Abstract
Pleistocene glaciations have profoundly affected patterns of genetic diversity within many species. Temperate alpine organisms likely experienced dramatic range shifts, given that much of their habitat was glaciated during this time. While the effects of glaciations are relatively well understood, the spatial locations of refugia and processes that gave rise to current patterns of diversity are less well known. We use a microsatellite data set to test hypotheses of population connectivity and refugial isolation in the web-toed salamanders (Hydromantes) of the Sierra Nevada. We reject models of refugia with subsequent expansion into either the high southern Sierra or low-elevation Owens Valley, in favor of a simple isolation model with no migration between current populations. We find no evidence of migration at even moderate spatial scales using a variety of analyses in the southern Sierra, and limited migration in the northern Sierra. These results suggest that divergence in isolation following fragmentation is the dominant process structuring genetic variation in these salamander species. In the context of anthropogenic climate change and habitat degradation, these results imply that salamanders and other low-vagility alpine organisms are at risk of decline as they are unlikely to migrate across large distances.
Similar content being viewed by others
Introduction
Major climatic changes have been shown to be a primary force structuring the genetic diversity within species (Hewitt, 2004). In particular, temperate zone species have been heavily influenced by Pleistocene glacial cycles, which markedly altered both the landscape and climate of temperate montane environments. These glacial cycles and resulting environmental changes have been a primary cause of divergence in many groups of species and have profoundly affected population demography and patterns of genetic variation within species (Hewitt, 2004), although demographic responses may differ both within and across groups of organisms (Burbrink et al., 2016). Temperate alpine organisms should be especially affected by such cycles because much of their current habitat was covered with ice during glacial periods. The presence of marked genetic structure within many alpine species suggests survival in ice-free glacial refugia, including adjacent lowland regions and nunataks within glaciated regions (Hewitt, 2004), but processes of refugial isolation and recolonization of alpine habitats remain poorly understood.
The Sierra Nevada mountains of California were extensively glaciated during the Pleistocene (Gillespie and Clark, 2011) and organisms there show evidence of a variety of histories, from very shallow genetic structure resulting from recent colonization (Schoville et al., 2011) to deep structure caused by persistence in multiple refugia (Schoville and Roderick, 2010). Distributional changes resulting from Pleistocene glacial cycles were likely involved in the divergence of many species, including several Sierra Nevada endemics (Rubidge et al., 2014). Much of the previous phylogeographic research in the Sierra Nevada focuses on organisms at lower elevations (for example, Kuchta et al., 2009), which may have responded to climatic changes differently than alpine species. The importance of glacial refugia on patterns of genetic diversity and species distributions in the Sierra Nevada has been shown in multiple studies (Eckert et al., 2008; Roberts and Hamann, 2015), yet precise locations of such refugia remain elusive. Furthermore, survival of populations in ice-free parts of the alpine zone (nunataks) has been shown for alpine taxa elsewhere (Westergaard et al., 2011). Such unglaciated regions occurred in the Sierra Nevada, including one relatively large area in the southern portion of the range (Gillespie and Clark, 2011). Phylogeographic studies that test models of gene flow across elevational zones of the Sierra Nevada would allow us to understand how glacial cycles have influenced the spatial distribution of genetic diversity and the recolonization process of different organisms.
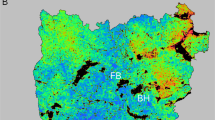
The web-toed salamanders of the Sierra Nevada (genus Hydromantes) provide an opportunity to test hypotheses of refugial isolation and persistence at high elevations. One species, H. platycephalus, is found throughout the high Sierra Nevada (primarily above 2000 m elevation) in areas of exposed granite with seeps or snowmelt as well as in scattered, riparian areas (1400–2000 m elevation) on the arid east side of the mountain range in the lower-elevation Owens Valley (Figure 1). This species has two major lineages in the northern and southern Sierra Nevada (HpN and HpSOV; Figure 1) that have been isolated for millions of years, as well as marked phylogeographic structure within each lineage (Rovito, 2010). The southern lineage (HpSOV) includes both high Sierra and Owens Valley populations. The second species, H. brunus, is confined to the Merced River basin on the western side of the Sierra Nevada (Figure 1) from ~200–900 m elevation in mossy talus habitat (Stebbins, 2003) and arose from within the northern lineage of H. platycephalus, probably as a result of glacial-mediated environmental divergence (Rovito, 2010). The two species differ in colour pattern and body proportions, habitat and season of surface activity (Stebbins, 2003). Although Rovito (2010) found H. platycephalus to comprise two distinct lineages, one of which is paraphyletic, he made no taxonomic changes because the inferred peripatric divergence of H. brunus would be expected to render H. platycephalus paraphyletic. Furthermore, recognizing the northern and southern lineages of H. platycephalus as distinct species would still leave the northern lineage paraphyletic with respect to H. brunus, and the two lineages of H. platycephalus are not distinguishable morphologically (Rovito, 2009).
Map of Sierra Nevada with sampling localities of Hydromantes platycephalus (white circles) and H. brunus (black circles). Circles are proportional to sample size. (a) Entire study area, covering most of the Sierra Nevada. Dashed line indicates a break between northern and southern lineages of H. platycephalus from Rovito (2010). Black rectangles 1b and 1c indicate spatial extent of (b, c), respectively. (b) Localities of H. brunus and nearby H. platycephalus locality at Bridalveil Falls, Yosemite National Park. (c) Owens Valley (PC1, PC2, ChC) and southern Sierra Nevada H. platycephalus localities. Locality codes correspond to those in Table 1.
