Abstract
The utilization of similar habitats by different species provides an ideal opportunity to identify genes underlying adaptation and acclimatization. Here, we analysed the gene expression of two closely related salamander species: Salamandra salamandra in Central Europe and Salamandra infraimmaculata in the Near East. These species inhabit similar habitat types: ‘temporary ponds’ and ‘permanent streams’ during larval development. We developed two species-specific gene expression microarrays, each targeting over 12 000 transcripts, including an overlapping subset of 8331 orthologues. Gene expression was examined for systematic differences between temporary ponds and permanent streams in larvae from both salamander species to establish gene sets and functions associated with these two habitat types. Only 20 orthologues were associated with a habitat in both species, but these orthologues did not show parallel expression patterns across species more than expected by chance. Functional annotation of a set of 106 genes with the highest effect size for a habitat suggested four putative gene function categories associated with a habitat in both species: cell proliferation, neural development, oxygen responses and muscle capacity. Among these high effect size genes was a single orthologue (14-3-3 protein zeta/YWHAZ) that was downregulated in temporary ponds in both species. The emergence of four gene function categories combined with a lack of parallel expression of orthologues (except 14-3-3 protein zeta) suggests that parallel habitat adaptation or acclimatization by larvae from S. salamandra and S. infraimmaculata to temporary ponds and permanent streams is mainly realized by different genes with a converging functionality.
Similar content being viewed by others
Introduction
Parallel evolution of phenotypic traits has been described for a broad range of organisms and is seen as an evidence for natural selection (Losos, 2011). Case studies of parallel phenotypic evolution can reveal genes and functions underlying the repeated evolution of adaptive traits (Schluter et al., 2004; Elmer and Meyer, 2011; Pfenninger et al., 2014). Evolutionary adaptation only takes place in phenotypically expressed traits that are exposed to natural selection. Hence, acclimatization, a plastic response in which gene expression and its phenotype change to accommodate variable environmental conditions within an organism’s lifespan, can be intimately linked with adaptation. The idea that phenotypic plasticity may play an important role in adaptive evolution has a long history (Pigliucci and Murren, 2003; West-Eberhard, 2003), but the underlying genetic mechanisms are still being explored. Although phenotypically plastic traits and a fixed adaptive divergence are difficult to distinguish, studies of parallel phenotypic divergence in relation to environmental factors provide the basis to identify how plasticity and adaptation may interact.
Phenotypic plasticity is mainly realized by gene expression regulation (Pigliucci, 1996). For the case of parallel evolution, adaptation to similar habitats may be realized through: (1) parallel adaptation of orthologous genes (that is, changes in the same genes associated with these habitats) or (2) changes in different genes with similar functions. Both scenarios have been found in studies on adaptation (Arendt and Reznick, 2008); for example, sticklebacks have reduced armour plating by the same genetic mechanism independently in different populations (Colosimo et al., 2005), whereas beetles have developed similar horn appendages using different genetic mechanisms (Moczek and Nagy, 2005) and populations of beech mice have developed similar colour morphs based on a different set of genes (Steiner et al., 2009). Similarly, acclimatization (plastic responses) to similar habitats may be realized through consistent expression of the same genes or through expression changes in different genes with similar functions (Whitehead et al., 2011, 2013). In this study, we aimed to distinguish between these two scenarios for parallel acclimatization or adaptation by investigating gene expression in two closely related fire salamander species using similar larval habitats: Salamandra salamandra in Central Europe and Salamandra infraimmaculata in the Near East. Many factors may influence whether local adaptation or phenotypic plasticity will evolve in response to environmental heterogeneity. A modelling study has shown that temporal variation in environmental conditions more readily favours phenotypic plasticity, whereas spatial variation in environmental conditions is more associated with local adaptation (Moran, 1992).
Although S. salamandra and S. infraimmaculata can be considered as terrestrial salamanders, females of each species deposit larvae into a variety of aquatic habitats, in which the larvae develop until metamorphosis. Across their ranges, both species utilize two broad categories of larval habitats: ‘temporary ponds’ with temporary (rain-dependent) and stagnant (lentic) water, and ‘permanent streams’ with permanent (spring-fed) and flowing (lotic) water. Temporary ponds are typically small stagnant rainwater-filled depressions in the landscape that dry up each year. Permanent streams are typically first-order spring-fed streams with a more or less constant water flow throughout the year. We expected that many environmental conditions (for example, physical water flow, hydroperiod, oxygen concentration, food sources and food availability) are systematically different between these two habitat categories in a similar manner across Central Europe and the Near East, even though the magnitude of the effects may vary between sites (see Reinhardt et al., 2013, 2015 for S. salamandra). These two habitats should therefore impose systematically different selection pressures. We further expected that salamander larvae will optimize survival, growth and development by adjusting their physiology, morphology and behaviour to local conditions through differential gene expression. We decided to address our hypothesis that habitat acclimatization in these two salamander species is realized through changes in the expression of different genes with similar functions, by investigating larvae sampled from these two habitat categories.
