Abstract
Animal models emulating essential hypertension are an informative means by which to elucidate the physiological mechanisms and gene–gene interactions underlying blood pressure (BP) regulation. We have localized earlier quantitative trait loci (QTLs) for BP on Chromosome (Chr) 2 of Dahl salt-sensitive (DSS) rats, but their chromosome delineations were too large for gene identification. To advance toward positional cloning of these QTLs, we constructed congenic strains that systematically dissect a Chr 2 segment with no overlaps. BP and cardiac functions were measured by telemetry and echocardiography. Six QTLs were delimited, each independently influencing BP. The intervals lodging two of them harbor 10–15 genes and undefined loci. These six QTLs can be grouped into two epistatic modules distinguishable by cardiac pathways/cascades. None of the genes known to exert physiological effects on BP in the segments harboring the six QTLs are leading candidates, as their protein products are the same in DSS rats and similar to those in their Milan normotensive counterparts. Specifically, the lack of an amino-acid alteration, coupled with a lack of difference in the α1-Na-K-ATPase activity, excluded ATPase, Na+/K+-transporting, α-1 polypeptide as a candidate gene for C2QTL6. The identification of the six QTLs will likely develop into a novel diagnostic and/or therapeutic target for essential hypertension and hypertension-associated diseases.
Similar content being viewed by others
Introduction
Essential hypertension is the most prominent disorder leading to fatal cardiovascular and renal diseases.1 In spite of our knowledge concerning the molecular bases of rare monogenic forms of hypertension,2 the pathogenesis of essential hypertension remains largely elusive. A major challenge in essential hypertension research is to identify genes that underlie physiological etiologies of blood pressure (BP) determination. The identification of quantitative trait loci (QTLs) contributing to BP control is considered the most direct means of confronting this challenge.3 Unfortunately, in contrast to several polygenic and complex traits, no major genes underlying essential hypertension have been consistently observed, and not a single QTL with genome-wide significance for essential hypertension has been detected in a general population.4, 5
Certain limitations are evident in directly studying general human populations, such as genetic heterogeneity, environmental complications and diagnostic variability,4, 5 including an influence of sex and age.6 To overcome these limitations and to aid in human hypertension research, inbred rodent, specifically rat, models that show polygenic hypertension have been used to elucidate the molecular basis for the high-BP phenotype and, consequentially, to gain insights into the physiological mechanisms underpinning BP regulation.7, 8 The Dahl salt-sensitive (DSS) rat model has been extensively used for this purpose.8 Our ultimate goal is to identify BP QTLs in DSS, predicating on functionally driven postulates and a cause–effect relationship.
Earlier, we defined three QTLs on Chromosome (Chr) 2.9 However, the chromosome segments harboring them were too large to allow their positional cloning. One effective means of limiting the number of genes to an amenable level is through fine congenic resolution. The current investigation had three objectives. The first was comprehensive fine congenic resolution of BP QTLs on Chr 2. The second was to consider phenotypes other than BP to provide potential testable hypotheses mechanistically linking QTLs to BP regulation. Finally, prominent candidate genes present in each QTL interval were screened for mutations.
As a result, a cluster of four epistatically interacting BP QTLs were defined by non-overlapping congenic strains. Two additional BP QTLs were also delimited. All of these QTLs can be modularized by epistasis into two classes ascertainable through distinct cardiac routes. No mutations were found in the coding domains of obvious candidate genes in the six QTL regions. Thus, it is likely that these QTLs may represent novel genes that were not recognized earlier to be involved in BP control.
Methods
Animals
Protocols for handling as well as maintaining animals were approved by our institutional animal committee (CIPA). The DSS, C2S.M7 and C2S.M8 congenic strains are the same as those used earlier.9 Congenic strains bear different chromosome segments of DSS that are replaced by the homologs of Milan normotensive (MNS) rats.