Previous analyses suggested a high degree of isolation between most populations of H. platycephalus but relatively close relationships between several low-elevation Owens Valley populations and nearby high Sierra populations (Rovito, 2010). The presence of populations with divergent mtDNA and nuclear haplotypes in the Owens Valley, coupled with the fact that the climate in this region was cooler and wetter (and thus more suitable for salamander presence compared to the hotter, more arid climate at present) prior to the Last Glacial Maximum (Koehler and Anderson, 1995), suggested that this area might have served as a glacial refugium for the species (Rovito, 2010). Together with the close relationship of H. brunus to high-elevation H. platycephalus, these results suggested relatively recent connectivity between low and high-elevation populations on both sides of the Sierra Nevada. Currently, habitat of both species is patchy across the landscape and between high and low elevation.
Did Hydromantes persist in multiple refugia and colonize high elevations from unglaciated low-elevation areas, or do the lower-elevation populations instead represent more recently founded populations? To test which demographic scenario best explains the present distribution of genetic diversity in these salamanders, we use microsatellites developed for H. platycephalus. Microsatellite loci typically have high mutation rates, making them especially useful for examining patterns of genetic diversity within populations and across landscapes over timescales of thousands or tens of thousands of years (Rubidge et al., 2014). We use analyses of population structure and connectivity to determine at which scale gene flow exists among populations and to define units for further demographic analyses. We then use coalescent analyses to test hypotheses of refugial locations, population connectivity and demography in the southern Sierra Nevada and Owens Valley. If the Owens Valley was indeed a Pleistocene refugium from which the high southern Sierra Nevada was recolonized, we expect Owens Valley populations to show increased levels of genetic variation as well as connectivity with high-elevation populations. We also predict that a model in which populations at high elevation were recolonized from low elevation would be favored over models of a high-elevation refugium, connectivity between all populations or simple isolation without migration.
Materials and methods
Sampling, DNA extraction and genotyping
DNA for microsatellite genotyping was collected primarily using buccal swabs. This method provides high-quality DNA for genetic analysis without harming salamanders (Pidancier et al., 2003). During a given sampling event, salamanders were retained until sampling was finished in order to ensure that individuals were sampled only once and were released at the point of capture. A total of 265 salamanders from 21 localities (Table 1 and Figure 1) were sampled. These samples were grouped based on geography for further analyses into three groups: H. brunus (Hb), northern Sierra Nevada H. platycephalus (HpN) and southern Sierra Nevada and Owens Valley H. platycephalus (HpSOV). On the basis of the phylogenetic results of Rovito (2010), we combined Hb and HpN populations for all further analyses (hereafter HbHpN) because of paraphyly of the northern lineage of H. platycephalus with respect to H. brunus, and analyzed HpSOV populations separately. Each of these groups is a well-supported clade in phylogenetic analyses (Rovito, 2010), and the high degree of phylogenetic divergence between the two groups suggests that it would be inadvisable to analyze microsatellite genotypes from both groups together because of the high probability of homoplasy over longer time periods.
The sampling design included populations spaced across nearly the entire range of H. platycephalus, as well as nearly all known populations of H. brunus, in order to test for population structure and connectivity at a regional scale. For the southern Sierra Nevada samples, two pairs of populations (at the Pine Creek (PC) and Sixty Lake Basin (SL) sites) were sampled in order to test for population structure at a local scale; 1.0 and 2.0 km separate these sites, respectively. For H. brunus, sites separated by 2.0 km at Hell Hollow (HH) were sampled in order to test for gene flow and population structure over short geographic distances. Swabs were air-dried in the field and stored at −80 °C in the lab prior to extraction. A guanidine–thiocyanate protocol (available upon request) was used to extract genomic DNA from buccal swabs.
A microsatellite library was developed by Genetic Identification Services (Chatsworth, CA, USA). From this library, 10 loci were selected based on variability across a panel of eight individuals (six H. platycephalus and two H. brunus) representing the geographic range of both species. Primer sequences and PCR conditions for these loci are given in Supplementary Table 1. The universal labeling method of Schuelke (2000) was used, with either M13 or T7 vector sequence included in forward primers. Two different fluorescent dyes were used: 6-carboxy-fluorescine with the M13-tagged primers and hexachloro-6-carboxy-fluorescine with the T7-tagged primers. PCR reactions included 0.6 μl of 10 mM reverse primer, 0.06 μl of forward primer with an M13 or T7 tail and 0.54 μl of the universal fluorescent-labeled primer. PCR consisted of an initial denaturing step at 95 °C for 30 s, followed by 24 cycles of 95 °C for 30 s, 30 s at the locus-specific annealing temperature and 72 °C for 45 s. Ten cycles of 95 °C for 30 s, 53 °C for 30 s and 72 °C for 45 s were then performed to incorporate the fluorescent dye into the PCR product, with a final annealing step of 72 °C for 12 min. Sigma Jumpstart Accutaq (Sigma-Aldrich, St Louis, MO, USA) was used for all reactions.
Genotyping was done on an ABI 3730 capillary sequencer (Applied Biosystems, Foster City, CA, USA). To reduce genotyping costs, PCR products for loci labeled with different coloured dyes and/or separated by allele size range were combined for genotyping. Genotypes were scored using Genemapper 3.7 (Applied Biosystems). Individuals with low-quality traces or ambiguous genotypes were re-run until they could be genotyped confidently. Individuals missing genotypes for three or more loci were excluded in analyses, leaving 135 and 123 individuals from the northern and southern Sierra Nevada, respectively.
The program MICRO-CHECKER (van Oosterhout et al., 2004) was used to test for the presence of null alleles. For the southern Sierra Nevada data set, four tests (four alleles from a single population each) out of 90 showed evidence of null alleles. For H. platycephalus from the northern Sierra Nevada, only one locus for a single population showed evidence of null alleles. For H. brunus, however, 15 of 50 null allele tests were significant. Because of the possibility of null alleles within this data set, approximate Bayesian computation (ABC) analyses (see below) were not done on the HbHpN lineage. We tested for deviations from Hardy–Weinberg equilibrium for each locus in each population using the exact test implemented in Arlequin 3.5 (Excoffier and Lischer, 2010). Significance values were adjusted for multiple comparisons using a Bonferroni correction. We also calculated allelic richness (Na), expected heterozygosity (HE) and genetic diversity (4NEμ) estimated from expected heterozygosity θH.