Some S. salamandra populations in Western Germany started utilizing temporary ponds after the last glaciation (Weitere et al., 2004; Steinfartz et al., 2007). The Kottenforst near Bonn in Western Germany is a forest hosting a particularly large and well-studied population of S. salamandra that has been shown to display ecological adaptation to both stream and pond habitats as well as habitat–habitat-specific population genetic diversification at a local scale (Steinfartz et al., 2007). This population represents a natural system to study the consequences of the ongoing adaptive divergence (Reinhardt et al., 2013; Caspers et al., 2014, 2015). In the Kottenforst (and indeed in many other cases), evolutionary adaptation and acclimatization processes take place simultaneously. This means that fixed genetic adaptation and plastic acclimatization effects on gene expression patterns cannot be easily disentangled. This question requires further study using, for example, common garden or transplant experiments. Here, we focus on the question of whether the same or different genes underlie phenotypic habitat adaptation.
Similar to S. salamandra in Germany, fire salamanders in Israel (S. infraimmaculata) have been intensively studied for their ecology (see, for example, Bar-David et al., 2007; Blank and Blaustein, 2014) and population genetics (Blank et al., 2013). Larval habitats of S. infraimmaculata in Israel are ecologically more diverse than those of S. salamandra and include a variety of temporary ponds, springs, temporary streams and water bodies in caves (Degani, 1996). A permanent or temporary hydroperiod of the larval habitat has been shown to be an important factor for population dynamics (Segev et al., 2010; Blank and Blaustein, 2014). Permanent streams are relatively rare in the range of S. infraimmaculata in Israel, but the first-order streams of the Tel Dan Nature Reserve resemble the permanent stream habitats of S. salamandra in Germany in that they have permanently flowing water, high oxygen concentrations and provide the larvae with stable food conditions.
A study of nuclear and mitochondrial genes revealed that S. salamandra and S. infraimmaculata diverged during the Pliocene ∼4 million years ago from an ancestral lineage, also including North-African S. algira (not included in this study) and have not been hybridizing since then (see Vences et al., 2014 for phylogenetic details). Considering the extensive time since common ancestry, we expect that parallel patterns in gene expression due to phylogenetic inheritance are unlikely. Because of their parallel use of larval habitats at the intraspecific level and shared phylogenetic ancestry, the larvae of S. salamandra and S. infraimmaculata provide an excellent case to study parallel habitat acclimatization and adaptation through gene expression analysis. For this purpose, we developed two species-specific microarrays (based on transcriptomes of larvae), both of which contain probes for >12 000 genes that are expressed at the larval stage. These two microarrays were overlapped by a subset of 8331 orthologous genes, and formed the basis for this study. The aim of the study was to identify whether parallel adaptation or acclimatization in S. salamandra and S. infraimmaculata was realized through the parallel expression of the same orthologous genes, or through the expression of different genes with a similar functionality. Based on their history of genetic isolation, we hypothesized that parallel adaptation or acclimatization mechanisms to temporary pond and permanent stream habitats in these two salamander species are realized through changes in the expression of different genes with similar functions, rather than parallel expression of orthologues.
Materials and methods
Sampling of salamander larvae of S. salamandra and S. infraimmaculata
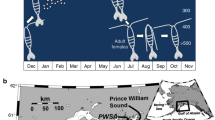
For the microarray experiments, we sampled four S. salamandra larvae per site from 4 temporary ponds and 6 permanent streams in Germany, and between 2 and 13 S. infraimmaculata larvae per site from 7 temporary ponds and 1 permanent stream in Northern Israel (see Table 1 for detailed numbers, see also Figures 1 and 2). Sampling in Germany took place in April 2013 and 2014 with permission from the nature reserve authority of the municipality of Bonn. Sampling in Israel took place during December–January 2013–2014 and December 2014 with permission from the Israel Nature and Park Authority. Freshly deposited larvae (with a ventral yolk patch) as well as large larvae with morphological signs of metamorphosis (gill reduction or increased colouration) were excluded from sampling to reduce maternal or metamorphosis effects on gene expression patterns, respectively. We applied a nonlethal sampling approach, as gene expression patterns from salamander larval tail-tip tissue were shown to be representative of whole-body gene expression patterns with largely overlapping gene functions for thermal response in S. salamandra (Czypionka et al., 2015). Salamander larvae were caught with nets, and a tail-tip tissue sample (~5-mm long) was taken from each larva. Tail-tip tissue in salamander larvae is known to have a regeneration capacity and can be harvested with little fitness consequences for the individual, as demonstrated for S. infraimmaculata in Israel (Segev et al., 2015). After sampling, the larvae were immediately released at the location of initial capture. Tail-tip tissue samples were stored in RNA-later at −20 °C until RNA extraction.
