Abstract
Range expansion has genetic consequences expected to result in differentiated wave-front populations with low genetic variation and potentially introgression from a local species. The northern expansion of Peromyscus leucopus in southern Quebec provides an opportunity to test these predictions using population genomic tools. Our results show evidence of recent and post-glacial expansion. Genome-wide variation in P. leucopus indicates two post-glacial lineages are separated by the St. Lawrence River, with a more recent divergence of populations isolated by the Richelieu River. In two of three transects we documented northern populations with low diversity in at least one genetic measure, although most relationships were not significant. Consistent with bottlenecks and allele surfing during northward expansion, we document a northern-most population with low nucleotide diversity, divergent allele frequencies and the most private alleles, and observed heterozygosity indicates outcrossing. Ancestry proportions revealed putative hybrids of P. leucopus and P. maniculatus. A formal test for gene flow confirmed secondary contact, showing that a reticulate population phylogeny between P. maniculatus and P. leucopus was a better fit to the data than a bifurcating model without gene flow. Thus, we provide the first genomic evidence of gene flow between this pair of species in natural populations. Understanding the evolutionary consequences of secondary contact is an important conservation concern as climate-induced range expansions are expected to result in new hybrid zones between closely related species.
Similar content being viewed by others
Introduction
Range expansion is a critical step in which evolutionary forces act to shape the genetic and phenotypic diversity in natural communities (Taylor and Keller, 2007). Expansion is generally thought of as a series of successive population founder events along an axis of colonization which leads to the most recently colonized area or ‘wave-front’ (Excoffier et al., 2009; Slatkin and Excoffier, 2012). Repeated bottlenecks during expansion are expected to result in genetically impoverished populations in the wave-front. This prediction is based on a model of expansion common to invasive species that lack extrinsic barriers to dispersal, where the rate of expansion is limited only by the traits intrinsic to the colonizing populations on the range margin (for example, reproduction rate, dispersal and so on). Many native species are also undergoing range expansion due to human-induced climate change (Hickling et al., 2006; Chen et al., 2011). However, in contrast to invasive species, populations undergoing climate-driven expansions may not suffer dramatic losses in genetic diversity (Pluess, 2011; Nullmeier and Hallatschek, 2013; Dai et al., 2014; Garnier and Lewis, 2016; Monzón et al., 2016). Genomic tools can provide the resolution needed to directly disentangle the evolutionary and demographic history of natural populations of non-model organisms (Shafer et al., 2015).
The process of expansion has both stochastic and deterministic elements that can be viewed as a spatial analog to genetic drift (Slatkin and Excoffier, 2012) and natural selection (Shine et al., 2011). Individuals colonizing a new area will likely carry mutations present at low frequency in the source population that can drift (‘surf’) to high frequency in subsequent generations in the newly colonized area. This spatial version of genetic drift, or ‘allele surfing’, can lead to genetic structuring and differentiation of wave-front populations (Hallatschek et al., 2007; Swaegers et al., 2015). Deleterious mutations can also surf due to low effective population size and accumulate as a genetic load of expansion (Peischl et al., 2013), a prediction empirically verified in humans (Henn et al., 2015; Peischl et al., 2016).
Another consequence of range expansions is hybridization. Introgression has long been recognized as an important potential source of genetic variation for adapting to a new environment (Lewontin and Birch, 1966; Pfennig et al., 2016). Although theory indicates that introgression is likely to occur during expasion (Currat et al., 2008; Excoffier et al., 2009), genomic studies (for example, White et al., 2013; Swaegers et al., 2015; Monzón et al., 2016) have tended to overlook the possibility of gene flow between closely related species (but see Mastrantonio et al., 2016). Species’ boundaries can be continuous and gene pools porous (Harrison, 1990; Wu, 2001); however, few empirical studies on the genomics of range expansion have looked for evidence of hybridization when not suspected a priori. As genome-wide markers become increasingly available, long standing questions regarding the frequency and consequences of hybridization and range expansion (for example, Seehausen, 2004; Lowe et al., 2015; Abbot et al., 2016; Canestrelli et al., 2016; Gompert and Buerkle, 2016) in natural population are increasingly tractable.
Hybridization between divergent lineages is more likely following habitat disturbances, for example during range expansion where low population density results in relatively few con-specific mates and less competition against hybrids (Seehausen, 2004). Mammalian examples include canids during colonization of North America (Reich et al., 1999; von Holdt et al., 2016), polar bears with ongoing climate change (Kelly et al., 2010; Cahill et al., 2015) and humans, that are thought to have hybridized with at least three hominin species while expanding out of Africa (Huerta-Sánchez et al., 2014; Fu et al., 2015; Mondal et al., 2016).
In many cases, interspecific gene flow can lower the mean fitness of a population through the introduction of mal-adapted alleles and deleterious epistasis (Lynch, 1991). Sometimes, however, outcrossing with a divergent lineage can lead to local adaptation through the introgression of pre-adapted alleles (Hedrick, 2013). One example is the Western Mediterranean mouse (Mus spretus), a rodent with populations that adapted to rodenticide through interspecific introgression of resistance genes from the house mouse, M. musculus (Song et al., 2011). Introgression of Denisovan DNA is thought to have resulted in adaptation to Arctic conditions in the Inuit from Greenland (Racimo et al., 2016), and in high altitude adaptation in Tibetans (Huerta-Sánchez et al., 2014). Paralleling the history of Tibetan people, genomic evidence indicates modern Tibetan Mastifs adapted to high altitude after hybridizing with gray wolves native to the Tibetan Plateau (Miao et al., 2016). Thus, interspecific gene flow on the wave-front may provide the pre-adapted genetic substrate needed to uncouple gene pools from a local adaptive maximum and obtain a higher global peak, further facilitating establishment and expansion.
P. leucopus populations in southern Quebec provide an opportunity to investigate the genomic consequences of a range expansion in a small mammal. P. leucopus is native to eastern North America (King, 1968), and one of many species undergoing range expansion which is likely due to anthropogenic climate change (Myers et al., 2009; Chen et al., 2011; Roy-Dufresne et al., 2013). Dispersal of individual P. leucopus varies widely, although average movement of individual mice has been estimated at about 330 m in Indiana (Khrone et al., 1984) and 930 m in Quebec (Gaitan and Millien, 2016). Over the past few decades, the northern distribution of this species has expanded at an estimated 10 km per year northward in southern Quebec (Roy-Dufresne et al., 2013). P. leucopus is sympatric with closely related P. maniculatus across most of its distribution and both species are key components of small mammal communities in North America. P. leucopus is typically found in deciduous forests (Vessey and Vessey, 2007), while P. maniculatus is more associated with conifer forests and, being adapted to colder winter temperatures (Pierce and Vogt, 1993), reaches higher latitudes where P. leucopus is absent. However, as it expands, P. leucopus is ecologically replacing P. maniculatus (Wan, 2014), becoming the dominant Peromyscus in some regions of southern Quebec (for example, Grant, 1976; Roy-Dufresne et al., 2013).
P. leucopus and P. maniculatus belong to sister clades which the fossil record suggest diverged ~500 000 years ago (Hibbard, 1968; but see Fiset et al., 2015). Both species have a more closely related sister species with which they can hybridize and produce fertile offspring, P. polionotus and P. gossypinus, respectively (Dice, 1937; Maddock and Dawson, 1974). After unsuccessful attempts to experimentally hybridize P. leucopus and P. maniculatus (Dice, 1933, Dice (1968) concluded that complete reproductive isolation had occurred. Decades later, through hormonally induced ovulation and artificial insemination, Dawson et al. (1972) and Maddock and Dawson (1974) were able to produce live hybrids, although they were deemed inviable. Following this work, it was assumed these two species were reproductively isolated across their entire range. However, Leo and Millien (2016) recently used microsatellite markers and found putative hybrids of both species in natural populations from southern Quebec.
We use individual-based, genome-wide data (Andrews et al., 2016) to investigate the demographic history of P. leucopus in southern Quebec during its northward expansion. We looked for the signal of range expansion and investigated the evidence of genetic bottlenecks and allele surfing, namely decreased genetic diversity, divergent allele frequencies and potentially introgression, among northern populations. We use this species-pair as a case study to examine whether sympatric species, reproductively isolated likely under reinforcement (Doty, 1972, 1973), are more likely to hybridize during range expansion.
