Abstract
The N-acetyltransferases (NATs) are xenobiotic-metabolizing enzymes involved in the metabolism of drugs, environmental toxins and the aromatic amine carcinogens present in cigarette smoke. Genetic variations in NAT2 have long been recognized as the cause of variable enzymatic activity or stability, leading to slow or rapid acetylation. In the present study, we genotyped three single-nucleotide polymorphisms (SNPs) from the NAT2 gene (rs1799929, rs1799930 and rs1799931), using TaqMan allelic discrimination, among 212 individuals from six major South Indian populations and compared the results with other available Indian and worldwide data. All three of the markers followed Hardy–Weinberg equilibrium and were highly polymorphic in the studied populations. The constructed haplotypes showed a high level of heterozygosity. All of the populations in the present study commonly shared only four haplotypes out of the eight possible three-site haplotypes. The haplotypes exhibited fairly high frequencies across multiple populations, where three haplotypes were shared by all six populations with a cumulative frequency ranging from 88.2% (Madiga) to 97.0% (Balija). We also observed a tribal-specific haplotype. A strong linkage disequilibrium (LD) between rs1799929 and rs1799930 was consistent in all of the studied populations, with the exception of the Madiga. A comparison of the genomic regions 20-kb up- and downstream of rs1799930 in a large number of worldwide samples showed a strong LD of this SNP with another NAT2 SNP, rs1112005, among the majority of the populations. Moreover, our lifestyle test (hunter–gatherer versus agriculturist) in comparison with the NAT2 variant suggested that two of the studied populations (Balija and Madiga) have likely shifted their diet more recently.
Similar content being viewed by others
Introduction
The N-acetyltransferases (NAT; EC 2.3.1.5) are involved in the initial biotransformation metabolism of aromatic amines and hydrazines and catalyzes the transfer of the acetyl group from acetyl CoA to the nitrogen of the substrate.1 NATs are polymorphic in the population and metabolize different types of carcinogens that have been directly implicated in tumor progression. The NAT2 gene (MIM no. - 243400), which codes for the NAT2 protein, is located on chromosome 8p22 and spans 9.9 kb. The human NAT2 gene has two exons, with a noncoding exon at the 5′ end and an uninterrupted coding region (exon 2) of 870 nucleotides that encodes a 290 amino-acid protein. The acetylation polymorphism (rapid or slow acetylator phenotype) was discovered more than 60 years ago following differences observed in tuberculosis patients to isoniazid toxicity.2 Genetic variations in the NAT2 gene have been known to result in individual acetylation polymorphisms. To date, 66 alleles have been described; however, many of these NAT2* alleles share sequence variations, and these sequence variations do not always lead to changes in the enzyme activity of the encoded protein (http://nat.mbg.duth.gr/Human%20NAT2%20alleles_2013.htm).