Construction of new congenic strains
The congenic strains9 used earlier were the basis for deriving new congenic substrains. The breeding and screening procedures in this process were similar to those reported earlier.9 Special attention was paid to produce congenic strains with no chromosome overlaps. For this work, 10 informative new congenic strains were produced (Figure 1) designated DSS.MNS-(D2Got93-D2Rat222)/Lt (abbreviated as C2S.M13), DSS.MNS-(D2Chm90-D2Rat38)/Lt (C2S.M15), DSS.MNS-(D2Chm381-D2Chm225)/Lt (C2S.M16), DSS.MNS-(D2Rat155-D2Chm419)/Lt (C2S.M17), DSS.MNS-(D2Chm254-D2Chm161)/Lt (C2S.M18), DSS.MNS-(D2Rat155-D2Chm419)/Lt (C2S.M19), DSS.MNS-(D2Chm161-D2Mgh10)/Lt (C2S.M21), DSS.MNS--(D2Chm153-D2Mgh10)/Lt (C2S.M22), DSS.MNS--(D2Chm324-D2Mgh10)/Lt (C2S.M23) and DSS.MNS-(D2Chm366-D2Rat52)/Lt (C2S.M24).
Fine congenic resolution of BP QTLs on Chr 2. The contigs and marker positions in megabases (Mb) within them are taken from NCBI (http://www.ncbi.nlm.nih.gov/mapview). Solid bars under congenic strains symbolize the Dahl salt-sensitive (DSS) chromosome fragments that have been replaced by those of the Milan normotensive rat (MNS). Hatched bars on ends of solid bars indicate the ambiguities of crossover breakpoints between markers. Pfn2; profilin 2; Npr1, natriuretic peptide receptor 1; Mme, Membrane metallo endopeptidase; Sucnr1, succinate receptor 1; Ctsk, cathepsin K; Prkab2, protein kinase, AMP-activated, beta 2 non-catalytic subunit; Atp1a1, ATPase, Na+/K+-transporting, α-1 polypeptide. D2Chm markers were newly produced from the rat genome sequence (http://www.ncbi.nlm.nih.gov/mapview). Mean arterial pressures (MAPs) for DSS and congenic strains are given at the bottom of the map to facilitate strain comparisons. ±s.e.m. for the strains range from 2 to 6; n, number of rats measured. C2S.M13, C2S.M15, C2S.M16, C2S.M17, C2S.M18, C2S.M19, C2S.M21, C2S.M22, C2S.M23 and C2S.L24 are newly generated from this study. Their full names are given in the Methods section under Construction of new congenic strains. C2S.M7 and C2S.M8 were produced earlier.9 The placements of the BP QTLs are marked in the right in parentheses. Below the QTL designation is the size of the QTL interval in megabases (Mb) or kilobases (kb) and the number of genes and undefined loci. P-values in italics represent significant differences.
Animal protocols, BP measurement and statistical analyses
Breeding protocols, dietary treatments, implantation and BP measurement schedules were virtually the same as those documented earlier.9 Briefly, male rats were weaned at 21 days of age and maintained on a low-salt diet (0.2% NaCl; Harlan Teklad 7034, Indianapolis, IN, USA), followed by a high-salt diet (2% NaCl; Harlan Teklad 94217) starting from 35 days of age until the end of the experiment. Telemetry probes were implanted at 56 days of age (namely 3 weeks from the time of the high-salt diet). BPs for most of the strains were measured at at least two different times to exclude seasonal as well as environmental effects. Thus, the BP data were pooled from separately reproducible measurements for each strain. BP of DSS rats was pooled from four different measurements. In the BP presentation, averaged 24-h readings are shown for the duration of our studies. As systolic and diastolic pressures were consistent (data not shown) with mean arterial pressures (MAPs) of all the strains, only their MAPs are presented.