Population structure analyses
We used the program STRUCTURE v.2.3.2 (Pritchard et al., 2000), which uses a Bayesian clustering algorithm to group individuals into a predefined number of populations (K) by minimizing deviations from linkage- and Hardy–Weinberg equilibrium within populations. We applied the method of Evanno et al. (2005) using the second derivative of the likelihood function (ΔK) as the selection criterion to estimate K. STRUCTURE was run 20 times for K, from 1 to the maximum number of sampling localities in each data set (northern data set Kmax=12; southern data set Kmax=9), with a burn-in of 10 000 steps and a run length of 100 000 steps. Structure Harvester v. 0.6 (Earl and vonHoldt, 2012) was used to calculate ΔK. Following the selection of K values, STRUCTURE was run for 1 × 107 generations with 1 × 106 generations of burn-in. Ten runs per data set for each value of K were performed, and the results were summarized using CLUMPP v1.1.2 (Jakobsson and Rosenberg, 2007) and visualized with DISTRUCT v1.1 (Rosenberg, 2004). In addition to the K-value selected by the ΔK method, we also ran analyses with a value of K=2 for the HpSOV data set (to test for structure between high and low-elevation sites) and K=3 and K=6 for the HbHpN data set based on a precipitous drop of ΔK between K=3 and K=4 and a plateau in the likelihood at K=6 (Supplementary Figure S1).
We used a graph theoretic approach implemented in Population Graphs (Dyer and Nason, 2004) to examine patterns of population connectivity within both data sets using the R package ‘gstudio’ (Dyer, 2012) in R v3.1.2 (R Development Core Team, 2015). This approach does not assume any a priori hierarchical population structure and works by eliminating links between populations in a model with connections between all populations to find the minimum model that sufficiently describes the genetic covariance among populations. The resulting population graph depicts both genetic variance within populations and patterns of population connectivity. We also performed Principal Coordinates Analysis using a covariance matrix of standardized distances calculated from each data set using the program GENALEX v.6.5 (Peakall and Smouse, 2012) to visualize genetic variation in multivariate space and look for clustering of individuals by locality, area or species.
We estimated the degree of population isolation using FST and tested for a pattern of isolation by distance (IBD). We calculated FST between all sampling sites for H. brunus and H. platycephalus from the northern Sierra Nevada using Genepop v4.1 (Rousset, 2008). We did not use the population grouping from STRUCTURE analysis for the northern Sierra Nevada because many sampling sites were grouped into only a few populations (see Results). Our intention was to examine patterns of IBD across the landscape, and using only a few populations from STRUCTURE would reduce our power to detect a pattern of IBD if it existed. For the H. platycephalus sites in the southern Sierra Nevada and Owens Valley, we used the population grouping from the results of STRUCTURE with K=6 (see Results), which grouped several pairs of geographically proximate populations together. We calculated genetic and geographic distances between populations in Genepop using FST/(1−FST) as a measure of genetic distance as well as a corrected measure (FST/(1−FST)*) that may be more appropriate when sample sizes are small. We used the Isolation by Distance Web Service v. 3.23 (Jensen et al., 2005) to perform a reduced major axis (RMA) regression of FST/(1−FST) against the log of geographic distance and tested the significance of the correlation using a Mantel test with 10 000 permutations.
ABC analyses and bottleneck tests
To infer the demographic history of H. platycephalus populations, we compared the fit of several alternative evolutionary models using ABC. In this approach, genetic data are simulated under each model by drawing values for the parameters that define each model (for example, population size, migration rate, and so on) at random from prior distributions. The observed data are then compared to the synthetic data and, based on similarity to the observed data, the best-fitting model and associated parameter values are estimated. Posterior distributions rather than explicit likelihoods are used to estimate each model and its parameters (Csilléry et al., 2010). We first proposed four models of population history (Figure 2) that differ in their assumptions about gene flow: (1) isolation without migration, (2) recent colonization of the high Sierra from the Owens Valley, (3) recent colonization of the Owens Valley from the high Sierra and (4) migration among geographic neighbors. For model 1, we grouped sampling sites by population according to the STRUCTURE results with K=6 and modeled a divergence history with major genetic breaks between lower-elevation Owens Valley populations and high Sierra populations occurring first and subsequent breaks occurring within these two groups by geography (Figure 2a). Within each group, we modeled the initial split as occurring between the most geographically distant population and the remaining populations, and repeated this to obtain a fully resolved tree. This divergence history was chosen to test alternate hypotheses of refugia in either the Owens Valley or high Sierra (Models 2–4). We also explored an alternative divergence history (Model 5; Supplementary Figure S2) reflecting the relationships between populations implied by the STRUCTURE results from K=2 and population graph results from the southern Sierra with a simultaneous initial split between groups of populations. The use of this alternative topology allowed us to account for uncertainty in the relationships between populations in models 1–4. After evaluating the effect of gene flow in the four main models, we conducted a second round of model selection focused on population size change. We iteratively contrasted the best of our four initial models to models with a population bottleneck in each population. By focusing on a single population bottleneck at a time, we minimized the overall number of free parameters, providing a stronger test of population size change.
Schematic of models used to test hypotheses of population connectivity and colonization from refugia in ABC analyses of the southern lineage of H. platycephalus. (a) Model 1: isolation without migration model, with initial break between low-elevation Owens Valley sites (PC1, PC2 and ChC) and high-elevation southern Sierra Nevada sites; (b) Model 2: Owens Valley refuge model, with colonization of high Sierra Nevada from Owens Valley; (c) Model 3: high Sierra refuge model, with recent colonization of Owens Valley from high Sierra Nevada; (d) model with gene flow between all neighboring populations. Grey circles indicate lowland Owens Valley populations; black circles indicate highland populations. Locality codes from Figure 1 and Table 1. Rectangle and ellipse around populations indicate potential refugia in high Sierra Nevada and Owens Valley, respectively.