Development of two microarrays for S. salamandra and S. infraimmaculata
Two separate microarrays for gene expression analysis of S. salamandra and S. infraimmaculata were designed following Czypionka et al. (2015). Transcriptome data were generated using paired-end total RNA sequencing (Illumina HiSeq2000, San Diego, CA, USA) of two larvae from each species representing both habitat types (Table 1). Contigs were assembled using Trinity (Grabherr et al., 2011) and scanned for open-reading frames using the software Transdecoder (Haas et al., 2013). Open-reading frames with no homology to known amphibian proteins based on BLASTP (Altschul and Lipman, 1990) against the NCBI (National Center for Biotechnology Information) non-redundant database were filtered out. Oligonucleotide probes were designed using the parent contigs of the filtered open-reading frames. Contigs with multiple open-reading frames were not used for probe design. The target-binding behaviour of surface-bound oligonucleotides varies greatly between different oligonucleotide sequences (Pozhitkov et al., 2006). For this reason, we assessed the target-binding behaviour of all probes on the two microarrays in a series of calibration experiments using a range of relevant mRNA concentrations following (Czypionka et al. 2012, 2015). We fitted a linear dose–response curve between the observed signal intensity and the amount of hybridized cRNA in the calibration experiment (signal intensity=slope × amount of cRNA+intercept). Probes with a nonlinear binding behaviour and unresponsive probes were excluded. Validated probes were spotted on Agilent microarrays (Santa Clara, CA, USA), finally representing 12 710 and 12 926 genes for S. salamandra (Agilent Design ID 069636) and S. infraimmaculata (Agilent Design ID 067672), respectively. For more details on the development of these two microarrays, please see the extended methods and results in the Supplementary Information.
Microarray analysis and data normalization
RNA was extracted from each tissue sample using a Trizol protocol and hybridized on the corresponding species-specific microarray using the Agilent Low Input Quick Amp Labelling kit for ‘One-Color Microarray-Based Gene Expression Analysis’. Fluorescent label signal intensities were quantified in an Agilent DNA Microarray Scanner (type C). Signal intensities were normalized using custom R code (R Core Team, 2010) that (1) corrected the signal intensities for probe-binding behaviour based on the dose–response curves from the calibration experiments, (2) merged the technical replicates of each probe on the array by taking their median and (3) performed between-array normalization by dividing the so-far normalized signal intensities by their 75 percentile, as recommended in the Agilent ‘One-Color Microarray-Based gene expression Analysis’ protocol.
Microarray differential gene expression and functional analysis
To explore the patterns of gene expression, we employed principal component analysis (PCA) in R. In addition, to identify genes that were significantly differentially expressed between habitat types, we employed the linear discriminant analysis of effect size (LEfSe) method (Segata et al., 2011) from the Galaxy platform (http://huttenhower.sph.harvard.edu/galaxy/). LEfSe is an algorithm that, based on high-dimensional quantitative markers (here genome-wide expression levels), identifies features characterizing the differences between two or more biological conditions. In the first step, it applies a nonparametric factorial Kruskal–Wallis rank-sum test to detect genes with a significant differential expression between habitat types (that is, it identifies violation of the null hypothesis of no difference between habitats). In the second step, biological significance is further restricted by using a set of pairwise tests among subclasses using the Wilcoxon rank-sum test. By applying LEfSe we effectively tested whether the expression values are significantly and consistently up- or downregulated in all subclasses (sites) of a class (habitat type) compared with all subclasses (sites) of the other class (habitat type). In the final step, LEfSe uses linear discriminant analysis to estimate the effect size of each differentially expressed gene. This method allowed us to perform a nonparametric high-dimensional comparative analysis of consistent gene expression differences between habitat types. We used default LEfSe parameters (including all-against-all multiclass analysis and α=0.05). Linear discriminant analysis scores were used for only effect size ranking.
To identify overrepresented functional categories in sets of differentially expressed candidate genes, we first performed Gene Ontology (GO) term enrichment analysis using Fisher’s exact tests with false discovery rate correction, as implemented in Blast2GO 2.8 using default parameters (Conesa et al., 2005). Annotation of genes was performed using BLASTX against the entire NCBI non-redundant database. However, GO annotations are often incomplete or biased, and the absence of annotation does not imply the absence of function (Thomas et al., 2012). In other words, different molecular solutions (in different species) to the same environmental challenge may not be annotated completely or identically in the GO system. For this reason, it is possible to miss certain functional similarities between nonmodel organisms using blast-based analyses. Therefore, we also examined the possible functional similarity independently of the GO system, using functional information from the biocurated UniProt Knowledgebase (August 2016, The_UniProt_Consortium, 2015). The ‘Function subsection’ and ‘Enzyme and pathway databases subsection’ of the Function section on the Uniprot Knowledgebase were manually retrieved for each candidate gene and assessed for the (co-)occurrence of putative biologically relevant functional keywords (that is, ‘epidermis’, ‘cell proliferation’, ‘immune’, ‘mitochondrion’/‘mitochondrial’, ‘muscle’, ‘neuron’/‘neural’, ‘oxygen’, ‘ribosome’/‘ribosomal’ and ‘stress’). Putative functionally similar genes with overlapping functional keywords were reported and discussed in addition to the results from GO term enrichment analysis.
Orthologous genes on these two microarrays were identified by reciprocal blast of the target gene sequences. This generated a list of 8331 genes (~65.5%) that we considered to represent orthologues shared between the two species. To evaluate whether the expression patterns of these orthologues were similar between species, we first compared the median normalized signal intensity across samples of temporary ponds with the median of the permanent stream samples. Second, we assessed for each orthologue whether it was congruently up- or downregulated in temporary ponds compared with permanent streams in both species. To test whether the number of orthologues with congruent expression patterns was more than expected by chance alone, we employed Fisher’s exact tests in R.
