also showed a healthier nutrient profile, with higher intake of vegetable protein; lower intakes of total fat, trans fatty acids, cholesterol and animal protein; and lower urinary sodium–potassium (Na/K) ratio, BMI and systolic BP compared with non-citrus consumers. These findings provide proof that metabolic phenotyping can discover novel dietary biomarkers that can be used to validate dietary assessment in large-scale epidemiologic data sets.
Acetaminophen and ibuprofen
Although the metabolism of commonly used analgesics such as acetaminophen and ibuprofen has been widely described,40, 41, 88, 89 within the INTERMAP Study, we showed that we could detect urinary metabolite signatures related to commonly used analgesics such as acetaminophen and ibuprofen,86 enabling the use of metabolic signatures to verify the self-reported data.87 We applied principal component analysis to the 1H NMR spectra of US participants and identified 413 urine samples containing acetaminophen and its metabolites (acetaminophen users) and then applied OPLS-DA analysis to a subset of 70 urine samples from acetaminophen users and 70 urine samples from acetaminophen non-users.86 The OPLS-DA loading coefficient plot showed that differentiation between acetaminophen users vs. non-users primarily resulted from the presence of acetaminophen and its metabolites acetaminophen glucuronide, acetaminophen sulfate and the acetylcysteine conjugate of acetaminophen. Similarly, a principal component analysis model was constructed to identify urine samples of ibuprofen users, and OPLS-DA was performed on a subset of urine samples. The OPLS-DA loading coefficient plot showed that participants who had ingested ibuprofen were differentiated from non-users by the presence of 2-hydroxy, carboxy and glucuronide conjugates of ibuprofen in the urinary NMR spectra.
We then used these metabolic signatures to verify the self-reported analgesic use of INTERMAP particpants.87 Urinary spectra of UK (Belfast, N=216) and US (Chicago, N=280) participants were inspected for the presence or absence of metabolites in spectral regions containing acetaminophen and ibuprofen.87 These spectra were used to construct prediction models (sensitivity >98%) based on self-reported analgesic use. The overall rates of concordance between questionnaire data and urinary spectra were high for both populations: 83.8% (95% confidence interval: 78.9, 88.7) in Belfast and 81.1% (95% confidence interval: 76.5, 85.7) in Chicago. Overall rates of underdetection of acetaminophen and ibuprofen were low (~1%) and were comparable for both Belfast and Chicago. We then applied these prediction models to 9260 urine spectra to evaluate reported analgesic use from a self-report questionnaire. High-level concordance was observed between self-reported analgesic use and 1H NMR-detected urinary acetaminophen and/or ibuprofen metabolites for all Western population samples, an overall concordance of 70.5% (95% confidence interval: 68.7, 72.2). Our findings demonstrated the efficacy of an objective 1H NMR-based method for validation of self-reported data on analgesic use, detecting an underreporting rate of ~15% in the INTERMAP Study. This MWA approach has demonstrated the potential of metabolic phenotyping in reducing recall bias and other biases in epidemiologic studies for a range of substances, including pharmaceuticals, dietary supplements and foods.
INTERMAP MWA study
INTERMAP is the first large-scale human population MWA study on diet and BP, using an exploratory analytical approach to investigate metabolic phenotype variation across and within 17 population samples in the East (China and Japan) and West (United Kingdom and United States) based on 1H NMR spectroscopy.13 Using a hierarchical clustering algorithm, we investigated the similarity/dissimilarity between populations based on their urinary profiles. East Asian and Western populations had well-differentiated metabolic phenotypes (Figure 3). Among the East Asian samples, Japan was differentiated from China and within China, North China was differentiated from the South despite a similar genetic background. Using OPLS-DA,70, 73 we reported significant differences among the metabolic profiles of these populations; discriminatory metabolites included gut microbial–host cometabolites (hippurate, phenylacetylglutamine and methylamines), amino acids (alanine, lysine and taurine), dietary-related metabolites (e.g., ethyl glucoside and trimethylamine-N-oxide), compounds related to energy metabolism (acetylcarnitine) and tricarboxylic acid cycle intermediates (succinate and citrate). Four discriminatory metabolites reflecting diet and gut microbial activities—alanine, formate, hippurate and N-methylnicotinate—were then quantified from the 1H NMR urinary spectral profiles. We found that alanine was highly correlated with 2-oxoglutarate (metabolic linkage via glutamate-pyruvate transaminase activity) and with formate (pyruvate/Co-A metabolism), and hippurate was highly correlated with N-methylnicotinate (renal transporter/secretion mechanisms). In multiple linear regression models, both formate and hippurate were inversely associated with systolic and diastolic BP, and alanine was positively associated with BP.
Hierarchical cluster analysis (HCA) of proton nuclear magnetic resonance (1H NMR) urine spectra, the INTERnational study of MAcro/micronutrients and blood Pressure (INTERMAP) Study.13 The HCA algorithm produces a dendrogram showing the overall similarity/dissimilarity between population samples. Similarity index is normalized to intercluster distance. Each branch of the dendrogram defines a subcluster; population samples within subclusters are more similar to each other than to those in other subclusters. The dendrogram shows clustering based on country and geographical location or gender. A full color version of this figure is available at the Hypertension Research journal online.
More detailed analysis was later performed using the Chinese population samples.81 We found that urinary metabolites were significantly different between the northern and southern Chinese, reflecting variations in dietary patterns as well as CVD risk between these two populations. Those that were higher in northern than southern Chinese populations included dimethylglycine, alanine, lactate, branched-chain amino acids (isoleucine, leucine, valine), N-acetyls of glycoprotein fragments (including uromodulin), N-acetyl neuraminic acid, pentanoic/heptanoic acid and methylguanidine; metabolites that were significantly higher in the south included gut microbial–host cometabolites (hippurate, 4-cresyl sulfate, phenylacetylglutamine, 2-hydroxyisobutyrate), succinate, creatine, scyllo-inositol, proline betaine and trans-aconitate. Compared with the south, northern Chinese had higher BMI, less favorable diet including lower Ca, Mg and P intakes, higher 24-h urinary Na excretion, higher urinary sodium–potassium ratio of excretion and higher BP (Table 1).90 The significant north–south differences in BP, BMI and diet90, 91, 92 are reflected in geographic variations in both CVD incidence and mortality rates, with higher rates in the north than the south.93, 94, 95 The INTERMAP MWA study indicates the likely importance of environmental influences (e.g., diet), endogenous metabolism and mammalian–gut microbial cometabolism in helping to explain north–south differences in Chinese CVD risk.
The INTERMAP Study confirmed that African Americans (AA) had higher systolic and diastolic BP compared with non-Hispanic white Americans (NHWAs),82 and this BP difference was due, in part, to less favorable multiple nutrient intake by AA, with lower intakes of fruits, vegetables and dairy products and lower intakes of vegetable protein, starch, fiber, K, Ca, Mg and P, compared with those of whites. In addition, there was greater obesity prevalence among black women compared with white women.82, 96 1H NMR spectra of the INTERMAP US participants showed that urinary metabolites that were significantly higher in AA compared with that in NHWA included creatinine, 3-hydroxyisovalerate, N-acetyls of glycoprotein fragments, dimethylglycine, lysine, N-acetyl neuraminic acid, leucine, dimethylamine, taurine and 2-hydroxy-isobutyrate; metabolites significantly higher in NHWA included trimethylamine, N-methylnicotinate, hippurate and succinate (Figure 4).82 The mean values of urinary hippurate (2.9 mmol per 24 h for AA men vs. 4.1 mmol per 24 h for NHWA men, P<5 × 10−9) and N-methylnicotinate (0.24 mmol per 24 h for AA men vs. 0.42 mmol per 24 h for NHWA men, P<3 × 10−10) were significantly lower in AA compared with NHWA. Multiple linear regression was used to examine these AA-NHWA differences in dietary and urinary metabolites in relation to BP; multiple foods, nutrients and metabolites accounted for part of the higher BP among AA.