Coalescent simulations were carried out using ABCtoolbox (Wegmann et al., 2010) and SIMCOAL v2.1.2 (Laval and Excoffier, 2004), with 1 000 000 synthetic data sets generated under each model. Uniform priors were set for population sizes, event times and mutation rates while truncated normal priors were set for each migration rate (prior values shown in Supplementary Table 2). Estimates of microsatellite mutation rate are available for Ambystoma tigrinum (Bulut et al., 2009), with a mean of 1.27 × 10−3 per generation (95% confidence interval 6.07 × 10−4–2.33 × 10−3) and other vertebrates with the mean rates ranging from 8 × 10−5 to 1.25 × 10−2 (Peery et al., 2012). Weighing the A. tigrinum estimate more heavily, we placed a uniform prior on the mean rate across 10 microsatellite loci (1 × 10−3 to 9 × 10−3), but allowed each simulated locus to vary from this mean according to a normal distribution truncated at zero (with s.d. 0.0005). In addition, we implemented a geometric stepwise mutation model with geometric parameter set to 0.5 to allow for the insertion or deletion of multiple repeats.
Summary statistics were calculated in Arlequin v3.5 (Excoffier and Lischer, 2010) for the simulated and observed microsatellite data sets. The mean and s.d. of the following statistics were calculated in each population and across all populations: the number of alleles (Na), the expected heterozygosity (H), the Garz–Williamson index the modified Garza–Williamson index, and the range of alleles (R). In addition, we calculated pairwise FST, the mean number of differences between populations (PI) and the genetic distance between populations based on the stepwise mutation model (DMUSQ). We implemented a rejection sampling approach coupled with feed-forward neural networks (nonlinear regression) to reduce the summary statistics into a smaller number of dimensions (Blum and François, 2010). We explored several tolerance levels for the rejection step using cross-validation (0.005, 0.01, 0.05 and 0.1) and chose to accept 1% of the simulations, as the tolerance level seemed to have little effect on the results. Competing models were compared based on their posterior probability (PP) and Bayes Factor scores (BF>3:1 were treated as substantial support for a more parameter-rich model). We used cross-validation to ensure that the models 1–4 could be correctly classified based on our model selection procedure. Finally, we estimated parameters in the best-fitting model using feed-forward neural networks and used cross-validation to assess the accuracy of our parameter estimates at several tolerance values (0.01, 0.05 and 0.1). These analyses were conducted in R using the ‘abc’ package (Csilléry et al., 2010).
To further assess the demographic history of Hydromantes populations, we examined demographic change within individual populations. The full likelihood method of Beaumont (1999) can be used to estimate a linear or exponential change in population size in a single population, assuming a stepwise mutation model. We used the software MSVAR 1.3, which implements a hierarchical Bayesian model to allow for variability among loci (Storz and Beaumont, 2002). We focused on those populations identified as clusters by STRUCTURE with large sample size (CC (Convict Creek)+TL (Tully Lake), SL1+SL2 and PC1+PC2), but because MSVAR is sensitive to underlying population structure (Chikhi et al., 2010), we also examined the CC population by itself. We chose the linear growth model and the following hyperpriors (log-scale) for the model parameters: means for ancestral (αN0) and current effective population (αN1) set to 3, mean mutation rate (αμ) set to −4.75, divergence time (αxa) set to 4, variance of the mean σN0=σN1=4, σμ=2, σxa=3, mean of the variance βN0=βN1=βμ=βxa=0 and variance of the variance τN0=τN1=τμ=τxa=0.5. For each data set, two separate MCMC chains were run for 25 × 109 steps with samples recorded every 1 000 000 steps. We assessed convergence using the Gelman–Rubin multivariate scale reduction factor (PRSF) and the effective sample size of each parameter across the independent chains, using the ‘CODA’ package in R (Plummer et al., 2006). For each parameter, the mode and 95% highest posterior density (HPD) intervals were estimated from the marginal posterior distribution using the ‘locfit’ package in R (Loader, 2013).
Results
Most populations of H. platycephalus in the northern Sierra Nevada have reduced genetic diversity (as measured by θH) compared to populations of H. brunus (Table 1). The Sonora Pass population in particular has low values of θH (0.35), allelic richness (2.5±1.08 (mean±s.d.)) and expected heterozygosity (0.26±0.26). The Owens Valley populations of the southern lineage of H. platycephalus have reduced genetic diversity (θH, HE) compared to those in the adjacent high southern Sierra Nevada. Unlike in many temperate zone species (Hewitt, 2004), there does not appear to be a clear-cut north–south pattern in genetic diversity in our results. Despite the generally lower genetic diversity of the northern lineage of H. platycephalus, the most northern population (SmL) has the highest value of θH and high allelic richness and southern Sierra populations at high elevation have genetic diversity comparable to that seen in H. brunus (Table 1). FST values between sites were generally lower within H. brunus than within northern H. platycephalus, or between the two species (Table 2).
Population structure analyses
For the HbHpN data set, the highest value of ΔK from STRUCTURE runs varying the value of K was for K=2 (ΔK=4.86) and the L(K) reached a plateau at K=6 (mean lnL=−4038.92), increasing little from K=6 to K=9 (Supplementary Figure S1). The ΔK for K=3 (4.44) was similar to that for K=2 but dropped precipitously at K=4 (ΔK=0.41). Although the STRUCTURE results for the northern Sierra Nevada with K=2 might be expected to show a clear division between the H. brunus and H. platycephalus, they instead show samples of H. platycephalus from the Merced drainage in Yosemite National Park (Bridalveil Falls and Half Dome) assigned with high PP to the same population as H. brunus (Figure 3a). Individuals of H. platycephalus from two high Sierra populations (EmL and SP) were classified with high probability (0.90) to a second population. Individuals from two other high Sierra populations of H. platycephalus were classified as admixed between these two populations. At K=6, some additional population structures are evident, although the only clearly demarcated populations are those from the northernmost part of the range of H. platycephalus. At K=3, all H. brunus individuals and H. platycephalus from Half Dome were grouped into a single population.
