Abstract
Celery (Apium graveolens L.) is one of the most important and widely grown vegetables in the Apiaceae family. Due to the lack of comprehensive genomic resources, research on celery has mainly utilized physiological and biochemical approaches, rather than molecular biology, to study this crop. Transcriptome sequencing has become an efficient and economic technology for obtaining information on gene expression that can greatly facilitate molecular and genomic studies of species for which a sequenced genome is not available. In the present study, 15 893 516 and 19 818 161 high-quality sequences were obtained by RNA-seq from two celery varieties ‘Ventura’ and ‘Jinnan Shiqin’, respectively. The obtained reads were assembled into 39 584 and 41 740 unigenes with mean lengths of 683 bp and 690 bp, respectively. A total of 1939 simple sequence repeat (SSR) markers were identified in ‘Ventura’ and 2004 SSRs in ‘Jinnan Shiqin’. Di-nucleotide repeats were the most common repeat motif, accounting for 55.49% and 54.84% in ‘Ventura’ and ‘Jinnan Shiqin’, respectively. A comparison of expressed genes between the two libraries, identified 338 differentially expressed genes (DEGs). Three hundred and three of the DEGs were annotated based on a sequence similarity search utilizing eight public databases. Additionally, the expression profile of eight annotated DEGs was characterized in response to abiotic stresses. The collective data generated in the present research represent a valuable resource for further genetic and molecular studies in celery.
Similar content being viewed by others
Introduction
Celery (Apium graveolens L.) originated in Europe, the Mediterranean region and Asia, west of the Himalayas. It has become one of the most important vegetables in the Apiaceae plant family.1 All parts of the celery plant have been reported to possess medicinal value and it is rich in flavonoids, carotenoids, coumarins and volatile oils.1,2 Research on celery has mainly focused on the physiology and biochemistry of the crop, rather than molecular biology, due to the lack of a sequenced genome and the existence of a large collection of expressed sequence tag data.3,4
Celery breeding can be accelerated and made more efficient utilizing marker assisted selection. Microsatellites, which are also known as simple sequence repeats (SSRs), are short tandem repetitive DNA sequences with a repeat length of 1–6 base pairs, and are often genotype-specific and are therefore used to characterize genotypic diversity.5 SSR markers are widely used in genetic linkage map construction, genetic diversity detection and molecular assisted breeding due to their high degree of polymorphism and codominant inheritance.6–8 Despite their value, few SSR markers have been reported in celery, which has hindered genetic research in this crop.
Next-generation sequencing has been widely employed to obtain transcriptome data in species without a sequenced genome. Transcriptome data contain a large amount of genetic information that can be used for many purposes including: investigations of alternative splicing;9 generation of gene sequences from non-model organisms that lack a reference genome;10 discovery of novel genes;10 analysis of gene expression;11 identification of SSRs and single-nucleotide polymorphism,12 etc. While RNA-seq technology has been successfully used in species such as potato,13 grapevine14 and peanut,15 to date, only one celery (Celery line ‘Q 111’ ) transcriptome has been reported.16 Despite the transcriptome report for celery, little is known about gene expression related to development or stress response.16
Here, transcriptome sequencing of two celery varieties, ‘Ventura’ and ‘Jinnan Shiqin’, the commonly grown celery varieties in the United States and China, respectively, was conducted. ‘Ventura’, a variety developed in the United States, is tall and compact with thick glossy leaves. ‘Jinnan Shiqin’, a variety developed in China, is tall, light green in color, has hypertrophic leaves and is less prone to premature bolting.17 The present study compared the genes comprising the transcriptomes of the two varieties and identified SSR markers. Subsequently, the expression of eight of the annotated differentially expressed genes (DEGs) was characterized in the two varieties in response to a variety of abiotic stresses. The collective data generated in the present research represents a valuable resource to enable further advancements in genetic and molecular studies in celery.