The median urinary proton nuclear magnetic resonance (1H NMR) spectrum of African Americans (AAs) and non-Hispanic white Americans (NHWAs).82 Top: median urinary 1H NMR spectrum of INTERnational study of MAcro/micronutrients and blood Pressure (INTERMAP) US AA and NHWA participants, based on the first urine collection (N=1455). Bottom: Manhattan plot indicating the significant spectral variables. Metabolites higher in AA individuals compared with NHWA individuals are shown in red, and metabolites higher in NHWA individuals compared with AA individuals are shown in blue. 1, leucine; 2, 3-hydroxyisovalerate; 3, 2-hydroxyisobutyrate; 4, N-acetyls of glycoprotein fragments; 5, N-acetyl neuraminic acid; 6, succinate; 7, dimethylamine; 8, trimethylamine; 9, dimethylglycine; 10, lysine; 11, creatinine; 12, hippurate; 13, N-methyl nicotinic acid.
Summary
In recent years, major advances have been made in the metabolic phenotyping of epidemiological samples. The advancement in the analytical techniques and the development of new statistical data analysis tools have enabled the identification of novel metabolic phenotypes associated with diet (including Na intake), xenobiotics and BP. The findings of the INTERMAP MWA study may provide insights into molecular pathways underlying complex biological processes such as adverse BP levels. We envisage that future studies will include the generation of testable hypotheses based on the findings from the INTERMAP MWA study. Moreover, the increasing number of population-based cohort studies, which also apply the MWA approach, will undoubtedly contribute to our understanding of the key mechanisms that are associated with CVD. As noted above, metabolomic data with hundreds to thousands of biomarkers being assayed are highly multivariate, colinear and noisy, with the potential for false-positive findings. It is also always a possibility but unlikely that phenotypes are specific to INTERMAP Study populations and not generalizable; replication studies are needed. Nonetheless, it is reasonable to state at this juncture that INTERMAP findings to date have demonstrated significant independent relationships of several nutrients/foods/eating patterns/metabolites to BP, thereby moving the field forward in exciting and unprecedented ways.
References
Van den Hoogen PC, Feskens EJ, Nagelkerke NJ, Menotti A, Nissinen A, Kromhout D . The relation between blood pressure and mortality due to coronary heart disease among men in different parts of the world. Seven countries study research group. N Engl J Med 2000; 342: 1–8.
Lim SS, Vos T, Flaxman AD, Danaei G, Shibuya K, Adair-Rohani H, Amann M, Anderson HR, Andrews KG, Aryee M, Atkinson C, Bacchus LJ, Bahalim AN, Balakrishnan K, Balmes J, Barker-Collo S, Baxter A, Bell ML, Blore JD, Blyth F, Bonner C, Borges G, Bourne R, Boussinesq M, Brauer M, Brooks P, Bruce NG, Brunekreef B, Bryan-Hancock C, Bucello C, Buchbinder R, Bull F, Burnett RT, Byers TE, Calabria B, Carapetis J, Carnahan E, Chafe Z, Charlson F, Chen H, Chen JS, Cheng AT, Child JC, Cohen A, Colson KE, Cowie BC, Darby S, Darling S, Davis A, Degenhardt L, Dentener F, Des Jarlais DC, Devries K, Dherani M, Ding EL, Dorsey ER, Driscoll T, Edmond K, Ali SE, Engell RE, Erwin PJ, Fahimi S, Falder G, Farzadfar F, Ferrari A, Finucane MM, Flaxman S, Fowkes FG, Freedman G, Freeman MK, Gakidou E, Ghosh S, Giovannucci E, Gmel G, Graham K, Grainger R, Grant B, Gunnell D, Gutierrez HR, Hall W, Hoek HW, Hogan A, Hosgood HD, Hoy D, Hu H, Hubbell BJ, Hutchings SJ, Ibeanusi SE, Jacklyn GL, Jasrasaria R, Jonas JB, Kan H, Kanis JA, Kassebaum N, Kawakami N, Khang YH, Khatibzadeh S, Khoo JP, Kok C, Laden F, Lalloo R, Lan Q, Lathlean T, Leasher JL, Leigh J, Li Y, Lin JK, Lipshultz SE, London S, Lozano R, Lu Y, Mak J, Malekzadeh R, Mallinger L, Marcenes W, March L, Marks R, Martin R, McGale P, McGrath J, Mehta S, Mensah GA, Merriman TR, Micha R, Michaud C, Mishra V, Mohd Hanafiah K, Mokdad AA, Morawska L, Mozaffarian D, Murphy T, Naghavi M, Neal B, Nelson PK, Nolla JM, Norman R, Olives C, Omer SB, Orchard J, Osborne R, Ostro B, Page A, Pandey KD, Parry CD, Passmore E, Patra J, Pearce N, Pelizzari PM, Petzold M, Phillips MR, Pope D, Pope CA III, Powles J, Rao M, Razavi H, Rehfuess EA, Rehm JT, Ritz B, Rivara FP, Roberts T, Robinson C, Rodriguez-Portales JA, Romieu I, Room R, Rosenfeld LC, Roy A, Rushton L, Salomon JA, Sampson U, Sanchez-Riera L, Sanman E, Sapkota A, Seedat S, Shi P, Shield K, Shivakoti R, Singh GM, Sleet DA, Smith E, Smith KR, Stapelberg NJ, Steenland K, Stockl H, Stovner LJ, Straif K, Straney L, Thurston GD, Tran JH, Van Dingenen R, van Donkelaar A, Veerman JL, Vijayakumar L, Weintraub R, Weissman MM, White RA, Whiteford H, Wiersma ST, Wilkinson JD, Williams HC, Williams W, Wilson N, Woolf AD, Yip P, Zielinski JM, Lopez AD, Murray CJ, Ezzati M, AlMazroa MA, Memish ZA . A comparative risk assessment of burden of disease and injury attributable to 67 risk factors and risk factor clusters in 21 regions, 1990–2010: a systematic analysis for the global burden of disease study 2010. Lancet 2012; 380: 2224–2260.
Ezzati M, Lopez AD, Rodgers A, Vander Hoorn S, Murray CJ, Comparative Risk Assessment Collaborating G.. Selected major risk factors and global and regional burden of disease. Lancet 2002; 360: 1347–1360.
World Health Organisation. Global Health Observatory (GHO) Data: Raise Blood Pressure. Available at: http://www.who.int/gho/ncd/risk_factors/blood_pressure_prevalence_text/en/ (last accessed 1 October 2016).