Materials and methods
Study system and DNA extraction
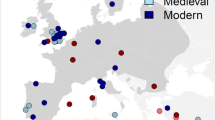
We used P. leucopus samples from 13 forested sites (n=229) and P. maniculatus from one forested site (‘Pm’, n=11) in southern Quebec, collected during the summer of 2013 and 2014. Sites were grouped into three transects, the northern (N), south-eastern (SE) and south-western transects (SW), separated by the St. Lawrence and Richelieu Rivers (Figure 1). Studies of P. leucopus using microsatellites and mitochondrial markers have indicated that the St. Lawrence and Richelieu Rivers inhibit gene flow, with differentiated populations on either side (Ledevin and Millien, 2013; Rogic et al., 2013; Fiset et al., 2015), while agricultural fields are relatively ineffective barriers (Marrotte et al., 2014). We thus suggest that our three transects represent distinct expansion routes that may provide independent genetic patterns of range expansion, taking the northern sites in each transect (for example, N4, SW5/SW4, SE4) as the approximate range margin. A search of capture records on Vertnet reveals that a putative P. leucopus individual has been caught further north of our northern-most sample site. However, this individual was a juvenile and visually identified, making it especially difficult to differentiate from P. maniculatus. In fact, according to the measurements taken by the collector (for example, ear and tail), this individual more closely resembles P. maniculatus (Lindquist et al., 2003). We extracted DNA from liver and muscle tissue using a 3-day phenol-chloroform protocol (Sambrook et al., 1989). Species were identified via PCR amplification of a mitochondrial COIII sequence, as described in Rogic et al. (2013).
RAD-seq library
Extracted DNA was sent to the Institut de biologie intégrative et des systèmes (IBIS) at Université Laval for individual-based library preparation using a modified version of the original genotyping-by-sequencing protocol (Elshire et al., 2011). DNA from each individual was digested with the rare cutting (1 cut /4096 bp) restriction enzyme PstI (CTGCAG) to seed markers, followed by MspI (CCGG), a frequent cutter (1 cut /256 bp) used to define read size. Next, a barcoded adapter with a complimentary end to the PstI site and a common adapter with a complimentary end to the MspI site were ligated to the DNA fragments. Following ligation, sets of 48 samples were pooled and PCR was applied to the barcoded DNA fragments using primers that hybridize to the adapters. Each library composed of 48 samples had 100 bp paired-end reads sequenced on a single Illumina HiSeq 2000 lane at the Genome Quebec Innovation Center.
SNP genotyping and population genetic summary statistics
We used the Stacks pipeline (Catchen et al., 2013a) to process raw data and identify single-nucleotide polymorphisms (SNPs). Raw reads were demultiplexed and quality filtered using the process_radtags script of Stacks 1.37. We followed the recommendation of recent studies (Nam et al., 2016; Shafer et al., 2017) and aligned to a reference genome for SNP calling. We aligned all processed reads to the scaffold-level assemblage of the P. maniculatus bairdii reference genome (NCBI assembly accession: GCF_000500345.1) with Bowtie2 V 2.2.8 (Langmead and Salzberg, 2012) using local alignment. We removed one P. maniculatus and six P. leucopus individuals that had relatively few reads sequenced. This left 223 P. leucopus and 10 P. maniculatus samples (Table 1). Scripts were executed using the ref_map.pl wrapper in Stacks (v. 1.44) to generate a catalog of RAD loci with a minimum depth of coverage of 4 reads (-m 4). The populations script was then used to obtain two SNP data sets for population genomic analysis, one consisting only of the 13 P. leucopus populations (n=223, ‘P. leucopus-only’ data set hereafter), and a second data set that included the 10 P. maniculatus individuals (n=233, ‘complete’ data set hereafter). Genome-wide markers were filtered so that each SNP had a minimum allele frequency (MAF) of at least 2.5% (—min_maf 0.025) and a heterozygosity of 0.50 or less to avoid paralogs (—max_obs_het 0.50). In addition, each locus had to be found in every population, that is 13 populations for the P. leucopus-only data set (-p 13) and 14 populations in the complete data set (-p 14), as well as in at least 60% of individuals in a given population (-r 0.60).
The following population genetic summary statistics were then calculated for each data set: nucleotide diversity (Pi), homozygosity (Homobs, Homexp) and heterozygosity (Hetobs, Hetexp), inbreeding coefficient (FIS), private alleles and differentiation (FST). Nucleotide diversity is related to expected heterozygosity and is an overall measure of genetic variation (Catchen et al., 2013b). FIS measures the degree of inbreeding due to non-random mating, or the change in observed homozygosity relative to the expected value. FST represents the amount of inbreeding due to random mating in a finite population and is used as a measure of population subdivision and genetic drift (Crow and Kimura, 1970; Summarized in Ewen et al., 2012). Thus, FIS measures changes in genotype frequencies while FST measures changes in allele frequencies. We applied an FST correction, using Fisher’s exact test to assess whether allele frequencies at a given locus are statistically different from zero. Loci with FST estimates that fail to reach statistical significance according to the P-value (α=0.05) are set to zero (—fst_correction P-value). To estimate ancestry proportions and construct a population tree, the complete data set was thinned to the first SNP of each RAD locus to reduce linkage disequilibrium (—write_single_snp). The ordination methods used (discriminant analysis of principal components (DAPC) and principal component analysis (PCA)) do not make assumptions of independence and thus all SNPs identified in each RAD locus were used in those analyses.
To examine patterns of genetic diversity, we followed White et al. (2013) and regressed measures of heterozygosity and nucleotide diversity onto ‘relative expansion distance’. For sites on the northern transect which are aligned in a north-eastern direction (Figure 1), we use distance to the southern-most site. We hypothesize that northward range expansion in this transect is likely occurring in a north-eastern direction and used the Euclidian distances (km) of sites N2, N3 and N4 to site N1 (Table 1). South of the St. Lawrence River, the Richelieu River is aligned in a north-south direction with no major barrier to direct northward expansion. For the south-western and south-eastern transects, we therefore used the distance to the southern-most latitude, or the Euclidian distance to the latitude of the southern-most site (that is, SW1 and SE1). We regressed genetic diversity measures for each transect separately. To evaluate evidence of increased drift among northern populations, we looked for a northward gradient of increasing FST. We used the two-dimensional isolation by distance model of Rousset (1997) and regressed pair-wise FST estimates (FST/1−FST) of 10 populations and their southern-most site as a function of the natural log of relative expansion distance. We excluded southern-most sites (FST=0) and pooled our 10 populations.
Genotype imputation
Next generation sequencing, especially the RAD-seq protocol, produces a patchy genotype matrix with a considerable amount of missing information. Imputation of missing data has been shown to aid allele frequency estimates and improve the power of genomic studies (Li et al., 2009). For this reason, we used the software LinkImpute (Money et al., 2015) to estimate missing genotypes in PLINK files (Purcell et al., 2007) for all data analyses. LinkImpute uses a k-nearest neighbor genotype imputation method and was designed for RAD-seq data from non-model organisms.
Genomic structure and admixture
To summarize overall genetic variation among individuals, we performed a PCA on both data sets (with and without P. maniculatus samples) using the package LEA (Frichot and François, 2015) in R. To look for major lineages within P. leucopus, we further analyzed P. leucopus population structure by transforming genomic variation to PCs and applying a discriminant analysis (Jombart et al., 2010) in the R package adegenet (Jombart, 2008). DAPC describes genomic clusters using synthetic variables and focuses on the genetic variation observed between groups, while minimizing within-group variance. Genomic clusters (K) were identified using a k-means algorithm and the number of clusters evaluated with the Bayesian information criterion (BIC; Supplementary Figure 1a). DAPC was performed on the two K values with the lowest BIC (K=2 and K=3). We used alpha-score optimization to examine the trade-offs between over-fitting and power to discriminate. This procedure identifies the PCs with the highest alpha scores using spline interpolation. The PC with the highest (‘optimal’) alpha-score provides an approximation and with other integers of similar alpha scores a range from which it is adequate to choose the number of PCs to retain (Jombart and Collins, 2015). We performed this procedure on 110, 10 and 6 PCs (Supplementary Figure 2), which showed that the first six PCs have very similar high alpha scores. We thus kept the first six PCs (12.1% of variance) for DAPC presented here. Individual ancestry proportions of P. leucopus and P. maniculatus were estimated using a model-based method implemented in ADMIXTURE using default settings (Alexander et al., 2009; Alexander and Lange, 2011). ADMIXTURE uses cross-validation to evaluate models with K ancestral source populations and maximum-likelihood (ML) algorithms to estimate ancestry proportions. We define a hybrid as any individual with visible hetero-specific ancestry.