Materials and methods
Plant material and RNA isolation
Two celery varieties (A. graveolens L. cvs. ‘Ventura’ and ‘Jinnan Shiqin’) were grown in plastic pots (17×17×20 cm) containing a soil: vermiculite mixture (3:1) in a controlled-environment growth chamber at 25/16 °C for day/night temperatures under a photoperiod of 16/8 h light/dark photoperiod with 3000 lux of light intensity. Leaves of 2-month-old plants were collected and immediately frozen in liquid nitrogen and stored at −80 °C. Total RNA was extracted from pooled samples of leaves using a total RNA kit (RNAsimply; Tiangen, Beijing, China) according to the manufacturer’s instructions. The quantity and quality of RNA samples were determined by gel electrophoresis and the use of a Nanodrop ND 1000 spectrophotometer (Nanodrop Technologies Inc., Rockland, DE, USA). The standards applied were OD260/OD230≥1.8 and 1.8≤OD260/OD280≤2.2. At least 20 μg RNA was pooled in equimolar fashion from each of three samples.
cDNA library construction and sequencing
After extracting the total RNA, mRNA in the samples was enriched using oligo magnetic adsorption and the resulting RNA was fragmented. The RNA fragments then served as a template for first strand cDNA synthesis using random hexamers and reverse transcriptase (Promega, Madison, WI, USA). Second strand cDNA was synthesized using DNA polymerase I and RNaseH, and then purified using a QiaQuick PCR extraction kit (Qiagen, Hilden, Germany). cDNA fragments of a suitable length (300–500 bp) were obtained by agarose gel electrophoresis and amplified by PCR to construct the final cDNA libraries for paired-end sequencing using the Illumina HiSeq 2000 (Biomarker Technologies Co., Ltd, Beijing, China). The transcriptome data were submitted to NCBI, under the accession number SRA109935.
De novo assembly and analysis
Raw sequences of each of the two datasets were assembled using the CLC Genomics Workbench software (CLC Bio, Aarhus, Denmark). Adaptor sequences were removed from each sequence prior to de novo assembly and low quantity reads (reads with more than 50% of bases with Q-value ≤20) were discarded. Clean reads were mapped back to the assembled contigs using Trinity software18 with the parameters set at a similarity of 90%. By performing pair-end joining and gap filling, contigs were assembled into transcripts and were subsequently clustered to obtain unigenes.
SSRs screening and data analysis
MISA software (http://pgrc.ipk-gatersleben.de/misa/) was used to identify potential SSRs within the unigenes.19 SSRs with one to six nucleotides in length were considered in the analysis. The minimum of contiguous repeat units were set as follows: 10 repeat units for mono-nucleotides, six for di-nucleotides and five for tri-, tetra-, penta- and hexa-nucelotides.
Analysis of gene expression
Gene expression of all unigenes in ‘Ventura’ and ‘Jinnan Shiqin’ were estimated by calculating read density as ‘reads per kilobase of exon model per million mapped reads’ (RPKM).20 The DEGs in the two libraries were identified based on the Audic and Claverie’s method.21 IDEG6 software22 (http://compgen.bio.unipd.it/bioinfo/software/) was used to identify the DEGs based on a statistical analysis of large matrices of multiconditional expression data. A threshold for false discovery rate of <0.001 and an absolute value of log 2 ratio >1 were used to determine significant differences in gene expression.
Functional annotation
A sequence similarity search was performed against eight public databases to identify and annotate the obtained unigenes. DEG sequences were compared with the Nr (non-redundant protein database, NCBI) and Nt (non-redundant nucleotide database, NCBI), Swiss-Prot, TrEMBL, GO (Gene Ontology, http://www.geneontology.org/), COG (Clusters of Orthologous Groups) and KEGG (Kyoto Encyclopedia of Genes and Genomes) databases.23–25 Blast2GO was employed to obtain the relevant GO terms and WEGO software was used to illustrate the distribution of gene classification.26,27
Stress treatments and RT-qPCR
Celery plants were grown in plastic pots (size 17×17×20 cm) at a density of four plants per pot, in a controlled-environment growth chamber at 25/16 °C day/night temperatures under 16/8 h light/dark photoperiod and 3000 lux of light intensity. After 2 months, 12 seedlings were transferred to growth chambers which were maintained at either 4 °C or 38 °C, which represented the low and high temperature stress treatments, respectively, under the same photoperiod and light intensity that was previously noted. The other twelve seedlings were irrigated with ddH2O (control), 200 mM NaCl (salt treatment) and 20 % PEG 6000 (drought treatment), respectively. Samples, consisting of young leaves, were collected at 0, 1, 2, 4, 8 and 24 h after the different treatments were initiated. The samples were frozen immediately in liquid nitrogen, and stored at −70 °C. Total RNA was extracted using the total RNA kit (RNAsimply; Tiangen). Reverse transcription into cDNA was conducted using the PrimeScript RT reagent Kit (TaKaRa, Dalian, China). RT-qPCR was performed using a MyiQ Single Color Real Time PCR Detection System (Bio-Rad, Hercules, CA, USA). The PCR conditions were 95 °C for 30 s, followed by 40 cycles of 95 °C for 5 s for denaturation and 60 °C for 20 s for annealing and extension. The experiments were repeated three times, and Agactin-1 was used as an internal control. Relative gene expression was calculated by the method.28 Primers specific to the genes being characterized were designed by ‘Primer Premier 6’ (http://www.premierbiosoft.com/crm/jsp/com/pbi/crm/clientside/ProductList.jsp) with the product lengths ranging from 70–200 bp for RT-qPCR (Supplementary Table 1).
