Abstract
Gene expression regulation has an important role in short-term acclimation and long-term adaptation to changing environments. However, the genetic architecture of gene expression has received much less attention than that of traditional phenotypic traits. In this study, we used a 5 × 5 full-factorial breeding design within each of two Atlantic salmon (Salmo salar) populations to characterize the genetic architecture of gene transcription. The two populations (LaHave and Sebago) are being used for reintroduction efforts into Lake Ontario, Canada. We used high-throughput quantitative real-time PCR to measure gene transcription levels for 22 genes in muscle tissue of Atlantic salmon fry. We tested for population differences in gene transcription and partitioned the transcription variance into additive genetic, non-additive genetic and maternal effects within each population. Interestingly, average additive genetic effects for gene transcription were smaller than those reported for traditional phenotypic traits in salmonids, suggesting that the evolutionary potential of gene transcription is lower than that of traditional traits. Contrary to expectations for early life stage traits, maternal effects were small. In general, the LaHave population had higher additive genetic effects for gene transcription than the Sebago population had, indicating that the LaHave fish have a higher adaptive potential to respond to the novel selection pressures associated with reintroduction into a novel environment. This study highlights not only the profound variation in gene transcription possible among salmonid populations but also the among-population variation in the underlying genetic architecture of such traits.
Similar content being viewed by others
Introduction
Acclimation via phenotypic plasticity and adaptation via selection are the two main mechanisms organisms use to cope with environmental stress, and they thus play a vital role in organism survival and population persistence in changing or novel environments (Hansen et al., 2012). Gene expression regulation has long been recognized as playing an important role in both acclimation and adaptation (Hochachka and Somero, 1984; Crawford and Powers, 1992). The ability of individual genotypes to express different phenotypes in response to environmental fluctuations and stress challenges is mediated by regulation of gene expression, specifically through quantitative changes in gene expression levels and selective expression of different isoforms (Schlichting and Smith, 2002; Schulte, 2004). Although gene expression is influenced by internal and external environmental factors (Hodgins-Davis and Townsend, 2009), genetic analysis has shown that gene expression variation is heritable, and gene expression levels are partially determined by genetic factors (Cheung and Spielman, 2002). Moreover, it was demonstrated over four decades ago that evolutionary differences among species may depend more on changes in gene expression regulation than changes in gene sequences (King and Wilson, 1975). Recently, novel genomic tools have shown that local adaptation among populations within a species is 10 times more likely to be driven by genetic variation in gene expression regulation than by genetic variation that affects amino-acid sequences (Fraser, 2013). Therefore, gene expression is not only a process whereby organisms acclimate to new or changing environments, but it also provides a source of variation that selection can act on.
Populations can exhibit different responses and tolerances to environmental stressors because of their separate evolutionary histories, resulting in divergent performance that is relevant for both commercial culture and conservation (Forsman, 2014; He et al., 2016). Population-level differences in tolerance of environmental toxins and temperature stress have been associated with differences in regulation and pre-existing differences in gene expression, reflecting local habitat conditions (for example, Whitehead et al., 2010; Gleason and Burton, 2015). Although there are many studies that report population differences in gene expression, comparisons of the genetic components of gene expression variance among populations have received much less attention.
The quantitative genetic definition of genetic architecture is the relative contribution of additive genetic, non-additive genetic and maternal effect variance components to observed phenotypic variance. All three components of genetic architecture are important for evolutionary processes due to their contributions to phenotypic variation. Additive genetic effects are critical for evolutionary potential, and their magnitude is directly related to the evolutionary rate of a trait in response to a selection pressure, following the breeder's equation (Falconer and Mackay, 1996). Non-additive genetic effects (for example, dominance and epistasis) can also contribute significantly to phenotypic variation (Evans and Neff, 2009; Gallardo et al., 2010), and non-additive effects can increase in response to extreme environments or stressors (Jinks et al., 1973; Aykanat et al., 2012a). Finally, maternal effects are significant contributors to variation in fitness-related traits at early life stages (Heath et al., 1999). Thus, populations of the same species, subject to different evolutionary histories (past selection pressures), can, in theory, have different genetic architectures for the same trait. Although gene expression is closely linked to phenotype and can be highly heritable, it is not clear how populations differ in additive genetic components of gene expression variation. Similarly, although maternal effects and non-additive genetic effects have important roles in individual and population performance, the contribution of those effects to gene expression variation among populations is unknown.