The NAT2 c.282C>T and c.481C>T polymorphisms are silent mutations that do not change the tyrosine at amino acid 94 (p.Tyr94Tyr) or the leucine at amino acid 161 (p.Leu161Leu), respectively, in the NAT2 protein. On the converse, the NAT2 c.191G>A and c.590G>A polymorphisms change the charged arginines to polar glutamines at residues 64 (p.Arg64Gln) and 197 (p.Arg197Gln), respectively. The NAT2 c.341T>C polymorphism alters the hydropathy profile of the protein by changing isoleucine to the more polar threonine at amino acid 114 (p.Ile114Thr).3 The NAT2 c.803G>A polymorphism is considered to be a conservative amino-acid change because it changes lysine to an arginine at position 268 (p.Arg268Lys). The NAT2 c.857G>A polymorphism changes glycine 286 to the more polar glutamic acid (p.Gly286Glu) in the C-terminal tail. Early genotyping studies have screened the c.481C>T, c.590G>A, c.857G>A and sometimes the c.191G>A polymorphisms and suggested their potential role in slowing down the acetylation phenotype.4 The c.481C>T single-nucleotide polymorphism (SNP; rs1799929: p.Leu161Leu) is a silent mutation designated as NAT2*11A, and it is not involved in altering the acetylator phenotype. The nucleotide substitutions at c.590G>A (rs1799930) and c.857G>A (rs1799931) are designated as NAT2*6 and NAT2*7, respectively, and the combination of these alleles causes the slow acetylator phenotype. Furthermore, a threefold decrease in clearance was reported between rapid acetylators and slow acetylators.5 Phenotype studies in the last five decades have discovered distinct ethnic differences for slow acetylator frequencies.6 Region wise, in the Caucasian and African populations, the frequency of the slow acetylation phenotype varies between 40 and 70%, whereas in the Asian population, in particular East and Southeast Asian populations, such as Japanese, Chinese, Korean and Thai, it ranges from 10 to 30%.7
Acetylation is a major biotransformation route for many arylamine and hydrazine drugs, as well as for a number of toxins and known carcinogens present in some diets, cigarette smoke and in the environment.8 Therefore, it is important to study the NAT2 gene to understand the molecular basis of biotransformation. Attempts have been made to study the NAT2 polymorphisms at the global level,9 and because of their unique population structuring,10 Indian populations may provide an immense opportunity to test various assumptions that were previously made.9 Therefore, in order to study the genetic variations of the NAT2 gene in six major Indian populations, three SNPs (rs1799929, rs1799930 and rs1799931) were selected for genotyping analysis. There are some potential limitations in assigning NAT2 alleles/haplotypes and deducing phenotypes solely from these three SNPs; however, in our preliminary analysis using HapMap GIH (Gujarati—a West Indian population) samples, we found these SNPs to be highly polymorphic and present in separate linkage disequilibrium (LD) blocks with many other potential SNPs. As GIH is the closest population group to the six ethnic groups analyzed in this study, it is likely that these three SNPs are the best fit for this study.
Materials and Methods
Subjects
The study populations included a total of 212 unrelated males belonging to six ethnic groups. Of these groups, Reddy belongs to the upper caste, Balija to a backward caste, Mala, Madiga and Sugali are from a scheduled tribe, whereas Muslims are a religious group. All of the populations, with the exception of Muslims (who are a religious group), speak a branch of the Dravidian linguistic group, which is found in the southern peninsula of India. The names of the populations, sample sizes, linguistic affiliations and place of inhabitation are presented in Table 1. All of the subjects were apparently normal healthy volunteers, and no diagnosis was performed on the individuals. All of the participants provided written informed consent. The procedures for the protection of the human subjects in this study were approved by the Institutional Ethical Review Committee of Narayana Medical College, Nellore, India. Intravenous blood samples (~3 ml each) were collected, and the DNA was isolated using a standard protocol.11
Genotyping
Three SNPs from the NAT2 gene c.481C>T (p.L161L, dbSNP rs1799929), c.590G>A (p.R197Q, dbSNP rs1799930) and c.857G>A (p.G286E, dbSNP rs1799931) were genotyped in the 212 individuals. The primers and probes for all of the SNPs used in this study were purchased from Applied Biosystems (Foster City, CA, USA). Each reaction contained 2.5 μl of TaqMan Universal PCR Master Mix, 0.125 μl of TaqMan SNP Genotyping Assay, 1.375 μl of distilled water and 1 μl of DNA (10 ng/μl), with a final reaction volume of 5 μl. For each SNP assay, a positive control for the wild-type, heterozygote and variant genotype was provided with at least two negative controls. Before analyzing the DNA, a pilot test was conducted to confirm the accuracy of the assay. After a successful pilot test, the sample analysis was carried out in 384-well optical reaction microplates (Applied Biosystems). The fluorescence was measured using an Applied Biosystems 7900HT Fast Real-Time PCR System and analyzed with its System SDS software, version 2.3.