Results
Transcriptome sequencing and de novo assembly
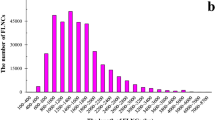
After removing adaptor sequences and discarding low quantity reads, 15 893 516 high-quality reads were obtained from the celery variety ‘Ventura’ and 19 818 161 high-quality reads from the variety ‘Jinnan Shiqin’. Using Trinity assembly software, the reads from ‘Ventura’ were assembled into 404 329 contigs and the reads from ‘Jinnan Shiqin’ into 516 744 contigs with mean lengths of 126 bp and 113 bp and a N50 length of 326 bp and 261 bp, respectively. By performing pair-end joining and gap filling, contigs in these two libraries were assembled into 50 924 and 57 760 transcripts with a mean length of 773 bp and 778 bp and N50 lengths of 1204 bp and 1193 bp for ‘Ventura’ and ‘Jinnan Shiqin’, respectively. The transcripts were then further clustered into unigenes. A total of 39 584 unigenes were obtained in ‘Ventura’ and 41 740 unigenes in ‘Jinnan Shiqin’ with a mean length of 683 bp and 690 bp and a N50 length of 1073 bp and 1088 bp, respectively. The size distribution of unigenes from ‘Ventura’ was quite similar with those obtained in ‘Jinnan Shiqin’ (Figure 1).
SSR markers
A total of 1939 SSRs were identified in ‘Ventura’ and 2004 SSRs in ‘Jinnan Shiqin’ using MISA software. According to the classification of Weber,29 the 1939 SSRs in ‘Ventura’ were divided into 1528 perfect, 273 imperfect and 138 compound repeat sequences, and the 2004 SSRs in ‘Jinnan Shiqin’ were divided into 1595 perfect, 272 imperfect and 137 compound repeat sequences. The lengths of the SSRs in the different groups varied greatly (Table 1) with the largest variation (12–64 bp) being present in di-nucleotide repeats and the smallest variation (15–20 bp) in penta-nucleotide repeats. Mono-, di- and tri-nucleotide repeats accounted for 98.1% and 98.3% of the SSRs in ‘Ventura’ and ‘Jinnan Shiqin’, respectively. Among the different types of SSRs, di-nucleotide repeats were the majority, accounting for 55.49% in ‘Ventura’ and 54.84% in ‘Jinnan Shiqin’, while other types (tetra-, penta- and hexa-nucleotide repeats) exhibited a frequency of less than 2%.
The number of each motif type in ‘Ventura’ and ‘Jinnan Shiqin’ was similar (Table 2). A/T SSRs occurred more frequently in mono-nucleotide repeats, almost 20-fold more than C/G SSRs. The most abundant type of di-nucleotide repeat, AG/CT, accounted for 45.54% and 44.51% of the dinucleotide SSRs in ‘Ventura’ and ‘Jinnan’ Shiqin, respectively. Among the tri-nucleotide SSRs, AAG/CTT, ATC/ATG, ACT/AGT and ACC/GGT, were most frequently observed, while only a small number of CCG/CGG repeats were identified. Overall, mono-, di- and tri-nucleotide SSRs had an A and T nucleotide base preference. Tetra-, penta- and hexa-nucleotide SSRS were only present in small numbers in the obtained transcriptome sequences from the two celery varieties and their base patterns were quite complex and were not subjected to detailed analysis.