Population splits and test of bifurcating model
We used the software TreeMix 1.13 (Pickrell and Pritchard, 2012) to build a ML tree and analyze the demographic histories of our P. leucopus populations. To construct a bifurcating tree, this method uses a Gaussian approximation of genetic drift and the covariance in allele frequencies between population pairs. We used PLINK 1.9 (Purcell and Chang, 2015) to stratify allele frequencies and then the plink2treemix.py script (Pickrell and Pritchard, 2012) to convert the stratified file (.frq) to TreeMix format. The ML tree was rooted using P. maniculatus. The amount of genetic drift estimated to have occurred in a lineage is proportional to the horizontal branch length on a given branch.
Unlike the clustering methods described above, TreeMix explicitly tests for the presence of gene flow by identifying population pairs that poorly fit a bifurcating evolutionary history, and then models gene flow events between these populations to increase the fit. Migration events are added in order of their statistical significance, such that the first gene flow event added to the tree is the one that most increases the likelihood of the model. The direction of gene flow is estimated based on asymmetries in the covariance matrix of allele frequencies, as shown on Figure 1 of Pickrell and Pritchard (2012). Here, modeled the first two ML trees, a tree based on a strict bifurcating model and a tree with one gene flow event.
Results
Data sets
Sequencing our 240 samples generated 1 117 579 057 raw paired-end reads. After removing seven individuals with few reads sequenced and filtering for population genomic analyses, we obtained 38 144 bi-allelic SNPs for the P. leucopus-only data set and 33 919 SNPs for the complete data set. Thinning the complete data set to the first SNP per RAD locus yielded 12 507 SNPs for analysis of ancestry proportions and construction of a ML tree.
Population genetic statistics
The mean observed heterozygosity (Hetobs) in P. leucopus is 0.145 and mean inbreeding coefficient (FIS) is 0.074 (s.d.=0.056, Table 1). Populations along the northern transect show a significant linear decrease in nucleotide diversity with relative expansion distance (P⩽0.002; adjusted R2: 0.993; n=4, Supplementary Figure 3b). In the south-western transect, populations from sites SW4 and SW5 show higher levels of diversity than sites further south. The most northern site in the south-eastern transect (SE4) also shows decreased diversity, although the relationship is not significant (Supplementary Figure 3 e and f). Interestingly, the northern-most population, N4, shows exceptionally high levels of observed heterozygosity (Hetobs=0.169) and low levels of inbreeding (FIS=0.026) relative to other sites in the northern transect. As a comparison, the population on Montreal island (N2) has an almost 8-fold higher FIS (FIS=0.204). This excess heterozygosity seen in the northern-most site also stands in contrast to the most proximal population to the south, N3, which shows a 6-fold higher inbreeding coefficient (FIS=0.157). Private alleles ranged from 0 (in 8 of 13 sites) to 14 (in the northern-most site N4).
Concordant with post-glacial expansion, we found a significant reduction in observed heterozygosity relative to the expected levels under Hardy–Weinberg equilibrium in two of three P. leucopus transects: the northern and south-western transects (one-tailed paired t-test; northern transect: P⩽0.031; south-western transect: P⩽0.017; south-eastern transect: P⩽0.104). The reduction in observed heterozygosity was also significant when all populations were pooled (P⩽0.003).
To further investigate the history of recent and post-glacial range expansion, we analyzed patterns of allele frequency divergence among populations. Overall, there was relatively low levels of differentiation among P. leucopus populations, with a mean FST of 0.037 (95% confidence interval [CI]: 0.033–0.041). However, 78 pair-wise FST estimates (Supplementary Table S1) between and within transects show relatively high divergence of P. leucopus south and north of the St. Lawrence River (Figure 2a). Population divergence within transects is also higher along the northern transect, consistent with populations having experienced greater amounts of drift. Populations south of the St. Lawrence River are less differentiated from each other than from populations north of this river, a pattern which supports our expectation of distinct post-glacial expansion routes separated by the St. Lawrence River.
(a) Summary of 78 pair-wise FST values of populations within and between transects. The plot summarizes data presented on Supplementary Table S1. (b) Isolation by distance model of P. leucopus (Rousset, 1997), plotting differentiation (FST/ (1−FST)) between 10 populations and their southern-most site against the natural log of the distance. The regression is y=0.0089x–0.0149.
Increased genetic drift due to allele surfing during range expansion is expected to result in wave-front populations with high FST. Consistent with this expectation, the northern-most site N4 shows exceptionally high FST (N4: FST=0.059, CI: 0.050–0.068) relative to the average. We detected a significant linear relationship between differentiation relative to southern-most sites and distance to this site (Figure 2b; P⩽0.006; Adjusted R2= 0.58). The slope of the regression is 0.0089, thus a rough approximation of the number of south to north effective migrants is 112 individuals. The linear increase in FST remained significant when site N4 is removed (P⩽0.02).
A substantially greater divergence is expected across species than between populations within species. Accordingly, FST estimates between P. leucopus and P. maniculatus are approximately an order of magnitude greater (mean FST=0.384; SD=0.018) than the mean FST observed within P. leucopus in southern Quebec.
Population genomic structure in P. leucopus
Overall genetic variation in P. leucopus summarized with PCA shows population structure is associated with geography. Lineages on either side of the St. Lawrence River are separated by PC1 (8.1% of variance) while individual populations tend to differentiate across PC2 (2.6% of variance; Figure 3). A gradient in population structure is observed within some populations, including SE4, SW4, N3 and especially N4. These populations cluster more widely across PC space relative to more southern populations, such as SE1, N1, SE2 and N2, where individuals tend to cluster more closely together. The northern site SW5 however did not display this gradient pattern while it is apparent in more southern sites, such as SW1 and SW2.
Analyzing between-group differences with DAPC supports two main lineages (K=2) north and south of the St. Lawrence River (Supplementary Figure 4a and b), with more subtle variation separating populations east and west of the Richelieu River (Figure 4). These results provide additional evidence that the Richelieu River and the preceding waterways split a lineage expanding northward from an eastern glacial refugium. Results also show evidence of migration across major barriers. Both PCA (Figure 3) and DAPC (Figure 4) show that one individual from the south-western transect (SW1) with genetic variation more typical of the genomic background found in the northern transect. Similarly, a few individuals from sites SE3 and SE4 have genotypes more similar to those found in the south-western transect.
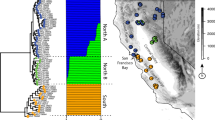
Population genomic structure inferred by DAPC on 6 retained PC eigenvalues (inset plot: 12.1% of variance) shows three (K=3) P. leucopus lineages separated by the St. Lawrence and Richelieu rivers. The outset plot shows each individual represented by a row, transects delimited by arrows, and color (red) denoting which of three clusters each individual is assigned to. Differences between individuals east and west of Richelieu River (clusters 2 and 3) are subtle compared to the differences to individuals on the northern transect (as shown by Supplementary Figure 4). A full color version of this figure is available at the Heredity journal online.
We used ADMIXTURE to analyze the ancestry of P. leucopus from southern Quebec and look for evidence of hybridization with P. maniculatus. After cross validating 12 potential ancestral populations, our results indicate up to nine relatively well-supported source populations contributing to ancestry (Supplementary Figure 1b). Ancestry proportions under the well-supported model of K=4 shows P. leucopus population genomic structure associated with extrinsic barriers to dispersal (rivers), as well as admixture in SW5 and SW4 from different transects (Figure 5), which is consistent with the relatively high nucleotide diversity and heterozygosity observed in these two sites. Supporting ordination, the most ancestral P. leucopus clusters (K=3) consist of individuals on either side of the St. Lawrence River (Supplementary Figure 5), while the Richelieu River separates more subtle sub-structure, consistent with a more recent divergence. We find additional sub-structure in the northern-most site, N4 and admixture in N3 (K=6).