Statistical analysis
The allele frequencies in each population were determined via direct counting. The Hardy–Weinberg equilibrium ratios were calculated using the software HWSIM, a DOS-based program.12 As phase-unknown genotypes were collected, the haplotype sites and frequencies were estimated using maximum likelihood with an expectation-maximization method in Arlequin.13 The LD (D’ and r2) was estimated using the program HaploView 3.12.14 For a worldwide comparison in a wider context, we also extracted the regions 20-kb up- and downstream of the SNP rs1799930 from a large pool of populations distributed worldwide.10,15 The physical map for the 20-kb regions up- and downstream of rs1799930 is shown in Supplementary Figure S1. The data were phased using BEAGLE.16 A homozygosity analysis was conducted using PLINK 1.07 (A sliding window of 25 SNPs was surveyed for the homozygosity).17 The threshold of homozygosity match was 0.99.
Results
The location, sequence and wild-type allele for the polymorphic sites, in addition to their NCBI reference IDs, are shown in Table 2. The wild-type and mutant alleles of all of the SNPs used in the present study (rs1799929, rs1799930 and rs1799931) were designated according to their respective nucleotides. All of the individual site allele frequencies for all of the populations are given in Table 3.
Consistent with our preliminary observations regarding the HapMap GIH populations, all three of the markers remained highly polymorphic in the studied populations. All of the SNPs of NAT2 followed Hardy–Weinberg proportions. The minor allele frequency (MAF) of rs1799929 was found to be greater than 23%, with a minimum of 23.5% (Madiga) and a maximum of 33.8% in the Balija populations. The rs1799930 site showed a minimum minor allele frequency of 30.9% (Balija) and a maximum of 44.4% (Sugali). A subsequent MAF analysis of the MAF data for rs1799930 in other world populations showed wide variations, with the highest frequencies in the GIH population (35.2%) followed by the MKK population (30.4%); the lowest frequencies were in the CHB (19%) and MEX (18%) HapMap populations. Similarly, the Indian populations also exhibited wide variations, ranging from 6.2% in the Tibeto-Burman to 42.3% in the Austroasiatic populations (Supplementary Table S1). It is interesting to note that, although the migrations of the Austroasiatic and Tibeto-Burman populations have been suggested to come from the East,18,19 the present study confirms that the East/Southeast Asian ancestry of both of the groups is derived from at least two distinct sources. Moreover, the rs1799931 SNP exhibited a lower MAF than the other two SNPs. The MAF of rs1799931 was found to be minimum (3%) in the Balija and maximum (11.8%) in the Madiga. The χ2 test, to determine departure from the single site Hardy–Weinberg expectations, was applied to each NAT2 polymorphic site in each population, and the results are shown in Table 3. The genotype frequencies for all of the polymorphisms meet the Hardy–Weinberg expectations in all of the populations.
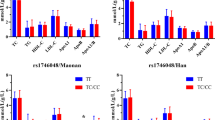
The haplotypes are coded according to the site order in Table 2. The haplotype-based analysis showed striking variations among the populations (Figure 1). Of the eight possible three-site haplotypes, only three haplotypes were shared by all of the populations, with a cumulative frequency ranging from 88.2% (Madiga) to 97.0% (Balija). The wild-type haplotype (CGG) was shared by all of the populations and its frequency ranged from 18.8% in Muslims to 35.3% in the Balija, where it is a major haplotype. In all of the populations, with the exception of the Balija, CAG is the major haplotype, with a frequency ranging from 35.5% (Mala) to 44.4% (Sugali); TGG is the second most prominent haplotype in Reddy (32.6%), Sugali (29.6%), Muslim 32.5% and Mala (30.0%) subjects, and the CGA haplotype was only found to be more than 5% in the Mala (10%) and Madiga (11.8%) populations, which belong to the Scheduled tribe group, suggesting that CGA is a tribal-specific haplotype.