Identification and selection of DEGs
In order to compare genes expression in the two celery libraries, unigenes were quantified as reads per kilobase of exon model RPKM. A total of 36 041 unigenes had detectable levels of expression in ‘Jinnan Shiqin’ and ‘Ventura’ (Figure 2a). The set of expressed genes were analyzed for differences in their level of expression in the two celery varities using IDE6 software.22 A threshold for false discovery rate (FDR) of <0.001 and an absolute value of log 2 ratio >1 were used to determine significant differences in gene expression. As graphically illustrated in Figure 2b, a total of 338 genes were found to be differentially expressed in ‘Jinnan Shiqin’ and ‘Ventura’. Among them, 147 genes (red dots) were upregulated and 191 genes (green dots) were downregulated in ‘Ventura’ compared to ‘Jinnan Shiqin’, while the black dots indicate no difference in expression (Figure 2b).
Comparison of levels of gene expression level in ‘Ventura’ and ‘Jinnan Shiqin’. (a) A scatter plot of RPKM logarithmic values in libraries derived from each of the varieties. Each dot represents the RPKM value of a specific gene. The greater the deviation from the diagonal slope indicates that the gene was expressed at a greater level in the corresponding variety. The x-axis represents ‘Ventura’ and the y-axis represents ‘Jinnan Shiqin’. (b) A scatter plot of the ratio of RPKM logarithmic numerical values of genes in the two varieties. This plot graphically represents genes that were differentially expressed in ‘Ventura’. Red dots represent genes that were upregulated, green dots represent genes that were downregulated and black dots represent genes where no significant difference was observed between the two varieties.
Functional annotation of DEGs
To identify the functions of the DEGs, sequence-based alignments were performed against the non-redundant database (Nr/Nt) at NCBI, Swiss-Prot/TrEMBL and the KEGG using the BLASTx algorithm with a threshold E value of 1×10−5. Domain-based alignments were executed in the InterPro and COG databases. A total of 303 DEGs were performed to against to eight public databases and allowed the assignment of several functional annotations (Table 3 and Supplementary Table 2). DEG sequences were searched against the Nr and Nt databases, resulting in 291 and 239 significant BLAST hits, respectively. Sequences were also blasted against the Swiss-Prot and TrEMBL databases in order to obtain more definitive annotations and as a result, 232 and 239 DEGs were annotated, respectively. Furthermore, 212 DEGs had matches in the InterPro database, resulting in 498 annotations.
To obtain a functional categorization of the DEGs, Gene Ontology (GO) analysis was utilized to classify the DEGs. A total of 156 DEGs were assigned to three gene ontology classes: biological process, cellular component and molecular function, with 509 functional terms (Figure 3). Biological process made up the majority of the GO annotations (204 genes or 40.1%), followed by molecular function (165 genes or 32.4%) and cellular component (140 genes or 27.5%). Together, metabolic and cellular processes accounted for more than 68% of the genes assigned to biological process. Among 10 different molecular function categories, ‘binding’ and ‘catalytic’ were the two most frequent terms identified for the genes that were part of the molecular function component. Within the cellular component, the major categories were ‘cell’ and ‘cell part’, which together accounted for 30% of the genes associated with this component. Additionally, a fraction of DEGs were associated other important processes, including transcription regulator, transport, biological regulation and response to stimulus.
GO classification of the DEGs in transcriptomes of ‘Ventura’ and ‘Jinnan Shiqin’. Numbers above each bar represent the actual number of DEGs that were classified in that particular component and descriptive term. y-axis on the right indicates the number of genes in a category as a percent of the total number of DEGs.
Based on the conserved domain-based alignment, 80 DEGs were further compared against COG database and 110 COG-annotated putative proteins were functionally classified into 21 function categories (Figure 4). ‘Post-translational modification, protein turnover, chaperone functions’ was the group with the most genes, followed by ‘General Functional Prediction’, ‘Translation, ribosomal structure and biogensis’ and ‘Signal Transduction metabolism’.
In order to identify the biological functions and pathways in celery, the DEGs were annotated by comparing them against the KEGG database to obtain the corresponding enzyme commission. By mapping enzyme commission numbers to the reference canonical pathways, a total of 285 DEGs were assigned to 53 KEGG pathways. Results showed that the most frequently represented pathways included ‘Photosynthesis’ (PATH:ko00195), ‘Photosynthesis-antenna proteins’ (PATH:ko00196), ‘Ribosome’ (PATH:ko03010) and ‘Plant-pathogen interaction’ (PATH:ko04626), representing 12, 9, 6 and 4 enzymes, respectively.