Levy D, Ehret GB, Rice K, Verwoert GC, Launer LJ, Dehghan A, Glazer NL, Morrison AC, Johnson AD, Aspelund T, Aulchenko Y, Lumley T, Kottgen A, Vasan RS, Rivadeneira F, Eiriksdottir G, Guo X, Arking DE, Mitchell GF, Mattace-Raso FU, Smith AV, Taylor K, Scharpf RB, Hwang SJ, Sijbrands EJ, Bis J, Harris TB, Ganesh SK, O'Donnell CJ, Hofman A, Rotter JI, Coresh J, Benjamin EJ, Uitterlinden AG, Heiss G, Fox CS, Witteman JC, Boerwinkle E, Wang TJ, Gudnason V, Larson MG, Chakravarti A, Psaty BM, van Duijn CM . Genome-wide association study of blood pressure and hypertension. Nat Genet 2009; 41: 677–687.
International Consortium for Blood Pressure Genome-Wide Association S, Ehret GB, Munroe PB, Rice KM, Bochud M, Johnson AD, Chasman DI, Smith AV, Tobin MD, Verwoert GC, Hwang SJ, Pihur V, Vollenweider P, O'Reilly PF, Amin N, Bragg-Gresham JL, Teumer A, Glazer NL, Launer L, Zhao JH, Aulchenko Y, Heath S, Sober S, Parsa A, Luan J, Arora P, Dehghan A, Zhang F, Lucas G, Hicks AA, Jackson AU, Peden JF, Tanaka T, Wild SH, Rudan I, Igl W, Milaneschi Y, Parker AN, Fava C, Chambers JC, Fox ER, Kumari M, Go MJ, van der Harst P, Kao WH, Sjogren M, Vinay DG, Alexander M, Tabara Y, Shaw-Hawkins S, Whincup PH, Liu Y, Shi G, Kuusisto J, Tayo B, Seielstad M, Sim X, Nguyen KD, Lehtimaki T, Matullo G, Wu Y, Gaunt TR, Onland-Moret NC, Cooper MN, Platou CG, Org E, Hardy R, Dahgam S, Palmen J, Vitart V, Braund PS, Kuznetsova T, Uiterwaal CS, Adeyemo A, Palmas W, Campbell H, Ludwig B, Tomaszewski M, Tzoulaki I, Palmer ND, Consortium CA, Consortium CK, KidneyGen C, EchoGen C, Consortium C-H, Aspelund T, Garcia M, Chang YP, O'Connell JR, Steinle NI, Grobbee DE, Arking DE, Kardia SL, Morrison AC, Hernandez D, Najjar S, McArdle WL, Hadley D, Brown MJ, Connell JM, Hingorani AD, Day IN, Lawlor DA, Beilby JP, Lawrence RW, Clarke R, Hopewell JC, Ongen H, Dreisbach AW, Li Y, Young JH, Bis JC, Kahonen M, Viikari J, Adair LS, Lee NR, Chen MH, Olden M, Pattaro C, Bolton JA, Kottgen A, Bergmann S, Mooser V, Chaturvedi N, Frayling TM, Islam M, Jafar TH, Erdmann J, Kulkarni SR, Bornstein SR, Grassler J, Groop L, Voight BF, Kettunen J, Howard P, Taylor A, Guarrera S, Ricceri F, Emilsson V, Plump A, Barroso I, Khaw KT, Weder AB, Hunt SC, Sun YV, Bergman RN, Collins FS, Bonnycastle LL, Scott LJ, Stringham HM, Peltonen L, Perola M, Vartiainen E, Brand SM, Staessen JA, Wang TJ, Burton PR, Soler Artigas M, Dong Y, Snieder H, Wang X, Zhu H, Lohman KK, Rudock ME, Heckbert SR, Smith NL, Wiggins KL, Doumatey A, Shriner D, Veldre G, Viigimaa M, Kinra S, Prabhakaran D, Tripathy V, Langefeld CD, Rosengren A, Thelle DS, Corsi AM, Singleton A, Forrester T, Hilton G, McKenzie CA, Salako T, Iwai N, Kita Y, Ogihara T, Ohkubo T, Okamura T, Ueshima H, Umemura S, Eyheramendy S, Meitinger T, Wichmann HE, Cho YS, Kim HL, Lee JY, Scott J, Sehmi JS, Zhang W, Hedblad B, Nilsson P, Smith GD, Wong A, Narisu N, Stancakova A, Raffel LJ, Yao J, Kathiresan S, O'Donnell CJ, Schwartz SM, Ikram MA, Longstreth WT Jr, Mosley TH, Seshadri S, Shrine NR, Wain LV, Morken MA, Swift AJ, Laitinen J, Prokopenko I, Zitting P, Cooper JA, Humphries SE, Danesh J, Rasheed A, Goel A, Hamsten A, Watkins H, Bakker SJ, van Gilst WH, Janipalli CS, Mani KR, Yajnik CS, Hofman A, Mattace-Raso FU, Oostra BA, Demirkan A, Isaacs A, Rivadeneira F, Lakatta EG, Orru M, Scuteri A, Ala-Korpela M, Kangas AJ, Lyytikainen LP, Soininen P, Tukiainen T, Wurtz P, Ong RT, Dorr M, Kroemer HK, Volker U, Volzke H, Galan P, Hercberg S, Lathrop M, Zelenika D, Deloukas P, Mangino M, Spector TD, Zhai G, Meschia JF, Nalls MA, Sharma P, Terzic J, Kumar MV, Denniff M, Zukowska-Szczechowska E, Wagenknecht LE, Fowkes FG, Charchar FJ, Schwarz PE, Hayward C, Guo X, Rotimi C, Bots ML, Brand E, Samani NJ, Polasek O, Talmud PJ, Nyberg F, Kuh D, Laan M, Hveem K, Palmer LJ, van der Schouw YT, Casas JP, Mohlke KL, Vineis P, Raitakari O, Ganesh SK, Wong TY, Tai ES, Cooper RS, Laakso M, Rao DC, Harris TB, Morris RW, Dominiczak AF, Kivimaki M, Marmot MG, Miki T, Saleheen D, Chandak GR, Coresh J, Navis G, Salomaa V, Han BG, Zhu X, Kooner JS, Melander O, Ridker PM, Bandinelli S, Gyllensten UB, Wright AF, Wilson JF, Ferrucci L, Farrall M, Tuomilehto J, Pramstaller PP, Elosua R, Soranzo N, Sijbrands EJ, Altshuler D, Loos RJ, Shuldiner AR, Gieger C, Meneton P, Uitterlinden AG, Wareham NJ, Gudnason V, Rotter JI, Rettig R, Uda M, Strachan DP, Witteman JC, Hartikainen AL, Beckmann JS, Boerwinkle E, Vasan RS, Boehnke M, Larson MG, Jarvelin MR, Psaty BM, Abecasis GR, Chakravarti A, Elliott P, van Duijn CM, Newton-Cheh C, Levy D, Caulfield MJ, Johnson T . Genetic variants in novel pathways influence blood pressure and cardiovascular disease risk. Nature 2011; 478: 103–109.