Atlantic salmon is an economically and culturally important species broadly distributed in North America and Europe (Parrish et al., 1998; COSEWIC, 2006). During the past two centuries, many Atlantic salmon populations have become endangered or extirpated (Parrish et al., 1998). In particular, Atlantic salmon was once an abundant fish in Lake Ontario and its tributaries but was extirpated by 1900 due to habitat degradation (Crawford, 2001). Because of its economic and cultural value, there have been increasing efforts since the 1980s to reintroduce Atlantic salmon into the revitalized Lake Ontario and its tributaries (Dimond and Smitka, 2005); however, those reintroduction attempts have been unsuccessful in producing self-sustaining populations (Stewart and Schaner, 2002; COSEWIC, 2006). Current reintroduction efforts for Atlantic salmon in Lake Ontario are using fish from two populations—LaHave and Sebago. The LaHave population fish contributed to the successful introduction of Atlantic salmon into Trout Lake, Ontario, whereas the Sebago population fish performed well in introductions in Maine (Dimond and Smitka, 2005). Reintroduced fish are subject to novel selective pressures after release and hence their adaptive potential is a key factor for their long-term establishment in new environments (He et al., 2016). Theoretically, higher additive genetic effects reflect higher adaptive potential in response to selection pressures; thus the population with higher additive genetic effects is a better candidate for reintroduction (Houde et al., 2015a).
In this study, we examined the genetic architecture of gene transcription for key muscle function-related genes in the two Atlantic salmon populations. Fish muscle accounts for 50–60% of the total body mass in salmonids (Jobling, 1993), and it plays an important role in swimming performance and energy regulation, which is vital for fish migration, foraging and predation avoidance in the wild (Álvarez and Metcalfe, 2007). Although there have been some studies published that document gene transcription data for muscle function in salmonids (for example, swimming effects (Palstra et al., 2013); feeding treatments (Bower et al., 2009; Garcia de la Serrana et al., 2015); flesh quality (Larsson et al., 2012)), none has investigated genetic architecture of gene transcription. The overall goals of this study were two-fold: first we characterize the genetic architecture of transcription at key genes associated with muscle function in early life history stage Atlantic salmon and, second, we test for population-level differences in that genetic architecture. Our understanding of the potential evolutionary response of transcription in non-model organisms is critical for both evolutionary theory and for conservation applications where adaptive potential is critical. Here, we test three predictions to address our goals. First, we predicted significant population effects on gene transcription because other studies have documented population effects on gene transcription in both wild and captive populations (for example, Whitehead and Crawford, 2005; Hutter et al., 2008). Second, we predicted substantial additive genetic effects (and hence adaptive potential) for gene transcription based on previous work (for example, Price et al., 2011; Aykanat, et al., 2012a; Tedeschi et al., 2016; Wheeler et al., 2016). Associated with that, we also expected to detect substantial non-additive genetic and maternal effects for gene transcription because those effects are strong early in life for more traditional phenotypic traits (for example, Aykanat et al., 2012b; Houde et al., 2013, 2015b). Third, we predicted that the two populations would differ in their relative additive genetic, non-additive genetic and maternal effect variance components (that is, genetic architecture). We expected to find significant differences in the genetic architecture of these two populations because of the substantial life history variation between the two populations (the LaHave population is an anadromous population and the Sebago population is a potoanadromous population), and previous work documented significant genetic architecture differences for more traditional phenotypic traits between these two populations (Houde et al., 2013, 2015b). To this end, we implemented a 5 × 5 full-factorial breeding design for each of LaHave and Sebago populations. We used a high-throughput quantitative real-time (qRT)-PCR system to quantify gene transcription at 22 gene loci in muscle for over 400 Atlantic salmon fry. We analyzed population and parental effects on gene transcription and partitioned transcription variance into additive genetic, non-additive genetic and maternal effect variance components within each population.
Materials and methods
Breeding design
We used parental fish from two Atlantic salmon populations (LaHave and Sebago), which were provided by the Ontario Ministry of Natural Resources and Forestry (OMNRF). Atlantic salmon typically spawn in the fall in rivers with the eggs and embryos developing in the gravel until they emerge as fry the following spring. Atlantic salmon juveniles remain in the freshwater tributaries for typically 2 years before migrating to the ocean (anadromous) or a larger freshwater waterbody (potoanadromous). The LaHave population is anadromous and originates from LaHave River, Nova Scotia, Canada; fertilized eggs from wild LaHave Atlantic salmon were originally received by OMNRF in 1995, and this group has been reared in captive breeding since then (Houde et al., 2015b). The Sebago population is potoanadromous and originates from Sebago Lake, Maine, USA; fertilized eggs from a hatchery supplemented river were received by OMNRF in 2006 (Houde et al., 2015b). Thus, LaHave was fourth-generation hatchery and Sebago was second-generation hatchery within the OMNRF facility at the time of this study. In early November 2011, we used five males and five females from each population where each male was crossed to each female in a full-factorial design to produce 25 families. The fertilized eggs were incubated in vertical stack incubators in replicate cells. After hatching, each family was reared in two replicate tanks. In May 2012 (187 days post fertilization), 3–5 fry per tank were humanely killed by overdose of MS222, and the whole fish was preserved in RNA later. All 50 families had representative fish from both replicate tanks except for one family of the LaHave population, which had one tank sampled because all the individuals from the other tank died. We collected the fish from this early developmental stage because fry are most commonly used for reintroduction of Atlantic salmon into Lake Ontario (Coghlan and Ringler, 2004); hence, factors affecting muscle function and growth at this stage may be disproportionally important for reintroduction success.