Results
Gene expression patterns of S. salamandra
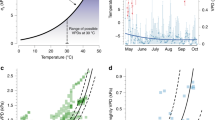
Larvae of S. salamandra (n=40) from 10 sites representing two habitat types (‘permanent stream’ and ‘temporary pond’) were analysed for the expression of 12 710 genes known to be expressed at the larval stage. PCA showed a clear clustering by habitat type (Figure 3). Individuals that belong to the ‘temporary pond’ habitat type clustered together at the bottom-right corner. Individuals that belong to the ‘permanent stream’ habitat type displayed a higher variance in the upper part of the PCA plot. Combined Kruskal–Wallis and pairwise Wilcoxon rank-sum tests as implemented in LEfSe revealed that 863 genes (6.8%) were significantly (α=0.05) and consistently (across sites) expressed differentially between stream and pond habitats. Among these 863 candidate habitat genes, 235 were consistently downregulated and 628 were consistently upregulated in ponds compared with streams. Blast2GO enrichment analysis of the 235 candidate habitat acclimatization genes that are downregulated in ponds versus the full 12 710 genes S. salamandra data set revealed no overrepresented GO terms. However, enrichment analysis of the 628 candidate habitat acclimatization genes that are upregulated in ponds revealed 33 specific overrepresented GO terms (Supplementary Table S3). Based on annotation information from the UniProt Knowledgebase, 30 candidate genes with the highest effect size (Linear Discriminant Analysis LDA >2.0) included 5 genes associated with cell replication or proliferation, 1 gene involved in neural development, 2 genes associated with oxygen responses and 3 genes involved in muscle capacity (Table 2, see Supplementary Table S1 for the full list of candidate genes).
(a) PCA plot of gene expression patterns from Salamandra salamandra in Germany. PC1 and PC2 explain 48.5% of the total variance. Open circles indicate permanent stream samples and closed circles indicate temporary pond samples. Ellipses indicate the 95% confidence intervals of the permanent stream and temporary pond habitat categories. A version with individual sample labels can be found in the Supplementary Figure S1. (b) PCA plot of gene expression patterns from Salamandra infraimmaculata larvae in Israel. PC1 and PC2 explain 43.3% of the total variance. Open circles indicate permanent stream samples and closed circles indicate temporary pond samples. Ellipses indicate the 95% confidence intervals of the permanent stream and temporary pond habitat categories. A version with individual sample labels can be found in the Supplementary Figure S2.
Gene expression patterns of S. infraimmaculata
Larvae of S. infraimmaculata (n=38) sampled from eight sites classified as the same two habitat types as utilized by S. salamandra were analysed for the expression of 12 926 genes known to be expressed by S. infraimmaculata at the larval stage. PCA again showed a clustering by habitat type (Figure 3). LEfSe analysis revealed 1353 genes (10.5%) that were expressed consistently (across sites) at significantly (α=0.05) different levels between the two habitat types (‘permanent stream’ and ‘temporary pond’). Of these 1353 candidate habitat genes, 711 were consistently downregulated and 642 were consistently upregulated in temporary ponds compared with the permanent stream. Blast2GO enrichment analysis of the 711 candidate habitat acclimatization genes that are downregulated in ponds versus the full 12 926 S. infraimmaculata gene set revealed no overrepresented GO terms. Blast2GO enrichment analysis of the 642 candidate habitat acclimatization genes that are upregulated in ponds again revealed no specific overrepresented GO terms. Based on annotation information from the UniProt Knowledgebase, 76 candidate habitat genes with the highest effect size (LDA >2.0) included 9 genes associated with cell replication and proliferation, 6 genes with an immune-related function, 4 genes involved in neural development, 3 genes associated with oxygen responses and 5 genes involved in muscle capacity (Table 3, see Supplementary Table S2 for the full candidate gene list).
Parallel habitat acclimatization in S. salamandra and S. infraimmaculata
All the 863 S. salamandra differentially expressed habitat-associated genes as well as the 1353 S. infraimmaculata differentially expressed habitat-associated genes are represented by orthologues on both microarrays, thus permitting direct comparisons between species. However, only 20 (or 2.3%) of them showed differential expression between habitats in both salamander species. Out of these 20 habitat-related orthologues, 9 (45.0%) displayed congruent expression changes (that is, they were expressed as the highest in the same habitat category in both species). This is not more than expected by chance (Fisher’s exact test, two-tailed P-value=0.729) considering an overall proportion of 48.9% parallel expression patterns among the remaining orthologues.
Investigation of the 106 candidate genes with the highest habitat effect size (LDA >2.0) from both species revealed a single case of orthology between the two candidate gene lists: two genes from either species blasted to ‘14-3-3 protein zeta’ (YWHAZ). These genes were downregulated in temporary ponds compared with permanent streams in both S. salamandra and S. infraimmaculata. Aside from this single overlapping orthologue, a functional comparison of the remaining (non-orthologous) candidate genes with the highest effect size for habitat in both salamander species revealed 30 genes that were functionally associated with the following four putative functional categories (putative traits): cell proliferation, neural development, oxygen response and muscle capacity (Tables 2 and 3).