STRUCTURE plots of population assignment for northern Sierra Nevada/H. brunus and southern Sierra Nevada/Owens Valley data sets. Shades of grey correspond to population assignments for different values of K. H. brunus populations (a) and H. platycephalus populations from the Owens Valley (b) are surrounded by a black box.
For HpSOV, ΔK had a single peak at K=6 (ΔK=86.79). The STRUCTURE analysis with K=6 assigned samples from CC and nearby Tully Lake to a single cluster (Figure 3b). Although CC is east of the Sierra Nevada crest, it is at substantially higher elevation than eastern Sierra sites and is located north of the Owens Valley; for these reasons, we treat it (together with Tully Lake) as a high Sierra population in all subsequent analyses. Paired, nearby sites at PC were placed in a cluster, as were those from Sixty Lake Basin. The other three sites were each classified as separate clusters. All individuals except one were assigned with high PP (>0.95) to a single cluster; the lone exception (from the Silliman Lakes locality) was assigned to its population of origin with PP=0.93. No evidence of admixture between populations is apparent. There was no division between Owens Valley and high Sierra Nevada samples at K=2 (Figure 3b). FST estimates were generally higher between Owens Valley and high Sierra populations than between populations in the high Sierra (Table 3).
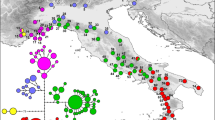
The population graph for the northern Sierra Nevada reveals two separate graphs with limited connectivity between populations (Figure 4). Four of the six H. brunus localities form one linear graph. The H. brunus localities from HH1 and HH2 are connected to localities of H. platycephalus in a second graph through the Half Dome site. Localities in the Yosemite area show the most connectivity to those from other parts of the range. H. brunus localities tend to have higher genetic variance than those from the high Sierra Nevada (Figure 4), in agreement with calculated values of genetic diversity (θH, Table 1). The population graph for the southern Sierra Nevada is even simpler, with three separate graphs and no locality with more than two connections. Each Owens Valley population (ChC and PC) is in a separate graph and the geographically proximate populations ChC and SL are in different graphs, in agreement with the STRUCTURE results.
Population graphs of (a) northern Sierra Nevada, and (b) southern Sierra Nevada data sets. Size of circle is proportional to genetic variance of population. Nodes connected by lines are part of the same graph. H. brunus localities are shown in black and H. platycephalus localities are shown in white. Some localities have been slightly offset for clarity.
The principal coordinate analysis results for both the HbHpN and HpSOV data sets largely mirrored those from STRUCTURE and Population Graphs analyses. The HbHpN results showed all individuals of H. brunus together in a single cluster, along with individuals of H. platycephalus from locality Half Dome (Supplementary Figure S3). Individuals of H. platycephalus from Bridalveil Falls, the geographically nearest locality to the distribution of H. brunus, are also closest to the H. brunus cluster. For the southern Sierra Nevada, individuals from Owens Valley populations separate from high Sierra populations and from each other in principal coordinates space. Individuals from localities FL and Silliman Lake are clustered together, as are individuals from TL and CC.
For the northern Sierra Nevada, the IBD analysis rejected the null model of no relationship between genetic distance and log geographic distance, both when using FST/(1−FST) (Mantel test Z=221.8, P=0.005) and when using the corrected FST/(1−FST)* (Mantel test Z=134.8, P<0.0001). reduced major axis regression estimated a positive slope for the relationship between genetic and geographic distance and indicated that geographic distance explains much of the variance in genetic distance (regular FST: slope=0.99, R2=0.34; corrected FST: slope=0.83, R2=0.56). For the southern Sierra, neither measure of genetic distance showed a significant relationship with geographic distance (regular FST: Mantel test Z=65.7, P=0.21, RMA slope=1.24, R2=0.05; corrected FST: Mantel test Z=37.0, P=0.37, RMA slope=0.69, R2=0.015).
ABC analyses and bottleneck tests
ABC model choice provides evidence of isolation without migration in the southern Sierra Nevada. Cross-validation of the model selection procedure indicates that ABC can assign data sets to each model accurately (model 1: 99.9%, model 2: 98%, model 3: 93%, model 4: 99%). On the basis of posterior model probabilities, the isolation without migration model is strongly favored (model 1: 0.9993) in comparison to the high Sierra colonization model (model 2: 0.0001), the Owens Valley colonization model (model 3: 0.0006) or the model of migration among geographic neighbors (model 4: 0.0000). Similarly, BF scores strongly favor the isolation without migration model (1 versus 2 BF=19073 1 versus 3 BF=1804, 1 versus 4=2765705). The primary model of isolation with no migration (model 1) was favored over the alternative model of population splitting (Supplementary Figure S1), and the use of the alternative model had little effect on parameter estimates (results not shown). The median parameter estimates based on the isolation without migration model (Table 4) suggest that populations in the southern high Sierra (SL, Silliman Lake and FL) diverged between 1.7 and 3.5 ka (τ1 and τ2 events), that populations in the Owens Valley (ChC and PC) also diverged during this period (2.7 ka, τ3 event), whereas the τ4 divergence of the CC/TL lineage was quite old (17.3 ka). The median estimate for the divergence time τ5 of all populations was 35.8 ka. Estimates for current effective size (Ne) range from 649 to 1098 among populations, while estimates for ancestral effective size were much larger (>28 × to >52 ×). Confidence intervals around ancestral effective size estimates are large, however, and cross-validation indicates a high prediction error associated with these estimates (Supplementary Table 3). The relaxation of tolerance levels (from 0.01 to 0.1) to include more simulated data points does not improve the prediction errors, suggesting that the data contain insufficient information to estimate ancestral population size. Including a bottleneck improves the fit of the observed data to the isolation without migration model for populations from CC/TL (PP of bottleneck model=0.725, BF=3.8), SL (PP of bottleneck model=0.807, BF=4.2) and FL (PP of bottleneck model=0.819, BF=4.5). There was no evidence of a bottleneck in either of the Owens Valley populations. The median parameter estimates suggest that the bottleneck times varied among populations (Table 5), although within a narrow time interval of 1.0–2.4 ka.