The inferred NAT2 haplotype frequencies in the different Indian populations. The haplotypes are coded according to the site order in Table 2.
To calculate the pairwise LD for all three of the NAT2 SNPs in the studied populations, we used the two most common LD measures, D′ and r2. A strong LD between rs1799929 and rs1799930 was observed in all of the populations, except the Madiga (D′=1.0; r2=0.19; Figure 2). None of the populations had significant LD between rs1799929 and rs1799931. Similarly, rs1799930 and rs1799931 did not reveal any strong LD because of low heterozygosity at the rs1799931 locus. Comparison of the 20-kb up- and downstream regions surrounding rs1799930 in large number of worldwide samples revealed the strong LD of this SNP with another NAT2 SNP (rs1112005) in the majority of the populations (Supplementary Figure S2). Interestingly, the homozygosity analysis of the Indian populations showed that the caste populations, who mostly depend upon agriculture and a vegetarian diet, did not have any homozygous blocks in this region (Table 4).
NAT2*4 refers to the NAT2 reference sequence (GenBank accession X14672). The NAT2*4 allele acts dominantly to result in rapid acetylation, and the presence of a mutant genotype (rs1799929, rs1799930 and rs1799931) leads to slow acetylation.20 On the basis of this assumption, the acetylator status of all of the samples in each population under analysis was determined. Samples possessing at least two mutant alleles were considered to be slow acetylators (Table 5). In several of the populations (Reddy, Mala, Sugali and Muslim), the prevalence of slow acetylators was greater than rapid acetylators, whereas the Balija and Madiga populations showed more rapid acetylators (52.9%) than slow acetylators (47.1%; Figure 3).
Discussion
The NAT2 enzyme is capable of N-acetylation, O-acetylation and N,O-acetylation, and it is implicated in the detoxification of a wide spectrum of naturally occurring xenobiotics, including carcinogens and drugs.21 The acetylator phenotype is determined by studying the acetylation of certain drugs, such as sulfadimidine, isoniazid (INH) dapsone or caffeine;22 therefore, NAT2 gene polymorphisms have been linked to human acetylation capacity, which alters susceptibility to cancer and adverse drug reactions.23 In the present study, we analyzed three SNPs: rs1799929 (p.Leu161Leu), which is a silent mutation that does not alter the acetylator phenotype, whereas the remaining two SNPs result in amino-acid substitutions (rs1799930: p.Arg197Gln; rs1799931: p.Gly286Glu) that lead to a significant decrease in acetylation capacity. Genotypic screening of these primary NAT2 polymorphisms in South Indian populations revealed high variations in their allele frequency spectra but without any deviation from Hardy–Weinberg equilibrium in any of the populations. Only three major haplotypes carry 88–97% of the chromosomes among all of the studied populations. A strong LD between rs1799929 and rs1799930 was found in five out of the six studied populations. Among the other Indian populations, the LD varied considerably. The SNP rs1799930 was not in LD with the majority of the SNPs in the Austroasiatic and Indo-European linguistic groups, whereas in the Dravidians, the same marker was in LD with many other markers, forming the largest LD block (Supplementary Figure S3). Notably, the Onge, Siddi and Tibeto-Burman groups did not have any other markers in LD with this SNP. The great variation in LD among the Indian populations is likely because of the heterogeneity caused by the combination of drift and selection due to various environmental and cultural pressures (Supplementary Figure S3).