Discussion
The parallel use of aquatic habitats by larvae of S. salamandra and S. infraimmaculata provides an excellent case to study the genes underlying habitat adaptation or acclimatization through gene expression analysis. The aim of this study was to identify whether parallel adaptation or acclimatization to temporary pond and permanent stream habitats was realized through the parallel expression of the same orthologous genes, or through the expression of different genes with a similar functionality. PCA of the expression of >12 000 annotated genes per species revealed a clear clustering of habitat types (Figure 3). This shows—without any kind of filtering of the data—that the environmental conditions are a dominant factor shaping gene expression in the larvae of both species, and underpins that the habitat categorizations used in this study are biologically meaningful.
Stream sites in the Near East are extremely rare and the stream habitat type of S. infraimmaculata was therefore represented by a single site (Tel Dan). The lack of replication of stream sites in this system is not ideal, as geographic divergence may influence inferences about habitat divergence. However, the variance of the S. infraimmaculata stream samples was augmented by (1) a higher sample size, (2) sampling in two separate years (winter 2013–2014 and December 2014) and (3) sampling two locations within the large stream system of Tel Dan (the Garden of Eden and the Wading Pool, ~200-m apart, with slightly diverging environmental conditions). The PCA (Figure 3b) indicates that the variance is relatively high among the TD samples, and that the overall pattern broadly resembles that of S. salamandra (Figure 3a). Furthermore, exclusion of Wolfsburg and Reitlingstal samples from the S. salamandra data set (thus removing most geographic variation from the data set) showed that the main gene expression pattern of separated pond and stream samples was hardly affected (Supplementary Figure S3). This finding suggests that geographic sampling effects have a relatively little effect on the observations in this study, and that the lack of replication in S. infraimmaculata stream sites is not a critical limitation for this study. Finally, we show that there is an agreement between the putative functional categories of differentially expressed genes in the two species, suggesting that similar traits are selected for by these environments, suggesting in turn that the lack of stream site replication for S. infraimmaculata may again not be a critical point.
Approximately 6.8–10.5% of the investigated genes were differentially expressed between habitats. The number of genes that are significantly differentially expressed between habitats is higher in S. infraimmaculata (1353 genes) than in S. salamandra (863 genes). We propose two nonmutually exclusive biological explanations that may contribute to this observation. First, S. infraimmaculata displays a higher ecological diversity utilizing more diverse larval habitats than S. salamandra (Degani, 1996). This may simply require more genes to be involved in the overall process of habitat acclimatization observed for this species. Second, populations of S. infraimmaculata might have had a longer evolutionary history for specific habitat adaptations to establish than S. salamandra. Based on a population-genetic study of mitochondrial D-loop haplotypes, S. infraimmaculata populations in the Near East indeed show a deeper phylogeographic structure than S. salamandra in Central Europe, where pleistocenic glaciations repeatedly caused the breakdown of population structures (Steinfartz et al., 2000). Accordingly, almost all of the ecological differentiation between populations in Central Europe has been established following the last glaciation, roughly 8000–10 000 years ago, as is the case for the evolution of pond reproduction found in the Kottenforst (Weitere et al., 2004; Steinfartz et al., 2007).
GO term enrichment analysis did not identify any habitat-related functional categories that were shared between the two salamander species, but this approach has limitations, and more detailed investigation is recommended for a biologically meaningful interpretation (Thomas et al., 2012). Detailed inspection of the annotated functional properties of the 106 habitat-associated candidate genes with the highest effect size revealed four putative phenotypic functional traits overlapping between the two salamander species: cell proliferation, neural development, oxygen response and muscle capacity (Table 2). Most of these traits make intuitive sense in the context of ‘temporary pond’ and ‘permanent stream’ habitats. Cell proliferation is a central process controlling the growth rate. Growth rate has been shown to be higher in S. salamandra larvae from temporary ponds, possibly as a survival mechanism to cope with the shorter hydroperiod in this habitat type (Weitere et al., 2004). Oxygen response processes are crucial for dealing with fluctuating environmental oxygen concentrations that are known to differ systematically between lentic ponds and lotic streams (see Introduction). The fact that most of the oxygen response genes are upregulated in temporary ponds supports such a physiological correlation, as small ponds inhabited by salamander larvae are known to have more highly fluctuating oxygen levels (Reinhardt et al., 2013). Muscle capacity may provide adaptation or acclimatization advantages to larvae in streams in terms of the increased swimming capacity, for example, to facilitate foraging and resist downstream drifting, a common risk for S. salamandra (Thiesmeier and Schuhmacher, 1990). For fish, it has been shown that swimming capacity is more geared towards powerful burst swimming in streams compared with lakes (Pavey et al., 2011). The fact that muscle capacity genes are mostly downregulated in temporary ponds suggests that swimming capacity is reduced in ponds compared with streams in both salamander species. The neural development trait is more challenging to interpret. Gene expression patterns suggest that neural development is upregulated (that is, possibly more advanced) in ponds compared with streams. We speculate that increased competition for food, aggression and cannibalism during periods of high density and food shortage in ponds requires a different neural performance, but it is unclear whether this could be reflected in the gene annotation functional comparison reported here.