RNA extraction and cDNA preparation
Although this study was designed to target muscle tissue, the fish were too small (average body mass: 0.34±0.11 g) to collect pure muscle tissue from preserved samples. Thus, to collect tissue samples for RNA isolation, we first cut the fish through the lateral line and then used the tissue between the dorsal and adipose fins for RNA extraction. Therefore, the tissue we used for RNA extraction in this study was primarily muscle; however, the sample also contained skin and some bone tissue. The tissue sample was mechanically homogenized in the presence of Isol-RNA Lysis Reagent (5 PRIME, Gaithersburg, MD, USA) following the manufacturer’s protocol. The concentration and purity of extracted RNA were checked using a NanoVue spectrophotometer (GE Healthcare Bio-Science Corp, Piscataway, NJ, USA). The quality of a subset of RNA samples was also checked using the Agilent RNA 6000 Nano Kit in an Agilent 2100 Bioanalyzer (Agilent Technologies, Mississauga, ON, Canada). About 2 μg RNA was treated with DNase I (Sigma-Aldrich, Oakville, ON, Canada) to remove contaminated DNA and then used for cDNA synthesis using the High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Burlington, ON, Canada). In total, we reverse transcribed 480 RNA samples. The quality of 192 cDNA samples was checked by PCR amplification of the elongation factor 1A (ef1ab) gene, and all showed the expected band on agarose gel.
Gene selection
The 22 candidate genes (Table 1) we chose for this study are involved in a variety of key processes in fish muscle. Of the 22 selected genes, 7 encode enzymes directly involved in energy generation by participating in fatty acid oxidation (acadl, acadm and cpt2), the citric acid cycle (cs), carbohydrate oxidation (dlat), anerobic glycolysis (ldha) and glycogenolysis (pygma). Five genes are involved in energy regulation through signal transduction (camk2g and prkag2), fatty acid transport (fabp3) and transcription regulation (ppara and tfam). Six genes encode muscle structural proteins that are involved in muscle contraction (dysi1, fmyhc, mlc2, smyhc1, tnni2 and tnnt1). The dysi1 gene was selected in the muscle contraction gene group because it is involved in muscle membrane repair (Han and Campbell, 2007). Four selected genes (frap1, myf5, myod1 and murf1) are growth-related genes. The myf5 and myod1 genes are members of the myogenic regulatory factors and regulate muscle cell differentiation and growth (Francetic and Li, 2011). The murf1 gene belongs to the ubiquitin-protein ligases, which are required for the ubiquitin proteasome system to degrade muscle proteins (Lecker, 2003). We also included the frap1 gene, which has multiple cellular signal transduction roles in mediation of stress response and is known to regulate cell growth and proliferation (Desai et al., 2002; Murakami et al., 2004).
Quantitative real-time PCR
The sequences of TaqMan primers and probes for three genes were obtained from journal publications ef1ab (Løvoll et al., 2011), fabp3 (Torstensen et al., 2009) and fmyhc (Hevrøy et al., 2006). Gene sequences for the other genes were downloaded from NCBI (http://www.ncbi.nlm.nih.gov/), and the primers and TaqMan MGB probes (Table 1) for those genes were designed using Primer Express 3.0 (Applied Biosystems). The primers were designed to amplify small amplicons (54–71 bp, Table 1) and are predicted to have 100% efficiency using the pcrEfficiency software (Mallona et al., 2011). The primers and probes were synthesized and spotted into through-holes of OpenArray chips by Applied Biosystems (Applied Biosystems).
OpenArray qRT-PCR was performed using a QuantStudio 12K Flex Real-Time PCR System following the manufacturer’s instructions. We prepared a 5 μl mixture for each cDNA sample, which contained 2.5 μl TaqMan OpenArray Real-Time PCR Master Mix (Applied Biosystems) and 1.2 μl cDNA. We used the 56 × 48 format OpenArray chip, which has 48 subarrays in each chip, and each subarray contains 64 through-holes. Each chip can be used to measure gene expression for 48 individual cDNA samples for all the targeted genes in duplicate. The 5 μl mixtures were prepared in 384-well plates and were then loaded into OpenArray chips using the OpenArray AccuFill System, and each qRT-PCR reaction was performed in a 33 nl volume.