Admixture proportions for two, four and nine putative ancestral populations for P. leucopus and P. maniculatus. Each column represents an individual and the length of colored segments denote the proportion of an individual’s genome inherited from one of K ancestral populations. K=4 was strongly supported by BIC (Supplementary Figure 1). A full color version of this figure is available at the Heredity journal online.
An admixture pattern consistent with range expansion (Falush et al., 2016) is observed in the two northern sites (for example, K=9, Figure 5). The second northern-most population, N3, is admixed with ancestry common to the south (for example, N1) and to the north (for example, N4). Falush et al., (2016) showed through simulations that this type of pattern would be consistent with gene flow, in this case from N1 and N4 to N3. However, such patterning of population genomic structure is also consistent with a bottleneck during expansion of the N4 lineage subsequent to the split from N3 (Excoffier and Ray, 2008; Falush et al., 2016), or perhaps introgression in N4 from an unsampled (‘ghost’) population (Currat et al., 2008; Falush et al., 2016). Indeed the relatively high observed levels of heterozygosity and low inbreeding (FIS) in N4 suggest the latter. Further sampling of P. leucopus and P. maniculatus north of the St. Lawrence River is warranted.
Ancestry proportions also show evidence of ancestral gene flow across the St. Lawrence River (for example, K=3, Supplementary Figure 5). Consistent with older admixture, the shared ancestry proportions are well homogenized within the populations. More recent admixture can be inferred when admixture proportions vary within a population because recombination has not had time to distribute genomic blocks across chromosomes. We find hierarchical structure in SW5 and SE1 (K=6), likely due to isolation. Site SW5 is situated in a park surrounded by the city of Longueuil, which has experienced a 58-fold increase in census human population during the most recently documented 143-year period (~286 mice generations), increasing from 3977 in 1871 to 231 409 in 2014 (Statistique Canada, 2014). Site SE1 is surrounded, other than to the north-east, by Lake Champlain and the Richelieu River. Finer population genetic structure is apparent in SW1 (K=9).
Admixture with P. maniculatus
Notably, our ancestry results show evidence of hybridization with P. maniculatus. A predicted consequence of expanding populations with low effective population size is potential introgression from closely related local species. ADMIXTURE results at K=2 show the presence of one putative P. leucopus hybrid in site SE3 with 15.5% of its ancestry inferred as coming from P. maniculatus (Figure 5). Two additional individuals have 1.6 and 1% inferred P. maniculatus ancestry, a percentage at least three orders of magnitude greater than in non-admixed individuals (0.001%). Given that most individuals in this site showed little to no interspecific ancestry, we suggest this result may be explained by a recent hybridization. Specifically, these ancestry proportions are consistent with ancestry expected in F3 (12.5%) and F6 (1.6%) hybrids backcrossed with P. leucopus. Our results also show that two out of the 10 P. maniculatus sampled share 18.8% and 7.5% ancestry proportions with P. leucopus. Further support for hybridization comes from PCA (McVean, 2009) on the complete data set, which shows the two species separated by PC1 (22% of explained variance) and the putative hybrids with more than 10% hetero-specific ancestry as intermediate genotypes compared to other individuals (Supplementary Figure 6).
Bifurcating P. leucopus population tree
In line with our analyses above, the ML tree supports an older divergence of P. leucopus on either side of the St. Lawrence River and a more recent split caused by the Richelieu River (Figure 6a). Interestingly, however, the ML tree places populations N2 and SW5 as basal lineages. The ML tree indicates that the inbred population on Montreal island (N2) is from an earlier split sharing common ancestor with a lineage that later diverged to expand north (N3 and N4), while another remained at a more southern latitude (N1). Consistent with being on the wave-front, the northern-most site shows substantial accumulated drift since splitting from N3, as indicated by the longer branch length. The ML tree indicates the population partially isolated by the city of Longueuil (SW5) is a basal P. leucopus lineage, one likely historically present in that area. Surprisingly, however, the ML tree indicates SW5 diverged from other P. leucopus in Quebec prior to the split of populations on either side of the Richelieu River. This would suggest that other south-western populations (for example, SW3) are more closely related to mice in the south-eastern transect than they are to mice in SW5, a hypothesis not supported by DAPC (Figure 4) and the ancestry proportions (K=4, Figure 5) that reveal shared genomic structure among individuals from south-western populations. Indeed, allele frequencies in SW5 are significantly less diverged from other south-western populations (FST=0.017) than they are relative to south-eastern populations (FST=0.026; one-tailed paired t-test: P⩽0.016). A possible explanation for the placement of SW5 as a unique lineage south of the St. Lawrence River is the complex evolutionary history of mice in this site, which show ancestral admixture from the northern and south-eastern transects (for example, K=4; Figure 5), as well as built-up sub-structure due to more recent isolation.
Population relationships inferred with TreeMix (a) under a bifurcating model without gene flow. Horizontal branch length represents amount of evolutionary change according to the drift parameter (related to FST). (b) The residual fit of model (a), a scenario without gene flow after divergence. The scale bar represents 10 × the average standard error of the entries in the sample covariance matrix (Pickrell and Pritchard, 2012). Residuals are qualified using the color palette. Positive residuals (green/blue/black) represent population pairs that are more closely related than presented on the tree, and thus candidates for admixture. Residuals below zero (yellow/orange/red) represent population pairs that are less closely related to each other than shown on the tree. (c) Population tree showing gene flow between P. maniculatus and P. leucopus from SE3. Direction of gene flow is shown by the arrow. The color of the arrow denotes the weight of migration (proportional to the amount of genetic ancestry contributed by the immigrant). A full color version of this figure is available at the Heredity journal online.
Test for introgression
Methods that calculate ancestry proportions, like ADMIXTURE, can give misleading inferences of population structure when sampling is incomplete or uneven (Falush et al., 2016; Wang, 2016). We therefore used TreeMix to formally test the bifurcating model of genealogical evolution. A residual fit of the bifurcating tree without gene flow (Figure 6b) shows that this model does not fit the evolutionary history of the Peromyscus populations in our study area, and instead supports a reticulate event between P. leucopus from SE3 and out-group P. maniculatus (Figure 6c). The direction of gene flow from demographically stationary P. maniculatus to colonizing P. leucopus (Fiset et al., 2015) is also consistent with predictions of introgression during range expansion (Currat et al., 2008). Allowing for two more migration events supports past gene flow from N1 to SW1 (Supplementary Figure 7a and b), and from SE1/SE2 to P. maniculatus (Supplementary Figure 7c and d). However, adding three migration events may be over-fitting as the topology of the ML tree is altered under this model.
Discussion
We used genomic tools to investigate the history of range expansion of P. leucopus in southern Quebec. We predicted allele surfing and genetic bottlenecks during recent northward expansion would lead to northern populations with reduced genetic diversity, divergent allele frequencies and potentially admixture. Consistent with these processes, we described a northern-most population with reduced nucleotide diversity, divergent allele frequencies, a high number of private alleles and heterozygosity levels that suggest admixture with an unsampled population. We detected two main lineages of P. leucopus on either side of the St. Lawrence River and additional sub-structure in populations east and west of the Richelieu River. We argue that this structure reflects the evolutionary history of post-glacial expansion of two lineages. We also documented gene flow between P. maniculatus and P. leucopus, undermining the notion of complete reproductive isolation between these two species.
Spatial genetic gradients in P. leucopus
We found clines of genetic diversity consistent with the northward range expansion of P. leucopus. Two of three transects showed northern populations with the least genetic diversity in at least one measure. The northern transect showed a significant linear decrease, although the overall difference in diversity was relatively small considering the distance between the southern-most and northern-most populations (~150 km). Contrasting our expectations, a positive trend was seen in the south-western transect due to higher levels of genetic diversity in admixed northern locations. Our isolation by distance model estimated gene flow (112 effective migrants) along a south-north axis. However, this estimate is based on a number of likely unrealistic assumptions (Whitlock and Mccauley, 1999) and likely differs among transects, and thus should be viewed only as a rough approximation and interpreted with caution.