The demographic analysis of population samples using MSVAR indicates weak population expansion from very small ancestral population sizes for the CC/TL and SL clusters, but no change in the PC cluster. In all cases, MCMC traces and diagnostics suggested convergence for both likelihood runs, good mixing (PSRF<1.1) and parameter estimates drawn from a large sample size (effective sample size>290). For the CC/TL cluster, estimates of the current population size (mode=16, 95% HPD: 0.02–14 443), ancestral population size (mode=1.72, 95% HPD: 1–3.17) and time of population size change (mode=1809 years, 95% HPD: 2.22–1 490 579) suggest an expansion following a bottleneck in the later Holocene (with wide uncertainty around the timing estimate). Analysis of CC alone suggested a slightly larger current effective population size (mode=30.51, 95% HPD: 0.04–21 979.91), ancestral population size (mode=1.47, 95% HPD: 1–2.77) and an earlier population size change event (mode=3620 years, 95% HPD: 4.66–2 464 325). For the SL cluster, estimates of the current population size (mode=32.33, 95% HPD: 0.04–26 018), ancestral population size (mode=1.11, 95% HPD: 1–2.03) and time of population size change (mode=3459 years, 95% HPD: 4.26–2 594 328) again suggest a Holocene bottleneck. For the PC cluster, estimates of the current population size (mode=3.93, 95% HPD: 0.002–6262,6), ancestral population size (mode=2.26, 95% HPD: 1–6.31) and time of population size change (mode=1222 years, 95% HPD: 0.59 −1 620 242) show no evidence of a bottleneck.
Discussion
Population responses to glaciation
We set out to test whether Hydromantes persisted in multiple refugia and colonized high-elevation sites from low-elevation sites in the Sierra Nevada. Multiple analyses converge in estimating very limited connectivity between populations of H. platycephalus even over relatively short geographic distances, especially in the southern Sierra Nevada. STRUCTURE results placed all southern Sierra localities that were separated by more than a few kilometers into different populations with no evidence of migration or admixture between populations. ABC analyses strongly favored a model that did not include migration between populations. The population graph for the southern Sierra Nevada also indicated limited population connectivity. In the northern Sierra Nevada, slightly more population connectivity was apparent, indicated by more connections between nodes of the population graph and STRUCTURE results that placed several sites within a single population.
The results of IBD analyses show that much of variance in genetic distance between localities of H. platycephalus and H. brunus in the northern Sierra Nevada is explained by geographic distance. A pattern of IBD, in which gene flow between populations decreases with spatial distance across the landscape, is in agreement with the signature of limited admixture seen in the STRUCTURE results. The population graph results also showed that there has been at least some gene flow between populations over time, especially between adjacent populations. Given a limited dispersal ability, based on a capture–recapture dispersal estimate from a related species (Herman, 2003), limited dispersal from population genetic analyses of microsatellite data within continuous habitat (Cabe et al., 2007) and across expanses of unsuitable habitat (Velo-Antón et al., 2013) of other terrestrial salamander species, and the apparently patchy distribution of its mossy talus habitat, it seems unlikely that the substantial gene flow is currently occurring among populations of H. brunus or among H. brunus and H. platycephalus. Past glaciations, however, likely pushed some populations of H. platycephalus in the northern Sierra Nevada to lower elevations, putting them in close proximity to H. brunus. Such distributional changes may have increased the spatial proximity of populations and environmental suitability, allowing for more gene flow across the landscape.
The results of ABC analyses, large FST values between populations, rejection of the IBD model and a lack of substantial connectivity in population graphs between sites in the southern Sierra Nevada support a scenario of very low (or no) migration between populations over moderate to large spatial scales. Our study design does indicate population connectivity over small spatial scales, however. In all three instances (HH, SL and PC), where two sites were sampled within 1–2 km, the two sites were grouped into a single population by STRUCTURE and connected in the population graph. At two of these sites (HH and SL), habitat appears fairly continuous, while at PC, salamanders are apparently restricted to a narrow area of riparian vegetation along each creek, and the two sampling sites are separated by rock and arid vegetation. The PC sites show that salamanders may be able to traverse short stretches of seemingly inhospitable habitat, and that dispersal occurs with enough frequency to maintain population connectivity over short scales.
Differences in glacial history between the northern and southern Sierra Nevada may partially account for differing levels of population structure and connectivity seen in the two lineages of Hydromantes. In the Sierra Nevada from the Yosemite region north, only the highest peaks of the range existed as ice-free nunataks at the LGM, with a highland ice field extending ~128 km along the Sierra Nevada with an average width of 64 km. Further south in the Sierra Nevada, higher ridges may have been ice-free and there may not have been an ice field extending across the high divides between river basins in the region of present-day Sequoia-Kings Canyon National Parks (Guyton, 1998). Maps of glacial ice extent at the LGM (Gillespie and Clark, 2011) show a relatively large unglaciated region in the high southern Sierra Nevada southwest of PC, and also show that sites FL and Silliman Lake were on the edge of glacial ice. Thus, populations in the high southern Sierra Nevada could have persisted either in high-elevation ice-free refugia or by moving short distances to track movement of ice during cold periods. Either persistence in local ice-free refugia or movement downslope over short distances (or both) would explain the lack of a signature of recolonization from a large glacial refugium seen in the ABC results and limited connectivity between populations in the STRUCTURE and population graphs results. By contrast, more extensive glacial ice covering current H. platycephalus populations in the northern Sierra Nevada may have led to recolonization and more extensive gene flow following deglaciation, accounting for the increased signature of gene flow and admixture in this lineage.