Repeated measures analysis of variance (ANOVA) followed by Dunnett's test, which corrects for multiple comparisons and unequal sample sizes, was used to compare a parameter in MAP between two groups, as reported earlier.9 During the BP comparison, ANOVA was first used to assess the inter-group differences. If the ANOVA was significant, Dunnett's test followed to identify which group was different from DSS rats and to determine the level of significance. A 2 × 2 ANOVA determined a QTL–QTL interaction (or lack thereof) by evaluating whether the observed effect of a congenic strain combining separate QTLs was significantly different from a predicted sum of effects from each individual QTL.9
Cardiac phenotypings and calculations
Cardiac functions for strains of interest were measured by transthoracic echocardiography and calculated accordingly, as reported recently.10 Male rats were fed a 2% (high) salt diet starting from 5 weeks of age and their cardiac phenotypes were measured at 11 weeks of age. Left ventricular (LV) short-axis view at the papillary muscle level was recorded; LV areas at both cardiac end diastole (EDA) and end systole (ESA) were measured with the papillary muscle being excluded and LV fractional area changing was calculated as (EDA−ESA)/EDA × 100%. M-mode spectrum was also obtained at this view to measure LV dimension, thickness of LV anterior wall and thickness of LV posterior wall at the end of cardiac diastole; LV mass corrected to small animals was calculated, as described earlier.10 Left-atrium (LA) M-mode was obtained at the level of the aortic valve in the parasternal long-axis view. LA dimensions at both end cardiac diastole (LADd) and systole (LADs) were measured; LA fractional shortening was calculated by (LADs−LADd)/LADs × 100%. Pulsed wave Doppler was used to visualize transmitral inflow in an apical four-chamber view. Peak velocity in early filling, E wave, E deceleration time (DT), deceleration rate (DR) and time interval between mitral opening and closure were measured. The pulsed wave Doppler was also used to study transaortic outflow. Aortic peak velocity, LV ejection time and the time interval from aortic closing to opening were measured; LV global myocardial performance index was calculated accordingly.10 Continuous wave Doppler spectrum at the conjunction of LV inflow and outflow was recorded in an apical five-chamber view. LV isovolumic relaxation time (IVRT) was measured and corrected (IVRTc) by the square root of R–R interval taken from simultaneously recorded ECG. Pulse wave Doppler was used to analyze transmitral inflow, transaortic outflow, aortic peak velocity, LV ejection time and the time interval from the aortic closing to opening. The LV global myocardial performance index was calculated accordingly.10 The average of three consecutive cardiac cycles was used for each measurement. A wave could not be obtained for 95% of rats because of their uncontrolled fast HR. Echocardiographic data are expressed as mean±s.d. A two-tailed P<0.05 was considered statistically significant because one DSS and one congenic strain were compared at a time.
Mutation screening for candidate genes
The coding region for a chosen candidate gene was first amplified by PCR either from genome DNAs or from cDNAs of DSS rats and a comparing congenic strain. When a mutation was detected in sequencing, the same nucleotide was re-sequenced after an independent PCR amplification.
Results
BP measurements
MAP data for congenic strains and their comparison with those of DSS rats are summarized below the representations of congenic strains in Figure 1. The actual readings of MAPs for the new strains are presented in Figure 2.
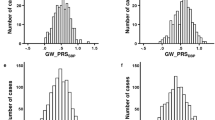
(a–c) Comparisons of mean arterial pressures (MAPs) between congenic strains and the DSS strain. Each time point on the graph represents an average of 24-h readings. Error bars represent s.e.m. n refers to the number of rats. DSS, Dahl salt-sensitive strain. The chromosome coverage for congenic strains is given in Figure 1.
QTL placements
The region originally thought to contain C2QTL29 turns out to lodge three QTLs, C2QTL2, C2QTL4 and C2QTL5, as the congenic strains delimiting them, namely C2S.M19, C2S.M18 and C2S.M21, carry no genomic overlaps among them (Figure 1). C2QTL6 is newly assigned. As the congenic strains delineating the QTL intervals all directly show a BP effect, the determination of C2QTL2, C2QTL4, C2QTL5 and C2QTL6 was definitive.