Various factors likely limited our power to detect latitudinal gradients, such as inter-site differences. For example, larger forest patches contain more individuals that are likely to harbor greater genetic diversity. The presence of partial barriers to dispersal between sites (for example, roadways) in the south-western transect (Rogic et al., 2013; Marrotte et al., 2014) may further blur any relationship between genetic diversity and expansion distance at the spatial scale and sampling design used here. Wave-front dynamics may also play a role, as simulations have shown that climate-driven range expansions maintain genetic diversity on the expanding range margin (Nullmeier and Hallatschek, 2013; Dai et al., 2014), although diversity may still decrease under models of rapid climate change (Garnier and Lewis, 2016).
Further limiting our ability to detect consistent spatial gradients in genetic diversity, our results indicate that some populations are partially isolated and are therefore unlikely to have undergone recent expansion. For example, the relatively high levels of inbreeding in mice on Montreal island (N2) suggest this population has been isolated for some time. It is also likely that the response of P. leucopus to climate change, in particular in the south-western transect, is an increase in abundance rather than northern expansion. For example, although P. maniculatus was the more commonly found Peromyscus species as recently as 40 years ago, a historical presence of P. leucopus at low abundance (Grant, 1976) may have contributed genetic variation during more recent migration from the south.
P. leucopus is one of many North American mammals (Lessa et al., 2003) that expanded north during interglacial events of the Pleistocene. Mitochondrial DNA data indicates that at the start of the current interglacial period, two P. leucopus lineages expanded north from eastern and western refugia ~17 000 and 15 000 bp, respectively (Rowe et al., 2006; Fiset et al., 2015). The genetic legacy of the older and more geographically extensive post-glacial expansion may be exposed as a reduction of observed heterozygosity relative to Hardy–Weinberg expectation in P. leucopus populations sampled in southern Quebec, a region covered by the Laurentide Ice Sheet 20 000 years ago. However, it is worth noting that reduced heterozygosity in a population at Hardy–Weinberg equilibrium can also be detected due to allele frequency differences among sub-populations, known as the ‘Wahlund effect’. P. leucopus in our study area consist of three meta-population lineages separated by geographic barriers. Because our comparison of observed and expected heterozygosity was based on estimates from individual sub-populations rather than meta-populations, the role of the Wahlund effect is likely relatively minor.
Population genetic structure in P. leucopus
Geographic isolation is generally thought to be a common precursor to generating genetic diversity and ultimately new species (Sobel et al., 2010). We found that genomic variation in P. leucopus is spatially structured and associated with geographical barriers. We identified two putative post-glacial lineages of P. leucopus north and south of St. Lawrence River. These lineages likely began diverging during the Late Pleistocene in western and eastern glacial refugia, respectively (Rowe et al., 2006; Fiset et al., 2015). More subtle genetic correlations can differentiate P. leucopus on either side of the south-north oriented Richelieu River. This supports a scenario in which ancestral populations from the eastern glacial lineage were separated during northward post-glacial expansion. Populations expanding from an eastern refugium would have been separated by what was then the ancient Champlain Sea left by the retreating ice sheet, that eventually became Lake George and Lake Champlain in the states of New York and Vermont, respectively, and the Richelieu River in Quebec (Rogic et al., 2013; Fiset et al., 2015). P. leucopus populations from the western refugium, on the other hand, colonized the Great Lakes area during post-glacial expansion (Rowe et al., 2006). Ongoing climate change is thought to be responsible for the rapid expansion of P. leucopus across the Upper Michigan Peninsula (Myers et al., 2009), where it is ecologically replacing P. maniculatus (Wan, 2014), a process which can be facilitated by hybridization (Rhymer and Simberloff, 1996). P. leucopus north of the St. Lawrence River belong to the western lineage and an extension of the expansion detected across the Upper Michigan Peninsula. The higher levels of drift and inbreeding observed north of the St. Lawrence River than south of it indeed suggest the north shore may represent a more recently established colonization route.
Populations on an expanding range margin are expected to become differentiated due to the spatial accumulation of genetic drift and selection experienced during expansion. During expansion into a new territory, low frequency variants may become established in newly colonized areas where they, in subsequent generations, can increase to high frequencies and lead to ‘genetic revolutions’ of wave-front population structure (Excoffier and Ray, 2008). Consistent with processes associated with expansion, including genetic bottlenecks, allele surfing and potentially introgression, we showed that the northern-most population had relatively low nucleotide diversity, the most differentiated allele frequencies, the highest number of private alleles and heterozygosity estimates that suggest outcrossing with an unsampled population. In addition, ancestry proportions showed a pattern of population structure consistent with a northward bottleneck (Falush et al., 2016), although similar population genomic structure may arise from admixture with an unsampled population (Falush et al., 2016) during expansion, which population genetic statistics suggest may have occurred. Additional sampling north of the St. Lawrence River will help unravel the evolutionary history this P. leucopus lineage.
We found evidence that some populations are at least partially isolated. For example, P. leucopus on Montreal island have higher levels of inbreeding relative to populations on the mainland. We also found population structure in a P. leucopus population in an area (city of Longueuil) which has had a 58-fold increase in census human population during the most recently documented 143-year period, or roughly equivalent to 286 generations of P. leucopus (Statistique Canada, 2014). Urbanization can have major effect on population genetic structure through isolation and likely imposes novel selective pressures. Munshi-South et al. (2016) used human population size and percent impervious surface cover in New York City to study the effects urbanization had on genomic variation of P. leucopus. They found that these proxies best explained the genome-wide variation in urban P. leucopus and Harris et al. (2013) identified candidate genes potentially under selection in an urban environment, many of which were associated with the immune system. The isolated P. leucopus populations identified here thus provide an opportunity to study different evolutionary processes such as adaptation in response to a recent human expansion, and the effects of inbreeding on fitness in an island population.
Population splits
Our results of the demographic history of P. leucopus in southern Quebec support a scenario of post-glacial and ongoing expansion. Our analysis of population splits showed putative glacial lineages as clades separated by the St. Lawrence River, and a post-glacial divergence of populations on either side of the Richelieu River. Our results also suggest that P. leucopus was historically present and that one lineage became isolated on Montreal island, splitting off from a lineage that remained on the mainland (N1) and one that expanded north (N3 and N4). Indeed our results showed that the most northern site has accumulated substantial genetic drift. We found evidence that suggest the population surrounded by recent urbanization (SW5) has been historically present and has had a complex evolutionary of ancient admixture and isolation. The placement of this population as basal relative to all others south of the St. Lawrence River may due to the complex evolutionary history of ancient admixture and recent isolation that has occurred in this lineage.
Secondary contact between P. leucopus and P. maniculatus
One of the most neglected consequences of range expansion is the potential for local introgression. We hypothesized that northern expansion of P. leucopus populations into territory historically occupied by P. maniculatus would lead to introgression. We used genome-wide data to test this prediction, and our results showed the presence of a few putative recombinant hybrids. However, these putative hybrids were identified using clustering methods that are sensitive to sampling design and can produce results that are easily misinterpreted (Novembre and Stephens, 2008; Schwartz and McKelvey, 2009; Falush et al., 2016; Puechmaille, 2016; Wang, 2016). Such clustering methods also do not directly test for admixture between divergent gene pools. To overcome this, we used TreeMix to explicitly test for gene flow and reject the bifurcating model of genealogical evolution, confirming secondary contact between P. leucopus and P. maniculatus.
Only five individuals showed hybrid ancestry, which is likely due to ecological differences between the two species. P. leucopus was sampled mostly from small woodlots scattered across an agricultural matrix where, as a generalist, P. leucopus thrives. In contrast, there is evidence of more introgression occurring in larger and more continuous forests where P. maniculatus and P. leucopus can be locally found (Leo and Millien, 2016). In particular, Leo and Millien (2016) sampled both species where P. maniculatus was more abundant and found putative P. leucopus hybrids with ancestry proportions composed mainly of P. maniculatus ancestry, a result consistent with predictions (Currat et al., 2008). However, the results of Leo and Millien (2016) come with the aforementioned caveat of clustering methods as well as less resolution provided by microsatellites. Taking a genomic approach and sampling broadly in sites where both species are found, including north of the St. Lawrence River, and testing for gene flow will help expose the frequency and extent of hybridization between these two species of Peromyscus in Quebec.