In addition, S. salamandra exhibited mostly an upregulation of ribosomal proteins in ponds versus streams (nine genes involved, see Supplementary Table S1). Ribosomal protein expression is known to be strongly associated with cell growth (Warner, 1999), and this upregulation in ponds may be associated with cell proliferation and growth rate as described above. Investigation of candidate genes in S. infraimmaculata further revealed genes involved in mitochondrial function (6 genes), environmental stress (5), immune functions (9) and epidermis development (3) that were associated with the habitat categories investigated in this study (see Supplementary Table S2). Even though these traits are not expressed similarly across habitats in these two salamander species, they do make intuitive sense in the context of habitat adaptation or acclimatization. This indicates that the selection pressures in the applied habitat categories may not be perfectly similar across the S. salamandra and S. infraimmaculata study systems. The temporary pond and permanent stream habitats were categorized such that they share many ecological properties across Central Europe and the Near East, but there are also some systematic differences between these two regions, for example, in terms of abiotic (for example, temperature regime, geological parameters) as well as biotic differences. Fire salamanders in Israel reach the southern range limit of any Salamandra species and are limited to limestone regions in the North of the country (Blank and Blaustein, 2012), whereas salamanders in Germany reach the northern range limit of any Salamandra species and experience quite different climatic and geologic conditions. The observation that genes associated with environmental stress, immune functions and epidermis development are expressed differently in response to the environment in both systems is therefore not very surprising.
In a subset of 8331 known orthologous genes, only 20 habitat-related orthologues were significantly differentially expressed between habitats in both species. These 20 overlapping orthologues did not show parallel expression patterns more than expected by chance, suggesting a stochastic result. However, one of the orthologues with a parallel expression pattern (14-3-3 protein zeta/YWHAZ) was ranked among the highest habitat effect size (LDA >2.0) genes of both species (Table 2). This gene encodes a protein that is involved in a range of signalling pathways: first, it inhibits cell cycle progression by sequestering cyclin-dependent kinases to the cytoplasm; second, it protects cells from environmental stresses by negatively regulating apoptosis and similar pathways; and third, it prevents nuclear import of proteins by blocking the nuclear localization signal of target proteins. The fact that this orthologue has a high habitat effect size and a parallel expression pattern across species suggests a case of gene reuse, where the same gene is used in the same way to adapt or acclimatize to different habitats. Aside from this single orthologue, we generally found that habitat adaptation or acclimatization through gene expression is realized by changes in the expression of different genes with a converging functionality in the case of larvae from S. salamandra and S. infraimmaculata.
Whether or not the observed gene expression changes of these habitat-related functional traits are the result of local adaptation or phenotypic plasticity could not be tested in the present field study. To tear these mechanisms apart, dedicated experiments should be carried out, such as a common environment and/or habitat transfer experiments. In the present study system, such an experiment would ideally include a full reciprocal transfer design involving multiple stream and pond sites. Performance over time would need to be monitored through growth parameters and gene expression, allowing comparison of transferred larvae with control individuals. The pond habitat is both temporally and spatially variable and may call for a higher degree of plasticity.
Our observation of different adaptation or acclimatization trajectories to similar habitats in these two salamander species supports a growing body of literature supporting a high degree of unpredictability of evolution, even under similar environmental conditions (Bailey et al., 2015; Pfenninger et al., 2015). Theoretically, and according to microbial evolutionary experiments, many mutations are beneficial at the start of the process of adaptation to a new environment (that is, far away from the fitness optimum) (Schoustra et al., 2009). In addition, early beneficial mutations have a particularly large influence on the further evolutionary trajectory through pleiotropic interactions, priority effects and so on (Lenormand et al., 2008). Parallel evolutionary processes may therefore often be unpredictable at the gene or lower pathway level, particularly for species that diverged a long time ago like S. salamandra and S. infraimmaculata (that is, 4 million years).
Conclusion
In our study of parallel habitat acclimatization or adaptation by larvae from S. salamandra and S. infraimmaculata to temporary pond and permanent stream habitats, we show that among the 106 highest habitat effect size genes in both species, there is only a single case of parallel expression of a shared orthologue. This may represent a case of gene reuse, where the same gene has been recruited independently for the same function in both species. In contrast, we identified 30 different genes converging on a similar functionality in four putative categories: cell proliferation, neural development, oxygen response and muscle capacity. We conclude that parallel habitat acclimatization in this system is mainly realized by different genes. A number of interesting candidate genes associated with these habitats call for further investigation. The applied temporary pond and permanent stream habitat categories appear to assert relatively consistent selection pressures, likely on multiple gene function categories, further suggesting that habitat selection in nature is usually composed of several selection pressures acting at the same time.
Data archiving
Raw signal intensity files, normalized gene expression files and tables with sample information for the S. salamandra and S. infraimmaculata data sets have been deposited at the NCBI Gene Expression Omnibus (GEO) repository under accession SuperSeries GSE101885. This includes the separate S. salamandra data set (accession number GSE100819) and S. infraimmaculata data set (accession number GSE101884).