The relative threshold cycle (CRT) value for each reaction was obtained using the ExpressionSuite Software v1.0.4 (Applied Biosystems). The CRT method is developed for OpenArray technology and takes individual PCR efficiency into account. Expression data (CRT value) that had a s.d. between the two technical replicates larger than 0.5 were removed from the analysis. For the remainder, we used the mean of the CRT values of the technical replicates for each individual. The expression level for each gene was normalized to ef1ab expression and the ΔCRT values (CRTtargeted gene – CRTef1ab) were used for all downstream analyses. We used ef1ab as the endogenous reference gene because it has been shown to be an excellent endogenous control compared to other common reference genes in Atlantic salmon muscle tissue (Olsvik et al., 2005); this gene exhibited similar CRT values in the two populations in this study (mean CRT values; 16.91 (LaHave) and 16.92 (Sebago)). The ratios of gene transcription between the two populations were calculated using the 2−ΔΔCT method under the assumption of 100% PCR efficiency (Pfaffl, 2001). To determine whether this assumption was valid, we used the LinRegPCR software (Ruijter et al., 2009) to estimate mean PCR efficiency based on fluorescent data generated in each PCR cycle for each of the analyzed genes for 95 samples across two chips (Supplementary Table S1). We found that the mean PCR efficiency for the primers was 96.03±9.68% (mean±s.d.).
Statistical analysis
To test for population effects on gene transcription between the two Atlantic salmon populations, we initially used the following model for each gene:

where Yijklmn is the normalized transcription level (relative to the ef1ab gene, CRTtargeted gene–CRTef1ab), μ is the mean value, Pi is the ith effect of population, Tj is the jth effect of tank, Sk is the kth effect of sire, Dl is the lth effect of dam, Im is the mth effect of interaction between sire and dam and eijklmn is the random residual. Population effect was included as a fixed effect; tank, sire, dam and the interaction between sire and dam was random effects. We used the step function in the lmerTest R package to perform backward elimination for model selection and statistical analysis for population effects; random factors with P-values of 0.1 or higher based on likelihood ratio test were removed from the model. The final models (Supplementary Table S2) were used to determine whether there were significant population effects on gene transcription based on an F-test (Kuznetsova et al., 2016).
To quantify the genetic architecture for all the genes within the two populations separately, we partitioned the total variance into sire, dam and dam-by-sire interaction components using the following model:

where tank (T), sire (S), dam (D) and the interaction between sire and dam (I) were treated as random effects. The significance for tank, sire, dam and the interaction was determined using the likelihood ratio test between the full model and a reduced one without the tested effect using the observLmer2 function in the fullfact R package (Houde and Pitcher, 2016). Gene transcription variance was partitioned into sire (VS), dam (VD) and the interaction between sire and dam (VI) separately. Additive (VA), non-additive (VNA) and maternal variance (VM) were calculated as follows: VA=4VS; VNA=4VI; VM=VD – VS (Lynch and Walsh, 1998). To compare the phenotypic variance in gene expression explained by each of the four factors in model (2) and genetic architecture of gene transcription between the two populations, we conducted Wilcoxon signed-rank test.
Results
Population effects on gene transcription
We measured transcription for 22 genes using qRT-PCR in 428 Atlantic salmon fry. Nine genes showed significantly different transcription between the two populations (Figure 1; Supplementary Table S2). The cpt2, myf5, myod1 and tfam genes showed significantly higher expression in the LaHave population, whereas the acadl, cs, ldha, mlc2 and pygma genes showed significantly higher expression in the Sebago population, with the expression difference ranging from 8 to 99% (Figure 1). Although the difference in gene expression between two populations for most genes is <30%, the expression of ldha and pygma for the Sebago population was close to double that of the LaHave population. Of the nine differentially expressed genes, five are involved in muscle energy generation, two genes are involved in muscle growth, one gene is involved in muscle energy regulation and one gene is involved in muscle contraction.
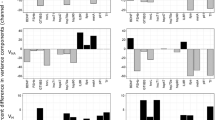
Differences in gene transcription between two Atlantic salmon (Salmo salar) populations (LaHave and Sebago) measured using qRT-PCR. The ratios of gene transcription between the two populations were calculated using the 2−ΔΔCT method under the assumption of 100% PCR efficiency. Gene names are described in Table 1. *P<0.05; **P<0.01.