By comparison, C2QTL1 was placed in the non-overlapping segment between C2S.M7 showing a BP effect and C2S.M13 lacking it. Similarly, C2QTL3 was assigned to the section between D2Chm90 and Mme by subtractive mapping. For example, C2S.M15 showed a BP effect, whereas C2S.M16 did not. This ‘subtractive’ comparative mapping cannot rule out the possibility that the congenic strain lacking a BP effect could harbor a QTL, which would require a combination with another putative QTL in the non-overlapping region carried by the strain evincing an effect. Thus, the placements of C2QTL1 and C2QTL3 are tentative.
Assessment of QTL interactions
The C2S.M17 congenic strain carried the non-overlapping segments dissected by C2S.M19 bearing C2QTL2 and C2S.M18 bearing C2QTL4. A 2 × 2 ANOVA9 showed an epistatic interaction (P<0.008) between C2QTL2 and C2QTL4. No similar analyses of epistasis can be conducted among C2QTL4, C2QTL5 and C2QTL6, because of a lack of necessary congenic strains (Figure 1). Nevertheless, C2S.M89 included C2QTL2, C2QTL4, C2QTL5 and C2QTL6, and its MAP (136 mm Hg) was not different (P>0.10) from that (125–144 mm Hg) of C2S.M19, C2S.M18, C2S.M21 and C2S.M24. This simple comparison suggests that all four QTLs, namely C2QTL2, C2QTL4, C2QTL5 and C2QTL6, may belong to the same epistatic module.7, 8
Appraisal of cardiac function of BP QTLs representing two separate epistatic modules
From our earlier9 and current work, all six QTLs could be functionally modularized into two epistatic groups that imply two different pathways/cascades.7, 8 Group 1 consists of C2QTL2, C2QTL4, C2QTL5 and C2QTL6 and group 2 is composed of C2QTL3. C2QTL1 belongs to either group 1 or 2,9 although its exact epistatic classification cannot be determined.
As the magnitude of BP effects from all six QTLs is indistinguishable (Figure 1), physiological parameters other than BP may differentiate these two epistatic groups with differing pathway/cascade implications. Cardiac function with detailed and extensive parameters10 can serve as discriminating phenotypes. For this purpose, we studied a representative of epistatic group 1 with the smallest congenic coverage, namely C2S.M21, lodging C2QTL5, and an epistatic group 2 QTL, namely C2QTL3, carried by C2S.M15. This choice of a strain with the shortest chromosome segment was made to maximize the possibility that the same QTL is responsible for both BP and cardiac function.
Table 1 summarizes the findings. MNS alleles of C2QTL5 reduced LV structural remodeling (enlargement and hypertrophy) and LV hemodynamic state. They also improved LV compliance (prolonged DT and decreased DR) and LV relaxation (shortened IVRT and IVRTc), thus ameliorating LV diastolic dysfunction. This was in accordance with improved left atrial structural and functional remodeling by MNS alleles of C2QTL5 in comparison with DSS. Furthermore, with reduced LV structural remodeling by MNS alleles of C2QTL5, LV global myocardial performance was significantly improved. By contrast, MNS alleles of C2QTL3 had no effects on these cardiac values, despite a similar reduction in BP (that is, −31 mm Hg) to that (that is, −34 mm Hg) of C2QTL5 (Figure 1). The MNS parental strain was no longer available, and no study was conducted on it.
Evaluation of candidate genes
All the possible genes and undefined loci (Locs) existing in the regions encamping all six QTLs were curated (Supplement 1). Notable candidate genes residing in segments containing six QTLs were selected for first-passage mutation screening for three reasons.