Although P. maniculatus was represented by only 10 individuals in our study, we believe our inference of interspecific admixture is robust. For example, a mammal population genomic study showed that sufficient genetic variation could be sampled from a population with as few as 10 individuals to construct a complete evolutionary history (Trask et al., 2011). The putative hybrids found by Leo and Millien (2016) were also sampled from a more evenly distributed data set of 69 P. leucopus and 84 P. maniculatus. Furthermore, the accuracy of hybrid identification increases with the number of markers and level of divergence (Vaha and Primmer, 2006), reaching 90% accuracy with fewer than 50 microsatellite markers in populations one third as diverged as the species in this study. Importantly, Vaha and Primmer (2006) also showed that unsampled source populations have negligible effects on hybrid identification, and McVean (2009) demonstrated that the positions of genotypes on PC space relative to non-admixed individuals can also identify hybrids (for example, Supplementary Figure S6), even when missing source populations.
Our results challenge the idea of complete isolation between P. leucopus and P. maniculatus, supporting an emerging view that reproductive isolation can vary depending on individual genotype and demographic context (Kozlowska et al., 2012; Chunco, 2014; Mandeville et al., 2015; Araripe et al., 2016; Gompert and Buerkle, 2016; Senerchia et al., 2016). The presence of hybrids on the range margin also supports the idea that pre-mating barriers (for example, Doty, 1972, 1973) between sympatric species may be altered during climate-driven range shifts; in essence a displacement of co-evolved genotypes driven by anthropogenic climate change (Crispo et al., 2011; Chunco, 2014).
Hybridization is expected to lead to costs in fitness due to intrinsic and extrinsic selective pressures. However, some of the fitness costs associated with hybridization during range expansion may be mitigated by evolutionary phenomena associated with expansion in small populations, including softened selection (Peischl et al., 2013), fixation of compensatory adaptive alleles (Poon and Otto, 2000), or a decrease in the rate of small-effect deleterious mutations (LaBar and Adami, 2016). Selection against, and purging of polymorphic incompatibility loci (Cutter, 2012) may also result in a rebound of hybrid fitness (Araripe et al., 2016). These processes may contribute to the conversion of a climate-driven wave-front of a single species to a moving hybrid zone (for example, Chunco, 2014; Taylor et al., 2014, 2015).
Conclusion
We showed that genetic variation in P. leucopus from southern Quebec is associated with geography and shaped by post-glacial and recent range expansion. Our results of genome-wide population structure and genetic diversity indicate Quebec was colonized by two putative glacial lineages, one of which was further isolated during post-glacial expansion by the Richelieu River. Consistent with the predicted consequences of range expansion (Currat et al., 2008; Excoffier and Ray, 2008; Excoffier et al., 2009), we found a northern-most population showing low nucleotide diversity, the lowest effective population size, divergent allele frequencies, the highest number of private alleles and heterozygosity levels that suggest introgression from an unsampled local population. Analysis of ancestry identified putative P. maniculatus and P. leucopus hybrids. Testing the bifurcating model genealogical evolution confirmed a reticulate phylogeny and past gene flow between P. maniculatus and P. leucopus in Quebec. This result adds to the rapidly increasing list of natural hybrids discovered between species-pairs previously thought to be isolated, supporting the idea that reproductive isolation between closely related species can vary based on genotype and demographic conditions. More generally, our study supports the view that species boundaries not only change over time but also vary across space.
Data archiving
The raw sequence data has been deposited at the NCBI Sequence Read Archive (SRA) repository and can be accessed under accession number PRJNA397983.
References
Abbott RJ, Barton NH, Good JM . (2016). Genomics of hybridization and its evolutionary consequences. Mol Ecol 25: 2325–2332.
Alexander DH, Lange K . (2011). Enhancements to the ADMIXTURE algorithm for individual ancestry estimation. BMC Bioinformatics 12: 1–6.
Alexander DH, Novembre J, Lange K . (2009). Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19: 1655–1664.
Andrews KR, Good JM, Miller MR, Luikart G, Hohenlohe PA . (2016). Harnessing the power of RADseq for ecological and evolutionary genomics. Nat Rev Genet 17: 81–92.
Araripe LO, Tao Y, Lemos B . (2016). Interspecific Y chromosome variation is sufficient to rescue hybrid male sterility and is influenced by the grandparental origin of the chromosomes. Heredity 116: 516–522.
Cahill J, Stirling I, Kistler L, Salamzade R, Ersmark E, Fulton TL et al. (2015). Genomic evidence of geographically widespread effect of gene flow from polar bears into brown bears. Mol Ecol 24: 1205–1217.
Canestrelli D, Porretta D, Lowe W, Bisconti R, Carere C, Nascetti G . (2016). The tangled evolutionary legacies of range expansion and hybridization. Trends Ecol Evol 31: 677–688.
Catchen J, Hohenlohe P, Bassham S, Amores A, Cresko W . (2013a). Stacks: an analysis tool set for population genomics. Mol Ecol 22: 3124–3140.
Catchen J, Bassham S, Wilson T, Currey M, O'Brien C, Yeates Q et al. (2013b). The population structure and recent colonization history of Oregon threespine stickleback determined using restriction‐site associated DNA‐sequencing. Mol Ecol 22: 2864–2883.
Chen I, Hill J, Ohlemuller R, Roy DThomas C . (2011). Rapid range sifts of species associated with high levels of climate warming. Science 333: 1024–1026.
Chunco AJ . (2014). Hybridization in a warmer world. Ecol Evol 4: 2019–2031.
Crispo E, Moore JS, Lee‐Yaw JA, Gray SM, Haller BC . (2011). Broken barriers: Human‐induced changes to gene flow and introgression in animals. BioEssays 33: 508–518.
Crow JF, Kimura M . (1970) An Introduction to Population Genetics Theory. Harper & Row Publishers: London and Evanston, NY, USA.
Currat M, Ruedi M, Petit RJ, Excoffier L . (2008). The hidden side of invasions: massive introgression by local genes. Evolution 62: 1908–1920.
Cutter A . (2012). The polymorphic prelude to Bateson–Dobzhansky–Muller incompatibilities. Trends Ecol Evol 27: 209–218.
Dai Q, Zhan X, Lu B, Fu J, Wang Q, Qi D . (2014). Spatial genetic structure patterns of phenotype-limited and boundary-limited expanding populations: a simulation study. PloS ONE 9: e85778.
Dawson WD, Mintz J, Maddock MB, Lewin AR . (1972). On hybridization between deermice (Peromyscus maniculatus and woodmice (P leucopus. ASB Bulletin 19: 64 (Abstract).
Dice LR . (1933). Fertility relationships between some of the species and subspecies of mice in the genus Peromyscus. J Mammal 14: 298–305.
Dice LR . (1937) Fertility Relations in the Peromyscus leucopus Group of Mice. Contrib. Lab. Vert. Genet. Univ. Michigan 4:1–3.
Dice LR . (1968) Biology of Peromyscus (Rodentia), Special 2nd edn. American Society of Mammalogists. pp 75–97.
Doty RL . (1972). Odor preferences of female Peromyscus maniculatus bairdi for male mouse odors of P. m. bairdi and P leucopus noveboracensis as a function of estrous state. J Comp Physio Psych 81: 191–197.
Doty RL . (1973). Reactions of deer mice (Peromyscus maniculatus and white-footed mice (Peromyscus leucopus to homospecific and heterospecific urine odors. J Comp Physio Psych 84: 296–303.
Elshire RJ, Glaubitz JC, Sun Q, Poland JA, Kawamoto K, Buckler ES et al. (2011). A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PloS ONE 6: e19379.
Ewen JG, Amstrong DP, Parker KAP, Seddon PJ (eds) (2012). Reintroduction biology: integrating science and management. John Wiley & Sons.
Excoffier L, Foll M, Petit RJ . (2009). Genetic consequences of range expansions. Annu Rev Ecol Evol Syst 40: 481–501.
Excoffier L, Ray N . (2008). Surfing during population expansions promotes genetic revolutions and structuration. Trends Ecol Evol 23: 347–351.
Falush D, van Dorp L, Lawson D . (2016). A tutorial on how (not) to over-interpret STRUCTURE/ADMIXTURE bar plots. bioRxiv 066431.
Fiset J, Tessier N, Millien V, Lapointe F . (2015). Phylogeographic structure of the white-footed mouse and the deer mouse, two Lyme disease reservoir hosts in Québec. PLoS One 10: e0144112.