Significant effects on gene transcription within each population
We tested for tank, sire, dam and the interaction between sire and dam effects on gene transcription for the 22 genes within each population. Ten genes showed significant tank effects in the LaHave population and 11 genes showed significant tank effects in the Sebago population (Table 2). The replicate groups within each family were created at the time of fertilization and incubated and reared in separate incubation cells and tanks. Thus the ‘tank’ effect is actually integrated over 187 days of rapid development, and although we did our best to keep the replicate condition identical, small environmental differences likely accumulate over time affecting transcriptional profiles. Seven genes showed significant tank effects in both populations. Among those seven genes, three genes are related to growth, two genes are related to muscle contraction and two genes encode enzymes involved in energy generation. Within the LaHave population, five genes showed significant dam effects and two genes showed significant sire effects. Within the Sebago population, four genes showed significant dam effects and one gene showed significant sire effects. The camk2g gene showed a significant sire effect in both populations with similar levels of explained variance, whereas all the other significant sire or dam effects on gene expression were population-specific with different levels of explained variance between the two populations. No genes showed significant sire-by-dam interaction effects in either population.
On average across all 22 genes, tank, sire, dam and sire-by-dam interaction effects explained 13.0, 4.2, 7.3 and 2.6% of the phenotypic variance in gene expression in the LaHave population, respectively, and 14.2, 1.5, 4.5 and 4.2% of phenotypic variance in the Sebago population, respectively (Table 3). The phenotypic variance in gene expression explained by sire effect was significantly higher in the LaHave population than in the Sebago population (Wilcoxon signed-rank test, P=0.01), whereas the phenotypic variation explained by tank, dam and sire-by-dam interaction effects was not significantly different.
Additive genetic, non-additive genetic and maternal effects
The two study populations exhibited both similarities and substantial differences in genetic variance components despite having been reared in a common environment (Table 2). Maternal and genetic effects collectively explained 30.3% of the total phenotypic variance in the LaHave population and 25.8% of the total variance in the Sebago population. Specifically, in the LaHave population, additive genetic effects explained 16.7% of the total variance and non-additive genetic effects explained 10.5% of the total variance (Table 3). In the Sebago population, additive genetic effects explained 6.2% of the total phenotypic variance and non-additive genetic effects explained 16.6% of the total variance. Maternal effects explained 3.1% of gene expression variance in the LaHave population and 3.0% of gene expression variance in the Sebago population.
Fourteen genes showed higher additive genetic effects in the LaHave population, whereas four genes showed higher additive genetic effects in the Sebago population (Table 2). Seven genes showed higher non-additive genetic effects in the LaHave population, whereas 10 genes showed higher non-additive genetic effects in the Sebago population (Table 2). Four genes showed no additive genetic effects and five genes showed no non-additive genetic effects in both populations. Half of the genes showed higher maternal effects in one population than in the other population (Table 2). Across all 22 genes, the two populations were significantly different in additive genetic effects (Wilcoxon signed-rank test, P=0.01), whereas there were no significant differences between the populations in non-additive genetic (P=0.37) and maternal effects (P=0.73).
Discussion
Population and individual differences in gene transcription have been reported in many studies (for example, Oleksiak et al., 2002; Storey et al., 2007; Hutter et al., 2008). We found that five of the seven metabolic enzyme genes involved in energy generation showed significant transcriptional differences between the two populations, after accounting for tank and family effects. The energy generation category genes had the highest proportion of significant population differences (Figure 1), perhaps reflecting a fundamental difference between the two populations. Differences in the kinetic properties and concentrations of metabolic enzymes among populations and species are thought to contribute to adaptation to temperature variation as they have an important role in homeostasis maintenance (Hochachka and Somero, 1984; Crawford and Powers, 1989; Crockett and Sidell, 1990). Higher temperature tolerance is especially important for salmonids as climate change-related temperature increase contributes to mortality in salmonids during migration to spawning sites (Eliason et al., 2011). The population differences in the transcription of metabolic enzyme genes presented here may underlie population differences in thermal tolerance, as other studies demonstrating differential expression of metabolic genes have been interpreted as evidence for local adaptation to different environmental temperatures (for example, the lactate dehydrogenase B gene in Fundulus heteroclitus) (Crawford and Powers, 1989, 1992).