First, these genes are well characterized, and consequently their coding realms are sufficiently documented to aid in recognizing sequence variations. Second, assayable tests can be performed on them so that a mutated protein product can be compared with that of the wild type for functional analyses. Finally, prior knowledge on their functional roles in BP regulation has been at least implicit. Thus, relevant insights can be translated into BP homeostasis through their biochemical, signaling or physiological mechanisms once a significant mutation is detected in the coding domain of such a gene. As preliminary gene profiling11 in the kidneys of the 5-week-old male rats did not reveal differentially expressed candidate genes (data not shown), their coding regions became the focus of the present investigation. Further justifications for emphasizing the structure of these genes emanate from the reports that mutations significantly altering amino acids exist in the genes potentially being BP QTLs, such as steroid 11-β-hydroxylase12 and α-Adducin.13
The chosen candidate genes and their sequence comparisons between DSS and MNS are shown in Table 2. In the interval harboring C2QTL1, transmembrane 4 superfamily member 4 (Tm4sf4) was implicated in actions mediated by the transforming growth factor-α signaling cascade,14 which can play a part in diverse cardiovascular functions.15 Profilin 2 (Pfn2)16 encodes an actin-binding protein regulating cytoskeleton dynamics. In the C2QTL2-containing interval, natriuretic peptide receptor 1 (Npr1) codes for a receptor in mediating the effect of natriuretic peptide (NP) and may cause salt-sensitive hypertension.17 In the C2QTL3-residing region, succinate receptor 1 (Sucnr1) encodes a receptor for an intermediate molecule in citric acid cycle, succinate, which can regulate BP.18 Membrane metallo endopeptidase (Mme) codes for a protease that degrades a wide assortment of vasoactive peptides.19 In the segment lodging C2QTL4, cathepsin K (Ctsk) codes for a kinin-degrading enzyme that can modulate BP.20 In the C2QTL5-dwelling region, protein kinase, AMP-activated, β-2 non-catalytic subunit (Prkab2) encodes a molecule that may play a role in the ischemic heart.21 In the C2QTL6-containing region, ATPase, Na+/K+-transporting, α-1 polypeptide (Atp1a1) mediates the Na+ and K+ transport and may play a role in BP regulation.22
From sequence analysis, some nucleotide variations in entire coding domains of these candidate genes were indeed found, but they did not result in changes in amino acids (Table 2).
Discussion
The major findings in this work are the following: (a) A cluster of epistatically interacting QTLs for BP have been conclusively resolved by non-overlapping congenic strains. Each of these QTLs can influence BP autonomous of one another and autonomous of the remaining genome. (b) Two representative QTLs from two separate epistatic modules show similar effects on BP, but distinct potencies on cardiac function, supporting the notion that differing physiological pathways/cascades likely underlie their mechanisms, leading to BP regulation. (c) Each of the known candidate genes from six QTL intervals codes for the same protein product in DSS as in MNS. Thus, these QTLs are likely either new genes earlier unknown for BP control, unidentified genes or candidate genes differentially regulated through expression or splicing.
Fine congenic resolution of BP QTLs
The building of congenic strains with no chromosome imbrications to one another is essential in conclusively delimiting the fragments lodging QTLs founded on a cause–effect relationship and separates those situated closely. From our comprehensive analyses, six BP QTLs have been defined to the genome segments between 0.926 and 5.64 Mb (Figure 1), paving the way for their positional cloning. This resolution is in sharp contrast to the outcome of the linkage analysis, in which only one QTL in a broad section was detected23 in the same region.