RT-qPCR analysis of selected DEGs in response to a variety of abiotic stresses
Among annotated genes, eight DEGs (Table 4) associated with stress response, were selected to examine their expression in the two celery varieties in response to a variety of abiotic stresses (cold, heat, drought and salt) (Figure 5 and Supplementary Table 3).
Cold treatment
After 24 h of a cold treatment, celery-10247 increased more than 400-fold in ‘Jinnan Shiqin’, but only 6-fold in ‘Ventura’. Throughout the cold treatment (0–24 h), celery-85905 exhibited little to no change in relative expression in either ‘Jinnan Shiqin’, or ‘Ventura’. Three genes (celery-75186, celery-20717 and celery-13876) expressed a pattern of increased expression in both celery varieties over the course of the cold treatment.
Heat treatment
Four genes (celery-89505, celery-28253, celery-20717 and celery-18591) were downregulated in response to heat stress in both ‘Ventura’ and ‘Jinanashiqin’. In contrast, two genes (celery-10247 and celery-75186) were upregulated in both ‘Ventura’ and ‘Jinanashiqin’, but to a higher relative level in ‘Jinnan Shiqin’.
Salt treatment
In response to the salt treatment, celery-89505, celery-28253 and celery-18591 were downregulated in ‘Jinnan Shiqin’ and celery-28253 was downregulated in ‘Ventura’. The gene, celery-13876 increased rapidly and reached a maximum at 1 h after salt treatment in ‘Ventura’ and ‘Jinnan Shiqin’.
Drought treatment
In response to the drought treatment, three genes (celery-85905, celery-42789 and celery-28253) exhibited a relatively low level of expression in both varieties, whereas celery-13876 and celery-10247 increased rapidly in ‘Jinnan Shiqin’ and remained elevated, whereas in ‘Ventura’, these genes initially increased but later decreased.
Discussion
De novo transcriptome sequencing and Trinity assembly
Molecular, genomic and molecular breeding research on celery has been restricted due to the limited availability of genomic and transcriptomic sequence data. High-throughput RNA-seq can generate massive amounts of data on whole-genome-wide transcripts and has become more cost efficient.30 In the present study, 39 584 and 41 740 unigenes were identified in the celery varieties ‘Ventura’ and ‘Jinnan Shiqin’, respectively. These unigenes can serve as a reference base for the future identification of trait-associated single-nucleotide polymorphism and SSR markers that can be used in marker-assisted breeding programs. The data can also serve as a resource for gene expression analyses, and the identification and characterization of celery genes with horticultural importance.31–33
Characteristics and evolutionary relationships of SSR markers
Among the SSRs identified in the two celery varieties, di-nucleotide repeats had the highest frequency, followed by mono- and tetra-nucleotide repeats. This distribution is similar to that reported in sesame,34 peach35 and kiwifruit.36 In contrast, tetra-nucleotide repeats dominate in species such as bread wheat,37 grape38 and sugarcane.39 The presence of a large number of short repeat sequences generally indicates a more rapid rate of evolution in a species, as long-repeat sequences frequently appear in species with a lower mutation rates.5,40 Celery contains a large number of short repeat motifs; therefore, it may be subject to more rapid rates of mutation and thus evolution. The difference between the percentages of different types of SSRs and the base preferences indicates that the two varieties are genetically very close.
An analysis of the types of motifs and the composition of their nucleotides in the two varieties indicated that AG/CT and A/T were the two most abundant types. The low frequency of C/G may be due to the methylation of C residues which results in the conversion to a T.41 A low frequency of CCG/CGG repeats in tetra-nucleotide SSRs in both varieties is consistent with other eucotyledonous plants such as Arabidopsis42 and soybean.43 CCG/CGG repeats, however, have been found to be dominant in some monocotyledonous plants, such as rice and maize.44 Monocotyledonous plants usually have a high G/C content and the preference of codon usage between different species may contribute to this difference.45,46 The SSRs identified in this study represent a valuable resource for the genetic analyses of different celery genotypes and marker-assisted selection in the future.