The genetic architecture underlying phenotypic traits critical for survival and reproduction is important as it is the basis for evolutionary response to changing environments. Thus the genetic architecture of gene transcription reflects the scope for evolutionary response to novel environments upon translocation or colonization. In the present study, we partitioned the additive genetic, non-additive genetic and maternal variance components of gene expression using a full-factorial breeding design. The average additive genetic variance components (16.7 in LaHave and 6.2% in Sebago) were comparable to studies in humans, which reported a mean heritability (h2) of genome-wide gene transcription of 0.017–0.234 depending on the tissue (Price et al., 2011; Wheeler et al., 2016). Our estimates were lower than a study focused on transcription of four cytokine genes in Chinook salmon (Oncorhynchus tshawytscha; mean h2=0.26) (Aykanat et al., 2012a) and another study on transcription of three heat shock protein genes in sea turtles (Caretta caretta; mean h2=0.58; Tedeschi et al., 2016). The differences in additive genetic effects between our study and previous studies in other species may be due to different strengths of selection acting on the selected genes, the animals experiencing different evolutionary histories or different experimental designs and analytical methods.
The majority of the genes included in our study exhibited lower additive genetic effects than the median heritability of 24 fitness-related traits in salmonids (Carlson and Seamons, 2008). This indicates that gene transcription may be under even stronger selection than more traditionally recognized ‘fitness-related’ traits. It is also possible that gene transcription may not follow classical quantitative genetic patterns; for example, if any of our selected genes were under simple transcriptional regulation, they would violate the assumption of polygenic control. However, given the nature and complexity of the function of the selected genes, we feel it is unlikely that they have a simple transcription regulatory network. The low additive genetic variance estimates suggest that the genes examined in the present study may be constrained in their evolutionary potential according to the breeder's equation (Falconer and Mackay, 1996). However, a few genes in our study (for example, camk2g and tnnt1) did exhibit high additive genetic variance, and may thus evolve more rapidly.
Populations subject to different selection pressures can exhibit different distributions of genetic variance components, as, for example, strong selection reduces additive genetic variance (Carlson and Seamons, 2008). We found that additive genetic variance was higher across the 22 genes examined in the LaHave population. One potential explanation is that the longer captive rearing of the LaHave population under relaxed selection pressures may have allowed this population to recover additive genetic variance. Another potential reason is their life history difference: the LaHave population is an anadromous population and the Sebago population is a potoanadromous population. Although this difference in life history is profound, it is consistent with natural levels of life history variation within this species. The LaHave population may experience higher rates of gene flow because of the river being connected to the ocean, and that gene flow may result in higher genetic variation in anadromous populations than landlocked populations in North America (King et al., 2001). However, we only examined one population from each life history type; thus, we cannot generalize across the range of life history variation in Atlantic salmon. Regardless of the reason, it is clear that our understanding of the evolutionary and ecological mechanisms that drive patterns of additive genetic variance in gene transcription need to be improved with further research.
Although the additive genetic effects of gene transcription have been investigated in a variety of studies (for example, Price et al., 2011; Aykanat, et al., 2012a; Tedeschi et al., 2016; Wheeler et al., 2016), non-additive genetic and maternal effects for gene transcription are seldom reported. In particular, dams usually have greater contribution to the phenotypes of their offspring at early life stages than do sires (Bernardo, 1996; Wolf and Wade, 2009). In our analyses, a greater number of genes showed significant dam effects than sire effects, and the dam variance component was higher than that of the sire. Aykanat et al. (2012a) also found that the dam variance component of gene transcription for four cytokine genes was higher than that of the sire in Chinook salmon. Videvall et al. (2016) found that the pattern of gene transcription in hybrids between two Arabidopsis lyrata populations was more similar to the maternal population than to the paternal population. We expected to find higher maternal effects on transcription for the genes we surveyed at young juvenile life stage. However, in our study, maternal effects were generally low, and the average maternal effect was smaller than the average additive and non-additive genetic variance components in both populations, suggesting that maternal effects for gene transcription decreased more rapidly than for more traditional phenotypic traits.
Our observed differences in the magnitude of additive genetic, non-additive genetic and maternal effects for gene transcription across the functional groups are perhaps not surprising, as different traits can exhibit dramatic variation in heritability. For example, a review of heritability estimates (h2) in salmonids found that 24 fitness-related traits exhibited a wide range of median h2, varying from 0.02 to 0.51 (Carlson and Seamons, 2008). In our study, we found that genes involved in energy regulation generally had higher additive genetic components than genes involved in energy generation and muscle growth. This likely reflects different function genes having experienced different selection pressures; however, regardless of the mechanism, our results indicate that the functional groups will have different responses to selection in the short term. Likewise, five of the seven energy generation-related genes had higher non-additive genetic variance components in the Sebago population, whereas all of muscle contraction genes had higher maternal effects in the Sebago population. Although individually no gene exhibited a significant sire-by-dam interaction, and maternal effects were general low, the pattern of population differences in genetic architecture indicates that genes within different functional categories can evolve independently as evolutionary processes are affected by non-additive genetic and maternal effects as well as by the more classically relevant additive genetic effects.