Distinction of separate epistatic modules for BP QTLs by cardiac pathways/cascades
The two epistatic modules were originally established because of the additivity of two QTLs.9 This genetic phenomenon implies two separate pathways/cascades7, 8 leading to BP regulation, but little evidence supported this notion. From our current work, C2QTL5 exemplifying epistatic module 1 and C2QTL3 as epistatic module 2 clearly show divergent cardiac influences (Table 1). As BP is a cardiovascular phenotype, mechanisms constituting it have to be mediated through cardiovascular physiological pathways/cascades. Pinpointing an associated phenotype such as diastolic function with BP for C2QTL5 furnishes a testable hypothesis, at least at the organ level, that a gene for C2QTL5 can be analyzed for mechanistic connections to BP. C2QTL5 belongs to 1 of 10 possible genes, namely olfactory receptor 390 (predicted) (Olr390), olfactory receptor pseudogene 391 (Olr391), chromodomain helicase DNA-binding protein 1-like (predicted) (Chd1l), flavin-containing monooxygenase 5 (Fmo5), Prkab2, phosphodiesterase 4D interacting protein (myomegalin) (Pde4dip), SEC22 vesicle trafficking protein-like 1 (Sec22l1), notch gene homolog 2 (Notch2), LOC690547 and regenerating islet-derived family, member 4 (Reg4). As none of these genes and loci are recognized for a direct BP effect, a novel gene modulating BP may emerge from identifying C2QTL5. It is likely that C2QTL5 determines both BP and diastolic function.
Candidate genes
Known candidate genes in all six QTL intervals carry no mutations that can change an amino acid (Table 2). These results, although not exclusive, downgrade them as candidate genes for the respective QTLs. A case in point is Atp1a1. No change in amino acids comparing DSS and MNS agrees with a lack of difference in the α1-Na-K-ATPase activity.24 A differential gene expression between DSS and MNS in the potential Atp1a1 transcription and splicing may not appear to be functionally relevant and worth examining. Thus, among all the candidate genes examined, at least Atp1a1 can be reasonably ruled out as a QTL. Most likely, the true genes for the six QTLs are those not renowned earlier for BP control.
Caveats
First, candidate genes were analyzed only for coding mutations. Differential gene regulations cannot be ruled out. Although no differential expressions for these genes under one condition (that is, data from rats at one age and examining one organ) were found (data not shown), other conditions may induce a differential gene expression, such as a varying amount of salt, in different organs, at various developmental stages and adult ages, or a combination of these factors. For instance, the level of Npr1 mRNA is different between DSS and MNS under various conditions, such as a high-fat diet (data not shown). Sequence variations, such as a differing length of TA repeat in the Npr1 promoter,25 also exist between the two strains (data not shown), but their relevance to controlling Npr1 expression cannot be established.
Moreover, differential splicing for any of these genes cannot be excluded and may depend on an analysis under the right conditions and in an appropriate organ. Work is underway to investigate every possible gene present in the C2QTL5 region for differential expression in the adult heart at levels of both gene expression and splicing because of the association of C2QTL5 with cardiac function (Table 1).
Second, coding sequences of genes (Supplement 1), other than those listed in Table 2, have not been completely determined, especially those predicated and unidentified, because of the uncertainty of the gene's exact composition. For instance, Chd1l residing in the C2QTL5 region is only a predicted gene and is supposed to encode a protein of 531 amino acids, but its size was 896 amino acids before 14 May 2008. Thus, before sequencing can commence for Chd1l, the actual coding size must be identified. For this purpose, a detailed molecular characterization is required at both the transcriptional and translational levels. The same pre-requisite applies to all the remaining possible genes of similar stature. Work is underway to sequence coding domains of all substantiated genes existing in the intervals lodging C2QTL5 and C2QTL1.
Third, the intervals harboring C2QTL2, C2QTL3, C2QTL4 and C2QTL6 contain too many genes to be thoroughly sequenced. A further reduction in the QTL-residing fragments is necessary. It is possible that more QTLs may appear from fine congenic resolution of these QTLs.
Prospect
Although screening for either coding mutations or differential gene expressions is a necessary first step in ferreting out potential genes responsible for a particular QTL, functional validation in vivo must ensue to prove that a mutated gene can truly affect BP. Evidently, if the QTL-residing interval can be narrowed by a congenic strain(s) to harbor only one gene, that gene must be the culprit for the QTL. Other appropriate gene manipulation approaches may aid in this endeavor, such as transgenesis and gene targeting.7
Extending beyond the gene level, mechanistic insights are essential to establish that a gene with no recognizable role in BP homeostasis can genuinely play a role in the biological process(es), biochemical pathway(s) or signaling cascade(s) leading to it. This functional information becomes primordial, especially when a significant and non-conserved mutation(s) is detected in the coding region of a gene with no prior physiological functions.