Ganesh SK, Tragante V, Guo W, Guo Y, Lanktree MB, Smith EN, Johnson T, Castillo BA, Barnard J, Baumert J, Chang YP, Elbers CC, Farrall M, Fischer ME, Franceschini N, Gaunt TR, Gho JM, Gieger C, Gong Y, Isaacs A, Kleber ME, Mateo Leach I, McDonough CW, Meijs MF, Mellander O, Molony CM, Nolte IM, Padmanabhan S, Price TS, Rajagopalan R, Shaffer J, Shah S, Shen H, Soranzo N, van der Most PJ, Van Iperen EP, Van Setten J, Vonk JM, Zhang L, Beitelshees AL, Berenson GS, Bhatt DL, Boer JM, Boerwinkle E, Burkley B, Burt A, Chakravarti A, Chen W, Cooper-Dehoff RM, Curtis SP, Dreisbach A, Duggan D, Ehret GB, Fabsitz RR, Fornage M, Fox E, Furlong CE, Gansevoort RT, Hofker MH, Hovingh GK, Kirkland SA, Kottke-Marchant K, Kutlar A, Lacroix AZ, Langaee TY, Li YR, Lin H, Liu K, Maiwald S, Malik R, Cardiogram M, Murugesan G, Newton-Cheh C, O'Connell JR, Onland-Moret NC, Ouwehand WH, Palmas W, Penninx BW, Pepine CJ, Pettinger M, Polak JF, Ramachandran VS, Ranchalis J, Redline S, Ridker PM, Rose LM, Scharnag H, Schork NJ, Shimbo D, Shuldiner AR, Srinivasan SR, Stolk RP, Taylor HA, Thorand B, Trip MD, van Duijn CM, Verschuren WM, Wijmenga C, Winkelmann BR, Wyatt S, Young JH, Boehm BO, Caulfield MJ, Chasman DI, Davidson KW, Doevendans PA, Fitzgerald GA, Gums JG, Hakonarson H, Hillege HL, Illig T, Jarvik GP, Johnson JA, Kastelein JJ, Koenig W, LifeLines Cohort S, Marz W, Mitchell BD, Murray SS, Oldehinkel AJ, Rader DJ, Reilly MP, Reiner AP, Schadt EE, Silverstein RL, Snieder H, Stanton AV, Uitterlinden AG, van der Harst P, van der Schouw YT, Samani NJ, Johnson AD, Munroe PB, de Bakker PI, Zhu X, Levy D, Keating BJ, Asselbergs FW . Loci influencing blood pressure identified using a cardiovascular gene-centric array. Hum Mol Genet 2013; 22: 1663–1678.
Huan T, Esko T, Peters MJ, Pilling LC, Schramm K, Schurmann C, Chen BH, Liu C, Joehanes R, Johnson AD, Yao C, Ying SX, Courchesne P, Milani L, Raghavachari N, Wang R, Liu P, Reinmaa E, Dehghan A, Hofman A, Uitterlinden AG, Hernandez DG, Bandinelli S, Singleton A, Melzer D, Metspalu A, Carstensen M, Grallert H, Herder C, Meitinger T, Peters A, Roden M, Waldenberger M, Dorr M, Felix SB, Zeller T, International Consortium for Blood Pressure G, Vasan R, O'Donnell CJ, Munson PJ, Yang X, Prokisch H, Volker U, van Meurs JB, Ferrucci L, Levy D . A meta-analysis of gene expression signatures of blood pressure and hypertension. PLoS Genet 2015; 11: e1005035.
Ehret GB, Munroe PB, Rice KM, Bochud M, Johnson AD, Chasman DI, Smith AV, Tobin MD, Verwoert GC, Hwang SJ, Pihur V, Vollenweider P, O'Reilly PF, Amin N, Bragg-Gresham JL, Teumer A, Glazer NL, Launer L, Zhao JH, Aulchenko Y, Heath S, Sober S, Parsa A, Luan J, Arora P, Dehghan A, Zhang F, Lucas G, Hicks AA, Jackson AU, Peden JF, Tanaka T, Wild SH, Rudan I, Igl W, Milaneschi Y, Parker AN, Fava C, Chambers JC, Fox ER, Kumari M, Go MJ, van der Harst P, Kao WH, Sjogren M, Vinay DG, Alexander M, Tabara Y, Shaw-Hawkins S, Whincup PH, Liu Y, Shi G, Kuusisto J, Tayo B, Seielstad M, Sim X, Nguyen KD, Lehtimaki T, Matullo G, Wu Y, Gaunt TR, Onland-Moret NC, Cooper MN, Platou CG, Org E, Hardy R, Dahgam S, Palmen J, Vitart V, Braund PS, Kuznetsova T, Uiterwaal CS, Adeyemo A, Palmas W, Campbell H, Ludwig B, Tomaszewski M, Tzoulaki I, Palmer ND, Aspelund T, Garcia M, Chang YP, O'Connell JR, Steinle NI, Grobbee DE, Arking DE, Kardia SL, Morrison AC, Hernandez D, Najjar S, McArdle WL, Hadley D, Brown MJ, Connell JM, Hingorani AD, Day IN, Lawlor DA, Beilby JP, Lawrence RW, Clarke R, Hopewell JC, Ongen H, Dreisbach AW, Li Y, Young JH, Bis JC, Kahonen M, Viikari J, Adair LS, Lee NR, Chen MH, Olden M, Pattaro C, Bolton JA, Kottgen A, Bergmann S, Mooser V, Chaturvedi N, Frayling TM, Islam M, Jafar TH, Erdmann J, Kulkarni SR, Bornstein SR, Grassler J, Groop L, Voight BF, Kettunen J, Howard P, Taylor A, Guarrera S, Ricceri F, Emilsson V, Plump A, Barroso I, Khaw KT, Weder AB, Hunt SC, Sun YV, Bergman RN, Collins FS, Bonnycastle LL, Scott LJ, Stringham HM, Peltonen L, Perola M, Vartiainen E, Brand SM, Staessen JA, Wang TJ, Burton PR, Soler Artigas M, Dong Y, Snieder H, Wang X, Zhu H, Lohman KK, Rudock ME, Heckbert SR, Smith NL, Wiggins KL, Doumatey A, Shriner D, Veldre G, Viigimaa M, Kinra S, Prabhakaran D, Tripathy V, Langefeld CD, Rosengren A, Thelle DS, Corsi AM, Singleton A, Forrester T, Hilton G, McKenzie CA, Salako T, Iwai N, Kita Y, Ogihara T, Ohkubo T, Okamura T, Ueshima H, Umemura S, Eyheramendy S, Meitinger T, Wichmann HE, Cho YS, Kim HL, Lee JY, Scott J, Sehmi JS, Zhang W, Hedblad B, Nilsson P, Smith GD, Wong A, Narisu N, Stancakova A, Raffel LJ, Yao J, Kathiresan S, O'Donnell CJ, Schwartz SM, Ikram MA, Longstreth WT Jr., Mosley TH, Seshadri S, Shrine NR, Wain LV, Morken MA, Swift AJ, Laitinen J, Prokopenko I, Zitting P, Cooper JA, Humphries SE, Danesh J, Rasheed A, Goel A, Hamsten A, Watkins H, Bakker SJ, van Gilst WH, Janipalli CS, Mani KR, Yajnik CS, Hofman A, Mattace-Raso FU, Oostra BA, Demirkan A, Isaacs A, Rivadeneira F, Lakatta EG, Orru M, Scuteri A, Ala-Korpela M, Kangas AJ, Lyytikainen LP, Soininen P, Tukiainen T, Wurtz P, Ong RT, Dorr M, Kroemer HK, Volker U, Volzke H, Galan P, Hercberg S, Lathrop M, Zelenika D, Deloukas P, Mangino M, Spector TD, Zhai G, Meschia JF, Nalls MA, Sharma P, Terzic J, Kumar MV, Denniff M, Zukowska-Szczechowska E, Wagenknecht LE, Fowkes FG, Charchar FJ, Schwarz PE, Hayward C, Guo X, Rotimi C, Bots ML, Brand E, Samani NJ, Polasek O, Talmud PJ, Nyberg F, Kuh D, Laan M, Hveem K, Palmer LJ, van der Schouw YT, Casas JP, Mohlke KL, Vineis P, Raitakari O, Ganesh SK, Wong TY, Tai ES, Cooper RS, Laakso M, Rao DC, Harris TB, Morris RW, Dominiczak AF, Kivimaki M, Marmot MG, Miki T, Saleheen D, Chandak GR, Coresh J, Navis G, Salomaa V, Han BG, Zhu X, Kooner JS, Melander O, Ridker PM, Bandinelli S, Gyllensten UB, Wright AF, Wilson JF, Ferrucci L, Farrall M, Tuomilehto J, Pramstaller PP, Elosua R, Soranzo N, Sijbrands EJ, Altshuler D, Loos RJ, Shuldiner AR, Gieger C, Meneton P, Uitterlinden AG, Wareham NJ, Gudnason V, Rotter JI, Rettig R, Uda M, Strachan DP, Witteman JC, Hartikainen AL, Beckmann JS, Boerwinkle E, Vasan RS, Boehnke M, Larson MG, Jarvelin MR, Psaty BM, Abecasis GR, Chakravarti A, Elliott P, van Duijn CM, Newton-Cheh C, Levy D, Caulfield MJ, Johnson T . Genetic variants in novel pathways influence blood pressure and cardiovascular disease risk. Nature 2011; 478: 103–109.