Conclusions
We used Atlantic salmon from two populations in a full-factorial breeding design to quantify gene transcription for 22 genes in muscle. We found that maternal and genetic effects combined explained 30.3 and 25.8% of the transcriptional variation for these genes in the LaHave and Sebago populations, respectively. Contrary to expectation for traditional phenotypic traits at this young fry life stage, our results also highlighted that maternal effects are lower than genetic effects for gene transcription, at least for the genes examined. Additionally, the evolutionary potential of gene transcription may be lower than traditional phenotypic traits because of the lower amount of additive genetic effects. Overall, the LaHave population exhibited higher additive genetic effects, suggesting a higher adaptive potential to selection pressures in Lake Ontario than the Sebago population. Although our study includes more genes than most previous transcription genetic architecture studies using qRT-PCR, it is limited in the breadth of gene function, tissue sampled and developmental stage for making strong inferences on patterns. Further research on the genetic architecture of gene transcription would be warranted for elucidating such patterns.
References
Álvarez D, Metcalfe NB . (2007). The tradeoff between catch-up growth and escape speed: variation between habitats in the cost of compensation. Oikos 116: 1144–1151.
Aykanat T, Heath JW, Dixon B, Heath DD . (2012a). Additive, non-additive and maternal effects of cytokine transcription in response to immunostimulation with Vibrio vaccine in Chinook salmon (Oncorhynchus tshawytscha. Immunogenetics 64: 691–703.
Aykanat T, Bryden CA, Heath DD . (2012b). Sex-biased genetic component distribution among populations: additive genetic and maternal contributions to phenotypic differences among populations of Chinook salmon. J Evol Biol 25: 682–690.
Bernardo J . (1996). Maternal effects in animal ecology. Am Zool 36: 83–105.
Bower NI, Taylor RG, Johnston IA . (2009). Phasing of muscle gene expression with fasting-induced recovery growth in Atlantic salmon. Front Zool 6: 18.
Carlson SM, Seamons TR . (2008). A review of quantitative genetic components of fitness in salmonids: implications for adaptation to future change. Evol Appl 1: 222–238.
Cheung VG, Spielman RS . (2002). The genetics of variation in gene expression. Nat Genet 32: 522–525.
Coghlan SM, Ringler NH . (2004). A comparison of Atlantic salmon embryo and fry stocking in the Salmon River, New York. North Am J Fish Manag 24: 1385–1397.
COSEWIC. (2006) COSEWIC assessment and Status Report on the Atlantic Salmon Salmo salar (Lake Ontario population) in Canada. Committee on the Status of Endangered Wildlife in Canada, Ottawa, pp vii+26.
Crawford SS . (2001). Salmonine introductions to the Laurentian Great Lakes: an historical review and evaluation of ecological effects. Canadian Special Publication of Fisheries and Aquatic Sciences 132. NRC Press: Ottawa, ON, Canada.
Crawford DL, Powers DA . (1989). Molecular basis of evolutionary adaptation at the lactate dehydrogenase-B locus in the fish Fundulus heteroclitus. Proc Natl Acad Sci USA 86: 9365–9369.
Crawford DL, Powers DA . (1992). Evolutionary adaptation to different thermal environments via transcriptional regulation. Mol Biol Evol 9: 806–813.
Crockett EL, Sidell BD . (1990). Some pathways of energy metabolism are cold adapted in Antarctic fishes. Physiol Zool 63: 472–488.
Desai BN, Myers BR, Schreiber SL . (2002). FKBP12-rapamycin-associated protein associates with mitochondria and senses osmotic stress via mitochondrial dysfunction. Proc Natl Acad Sci USA 99: 4319–4324.
Dimond P, Smitka J . (2005). Evaluation of selected strains of Atlantic salmon as potential candidates for the restoration of Lake Ontario. Trout Unlimited Canada Technical Report ON-012: Geulph, ON, Canada, pp 41.
Eliason EJ, Clark TD, Hague MJ, Hanson LM, Gallagher ZS, Jeffries KM et al(. (2011). Differences in thermal tolerance among sockeye salmon populations. Science 332: 109–112.
Evans ML, Neff BD . (2009). Non-additive genetic effects contribute to larval spinal deformity in two populations of Chinook salmon (Oncorhynchus tshawytscha. Aquaculture 296: 169–173.
Falconer DS, Mackay TFC . (1996). Introduction to Quantitative Genetics. Longmans Green: Harlow, Essex, UK.
Forsman A . (2014). Effects of genotypic and phenotypic variation on establishment are important for conservation, invasion, and infection biology. Proc Natl Acad Sci USA 111: 302–307.