Selection and functional annotation of DEGs
Another major outcome of transcriptome sequencing is the ability to identify and characterize expressed genes.47 ‘Jinnan Shiqin’ from China and ‘Ventura’ from USA are the two most popular varieties grown in their respective countries. They are similar in morphology, but ‘Jinnan Shiqin’ is preferred by Chinese farmers because it exhibits strong resistance to abiotic stresses. However, no genetic or molecular evidence has been reported to support this observation. The transcriptome data obtained in the present study identified differentially expressed unigenes in the two varieties. The potential function of the majority of the annotated DEGs was assigned to metabolic processes based on sequence similarity of genes in public databases. This information provides a basis for further studies of important compounds in celery, such as apigenin and flavonoids.
Gene expression analysis in abiotic stresses
Environmental stresses, such as drought, salinity, high and low temperature, seriously restrict plant growth and development, reduce yields, and if extreme, result in plant death. Genes that ameliorate the impact of abiotic stress represent a valuable resource for the genetic improvement of plants.48,49 Most stress-resistance traits, however, are often regulated by multiple genes.50,51 In the present study, eight of the identified DEGs, selected based on their association with stress response, also responded to stress stimulation in celery. This result is consistent with the annotation of these DEGs. Although the specific functions of these genes remains to be demonstrated, they appear to be involved in of the temporal response of celery to abiotic stress, as some responded within 1 h, while others responded much later in the course of the stress treatment. As previously mentioned, farmers in China prefer to grow the‘Jinnan Shiqin’ variety celery over ‘Ventura’ due to its ability to adapt to environmental stress. In the present study, celery-10247 exhibited increasing levels of expression in ‘Jinnan Shiqin’, but not in ‘Ventura’ in response to four different abiotic stresses (cold, heat, salt and drought). In cold treatment, the expression levels of all eight of the selected DEGs exhibited upregulation in ‘Jinnan Shiqin’, whereas only three of the genes genes were upregulated in ‘Ventura’. These genes may play a role in the differential ability of the two celery varieties to adapt to different environmental and climatic conditions. Further functional characterization is in progress to determine the specific functions of these genes in relation to cold tolerance in ‘Jinnan Shiqin’ celery.
References
Krishnamurthy K . 22 Celery. In: Chemistry of Spices. Cambridge, MA: CABI; 2008. p401.
Craig WJ . Health-promoting properties of common herbs. Am J Clin Nutr 1999; 70(3 Suppl): 491S–499S.
Raffo A, Sinesio F, Moneta E, Nardo N, Peparaio M, Paoletti F . Internal quality of fresh and cold stored celery petioles described by sensory profile, chemical and instrumental measurements. Eur Food Res Technol 2005; 222: 590–599.
Hostetler GL, Riedl KM, Schwartz SJ . Effects of food formulation and thermal processing on flavones in celery and chamomile. Food Chem 2013; 141: 1406–1411.
Toth G, Gaspari Z, Jurka J . Microsatellites in different eukaryotic genomes: survey and analysis. Genome Res 2000; 10: 967–981.
Rongwen J, Akkaya MS, Bhagwat AA, Lavi U, Cregan PB . The use of microsatellite DNA markers for soybean genotype identification. Theor Appl Genet 1995; 90: 43–48.
Gharghani A, Zamani Z, Talaie A et al. Genetic identity and relationships of Iranian apple (Malus × domestica Borkh.) cultivars and landraces, wild Malus species and representative old apple cultivars based on simple sequence repeat (SSR) marker analysis. Genet Resour Crop Evol 2009; 56: 829–842.
Chen XM, Luo YH, Xia XC et al. Chromosomal location of powdery mildew resistance gene Pm16 in wheat using SSR marker analysis. Plant Breeding 2005; 124: 225–228.
Marquez Y, Brown JW, Simpson C, Barta A, Kalyna M . Transcriptome survey reveals increased complexity of the alternative splicing landscape in Arabidopsis. Genome Res 2012; 22: 1184–1195.
Garg R, Patel RK, Jhanwar S et al. Gene discovery and tissue-specific transcriptome analysis in chickpea with massively parallel pyrosequencing and web resource development. Plant Physiol 2011; 156: 1661–1678.
Yuan Y, Song L, Li M et al. Genetic variation and metabolic pathway intricacy govern the active compound content and quality of the Chinese medicinal plant Lonicera japonica thunb. BMC Genomics 2012; 13: 195.