Gillespie CD, Hurvitz KA,, Centers for Disease C Prevention.. Prevalence of hypertension and controlled hypertension—United States, 2007–2010. Morb Mort Wkly Rep Surveill Summ 2013; 62 (Suppl 3): 144–148.
van den Berg N, Meinke-Franze C, Fiss T, Baumeister SE, Hoffmann W . Prevalence and determinants of controlled hypertension in a German population cohort. BMC Public Health 2013; 13: 594.
Stamler J, Elliott P, Dennis B, Dyer AR, Kesteloot H, Liu K, Ueshima H, Zhou BF . INTERMAP: background, aims, design, methods, and descriptive statistics (nondietary). J Hum Hypertens 2003; 17: 591–608.
Holmes E, Loo RL, Stamler J, Bictash M, Yap IKS, Chan Q, Ebbels T, De Iorio M, Brown IJ, Veselkov KA, Daviglus ML, Kesteloot H, Ueshima H, Zhao L, Nicholson JK, Elliott P . Human metabolic phenotype diversity and its association with diet and blood pressure. Nature 2008; 453: 396–U350.
Dennis B, Stamler J, Buzzard M, Conway R, Elliott P, Moag-Stahlberg A, Okayama A, Okuda N, Robertson C, Robinson F, Schakel S, Stevens M, Van HN, Zhao L, Zhou BF . INTERMAP: the dietary data—process and quality control. J Hum Hypertens 2003; 17: 609–622.
Chan Q, Stamler J, Griep LM, Daviglus ML, Horn LV, Elliott P . An update on nutrients and blood pressure. J Atherosclerosis Thromb 2015; 23: 276–289.
Elliott P, Stamler J, Dyer AR, Appel L, Dennis B, Kesteloot H, Ueshima H, Okayama A, Chan Q, Garside DB, Zhou B . Association between protein intake and blood pressure: the INTERMAP study. Archiv Intern Med 2006; 166: 79–87.
Stamler J, Brown IJ, Daviglus ML, Chan Q, Kesteloot H, Ueshima H, Zhao L, Elliott P . Glutamic acid, the main dietary amino acid, and blood pressure: the INTERMAP study (international collaborative study of macronutrients, micronutrients and blood pressure). Circulation 2009; 120: 221–228.
Aljuraiban GS, Griep LM, Chan Q, Daviglus ML, Stamler J, Van Horn L, Elliott P, Frost GS . Total, insoluble and soluble dietary fibre intake in relation to blood pressure: the INTERMAP study. Br J Nutr 2015; 114: 1480–1486.
Chan Q, Stamler J, Brown IJ, Daviglus ML, Van Horn L, Dyer AR, Oude Griep LM, Miura K, Ueshima H, Zhao L, Nicholson JK, Holmes E, Elliott P . Relation of raw and cooked vegetable consumption to blood pressure: the INTERMAP study. J Hum Hypertens 2014; 28: 353–359.
Miura K, Stamler J, Nakagawa H, Elliott P, Ueshima H, Chan Q, Brown IJ, Tzoulaki I, Saitoh S, Dyer AR, Daviglus ML, Kesteloot H, Okayama A, Curb JD, Rodriguez BL, Elmer PJ, Steffen LM, Robertson C, Zhao L . Relationship of dietary linoleic acid to blood pressure—the international study of macro-micronutrients and blood pressure study. Hypertension 2008; 52: 408–414.
Miura K, Stamler J, Brown IJ, Ueshima H, Nakagawa H, Sakurai M, Chan Q, Appel LJ, Okayama A, Okuda N, Curb JD, Rodriguez BL, Robertson C, Zhao L, Elliott P . Relationship of dietary monounsaturated fatty acids to blood pressure: the international study of macro/micronutrients and blood pressure. J Hypertens 2013; 31: 1144–1150.
Ueshima H, Stamler J, Elliott P, Chan Q, Brown IJ, Carnethon MR, Daviglus ML, He K, Moag-Stahlberg A, Rodriguez BL, Steffen LM, Van Horn L, Yarnell J, Zhou B . Food omega-3 fatty acid intake of individuals (total, linolenic acid, long-chain) and their blood pressure INTERMAP study. Hypertension 2007; 50: 313–319.
Elliott P, Kesteloot H, Appel LJ, Dyer AR, Ueshima H, Chan Q, Brown IJ, Zhao L, Stamler J . Dietary phosphorus and blood pressure: international study of macro- and micro-nutrients and blood pressure. Hypertension 2008; 51: 669–675.
Tzoulaki I, Brown IJ, Chan Q, Van Horn L, Ueshima H, Zhao L, Stamler J, Elliott P . Relation of iron and red meat intake to blood pressure: cross sectional epidemiological study. BMJ 2008; 337: a258.
Brown IJ, Elliott P, Robertson CE, Chan Q, Daviglus ML, Dyer AR, Huang CC, Rodriguez BL, Sakata K, Ueshima H, Van HL, Zhao L, Stamler J . Dietary starch intake of individuals and their blood pressure: the international study of macronutrients and micronutrients and blood pressure. J Hypertens 2009; 27: 231–236.