References
Altschul SF, Lipman DJ . (1990). Protein database searches for multiple alignments. Proc Natl Acad Sci USA 87: 5509–5513.
Arendt J, Reznick D . (2008). Convergence and parallelism reconsidered: what have we learned about the genetics of adaptation? Trends Ecol Evol 23: 26–32.
Bailey SF, Rodriguez N, Kassen R . (2015). The effect of selection environment on the probability of parallel evolution. Mol Biol Evol 32: 1436–1448.
Bar-David S, Segev O, Peleg N, Hill N, Templeton AR, Schultz CB et al. (2007). Long-distance movements by fire salamanders (Salamandra infraimmaculata and implications for habitat fragmentation. Isr J Ecol Evol 53: 143–159.
Blank L, Blaustein L . (2012). Using ecological niche modeling to predict the distributions of two endangered amphibian species in aquatic breeding sites. Hydrobiologia 693: 157–167.
Blank L, Blaustein L . (2014). A multi-scale analysis of breeding site characteristics of the endangered fire salamander (Salamandra infraimmaculata at its extreme southern range limit. Hydrobiologia 726: 229–244.
Blank L, Sinai I, Bar-David S, Peleg N, Segev O, Sadeh A et al. (2013). Genetic population structure of the endangered fire salamander (Salamandra infraimmaculata at the southernmost extreme of its distribution. Anim Conserv 16: 412–421.
Caspers BA, Krause ET, Hendrix R, Kopp M, Rupp O, Rosentreter K et al. (2014). The more the better – polyandry and genetic similarity are positively linked to reproductive success in a natural population of terrestrial salamanders (Salamandra salamandra. Mol Ecol 23: 239–250.
Caspers BA, Steinfartz S, Krause ET . (2015). Larval deposition behaviour and maternal investment of females reflect differential habitat adaptation in a genetically diverging salamander population. Behav Ecol Sociobiol 69: 407–413.
Colosimo PF, Hosemann KE, Balabhadra S, Villarreal G, Dickson M, Grimwood J et al. (2005). Widespread Parallel evolution in sticklebacks by repeated fixation of ectodysplasin alleles. Science 307: 1928–1933.
Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M . (2005). Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21: 3674–3676.
Czypionka T, Cheng J, Pozhitkov A, Nolte AW . (2012). Transcriptome changes after genome-wide admixture in invasive sculpins (Cottus. Mol Ecol 21: 4797–4810.
Czypionka T, Krugman T, Altmüller J, Blaustein L, Steinfartz S, Templeton AR et al. (2015). Ecological transcriptomics – a non-lethal sampling approach for endangered fire salamanders. Methods Ecol Evol 6: 1417–1425.
Degani G . (1996) Salamandra Salamandra: At the Southern Limit of Its Distribution. Laser Pages Publishing: Israel.
Elmer KR, Meyer A . (2011). Adaptation in the age of ecological genomics: insights from parallelism and convergence. Trends Ecol Evol 26: 298–306.
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I et al. (2011). Trinity: reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nat Biotechnol 29: 644.
Haas BJ, Papanicolaou A, Yassour M, Grabherr M, Blood PD, Bowden J et al. (2013). De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat Protoc 8: 1494–1512.
Lenormand T, Roze D, Rousset F . (2008). Stochasticity in evolution. Trends Ecol Evol 24: 157–166.
Losos JB . (2011). Convergence, adaptation, and constraint. Evolution 65: 1827–1840.
Moczek AP, Nagy LM . (2005). Diverse developmental mechanisms contribute to different levels of diversity in horned beetles. Evol Dev 7: 175–185.
Moran NA . (1992). The evolutionary maintenance of alternative phenotypes. Am Nat 139: 971–989.
Pavey SA, Sutherland BJ, Leong J, Robb A, von Schalburg K, Hamon TR et al. (2011). Ecological transcriptomics of lake-type and riverine sockeye salmon (Oncorhynchus nerka. BMC Ecol 11: 1–18.
Pfenninger M, Lerp H, Tobler M, Passow C, Kelley JL, Funke E et al. (2014). Parallel evolution of Cox genes in H2S-tolerant fish as key adaptation to a toxic environment. Nat Commun 5: 3873.
Pfenninger M, Patel S, Arias-Rodriguez L, Feldmeyer B, Riesch R, Plath M . (2015). Unique evolutionary trajectories in repeated adaptation to hydrogen sulphide-toxic habitats of a neotropical fish (Poecilia mexicana. Mol Ecol 24: 5446–5459.
Pigliucci M . (1996). How organisms respond to environmental changes: from phenotypes to molecules (and vice versa). Trends Ecol Evol 11: 168–173.
Pigliucci M, Murren CJ . (2003). Perspective: genetic assimilation and a possible evolutionary paradox: can macroevolution be so fast as to pass us by? Evolution 57: 1455–1464.
Pozhitkov A, Noble PA, Domazet-Lošo T, Nolte AW, Sonnenberg R, Staehler P et al. (2006). Tests of rRNA hybridization to microarrays suggest that hybridization characteristics of oligonucleotide probes for species discrimination cannot be predicted. Nucleic Acids Res 34: e66–e66.