Frichot E, François O . (2015). LEA: An R package for landscape and ecological association studies. Methods Ecol Evol 6: 925–929.
Fu Q, Hajdinjak M, Moldovan O, Constantin S, Mallick S, Skoglund P et al. (2015). An early modern human from Romania with a recent Neanderthal ancestor. Nature 524: 216–219.
Gaitan J, Millien V . (2016). Stress level, parasite load, and movement pattern in a small mammal reservoir host for Lyme disease. Can J Zoology 94: 565–573.
Garnier J, Lewis MA . (2016). Expansion under climate change: the genetic consequences. Bull Math Biol 78: 2165–2185.
Grant PR . (1976). An 11-year study of small mammal populations at Mont St. Hilaire, Quebec. Can J Zool 54: 2156–2173.
Hallatschek O, Hersen P, Ramanathan S, Nelson DR . (2007). Genetic drift at expanding frontiers promotes gene segregation. Proc Natl Acad Sci USA 104: 19926–19930.
Harris SE, Munshi-South J, Obergfell C, O’Neill R . (2013). Signatures of rapid evolution in urban and rural transcriptomes of white-footed mice (Peromyscus leucopus in the New York metropolitan area. PLoS One 8: e74938.
Harrison RG . (1990). Hybrid zones: windows on evolutionary process. Oxf Surv Evol Biol 7: 69–128.
Hedrick P . (2013). Adaptive introgression in animals: examples and comparison to new mutation and standing variation as sources of adaptive variation. Mol Ecol 22: 4606–4618.
Henn B, Botigué L, Peischl S, Dupanloup I, Lipatov M, Maples BK et al. (2015). Distance from sub-Saharan Africa predicts mutational load in diverse human genomes. Proc Natl Acad Sci USA 113: E440–E449.
Hibbard CW . (1968). Paleontology. In: JA King (ed). Biology of Peromyscus (Rodentia), Special publication (American Society of Mammalogists) no. 2, pp 6–26.
Hickling R, Roy DB, Hill JK, Fox R, Thomas CD . (2006). The distributions of a wide range of taxonomic groups are expanding polewards. Global Change Biol 12: 450–455.
Huerta-Sánchez E, Jin X, Asan, Bianba Z, Peter B, Vinckenbosch N et al. (2014). Altitude adaptation in Tibetans caused by introgression of Denisovan-like DNA. Nature 512: 194–197.
Jombart T . (2008). adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24: 1403–1405.
Jombart T, Collins CA . (2015) tutorial for discriminant analysis of principal components (DAPC) using adegenet 2.0.0. Imperial College London, MRC Centre for Outbreak Analysis and Modelling: London.
Jombart T, Devillard S, Balloux F . (2010). Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics 11: 94.
Kelly B, Whiteley A, Tallmon D . (2010). The Arctic melting pot. Nature 468: 891–891.
King J . (1968) Biology of Peromyscus (Rodentia), Special publication (American Society of Mammalogists) no. 2.
Kozlowska J, Ahmad A, Jahesh E, Cutter A . (2012). Genetic variation for postzygotic reproductive isolation between Caenorhabditis briggsae and Caenorhabditis SP. 9. Evolution 66: 1180–1195.
Krohne DT, Dubbs BA, Baccus R . (1984). An analysis of dispersal in an unmanipulated population of Peromyscus leucopus. Am Midl Nat 112: 146–156.
LaBar T, Adami C . (2016). Evolution of drift robustness in small populations of digital organisms. bioRxiv 071894.
Langmead B, Salzberg S . (2012). Fast gapped-read alignment with Bowtie 2. Nat Methods 9: 357–359.
Ledevin R, Millien V . (2013). Congruent morphological and genetic differentiation as a signature of range expansion in a fragmented landscape. Ecol Evol 3: 4172–4182.
Leo SS, Millien V . (2016). Microsatellite markers reveal low frequency of natural hybridization between the white-footed mouse (Peromyscus leucopus and deer mouse (Peromyscus maniculatus in southern Quebec, Canada. Genome 60: 454–463.
Lessa EP, Cook JA, Patton JL . (2003). Genetic footprints of demographic expansion in North America, but not Amazonia, during the Late Quaternary. Proc Natl Acad Sci USA 100: 10331–10334.
Lewontin R, Birch L . (1966). Hybridization as a source of variation for adaptation to new environments. Evolution 20: 315–336.
Li Y, Willer C, Sanna S, Abecasis G . (2009). Genotype imputation. Ann Rev Genomics Hum Genetics 10: 387–406.
Lindquist ES, Aquadro CF, McClearn D, McGowan KJ . (2003). Field identification of the mice Peromyscus leucopus noveboracensis and P maniculatus gracilis in central New York. Canadian FieldNaturalist 117: 184–189.
Lowe W, Muhlfeld C, Allendorf F . (2015). Spatial sorting promotes the spread of maladaptive hybridization. Trends Ecol Evol 30: 456–462.
Lynch M . (1991). The genetic interpretation of inbreeding depression and outbreeding depression. Evolution 45: 622–629.
Maddock MB, Dawson WD . (1974). Artificial insemination of deermice (Peromyscus maniculatus) with sperm from other rodent species. Development 31: 621–634.
Mandeville E, Parchman T, McDonald D, Buerkle C . (2015). Highly variable reproductive isolation among pairs of Catostomus species. Mol Ecol 24: 1856–1872.
Marrotte R, Gonzalez A, Millien V . (2014). Landscape resistance and habitat combine to provide an optimal model of genetic structure and connectivity at the range margin of a small mammal. Mol Ecol 23: 3983–3998.
Mastrantonio V, Porretta D, Urbanelli S, Crasta G, Nascetti G . (2016). Dynamics of mtDNA introgression during species range expansion: insights from an experimental longitudinal study. Sci Rep 6: 30355.
McVean G . (2009). A genealogical interpretation of principal components analysis. PLoS Genet 5: e1000686.
Miao B, Wang Z, Li Y . (2016). Genomic analysis reveals hypoxia adaptation in the Tibetan mastiff by introgression of the grey wolf from the Tibetan plateau. Mol Biol Evol 34: 734–743.
Mondal M, Casals F, Xu T, Dall'Olio GM, Pybus M, Netea MG . (2016). Genomic analysis of Andamanese provides insights into ancient human migration into Asia and adaptation. Nat Genet 48: 1066–1070.
Money D, Gardner K, Migicovsky Z, Schwaninger H, Zhong GY, Myles S . (2015). LinkImpute: fast and accurate genotype imputation for nonmodel organisms. G3 5: 2383–2390.
Monzón JD, Atkinson EG, Henn BM, Benach JL . (2016). Population and evolutionary genomics of Amblyomma americanum, an expanding arthropod disease vector. Genome Biol Evol 8: 1351–1360.
Munshi-South J, Zolnik C, Harris S . (2016). Population genomics of the Anthropocene: urbanization is negatively associated with genome-wide variation in white-footed mouse populations. Evol Appl 9: 546–564.
Myers P, Lundrigan BL, Hoffman SM, Haraminac AP, Seto SH . (2009). Climate‐induced changes in the small mammal communities of the Northern Great Lakes region. Global Change Biol 15: 1434–1454.
Nam K, Jeong H, Nam J . (2016). Pseudo-reference-based assembly of vertebrate transcriptomes. Genes 7: 10.
Novembre J, Stephens M . (2008). Interpreting principal component analyses of spatial population genetic variation. Nat Genet 40: 646–649.
Nullmeier J, Hallatschek O . (2013). The coalescent in boundary-limited range expansions. Evolution 67: 1307–1320.
Peischl S, Dupanloup I, Kirkpatrick M, Excoffier L . (2013). On the accumulation of deleterious mutations during range expansions. Mol Ecol 22: 5972–5982.
Peischl S, Dupanloup I, Foucal A, Jomphe M, Bruat V, Grenier JC et al. (2016). Relaxed selection during a recent human expansion. bioRxiv. 064691.
Pfennig KS, Kelly AL, Pierce AA . (2016). Hybridization as a facilitator of species range expansion. Proc R Soc B 283: 20161329–20161338.
Pierce S, Vogt F . (1993). Winter acclimatization in Peromyscus maniculatus gracilis P leucopus noveboracensis, and P l. leucopus. J Mammal 74: 665–677.