Perspective
A systematic fine congenic resolution yielded six QTLs, each of which can independently influence BP. These QTLs appear to be organized into two discrete epistatic modules discernible by their diastolic pathways/cascades. The QTL intervals for two of them contain 10 and 15 possible genes, respectively. Sequence comparisons revealed no coding mutations in prominent candidate genes, although differential gene regulation for them, with the combination of the right organ, time and dietary treatment, cannot be ruled out. It is probable that the discovery of these QTLs may unravel novel genes and their underlying physiological mechanisms that can generate innovative diagnostic tools and therapeutic targets for essential hypertension and possibly for hypertension-related diseases.
Conflict of interest
JCT holds the Pfizer and Canadian Institutes of Health Research Chair in Atherosclerosis, and SN the Paul-David Chair in Cardiovascular Electrophysiology.
References
Kearney PM, Whelton M, Reynolds K, Muntner P, Whelton PK, He J . Global burden of hypertension: analysis of worldwide data. Lancet 2005; 365: 217–223.
Lifton RP . Genetic dissection of human blood pressure variation: common pathways from rare phenotypes. Harvey Lect 2004; 100: 71–101.
Flint J, Valdar W, Shifman S, Mott R . Strategies for mapping and cloning quantitative trait genes in rodents. Nat Rev Genet 2005; 6: 271–286.
Caulfield M, Munroe P, Pembroke J, Samani N, Dominiczak A, Brown M, Benjamin N, Webster J, Ratcliffe P, O’Shea S, Papp J, Taylor E, Dobson R, Knight J, Newhouse S, Hooper J, Lee W, Brain N, Clayton D, Lathrop GM, Farrall M, Connell J . Genome-wide mapping of human loci for essential hypertension. Lancet 2003; 361: 2118–2123.
The Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 2007; 447: 661–678.
Seda O, Tremblay J, Gaudet D, Brunelle PL, Gurau A, Merlo E, Pilote L, Orlov SN, Boulva F, Petrovich M, Kotchen TA, Cowley Jr AW, Hamet P . Systematic, genome-wide, sex-specific linkage of cardiovascular traits in French Canadians. Hypertension 2008; 51: 1156–1162.
Deng AY . Positional cloning of quantitative trait loci for blood pressure: how close are we? A critical perspective. Hypertension 2007; 49: 740–747.
Deng AY . Genetic basis of polygenic hypertension. Hum Mol Genet 2007; 16: R195–R202.
Dutil J, Eliopoulos V, Tremblay J, Hamet P, Charron S, Deng AY . Multiple quantitative trait loci for blood pressure interacting epistatically and additively on dahl rat chromosome 2. Hypertension 2005; 45: 557–564.
Deng AY, Nattel S, Shi Y, L’Heureux N, Cardin S, Ménard A, Roy J, Tardif J . Distinct genomic replacements from Lewis correct diastolic dysfunction, attenuate hypertension and reduce left ventricular hypertrophy in Dahl salt-sensitive rats. J Hypertens 2008; 26: 1935–1943.
Moujahidine M, Lambert R, Dutil J, Palijan A, Sivo Z, Ariyarajah A, Deng AY . Combining congenic coverage with gene profiling in search of candidates for blood pressure quantitative trait loci in Dahl rats. Hypertens Res 2004; 27: 203–212.
Cicila GT, Rapp JP, Wang JM, St Lezin E, Ng SC, Kurtz TW . Linkage of 11 beta-hydroxylase mutations with altered steroid biosynthesis and blood pressure in the Dahl rat. Nat Genet 1993; 3: 346–353.