Yang SS, Tu ZJ, Cheung F et al. Using RNA-Seq for gene identification, polymorphism detection and transcript profiling in two alfalfa genotypes with divergent cell wall composition in stems. BMC Genomics 2011; 12: 199.
Zhang R, Marshall D, Bryan GJ, Hornyik C . Identification and characterization of miRNA transcriptome in potato by high-throughput sequencing. PloS ONE 2013; 8: e57233.
Venturini L, Ferrarini A, Zenoni S et al. De novo transcriptome characterization of Vitis vinifera cv. Corvina unveils varietal diversity. BMC Genomics 2013; 14: 41.
Chen X, Zhu W, Azam S et al. Deep sequencing analysis of the transcriptomes of peanut aerial and subterranean young pods identifies candidate genes related to early embryo abortion. Plant Biotechnol J 2013; 11: 115–127.
Fu N, Wang Q, Shen HL . De novo assembly, gene annotation and marker development using illumina paired-end transcriptome sequences in celery (Apium graveolens L.). PloS ONE 2013; 8: e57686.
Li H . A celery variety—‘Jinnan Shiqin’. Sci Technol Tianjin Agric For 2004; (04): 21.
Grabherr MG, Haas BJ, Yassour M et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 2011; 29: 644–652.
Thiel T, Michalek W, Varshney RK, Graner A . Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). TAG Theor Appl Genet 2003; 106: 411–422.
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B . Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 2008; 5: 621–628.
Audic S, Claverie JM . The significance of digital gene expression profiles. Genome Res 1997; 7: 986–995.
Romualdi C, Bortoluzzi S, d’Alessi F, Danieli GA . IDEG6: a web tool for detection of differentially expressed genes in multiple tag sampling experiments. Physiol Genomics 2003; 12: 159–162.
Ashburner M, Ball CA, Blake JA et al. Gene Ontology: tool for the unification of biology. Nat Genet 2000; 25: 25–29.
Tatusov RL, Fedorova ND, Jackson JD et al. The COG database: an updated version includes eukaryotes. BMC Bioinformatics 2003; 4: 41.
Kanehisa M, Goto S . KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res 2000; 28: 27–30.
Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M . Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics (Oxford, England) 2005; 21: 3674–3676.
Ye J, Fang L, Zheng H et al. WEGO: a web tool for plotting GO annotations. Nucleic Acids Res 2006; 34: W293–W297.
Pfaffl MW . A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 2001; 29: e45.
Weber JL . Informativeness of human (dC–dA)n.(dG–dT)n polymorphisms. Genomics 1990; 7: 524–530.
Wang Z, Gerstein M, Snyder M . RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 2009; 10: 57–63.
Nicolai M, Pisani C, Bouchet JP, Vuylsteke M, Palloix A . Discovery of a large set of SNP and SSR genetic markers by high-throughput sequencing of pepper (Capsicum annuum). Genet Mol Res 2012; 11: 2295–2300.
Strozycki PM, Szymanski M, Szczurek A, Barciszewski J, Figlerowicz M . A new family of ferritin genes from Lupinus luteus—comparative analysis of plant ferritins, their gene structure, and evolution. Mol Biol Evol 2010; 27: 91–101.
Bachlava E, Taylor CA, Tang S et al. SNP discovery and development of a high-density genotyping array for sunflower. PloS ONE 2012; 7: e29814.
Wei W, Qi X, Wang L et al. Characterization of the sesame (Sesamum indicum L.) global transcriptome using Illumina paired-end sequencing and development of EST-SSR markers. BMC Genomics 2011; 12: 1–13.
Jung S, Abbott A, Jesudurai C, Tomkins J, Main D . Frequency, type, distribution and annotation of simple sequence repeats in Rosaceae ESTs. Funct Integr Genomics 2005; 5: 136–143.
Fraser LG, Harvey CF, Crowhurst RN, Silva HN . EST-derived microsatellites from Actinidia species and their potential for mapping. Theor Appl Genet 2004; 108: 1010–1016.
Gupta PK, Rustgi S, Sharma S, Singh R, Kumar N, Balyan HS . Transferable EST-SSR markers for the study of polymorphism and genetic diversity in bread wheat. Mol Genet Genomics 2003; 270: 315–323.
Scott KD, Eggler P, Seaton G et al. Analysis of SSRs derived from grape ESTs. Theor Appl Genet 2000; 100: 723–726.