The frequency of c.481C>T was found to be 7–9% in African24 and 3.8% in Thai populations.25 A higher frequency of c.481C>T was reported (30%) in Brazilians, Spanish, Iranians, Emiratis and African Americans.26–30 A strong LD between NAT2*11A (c.481C>T) and 2*6 (c.590G>A) has been reported in Indian31 and Brazilian admixed populations;32 therefore, the c.481T variant is sufficient for detecting NAT2*11A normal acetylators.33 The low frequency of the c.857G>A polymorphism in many populations reduces the predictive power of slow acetylators. A recent study using the SNPs rs1801280 and rs1799930 showed five different phenotype categories (corresponding to the genotypes NAT2*4/*4, NAT2*4/*5 or *4/*6, NAT2*5/*5, NAT2*5/*6 and NAT2*6/*6) that corresponded to a higher or lower acetylation capacity in vivo.34 Using accurate and reliable estimates for the NAT2 haplotype frequencies and the individual haplotype phases, no LD was observed between NAT2*5 and 2*6 in the Spanish, Korean and Black South African samples.35 Analyses of seven NAT2 SNPs revealed interethnic variability for some of the SNPs but intraethnic variability for other SNPs; c.282C>T, c.806A>G and c.857G>A have not shown any major intraethnic variability.36
The adaptation of agricultural practices from a hunter–gatherer lifestyle has allowed humans to shift their diet towards less protein and more starchy foods. This shift in diet has created various selective pressures on the human genome, which acts on the genetic variations in different human populations. The relationship between human dietary habits and NAT2 has been hypothesized in several studies.9,37 It was suggested that the hunter–gatherers carried rapid acetylator phenotypes, whereas agriculturists possess the slow acetylator phenotype.38,39 In our analysis, all of the populations predominantly carried the slow acetylator phenotype, with the exception of the Balija and Madiga (Figure 3), suggesting that the dietary shift of these two populations from the hunter–gatherer lifestyle likely happened recently. The 20-kb up- and downstream genomic analysis also showed a similar pattern, where the tribal and lower caste populations showed a variable number of homozygous blocks in the analyzed region (Table 5), suggesting their association with the rapid acetylator phenotype.
Therefore, our analyses of the NAT2 gene polymorphisms in a large body of data have provided strong support for evidence of a high diversity of slow acetylator phenotypes in the majority of the world’s populations, which has been shaped by natural selection40 and population-specific selective pressures acting on this locus.38 Moreover, we also contribute to the knowledge regarding the distribution of NAT2 gene polymorphisms among the various ethnic groups of India. We observed that because of the vast amount of genetic, cultural and phenotypic variations found in Indian populations, all of the studied SNPs have great variations in the LD among the Indian populations, and a complete study of the NAT2 gene in these populations will help in drawing more inferences about the processes that have led to the current gene frequency patterns.
References
Riddle B, Jencks WP . Acetyl-coenzyme A: arylamine N-acetyltransferase. Role of the acetyl-enzyme intermediate and the effects of substituents on the rate. J Biol Chem 1971; 246: 3250–3258.
Hughes HB, Biehl JP, Jones AP, Schmidt LH . Metabolism of isoniazid in man as related to the occurrence of peripheral neuritis. Am Rev Tuberc 1954; 70: 266–273.
Vatsis KP, Martell KJ, Weber WW . Diverse point mutations in the human gene for polymorphic N-acetyltransferase. Proc Natl Acad Sci USA 1991; 88: 6333–6337.
Hickman D, Risch A, Camilleri JP, Sim E . Genotyping human polymorphic arylamine N-acetyltransferase: identification of new slow allotypic variants. Pharmacogenetics 1992; 2: 217–226.
Dorne JL, Walton K, Renwick AG . Polymorphic CYP2C19 and N-acetylation: human variability in kinetics and pathway-related uncertainty factors. Food Chem Toxicol 2003; 41: 225–245.
Evans DA . N-acetyltransferase. Pharmacol Ther 1989; 42: 157–234.
Meyer UA, Zanger UM . Molecular mechanisms of genetic polymorphisms of drug metabolism. Annu Rev Pharmacol Toxicol 1997; 37: 269–296.
Hein DW . Molecular genetics and function of NAT1 and NAT2: role in aromatic amine metabolism and carcinogenesis. Mutat Res 2002; 506–507: 65–77.