Brown IJ, Stamler J, Van Horn L, Robertson CE, Chan Q, Dyer AR, Huang CC, Rodriguez BL, Zhao L, Daviglus ML, Ueshima H, Elliott P . Sugar-sweetened beverage, sugar intake of individuals, and their blood pressure: international study of macro/micronutrients and blood pressure. Hypertension 2011; 57: 695–701.
Sakurai M, Stamler J, Miura K, Brown IJ, Nakagawa H, Elliott P, Ueshima H, Chan Q, Tzoulaki I, Dyer AR, Okayama A, Zhao L . Relationship of dietary cholesterol to blood pressure: the INTERMAP study. J Hypertens 2011; 29: 222–228.
Stamler J, Brown IJ, Daviglus ML, Chan QEN, Miura K, Okuda N, Ueshima H, Zhao LC, Elliott P . Dietary glycine and blood pressure: the international study on macro/micronutrients and blood pressure. Am J Clin Nutr 2013; 98: 136–145.
Oude Griep LM, Stamler J, Chan Q, Van Horn L, Steffen LM, Miura K, Ueshima H, Okuda N, Zhao L, Daviglus ML, Elliott P . Association of raw fruit and fruit juice consumption with blood pressure: the INTERMAP study. Am J Clin Nutr 2013; 97: 1083–1091.
Campanella GCQ, Daviglus ML, Van Horn L, Miura K, Ueshima H, Zhao L, Stamler J, Nicholson JK, Elliott P, Chadeau-Hyam M . Relation of diet-induced metabolic acidosis to blood pressure. Circulation 2014; 129: AP179.
Park M, Jung SJ, Yoon S, Yun JM, Yoon HJ . Association between the markers of metabolic acid load and higher all-cause and cardiovascular mortality in a general population with preserved renal function. Hypertens Res 2015; 38: 433–438.
Kesteloot H, Tzoulaki I, Brown IJ, Chan Q, Wijeyesekera A, Ueshima H, Zhao L, Dyer AR, Unwin RJ, Stamler J, Elliott P . Relation of urinary calcium and magnesium excretion to blood pressure: the international study of macro- and micro-nutrients and blood pressure and the international cooperative study on salt, other factors, and blood pressure. Am J Epidemiol 2011; 174: 44–51.
Yamori Y, Sagara M, Mizushima S, Liu LJ, Ikeda K, Nara Y, Grp CS . An inverse association between magnesium in 24-h urine and cardiovascular risk factors in middle-aged subjects in 50 cardiac study populations. Hypertens Res 2015; 38: 219–225.
Laragh JH, Baer L, Brunner HR, Buhler FR, Vaughan JE . Renin, angiotensin and aldosterone system in pathogenesis and management of hypertensive vascular disease. Am J Med 1972; 52: 633–652.
Weir MR, Dzau VJ . The renin–angiotensin–aldosterone system: a specific target for hypertension management. Am J Hypertens 1999; 12: 205S–213S.
Manrique C, Lastra G, Gardner M, Sowers JR . The renin–angiotensin–aldosterone system in hypertension: roles of insulin resistance and oxidative stress. Med Clin N Am 2009; 93: 569–582.
Grassi G . Role of the sympathetic nervous system in human hypertension. J Hypertens 1998; 16: 1979–1987.
DiBona GF . Sympathetic nervous system and the kidney in hypertension. Curr Opin Nephrol Hypertens 2002; 11: 197–200.
Burton AC . Relation of structure to function of the tissues of the wall of blood vessels. Physiol Rev 1954; 34: 619–642.
Nicholson JK, Wilson ID . High-resolution proton magnetic-resonance spectroscopy of biological-fluids. Prog Nucl Mag Res Sp 1989; 21: 449–501.
Lindon JC, Holmes E, Nicholson JK . Toxicological applications of magnetic resonance. Prog Nucl Mag Res Sp 2004; 45: 109–143.
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI . An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006; 444: 1027–1031.
Qin J, Li Y, Cai Z, Li S, Zhu J, Zhang F, Liang S, Zhang W, Guan Y, Shen D, Peng Y, Zhang D, Jie Z, Wu W, Qin Y, Xue W, Li J, Han L, Lu D, Wu P, Dai Y, Sun X, Li Z, Tang A, Zhong S, Li X, Chen W, Xu R, Wang M, Feng Q, Gong M, Yu J, Zhang Y, Zhang M, Hansen T, Sanchez G, Raes J, Falony G, Okuda S, Almeida M, LeChatelier E, Renault P, Pons N, Batto JM, Zhang Z, Chen H, Yang R, Zheng W, Li S, Yang H, Wang J, Ehrlich SD, Nielsen R, Pedersen O, Kristiansen K, Wang J . A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 2012; 490: 55–60.
Mazidi M, Rezaie P, Kengne AP, Mobarhan MG, Ferns GA . Gut microbiome and metabolic syndrome. Diabetes Metab Syndr 2016; 10: S150–S157.
Mell B, Jala VR, Mathew AV, Byun J, Waghulde H, Zhang Y, Haribabu B, Vijay-Kumar M, Pennathur S, Joe B . Evidence for a link between gut microbiota and hypertension in the dahl rat. Physiol Genom 2015; 47: 187–197.
Tang WHW, Wang ZN, Kennedy DJ, Wu YP, Buffa JA, Agatisa-Boyle B, Li XMS, Levison BS, Hazen SL . Gut microbiota-dependent trimethylamine n-oxide (tmao) pathway contributes to both development of renal insufficiency and mortality risk in chronic kidney disease. Circ Res 2015; 116: 448–455.
Koeth RA, Wang ZE, Levison BS, Buffa JA, Org E, Sheehy BT, Britt EB, Fu XM, Wu YP, Li L, Smith JD, DiDonato JA, Chen J, Li HZ, Wu GD, Lewis JD, Warrier M, Brown JM, Krauss RM, Tang WHW, Bushman FD, Lusis AJ, Hazen SL . Intestinal microbiota metabolism of l-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat Med 2013; 19: 576–585.
Menni C, Graham D, Kastenmuller G, Alharbi NH, Alsanosi SM, McBride M, Mangino M, Titcombe P, Shin SY, Psatha M, Geisendorfer T, Huber A, Peters A, Wang-Sattler R, Xu T, Brosnan MJ, Trimmer J, Reichel C, Mohney RP, Soranzo N, Edwards MH, Cooper C, Church AC, Suhre K, Gieger C, Dominiczak AF, Spector TD, Padmanabhan S, Valdes AM . Metabolomic identification of a novel pathway of blood pressure regulation involving hexadecanedioate. Hypertension 2015; 66: 422–429.
Nikolic SB, Sharman JE, Adams MJ, Edwards LM . Metabolomics in hypertension. J Hypertens 2014; 32: 1159–1169.
Zheng Y, Yu B, Alexander D, Mosley TH, Heiss G, Nettleton JA, Boerwinkle E . Metabolomics and incident hypertension among blacks: the atherosclerosis risk in communities study. Hypertension 2013; 62: 398–403.
Nicholson JK, Lindon JC, Holmes E . 'Metabonomics': understanding the metabolic responses of living systems to pathophysiological stimuli via multivariate statistical analysis of biological NMR spectroscopic data. Xenobiotica 1999; 29: 1181–1189.