Cordeiro GM, Casu R, McIntyre CL, Manners JM, Henry RJ . Microsatellite markers from sugarcane (Saccharum spp.) ESTs cross transferable to erianthus and sorghum. Plant Sci 2001; 160: 1115–1123.
Harr B, Schlotterer C . Long microsatellite alleles in Drosophila melanogaster have a downward mutation bias and short persistence times, which cause their genome-wide underrepresentation. Genetics 2000; 155: 1213–1220.
Schorderet DF, Gartler SM . Analysis of CpG suppression in methylated and nonmethylated species. Proc Natl Acad Sci USA 1992; 89: 957–961.
Depeiges A, Goubely C, Lenoir A et al. Identification of the most represented repeated motifs in Arabidopsis thaliana microsatellite loci. Theor Appl Genet 1995; 91: 160–168.
Xin D, Sun J, Wang J et al. Identification and characterization of SSRs from soybean (Glycine max) ESTs. Mol Biol Rep 2012; 39: 9047–9057.
Morgante M, Hanafey M, Powell W . Microsatellites are preferentially associated with nonrepetitive DNA in plant genomes. Nat Genet 2002; 30: 194–200.
Zhou M, Tong CF, Shi JS . A preliminary analysis of synonymous codon usage in poplar species. Zhi Wu Sheng Li Yu Fen Zi Sheng Wu Xue Xue Bao 2007; 33: 285–293.
Morton BR, Wright SI . Selective constraints on codon usage of nuclear genes from Arabidopsis thaliana. Mol Biol Evol 2007; 24: 122–129.
Tao X, Gu YH, Wang HY et al. Digital gene expression analysis based on integrated de novo transcriptome assembly of sweet potato [Ipomoea batatas (L.) Lam]. PloS ONE 2012; 7: e36234.
Ashraf M . Inducing drought tolerance in plants: recent advances. Biotechnol Adv 2010; 28: 169–183.
Ashraf M, Foolad MR . Crop breeding for salt tolerance in the era of molecular markers and marker-assisted selection. Plant Breeding 2013; 132: 10–20.
Yue B, Xue W, Xiong L et al. Genetic basis of drought resistance at reproductive stage in rice: separation of drought tolerance from drought avoidance. Genetics 2006; 172: 1213–1228.
Yang Q, Chen ZZ, Zhou XF et al. Overexpression of SOS (Salt Overly Sensitive) genes increases salt tolerance in transgenic Arabidopsis. Mol Plant 2009; 2: 22–31.
Acknowledgements
The research was supported by the National Natural Science Foundation of China (31272175), New Century Excellent Talents in University (NCET-11-0670), Jiangsu Natural Science Foundation (BK20130027), Graduate Educated Innovation Project of Jiangsu Province (CXZZ13_0297) and Priority Academic Program Development of Jiangsu Higher Education Institutions and Jiangsu Shuangchuang Project.
Author information
Authors and Affiliations
Contributions
ASX conceived and designed the experiments. MYL, FW, QJ and JM performed the experiments. MYL, FW, JQ and ASX analyzed the data. ASX contributed reagents/materials/analysis tools. MYL wrote the paper. MYL and ASX revised the paper. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0
About this article
Cite this article
Li, MY., Wang, F., Jiang, Q. et al. Identification of SSRs and differentially expressed genes in two cultivars of celery (Apium graveolens L.) by deep transcriptome sequencing. Hortic Res 1, 10 (2014). https://doi.org/10.1038/hortres.2014.10
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/hortres.2014.10
This article is cited by
-
AgMYB5, an MYB transcription factor from celery, enhanced β-carotene synthesis and promoted drought tolerance in transgenic Arabidopsis
BMC Plant Biology (2023)
-
Integrative genome, transcriptome, microRNA, and degradome analysis of water dropwort (Oenanthe javanica) in response to water stress
Horticulture Research (2021)
-
The Accumulation of Lutein and β-Carotene and Transcript Profiling of Genes Related to Carotenoids Biosynthesis in Yellow Celery
Molecular Biotechnology (2021)
-
The genome sequence of celery (Apium graveolens L.), an important leaf vegetable crop rich in apigenin in the Apiaceae family
Horticulture Research (2020)
-
Effect of Elevated CO2 on Ascorbate Accumulation and the Expression Levels of Genes Involved in Ascorbate Metabolism in Celery
Journal of Plant Growth Regulation (2020)