Pluess AR . (2011). Pursuing glacier retreat: genetic structure of a rapidly expanding Larix decidua population. Mol Ecol 20: 473–485.
Poon A, Otto S . (2000). Compensating for our load of mutations: freezing the meltdown of small populations. Evol 54: 1467–1479.
Puechmaille SJ . (2016). The program structure does not reliably recover the correct population structure when sampling is uneven: subsampling and new estimators alleviate the problem. Mol Ecol Resour 16: 608–627.
Purcell S, Chang C . (2015). PLINK 1.9. Available at https://www.cog-genomics.org/plink2.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D et al. (2007). PLINK: a tool set for whole-genome association and population-based linkage analyses. The Am J Hum Gene 81: 559–575.
Pickrell JK, Pritchard JK . (2012). Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet 8: e1002967.
Racimo F, Gokhman D, Fumagalli M, Hansen T, Moltke I, Albrechtsen A et al. (2016). Archaic adaptive introgression in TBX15/WARS2. Molecular Biology and Evolution 34: 509–524.
Reich D, Wayne R, Goldstein D . (1999). Genetic evidence or a recent origin by hybridization of red wolves. Mol Ecol 8: 139–144.
Rhymer JM, Simberloff D . (1996). Extinction by hybridization and introgression. Ann Rev Ecol Syst 27: 83–109.
Rogic A, Tessier N, Legendre P, Lapointe F, Millien V . (2013). Genetic structure of the white-footed mouse in the context of the emergence of Lyme disease in southern Québec. Ecol Evol 3: 2075–2088.
Rousset F . (1997). Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145: 1219–1228.
Rowe KC, Heske EJ, Paige KN . (2006). Comparative phylogeography of eastern chipmunks and white‐footed mice in relation to the individualistic nature of species. Mol Ecol 15: 4003–4020.
Roy-Dufresne E, Logan T, Simon J, Chmura G, Millien V . (2013). Poleward expansion of the white-footed mouse (Peromyscus leucopus under climate change: implications for the spread of Lyme disease. PLoS One 8: e80724.
Sambrook J, Fritsch EF, Maniatis T . (1989) Molecular cloning: a laboratory manual. Chapter 6 - Preparation and Analysis of Eukaryotic Genomic DNA. Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY.
Schwartz MK, McKelvey KS . (2009). Why sampling scheme matters: the effect of sampling scheme on landscape genetic results. Conservation Genetics 10: 441–452.
Seehausen O . (2004). Hybridization and adaptive radiation. Trends Ecol Evol 19: 198–207.
Senerchia N, Felber F, North B, Sarr A, Guadagnuolo R, Parisod C . (2016). Differential introgression and reorganization of retrotransposons in hybrid zones between wild wheats. Mol Ecol 11: 2518–2528.
Shafer A, Gattepaille LM, Stewart RE, Wolf JB . (2015). Demographic inferences using short‐read genomic data in an approximate Bayesian computation framework: in silico evaluation of power, biases and proof of concept in Atlantic walrus. Mol Ecol 24: 328–345.
Shafer A, Peart CR, Tusso S, Maayan I, Brelsford A, Wheat CW et al. (2017). Bioinformatic processing of RAD‐seq data dramatically impacts downstream population genetic inference. Methods in Ecol Evol 8: 907–917.
Shine R, Brown G, Phillips B . (2011). An evolutionary process that assembles phenotypes through space rather than through time. Proceedings of the Natl Acad Sci 108: 5708–5711.
Slatkin M, Excoffier L . (2012). Serial Founder Effects During Range Expansion: A Spatial Analog of Genetic Drift. Genetics 191: 171–181.
Sobel JM, Chen GF, Watt LR, Schemske DW . (2010). The biology of speciation. Evolution 64: 295–315.
Song Y, Endepols S, Klemann N, Richter D, Matuschka F, Shih C et al. (2011). Adaptive Introgression of Anticoagulant Rodent Poison Resistance by Hybridization between Old World Mice. Current Biology 21: 1296–1301.
Statistique Canada. (2014). Recensements du Canada. Available at http://www.stat.gouv.qc.ca/statistiques/population-demographie/structure/Tableau_top_10.htm.
Swaegers J, Mergeay J, Van Geystelen A, Therry L, Larmuseau MH, Stoks R . (2015). Neutral and adaptive genomic signatures of rapid poleward range expansion. Mol Ecol 24: 6163–6176.
Taylor DR, Keller SR . (2007). Historical range expansion determines the phylogenetic diversity introduced during contemporary species invasion. Evolution 61: 334–345.
Taylor S, Curry R, White T, Ferretti V, Lovette I . (2014). Spatiotemporally consistent genomic signatures of reproductive isolation in a moving hybrid zone. Evolution 68: 3066–3081.
Taylor SA, Larson EL, Harrison RG . (2015). Hybrid zones: windows on climate change. Trends Ecol Evol 30: 398–406.
Trask JA, Malhi RS, Kanthaswamy S, Johnson J, Garnica WT, Malladi VS et al. (2011). The effect of SNP discovery method and sample size on estimation of population genetic data for Chinese and Indian rhesus macaques (Macaca mulatta. Primates 52: 129–138.
Vaha JP, Primmer CR . (2006). Efficiency of model-based Bayesian methods for detecting hybrid individuals under different hybridization scenarios and with different numbers of loci. Mol Ecol 15: 63–72.
Vessey SH, Vessey KB . (2007). Linking behavior, life history and food supply with the population dynamics of white‐footed mice (Peromyscus leucopus. Integr Zoology 2: 123–130.
von Holdt B, Kays R, Pollinger J, Wayne R . (2016). Admixture mapping identifies introgressed genomic regions in North American canids. Mol Ecol 25: 2443–2453.
Wan JJ . (2014). Ecological Replacement of Peromyscus maniculatus gracilis by Peromyscus leucopus in northern Michigan. PhD dissertation, University of Michigan.
Wang J . (2017). The computer program Structure for assigning individuals to populations: easy to use but easier to misuse. Mol Ecol Resour. doi:10.1111/1755-0998.12650.
White T, Perkins S, Heckel G, Searle J . (2013). Adaptive evolution during an ongoing range expansion: the invasive bank vole (Myodes glareolus in Ireland. Mol Ecol 22: 2971–2985.
Whitlock MC, Mccauley DE . (1999). Indirect measures of gene flow and migration: FST≠ 1/(4Nm+ 1). Heredity 82: 117–125.
Wu CI . (2001). The genic view of the process of speciation. J Evoly Biol 6: 851–865.
Acknowledgements
We thank two anonymous reviewers for their useful comments and suggestions on the manuscript. This research was funded through NSERC Discovery Grants to VM and to JTD and a Canada Research Chair to RDHB. Work on this publication was also supported by a McGill Biology Department University Research Travel Award to AG and by the National Institute of General Medical Sciences of the National Institutes of Health under award number R15GM099055 to JM-S. The content is solely the responsibility of the authors and does not represent the official views of the National Institutes of Health. We thank Brian Boyle from IBIS for his assistance during the library preparation and sharing his considerable expertise. We also thank Antoine Paccard and Frederique Chain for bioinformatic support and Sarah T. Leo for her help during molecular laboratory work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Supplementary information
Rights and permissions
About this article
Cite this article
Garcia-Elfring, A., Barrett, R., Combs, M. et al. Admixture on the northern front: population genomics of range expansion in the white-footed mouse (Peromyscus leucopus) and secondary contact with the deer mouse (Peromyscus maniculatus). Heredity 119, 447–458 (2017). https://doi.org/10.1038/hdy.2017.57
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2017.57
This article is cited by
-
It’s about time: small mammal communities and Lyme disease emergence
Scientific Reports (2023)
-
Genome-wide assessment of population structure in Florida’s coastal seaside sparrows
Conservation Genetics (2022)
-
Introgression at the emerging secondary contact zone of magpie Pica pica subspecies (Aves: Corvidae): integrating data on nuclear and mitochondrial markers, vocalizations, and field observations
Organisms Diversity & Evolution (2022)
-
Harmonizing hybridization dissonance in conservation
Communications Biology (2020)
-
Paternal leakage and mtDNA heteroplasmy in Rhipicephalus spp. ticks
Scientific Reports (2019)