Bictash M, Ebbels TM, Chan Q, Loo RL, Yap IK, Brown IJ, de Iorio M, Daviglus ML, Holmes E, Stamler J, Nicholson JK, Elliott P . Opening up the ‘black box’: metabolic phenotyping and metabolome-wide association studies in epidemiology. J Clin Epidemiol 2010; 63: 970–979.
Nicholson JK, Holmes E, Lindon JC., Chapter 1—metabonomics and metabolomics techniques and their applications in mammalian systems. In Lindon JC, Nicholson JK, Holmes E (eds). The Handbook of Metabonomics and Metabolomics. Elsevier Science BV: Amsterdam, The Netherlands. 2007, pp 1–33.
Grainger DJ., Chapter 12—metabolite profiling and cardiovascular disease. In Lindon JC, Nicholson JK, Holmes E (eds). The Handbook of Metabonomics and Metabolomics. Elsevier Science BV: Amsterdam, The Netherlands. 2007, pp 327–343.
Viant MR, Ludwig C, Gunther UL . 1D and 2D NMR Spectroscopy: from metabolic fingerprinting to profiling. In: Griffiths WJ (ed.). Metabolomics, Metabonomics and Metabolite Profiling Ch. 2. The Royal Society of Chemistry: Cambridge, UK. 2008, pp 44–70.
Dumas ME, Maibaum EC, Teague C, Ueshima H, Zhou B, Lindon JC, Nicholson JK, Stamler J, Elliott P, Chan Q, Holmes E . Assessment of analytical reproducibility of 1 H NMR spectroscopy based metabonomics for large-scale epidemiological research: the INTERMAP study. Anal Chem 2006; 78: 2199–2208.
Smith LM, Maher AD, Want EJ, Elliott P, Stamler J, Hawkes GE, Holmes E, Lindon JC, Nicholson JK . Large-scale human metabolic phenotyping and molecular epidemiological studies via 1 h NMR spectroscopy of urine: investigation of borate preservation. Anal Chem 2009; 81: 4847–4856.
Barton RH, Nicholson JK, Elliott P, Holmes E . High-throughput h-1 NMR-based metabolic analysis of human serum and urine for large-scale epidemiological studies: Validation study. Int J Epidemiol 2008; 37: 31–40.
Kaspar H, Dettmer K, Chan Q, Daniels S, Nimkar S, Daviglus ML, Stamler J, Elliott P, Oefner PJ . Urinary amino acid analysis: a comparison of ITRAQ (R)-LC-MS/MS, GC-MS, and amino acid analyzer. J Chromatogr B 2009; 877: 1838–1846.
Wijeyesekera A, Clarke PA, Bictash M, Brown IJ, Fidock M, Ryckmans T, Yap IK, Chan Q, Stamler J, Elliott P, Holmes E, Nicholson JK . Quantitative UPLC-MS/MS analysis of the gut microbial co-metabolites phenylacetylglutamine, 4-cresyl sulphate and hippurate in human urine: Intermap study. Anal Methods 2012; 4: 65–72.
Ley RE, Turnbaugh PJ, Klein S, Gordon JI . Microbial ecology: human gut microbes associated with obesity. Nature 2006; 444: 1022–1023.
Poesen R, Viaene L, Verbeke K, Augustijns P, Bammens B, Claes K, Kuypers D, Evenepoel P, Meijers B . Cardiovascular disease relates to intestinal uptake of p-cresol in patients with chronic kidney disease. BMC Nephrol 2014; 15: 87–87.
Ramezani A, Raj DS . The gut microbiome, kidney disease, and targeted interventions. J Am Soc Nephrol 2014; 25: 657–670.
Ebbels TMD, De Iorio M . Statistical data analysis in metabolomics. In: Stumpf M, Balding DJ, Girolami M (eds). Handbook of Statistical Systems Biology. Ch. 8. John Wiley & Sons, Ltd: Chichester, UK. 2011, pp 163–180.
Trygg J, Lundstedt T., Chapter 6—chemometrics techniques for metabonomics. In Lindon JC, Nicholson JK, Holmes E (eds). The Handbook of Metabonomics and Metabolomics. Elsevier Science: Amsterdam, The Netherlands. 2007, pp 171–199.
De Iorio M, Ebbels TMD, Stephens DA . Statistical techniques in metabolic profiling. In: Balding DJ, Bishop M, Cannings C (eds). Handbook of Statistical Genetics. Ch. 11. John Wiley & Sons, Ltd: Chichester, UK. 2008, pp 347–373.
Want E, Masson P . Processing and analysis of GC/LC-ms-based metabolomics data. Methods Mol Biol (Clifton, NJ) 2011; 708: 277–298.
Ebbels TM, Lindon JC, Coen M . Processing and modeling of nuclear magnetic resonance (NMR) metabolic profiles. Methods Mol Biol (Clifton, NJ) 2011; 708: 365–388.
Ebbels TMD, Cavill R . Bioinformatic methods in NMR-based metabolic profiling. Prog Nucl Mag Res Sp 2009; 55: 361–374.
Trygg J, Wold S . Orthogonal projections to latent structures (O-PLS). J Chemometr 2002; 16: 119–128.
Wold S, Sjöström M, Eriksson L . Pls-regression: a basic tool of chemometrics. Chemometr Intell Lab Syst 2001; 58: 109–130.
Fonville JM, Richards SE, Barton RH, Boulange CL, Ebbels TMD, Nicholson JK, Holmes E, Dumas ME . The evolution of partial least squares models and related chemometric approaches in metabonomics and metabolic phenotyping. J Chemometr 2010; 24: 636–649.
Cloarec O, Dumas ME, Craig A, Barton RH, Trygg J, Hudson J, Blancher C, Gauguier D, Lindon JC, Holmes E, Nicholson J . Statistical total correlation spectroscopy: an exploratory approach for latent biomarker identification from metabolic 1 H NMR data sets. Anal Chem 2005; 77: 1282–1289.
Smith LM, Maher AD, Cloarec O, Rantalainen M, Tang H, Elliott P, Stamler J, Lindon JC, Holmes E, Nicholson JK . Statistical correlation and projection methods for improved information recovery from diffusion-edited NMR spectra of biological samples. Anal Chem 2007; 79: 5682–5689.
Sands CJ, Coen M, Ebbels TM, Holmes E, Lindon JC, Nicholson JK . Data-driven approach for metabolite relationship recovery in biological 1 H NMR data sets using iterative statistical total correlation spectroscopy. Anal Chem 2011; 83: 2075–2082.
Robinette SL, Veselkov KA, Bohus E, Coen M, Keun HC, Ebbels TMD, Beckonert O, Holmes EC, Lindon JC, Nicholson JK . Cluster analysis statistical spectroscopy using nuclear magnetic resonance generated metabolic data sets from perturbed biological systems. Anal Chem 2009; 81: 6581–6589.
Posma JM, Garcia-Perez I, De Iorio M, Lindon JC, Elliott P, Holmes E, Ebbels TM, Nicholson JK . Subset optimization by reference matching (STORM): an optimized statistical approach for recovery of metabolic biomarker structural information from 1 H NMR spectra of biofluids. Anal Chem 2012; 84: 10694–10701.
Crockford DJ, Maher AD, Ahmadi KR, Barrett A, Plumb RS, Wilson ID, Nicholson JK . H-1 NMR and UPLC-MS statistical heterospectroscopy: characterization of drug metabolites (xenometabolome) in epidemiological studies. Anal Chem 2008; 80: 6835–6844.