R Core Team. (2010) R: A Language and Environment for Statistical Computing.: R Foundation for Statistical Computing: Vienna, Austria.
Reinhardt T, Steinfartz S, Paetzold A, Weitere M . (2013). Linking the evolution of habitat choice to ecosystem functioning: direct and indirect effects of pond-reproducing fire salamanders on aquatic-terrestrial subsidies. Oecologia 173: 281–291.
Reinhardt T, Steinfartz S, Weitere M . (2015). Inter-annual weather variability can drive the outcome of predator prey match in ponds. Amphibia-Reptilia 36: 97–109.
Schluter D, Clifford Elizabeth A, Nemethy M, McKinnon Jeffrey S . (2004). Parallel evolution and inheritance of quantitative traits. Am Nat 163: 809–822.
Schoustra SS, Batallion T, Gifford DR, Kassen R . (2009). The properties of adaptive walks in evolving populations of fungus. PLoS Biol 7: e1000250.
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS et al. (2011). Metagenomic biomarker discovery and explanation. Genome Biol 12: 1–18.
Segev O, Hill N, Templeton AR, Blaustein L . (2010). Population size, structure and phenology of an endangered salamander at temporary and permanent breeding sites. J Nat Conserv 18: 189–195.
Segev O, Polevikove A, Blank L, Goedbloed D, Küpfer E, Gershberg A et al. (2015). Effects of tail clipping on larval performance and tail regeneration rates in the Near Eastern Fire Salamander, Salamandra infraimmaculata. PLoS One 10: e0128077.
Steiner CC, Römpler H, Boettger LM, Schöneberg T, Hoekstra HE . (2009). The genetic basis of phenotypic convergence in beach mice: similar pigment patterns but different genes. Mol Biol Evol 26: 35–45.
Steinfartz S, Veith M, Tautz D . (2000). Mitochondrial sequence analysis of Salamandra taxa suggests old splits of major lineages and postglacial recolonization of Central Europe from distinct source populations of S. salamandra. Mol Ecol 9: 397–410.
Steinfartz S, Weitere M, Tautz D . (2007). Tracing the first step to speciation: ecological and genetic differentiation of a salamander population in a small forest. Mol Ecol 16: 4550–4561.
The_UniProt_Consortium. (2015). UniProt: a hub for protein information. Nucleic Acids Res 43: D204–D212.
Thiesmeier B, Schuhmacher H . (1990). Causes of larval drift of the fire salamander, Salamandra salamandra terrestris, and its effects on population dynamics. Oecologia 82: 259–263.
Thomas PD, Wood V, Mungall CJ, Lewis SE, Blake JA,, Gene Ontology Consortium. (2012). On the use of gene ontology annotations to assess functional similarity among orthologs and paralogs: a short report. PLoS Comput Biol 8: e1002386.
Vences M, Sanchez E, Hauswaldt JS, Eikelmann D, Rodriguez A, Carranza S et al. (2014). Nuclear and mitochondrial multilocus phylogeny and survey of alkaloid content in true salamanders of the genus Salamandra (Salamandridae). Mol Phylogenet Evol 73: 208–216.
Warner JR . (1999). The economics of ribosome biosynthesis in yeast. Trends Biochem Sci 24: 437–440.
Weitere M, Tautz D, Neumann D, Steinfartz S . (2004). Adaptive divergence vs. environmental plasticity: tracing local genetic adaptation of metamorphosis traits in salamanders. Mol Ecol 13: 1665–1677.
West-Eberhard MJ . (2003) Developmental Plasticity and Evolution. Oxford University Press: Oxford, UK.
Whitehead A, Pilcher W, Champlin D, Nacci D . (2011). Common mechanism underlies repeated evolution of extreme pollution tolerance. Proc R Soc Lond B Biol Sci 279: 427–433.
Whitehead A, Zhang S, Roach JL, Galvez F . (2013). Common functional targets of adaptive micro- and macro-evolutionary divergence in killifish. Mol Ecol 22: 3780–3796.
Acknowledgements
We thank Timm Reinhardt for help in the field and Elke Blohm-Sievers for laboratory technical support. We are grateful to the Israel Nature and Parks Authority and local authorities in Germany for sampling permissions (INPA permits 2013/40174 and 2014/40672). This research was funded by a DFG DIP Grant (BL 1271/1-1 | STE 1130/8-1) awarded to LB, SS, ART and AWN, and by support from the Max Planck Society to AWN.
Author contributions
SS and DJG designed the study; DJG, TC, EK and OS collected the samples; TC, JA and AWN developed the microarrays; DJG performed primary microarray analysis; DJG, TC, AR, LB, ART, AWN and SS contributed to data analysis and writing.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Rights and permissions
About this article
Cite this article
Goedbloed, D., Czypionka, T., Altmüller, J. et al. Parallel habitat acclimatization is realized by the expression of different genes in two closely related salamander species (genus Salamandra). Heredity 119, 429–437 (2017). https://doi.org/10.1038/hdy.2017.55
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2017.55