Sabbagh A, Darlu P, Crouau-Roy B, Poloni ES . Arylamine N-acetyltransferase 2 (NAT2) genetic diversity and traditional subsistence: a worldwide population survey. PLoS ONE 2011; 6: e18507.
Reich D, Thangaraj K, Patterson N, Price AL, Singh L . Reconstructing Indian population history. Nature 2009; 461: 489–494.
Sambrook J, Fitsch EF, Maniatis T . Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Press: Cold Spring Harbor, 1989.
Cubells JF, Kobayashi K, Nagatsu T, Kidd KK, Kidd JR, Calafell F et al. Population genetics of a functional variant of the dopamine beta-hydroxylase gene (DBH). Am J Med Genet 1997; 74: 374–379.
Schneider S, Roessli D, Excoffier L . Arlequin v.2.0: a Software for Population Genetics Data Analysis. Genetics and Biometry Laboratory, University of Geneva: Geneva, Switzerland, 2000.
Barrett JC, Fry B, Maller J, Daly MJ . Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 2005; 21: 263–265.
Tanaka T . [HapMap project]. Nihon Rinsho 2009; 67: 1068–1071.
Browning BL, Browning SR . A fast, powerful method for detecting identity by descent. Am J Hum Genet 2011; 88: 173–182.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 2007; 81: 559–575.
Metspalu M, Kivisild T, Metspalu E, Parik J, Hudjashov G, Kaldma K et al. Most of the extant mtDNA boundaries in south and southwest Asia were likely shaped during the initial settlement of Eurasia by anatomically modern humans. BMC Genet 2004; 5: 26.
Chaubey G, Metspalu M, Choi Y, Magi R, Romero IG, Soares P et al. Population genetic structure in Indian Austroasiatic speakers: the role of landscape barriers and sex-specific admixture. Mol Biol Evol 2011; 28: 1013–1024.
Cascorbi I, Drakoulis N, Brockmoller J, Maurer A, Sperling K, Roots I . Arylamine N-acetyltransferase (NAT2) mutations and their allelic linkage in unrelated Caucasian individuals: correlation with phenotypic activity. Am J Hum Genet 1995; 57: 581–592.
Lee MS, Su L, Christiani DC . Synergistic effects of NAT2 slow and GSTM1 null genotypes on carcinogen DNA damage in the lung. Cancer Epidemiol Biomarkers Prev 2010; 19: 1492–1497.
Sabbagh A, Darlu P . SNP selection at the NAT2 locus for an accurate prediction of the acetylation phenotype. Genet Med 2006; 8: 76–85.
Hein DW, Doll MA . Accuracy of various human NAT2 SNP genotyping panels to infer rapid, intermediate and slow acetylator phenotypes. Pharmacogenomics 2012; 13: 31–41.
Dandara C, Masimirembwa CM, Magimba A, Kaaya S, Sayi J, Sommers DK et al. Arylamine N-acetyltransferase (NAT2) genotypes in Africans: the identification of a new allele with nucleotide changes 481C>T and 590G>A. Pharmacogenetics 2003; 13: 55–58.
Kukongviriyapan V, Prawan A, Tassaneyakul W, Aiemsa-Ard J, Warasiha B . Arylamine N-acetyltransferase-2 genotypes in the Thai population. Br J Clin Pharmacol 2003; 55: 278–281.
Bell DA, Taylor JA, Butler MA, Stephens EA, Wiest J, Brubaker LH et al. Genotype/phenotype discordance for human arylamine N-acetyltransferase (NAT2) reveals a new slow-acetylator allele common in African-Americans. Carcinogenesis 1993; 14: 1689–1692.
Woolhouse NM, Qureshi MM, Bastaki SM, Patel M, Abdulrazzaq Y, Bayoumi RA . Polymorphic N-acetyltransferase (NAT2) genotyping of Emiratis. Pharmacogenetics 1997; 7: 73–82.