Bianchi G, Tripodi G, Casari G, Salardi S, Barber BR, Garcia R, Leoni P, Torielli L, Cusi D, Ferrandi M . Two point mutations within the adducin genes are involved in blood pressure variation. Proc Natl Acad Sci USA 1994; 91: 3999–4003.
Qiu J, Liu Z, Da L, Li Y, Xuan H, Lin Q, Li F, Wang Y, Li Z, Zhao M . Overexpression of the gene for transmembrane 4 superfamily member 4 accelerates liver damage in rats treated with CCl4. J Hepatol 2007; 46: 266–275.
Prabhu SD . Cytokine-induced modulation of cardiac function. Circ Res 2004; 95: 1140–1153.
Sohn RH, Goldschmidt-Clermont PJ . Profilin: at the crossroads of signal transduction and the actin cytoskeleton. Bioessays 1994; 16: 465–472.
Lopez MJ, Wong SK, Kishimoto I, Dubois S, Mach V, Friesen J, Garbers DL, Beuve A . Salt-resistant hypertension in mice lacking the guanylyl cyclase-A receptor for atrial natriuretic peptide. Nature 1995; 378: 65–68.
He W, Miao FJ, Lin DC, Schwandner RT, Wang Z, Gao J, Chen JL, Tian H, Ling L . Citric acid cycle intermediates as ligands for orphan G-protein-coupled receptors. Nature 2004; 429: 188–193.
Quaschning T, Galle J, Wanner C . Vasopeptidase inhibition: a new treatment approach for endothelial dysfunction. Kidney Int Suppl 2003; 84: S54–S57.
Lecaille F, Vandier C, Godat E, Hervé-Grépinet V, Brömme D, Lalmanach G . Modulation of hypotensive effects of kinins by cathepsin K. Arch Biochem Biophys 2007; 459: 129–136.
Li J, Coven DL, Miller EJ, Hu X, Young ME, Carling D, Sinusas AJ, Young LH . Activation of AMPK {alpha}- and {gamma}-isoform complexes in the intact ischemic rat heart. Am J Physiol Heart Circ Physiol 2006; 291: H1927–H1934.
Pritchard TJ, Parvatiyar M, Bullard DP, Lynch RM, Lorenz JN, Paul RJ . Transgenic mice expressing Na+-K+-ATPase in smooth muscle decreases blood pressure. Am J Physiol Heart Circ Physiol 2007; 293: H1172–H1182.
Deng AY, Dene H, Rapp JP . Mapping of a quantitative trait locus for blood pressure on rat chromosome 2. J Clin Invest 1994; 94: 431–436.
Orlov SN, Dutil J, Hamet P, Deng AY . Replacement of (alpha)1-Na-K-ATPase of Dahl rats by Milan rats lowers blood pressure but does not affect its activity. Physiol Genomics 2001; 7: 171–177.
Tremblay J, Hum DH, Sanchez R, Dumas P, Pravenec M, Krenova D, Kren V, Kunes J, Pausova Z, Gossard F, Hamet P . TA repeat variation, Npr1 expression, and blood pressure: impact of the Ace locus. Hypertension 2003; 41: 16–24.
Acknowledgements
We appreciate the assistance of the Cardiovascular Health Network (RSCV) of the Quebec Health Research fund (FRSQ) for echocardiographic assessment. This work was supported by grants from the Canadian Institutes of Health Research (AYD, JT and SN). CX is supported by grants of the 973 program (2006CB708502) and the National Nature Science Foundation of China (30660076).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Hypertension Research website (http://www.nature.com/hr)
Supplementary information
Rights and permissions
About this article
Cite this article
Chauvet, C., Ménard, A., Tremblay, J. et al. Cardiac pathways distinguish two epistatic modules enacting BP quantitative trait loci and candidate gene analysis. Hypertens Res 32, 631–637 (2009). https://doi.org/10.1038/hr.2009.70
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hr.2009.70