Rovito (2010) showed that some populations in the Owens Valley south of those sampled for this study had unique, divergent haplotypes for both mtDNA and some nuclear loci. The presence of these divergent haplotypes suggested that the Owens Valley may have been a refugium from which the high southern Sierra Nevada was recolonized following deglaciation. Those populations, from the southernmost portion of the Owens Valley, had too few tissue samples available and were not included in this study. At present, we can state that the northern portion of the Owens Valley was apparently not a glacial refugium for high-elevation populations of H. platycephalus. We cannot, however, rule out the possibility that unsampled populations in the southern Owens Valley may be within an area that served as a glacial refugium; additional sampling of these populations would be necessary to test this hypothesis.
The timing of more recent splits both within the high southern Sierra Nevada and within the Owens Valley appears to correlate well with a period of neoglacial cooling following the hot and dry conditions of the early to mid-Holocene (Koehler and Anderson, 1995). It may be that salamanders were able to colonize new areas under the cooler and wetter conditions of this period; our isolation without migration model does not distinguish between vicariance of previously continuous populations and the establishment of new populations via dispersal. If warm, dry conditions caused vicariance of salamander populations, we would expect the population divergence times to fall earlier in the Holocene. The confidence intervals on divergence time between highland and lower-elevation populations or on the divergence of the CC/TL population from others in the high southern Sierra (Table 2) are too wide to permit interpretation of the possible effect of climate on population divergence. The relatively narrow time interval (1400 years) within which bottlenecks occurred in three populations suggests that climatic changes could have been responsible for reductions in population size, since multiple populations were affected.
Although we attempted to provide conservative estimates of microsatellite mutation rates, a misspecification of these rates could confound our ABC parameter estimates, including estimates of population size and divergence time. Thus, all time estimates should be interpreted with some caution. In particular, the mutation rate estimate from Bulut et al. (2009) included monomorphic loci and was calculated over only a few generations. If the mutation rate in Hydromantes is actually faster, our estimates of divergence time would be too old and our population size estimates would be too high. However, most vertebrate microsatellite mutation rates are slower; thus, our expectation is that population divergence is well placed within the context of Holocene environmental change. The relative changes in population size we observe, however, should be unaffected by different mutation rates and therefore our detection of bottlenecks should be fairly robust.
Conservation implications
Both H. platycephalus and H. brunus are species of conservation concern in California (Jennings and Hayes, 1994) and our results are relevant to management of these species. The only connectivity between populations within the southern Sierra Nevada is over very short spatial scales (⩽2 km), suggesting that populations separated by more than a few kilometers can be considered as effectively isolated from one another and treated as separate units for management purposes. While less population structure is evident within H. brunus, this is likely due to more recent population fragmentation than to ongoing gene flow because of the lack of intervening suitable habitat. In general, populations do not appear to have reduced genetic diversity (measured by NE, θH, allelic richness or HE), although genetic diversity is somewhat lower in the Owens Valley and northern high Sierra Nevada. Populations of H. brunus, which is of greater concern due to its small range and limited number of known localities, have relatively high genetic diversity and perhaps more historical connectivity than those in either the high Sierra or the Owens Valley. Such isolation could affect population-level responses to anthropogenic climate change that depend on either adaptation to new environmental conditions or movement across the landscape.
Data archiving
Microsatellite genotypes for all individuals in Genepop and Structure formats, and input files for ABC and MSVAR analyses, are available from the Dryad Digital Repository: http://dx.doi.org/10.5061/dryad.c197n.
References
Beaumont MA . (1999). Detecting population expansion and decline using microsatellites. Genetics 153: 2013–2029.
Blum MGB, François O . (2010). Non-linear regression models for approximate Bayesian computation. Stat Comput 20: 63–73.
Bulut Z, McCormick CR, Gopurenko D, Williams RN, Bos DH, DeWoody JA . (2009). Microsatellite mutation rates in the eastern tiger salamander (Ambystoma tigrinum tigrinum differ 10-fold across loci. Genetica 136: 501–504.
Burbrink FT, Chan YL, Myers EA, Ruane S, Tilston Smith B, Hickerson MJ . (2016). Asynchronous demographic responses to Pleistocene climate change in Eastern Nearctic vertebrates. Ecol Lett 19: 1457–1467.
Cabe PR, Page RB, Hanlon TJ, Aldrich ME, Conners L, Marsh DM . (2007). Fine-scale population differentiation and gene flow in a terrestrial salamander (Plethodon cinereus living in continuous habitat. Heredity 98: 53–60.
Chikhi L, Sousa VC, Luisi P, Goossens B, Beaumont MA . (2010). The confounding effects of population structure, genetic diversity and the sampling scheme on the detection and quantification of population size changes. Genetics 186: 983–995.
Csilléry K, Blum MGB, Gaggiotti OE, Francois O . (2010). Approximate Bayesian Computation (ABC) in practice. Trends Ecol Evol 25: 410–418.
Dyer RJ . (2012) Gstudio: a Package for the Spatial Analysis of Population Genetic Marker Data. Virginia Commonwealth University: Richmond, VA, USA.
Dyer RJ, Nason JD . (2004). Population graphs: the graph theoretic shape of genetic structure. Mol Ecol 13: 1713–1727.
Earl DA, vonHoldt BM . (2012). STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Cons Gen Res 4: 359–361.
Eckert AJ, Tearse BR, Hall BD . (2008). A phylogeographical analysis of the range disjunction for foxtail pine (Pinus balfouriana, Pinaceae): the role of Pleistocene glaciation. Mol Ecol 17: 1983–1997.
Evanno G, Regnaut S, Goudet J . (2005). Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14: 2611–2620.
Excoffier L, Lischer HEL . (2010). Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linus and Windows. Mol Ecol Res 10: 564–567.
Gillespie AR, Clark DH . (2011). Glaciations of the Sierra Nevada, California, USA. Devel Quaternary Sci 15: 447–462.