Bakayev VV, Mohammadi F, Bahadori M, Sheikholslami M, Javeri A, Masjedi MR et al. Arylamine N-acetyltransferase 2 slow acetylator polymorphisms in unrelated Iranian individuals. Eur J Clin Pharmacol. 2004; 60: 467–471.
Taja-Chayeb L, Agundez JA, Miguez-Munoz C, Chavez-Blanco A, Duenas-Gonzalez A . Arylamine N-acetyltransferase 2 genotypes in a Mexican population. Genet Mol Res 2012; 11: 1082–1092.
Agundez JA, Olivera M, Martinez C, Ladero JM, Benitez J . Identification and prevalence study of 17 allelic variants of the human NAT2 gene in a white population. Pharmacogenetics 1996; 6: 423–428.
Malik MA, Upadhyay R, Modi DR, Zargar SA, Mittal B . Association of NAT2 gene polymorphisms with susceptibility to esophageal and gastric cancers in the Kashmir Valley. Arch Med Res 2009; 40: 416–423.
Talbot J, Magno LA, Santana CV, Sousa SM, Melo PR, Correa RX et al. Interethnic diversity of NAT2 polymorphisms in Brazilian admixed populations. BMC Genet 2010; 11: 87.
Lin HJ, Han CY, Lin BK, Hardy S . Slow acetylator mutations in the human polymorphic N-acetyltransferase gene in 786 Asians, blacks, Hispanics, and whites: application to metabolic epidemiology. Am J Hum Genet 1993; 52: 827–834.
Ruiz JD, Martinez C, Anderson K, Gross M, Lang NP, Garcia-Martin E et al. The differential effect of NAT2 variant alleles permits refinement in phenotype inference and identifies a very slow acetylation genotype. PLoS ONE 2012; 7: e44629.
Sabbagh A, Darlu P . Inferring haplotypes at the NAT2 locus: the computational approach. BMC Genet 2005; 6: 30.
Garcia-Martin E . Interethnic and intraethnic variability of NAT2 single nucleotide polymorphisms. Curr Drug Metab 2008; 9: 487–497.
Luca F, Bubba G, Basile M, Brdicka R, Michalodimitrakis E, Rickards O et al. Multiple advantageous amino acid variants in the NAT2 gene in human populations. PLoS ONE 2008; 3: e3136.
Sabbagh A, Langaney A, Darlu P, Gerard N, Krishnamoorthy R, Poloni ES . Worldwide distribution of NAT2 diversity: implications for NAT2 evolutionary history. BMC Genet 2008; 9: 21.
Patin E, Harmant C, Kidd KK, Kidd J, Froment A, Mehdi SQ et al. Sub-Saharan African coding sequence variation and haplotype diversity at the NAT2 gene. Hum Mutat 2006; 27: 720.
Magalon H, Patin E, Austerlitz F, Hegay T, Aldashev A, Quintana-Murci L et al. Population genetic diversity of the NAT2 gene supports a role of acetylation in human adaptation to farming in Central Asia. Eur J Hum Genet 2008; 16: 243–251.
Acknowledgements
LS is supported by UGC-BSR Research Fellowships in Science for Meritorious Students (RFSMS) Grant. GC is supported by the European Union European Regional Development Fund through the Centre of Excellence in Genomics to Estonian Biocentre and Estonian Institutional Grant IUT24-1. The genome-wide analysis was carried out in part in the High Performance Computing Center of University of Tartu.
Author information
Authors and Affiliations
Contributions
LS, RMP, AKM, BLVKS and RM conceived the study and contributed reagents and tools. LS performed the experiments. LS, GC and BLVKS analyzed the data and drafted the final manuscript; all of the authors read, revised and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflicts of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Lakkakula, S., Mohan Pathapati, R., Chaubey, G. et al. NAT2 genetic variations among South Indian populations. Hum Genome Var 1, 14014 (2014). https://doi.org/10.1038/hgv.2014.14
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/hgv.2014.14