Zou X, Holmes E, Nicholson JK, Loo RL . Statistical homogeneous cluster spectroscopy (SHOCSY): an optimized statistical approach for clustering of (1)H NMR spectral data to reduce interference and enhance robust biomarkers selection. Anal Chem 2014; 86: 5308–5315.
Zou X, Holmes E, Nicholson JK, Loo RL . Automatic spectroscopic data categorization by clustering analysis (ASCLAN): a data-driven approach for distinguishing discriminatory metabolites for phenotypic subclasses. Anal Chem 2016; 88: 5670–5679.
Yap IK, Brown IJ, Chan Q, Wijeyesekera A, Garcia-Perez I, Bictash M, Loo RL, Chadeau-Hyam M, Ebbels T, De Iorio M, Maibaum E, Zhao L, Kesteloot H, Daviglus ML, Stamler J, Nicholson JK, Elliott P, Holmes E . Metabolome-wide association study identifies multiple biomarkers that discriminate north and south Chinese populations at differing risks of cardiovascular disease: INTERMAP study. J Proteome Res 2010; 9: 6647–6654.
Stamler J, Brown IJ, Yap IK, Chan Q, Wijeyesekera A, Garcia-Perez I, Chadeau-Hyam M, Ebbels TM, De Iorio M, Posma J, Daviglus ML, Carnethon M, Holmes E, Nicholson JK, Elliott P . Dietary and urinary metabonomic factors possibly accounting for higher blood pressure of black compared with white Americans: results of international collaborative study on macro-/micronutrients and blood pressure. Hypertension 2013; 62: 1074–1080.
Chadeau-Hyam M, Ebbels TM, Brown IJ, Chan Q, Stamler J, Huang CC, Daviglus ML, Ueshima H, Zhao L, Holmes E, Nicholson JK, Elliott P, De Iorio M . Metabolic profiling and the metabolome-wide association study: significance level for biomarker identification. J Proteome Res 2010; 9: 4620–4627.
Teague C, Holmes E, Maibaum E, Nicholson J, Tang H, Chan Q, Elliott P, Stamler J, Ueshima H, Zhou B, Wilson I . Ethyl glucoside in human urine following dietary exposure: detection by 1H NMR spectroscopy as a result of metabonomic screening of humans. Analyst 2004; 129: 259–264.
Heinzmann SS, Brown IJ, Chan Q, Bictash M, Dumas ME, Kochhar S, Stamler J, Holmes E, Elliott P, Nicholson JK . Metabolic profiling strategy for discovery of nutritional biomarkers: proline betaine as a marker of citrus consumption. Am J Clin Nutr 2010; 92: 436–443.
Holmes E, Loo RL, Cloarec O, Coen M, Tang H, Maibaum E, Bruce S, Chan Q, Elliott P, Stamler J, Wilson ID, Lindon JC, Nicholson JK . Detection of urinary drug metabolite (xenometabolome) signatures in molecular epidemiology studies via statistical total correlation (NMR) spectroscopy. Anal Chem 2007; 79: 2629–2640.
Loo RL, Chan Q, Brown IJ, Robertson CE, Stamler J, Nicholson JK, Holmes E, Elliott P, Group IR. A comparison of self-reported analgesic use and detection of urinary ibuprofen and acetaminophen metabolites by means of metabonomics: the INTERMAP study. Am J Epidemiol 2012; 175: 348–358.
Spraul M, Hofmann M, Dvortsak P, Nicholson JK, Wilson ID . High-performance liquid chromatography coupled to high-field proton nuclear magnetic resonance spectroscopy: application to the urinary metabolites of ibuprofen. Anal Chem 1993; 65: 327–330.
Nicholls AW, Farrant RD, Shockcor JP, Unger SE, Wilson ID, Lindon JC, Nicholson JK . NMR and HPLC-NMR spectroscopic studies of futile deacetylation in paracetamol metabolites in rat and man. J Pharm Biomed Anal 1997; 15: 901–910.
Zhao L, Stamler J, Yan LL, Zhou B, Wu Y, Liu K, Daviglus ML, Dennis BH, Elliott P, Ueshima H, Yang J, Zhu L, Guo D, Group IR.. Blood pressure differences between northern and southern Chinese: role of dietary factors: the international study on macronutrients and blood pressure. Hypertension 2004; 43: 1332–1337.
Huang Z, Wu X, Stamler J, Rao X, Tao S, Friedewald WT, Liao Y, Tsai R, Stamler R, He H, Zhou B, Taylor J, Li Y, Xiao Z, Williams D, Cen R, Zhang H . A north–south comparison of blood pressure and factors related to blood pressure in the People's Republic of China: a report from the PRC-USA collaborative study of cardiovascular epidemiology. J Hypertens 1994; 12: 1103–1112.
Wang D, He Y, Li Y, Luan D, Yang X, Zhai F, Ma G . Dietary patterns and hypertension among Chinese adults: a nationally representative cross-sectional study. BMC Public Health 2011; 11: 925.
Wu Z, Yao C, Zhao D, Wu G, Wang W, Liu J, Zeng Z, Wu Y . Sino-Monica Project: a collaborative study on trends and determinants in cardiovascular diseases in China, part I: morbidity and mortality monitoring. Circulation 2001; 103: 462–468.
He J, Klag MJ, Wu Z, Whelton PK . Stroke in the People's Republic of China. I. Geographic variations in incidence and risk factors. Stroke 1995; 26: 2222–2227.
Xu G, Ma M, Liu X, Hankey GJ . Is there a stroke belt in China and why? Stroke 2013; 44: 1775–1783.
Chan Q, Stamler J, Elliott P . Dietary factors and higher blood pressure in African-Americans. Curr Hypertens Rep 2015; 17: 10.
Acknowledgements
The study is supported by Grants R01 HL50490 and R01 HL084228 from the National Heart, Lung and Blood Institute, National Institutes of Health (Bethesda, MD, USA) and by national agencies in China, Japan and the United Kingdom. The sponsors had no role in the design or conduct of the study; the collection, management, analysis or interpretation of the data; or the preparation, review or approval of the manuscript. PE is supported, in part, by the National Institute for Health Research (NIHR) Biomedical Research Centre, Imperial College Healthcare NHS Trust and Imperial College London. We thank all the INTERMAP staff for their invaluable efforts; a partial listing of these colleagues is given in Stamler et al.12 in this paper.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Chan, Q., Loo, R., Ebbels, T. et al. Metabolic phenotyping for discovery of urinary biomarkers of diet, xenobiotics and blood pressure in the INTERMAP Study: an overview. Hypertens Res 40, 336–345 (2017). https://doi.org/10.1038/hr.2016.164
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hr.2016.164
Keywords
This article is cited by
-
Proportional changes in the gut microbiome: a risk factor for cardiovascular disease and dementia?
Hypertension Research (2019)
-
Serum Concentrations of Citrate, Tyrosine, 2- and 3- Hydroxybutyrate are Associated with Increased 3-Month Mortality in Acute Heart Failure Patients
Scientific Reports (2019)
-
Gut microbiota and inflammation in chronic kidney disease and their roles in the development of cardiovascular disease
Hypertension Research (2019)