Guyton B . (1998) Glaciers of California. University of California Press: Berkeley, CA, USA.
Herman AE . (2003). Aspects of the Ecology of the Shasta Salamander, Hydromantes shastae, near Samwell Cave, Shasta County, California. MSc thesis, Humboldt State University, Humboldt, CA, USA..
Hewitt GM . (2004). Genetic consequences of climatic oscillations in the Quaternary. Philos Trans R Soc B 359: 183–195.
Jakobsson M, Rosenberg NA . (2007). CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23: 1801–1806.
Jennings MR, Hayes MP . (1994) Amphibian and Reptile Species of Special Concern in California. California Department of Fish and Game, Inland Fisheries Division: Rancho Cordova, CA, USA.
Jensen J, Bohonak A, Kelley S . (2005). Isolation by distance, web service. BMC Genet 6: 13.
Koehler PA, Anderson RS . (1995). 30,000 years of vegetation changes in the Alabama Hills, Owens Valley, California. Quaternary Res 43: 238–248.
Kuchta SR, Parks DS, Mueller RM, Wake DB . (2009). Closing the ring: historical biogeography of the salamander ring species Ensatina eschscholtzii. J Biogeogr 36: 985–995.
Laval G, Excoffier L . (2004). SIMCOAL 2.0: a program to simulate genomic diversity over large recombining regions in a subdivided population with a complex history. Bioinformatics 20: 2485–2487.
Loader K . (2013) locfit: Local Regression, Likelihood and Density Estimation. Version 1.5: Merck, Kenilworth, NJ, USA.
Peakall R, Smouse PE . (2012). GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28: 2537–2539.
Peery MZ, Kirby R, Reid BN et al. (2012). Reliability of genetic bottleneck tests for detecting recent population declines. Mol Ecol 21: 3403–3418.
Pidancier N, Miquel C, Miaud C . (2003). Buccal swabs as a non-destructive tissue sampling method for DNA analysis in amphibians. Herpetol J 13: 175–178.
Plummer M, Best N, Cowles K, Vines K . (2006). CODA: convergence diagnosis and output analysis for MCMC. R News 6: 7–11.
Pritchard JK, Stephens M, Donnelly P . (2000). Inference of population structure using multilocus genotype data. Genetics 155: 945–959.
R Development Core Team. (2015) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria.
Roberts DR, Hamann A . (2015). Glacial refugia and modern genetic diversity of 22 western North American tree species. Proc R Soc Lond B Biol 282: 20142903.
Rosenberg NA . (2004). DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4: 137–138.
Rousset F . (2008). genepop'007: a complete re-implementation of the genepop software for Windows and Linux. Mol Ecol Res 8: 103–106.
Rovito SM . (2009) Lineage divergence and species formation in plethodontid salamanders. PhD Thesis, University of California: Berkeley, CA, USA.
Rovito SM . (2010). Lineage divergence and speciation in the Web-toed Salamanders (Plethodontidae: Hydromantes of the Sierra Nevada, California. Mol Ecol 19: 4554–4571.
Rubidge EM, Patton JL, Moritz C . (2014). Diversification of the Alpine Chipmunk, Tamias alpinus, an alpine endemic of the Sierra Nevada, California. BMC Evol Biol 14: 34.
Schoville SD, Roderick GK . (2010). Evolutionary diversification of cryophilic Grylloblatta species (Grylloblattodea: Grylloblattidae) in alpine habitats of California. BMC Evol Biol 10: 163.
Schoville SD, Stuckey M, Roderick GK . (2011). Pleistocene origin and population history of a neoendemic alpine butterfly. Mol Ecol 20: 1233–1247.
Schuelke M . (2000). An economic method for the fluorescent labeling of PCR fragments. Nat Biotechnol 18: 233–234.
Stebbins RC . (2003) Western Reptiles and Amphibians. 3rd edn. Houghton Mifflin Company: New York, NY, USA.
Storz JF, Beaumont MA . (2002). Testing for genetic evidence of population expansion and contraction: an empirical analysis of microsatellite DNA variation using a hierarchical Bayesian model. Evolution 56: 154–166.
van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P . (2004). MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4: 535–538.
Velo-Antón G, Parra JL, Parra-Olea G, Zamudio KR . (2013). Tracking climate change in a dispersal-limited species: reduced spatial and genetic connectivity in a montane salamander. Mol Ecol 22: 3261–3278.
Wegmann D, Leuenberger C, Neuenschwander S, Excoffier L . (2010). ABCtoolbox: a versatile toolkit for approximate Bayesian computations. BMC Bioinformatics 11: 116–116.
Westergaard KB, Alsos IG, Popp M, Engelskjøn T, Flatberg KI, Brochmann C . (2011). Glacial survival may matter after all: nunatak signatures in the rare European populations of two west-arctic species. Mol Ecol 20: 376–393.
Acknowledgements
SMR was funded by a Graduate Research Fellowship from the National Science Foundation (NSF), a Berkeley Fellowship and a postdoctoral fellowship from the NSF (DEB 1026396). SDS was supported by a postdoctoral fellowship from NSF (OISE-0965038). Fieldwork and microsatellite genotyping were funded by a grant from the California Department of Fish and Game and the UC Davis Wildlife Health Center, an NSF Doctoral Dissertation Improvement grant and the Museum of Vertebrate Zoology. Collecting and research permits were provided by the California Department of Fish and Game, Yosemite, and Sequoia-Kings Canyon National Parks. We thank C Moritz for extensive comments and discussion, D Wake for discussion and comments on the manuscript and D Giuliani, T Tunstall, K Tsao and V Vredenburg for help in the field. All research was conducted under UC Berkeley Animal Care and Use protocol #R278.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Rights and permissions
About this article
Cite this article
Rovito, S., Schoville, S. Testing models of refugial isolation, colonization and population connectivity in two species of montane salamanders. Heredity 119, 265–274 (2017). https://doi.org/10.1038/hdy.2017.31
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2017.31