Francetic T, Li Q . (2011). Skeletal myogenesis and Myf5 activation. Transcription 2: 109–114.
Fraser HB . (2013). Gene expression drives local adaptation in humans. Genome Res 23: 1089–1096.
Gallardo JA, Lhorente JP, Neira R . (2010). The consequences of including non-additive effects on the genetic evaluation of harvest body weight in Coho salmon (Oncorhynchus kisutch. Genet Sel Evol 42: 19.
Garcia de la serrana D, Devlin RH, Johnston IA . (2015). RNAseq analysis of fast skeletal muscle in restriction-fed transgenic coho salmon (Oncorhynchus kisutch: an experimental model uncoupling the growth hormone and nutritional signals regulating growth. BMC Genomics 16: 564.
Gleason LU, Burton RS . (2015). RNA-seq reveals regional differences in transcriptome response to heat stress in the marine snail Chlorostoma funebralis. Mol Ecol 24: 610–627.
Han R, Campbell KP . (2007). Dysferlin and muscle membrane repair. Curr Opin Cell Biol 19: 409–416.
Hansen MM, Olivieri I, Waller DM, Nielsen EE, GeM Working Group. (2012). Monitoring adaptive genetic responses to environmental change. Mol Ecol 21: 1311–1329.
He X, Johansson ML, Heath DD . (2016). Role of genomics and transcriptomics in selection of reintroduction source populations. Conserv Biol 30: 1010–1018.
Heath DD, Fox CW, Heath JW . (1999). Maternal effects on offspring size: variation through early development of Chinook salmon. Evolution 53: 1605–1611.
Hevrøy EM, Jordal A-EO, Hordvik I, Espe M, Hemre G-I, Olsvik PA . (2006). Myosin heavy chain mRNA expression correlates higher with muscle protein accretion than growth in Atlantic salmon, Salmo salar. Aquaculture 252: 453–461.
Hochachka PW, Somero GN . (1984) Biochemical Adaptation. Princeton University Press: Princeton, NJ, USA.
Hodgins-Davis A, Townsend JP . (2009). Evolving gene expression: from G to E to GxE. Trends Ecol Evol 24: 649–658.
Houde ALS, Garner SR, Neff BD . (2015xa). Restoring species through reintroductions: strategies for source population selection. Restor Ecol 23: 746–753.
Houde ALS, Black CA, Wilson CC, Pitcher TE, Neff BD . (2015b). Genetic and maternal effects on juvenile survival and fitness-related traits in three populations of Atlantic salmon. Can J Fish Aquat Sci 72: 751–758.
Houde ALS, Pitcher TE . (2016). Fullfact: an R package for the analysis of genetic and maternal variance components from full factorial mating designs. Ecol Evol 6: 1656–1665.
Houde AL, Wilson CC, Neff BD . (2013). Genetic architecture of survival and fitness-related traits in two populations of Atlantic salmon. Heredity 111: 513–519.
Hutter S, Saminadin-Peter SS, Stephan W, Parsch J . (2008). Gene expression variation in African and European populations of Drosophila melanogaster. Genome Biol 9: R12.
Jinks J, Perkins JM, Pooni H . (1973). The incidence of epistasis in normal and extreme environments. Heredity 31: 263–269.
Jobling M . (1993) Bioenergetics: feed intake and energy partitioning. In: Rankin JC, Jensen FB (eds). Fish Ecophysiology, Chapman & Hall Fish and Fisheries Series. Springer: Netherlands, pp 1–44.
King TL, Kalinowski ST, Schill WB, Spidle AP, Lubinski BA . (2001). Population structure of Atlantic salmon (Salmo salar L.): a range-wide perspective from microsatellite DNA variation. Mol Ecol 10: 807–821.
King MC, Wilson AC . (1975). Evolution at two levels in humans and chimpanzees. Science 188: 107–116.
Kuznetsova A, Brockhoff PB, Christensen RHB . (2016). lmerTest: tests in linear mixed effects models. R package version. 2, pp 0-32.
Larsson T, Mørkøre T, Kolstad K, Østbye T-K, Afanasyev S, Krasnov A . (2012). Gene expression profiling of soft and firm Atlantic salmon fillet. PLoS One 7: e39219.
Lecker SH . (2003). Ubiquitin-protein ligases in muscle wasting: multiple parallel pathways? Curr Opin Clin Nutr Metab Care 6: 271–275.
Løvoll M, Austbø L, Jørgensen JB, Rimstad E, Frost P . (2011). Transcription of reference genes used for quantitative RT-PCR in Atlantic salmon is affected by viral infection. Vet Res 42: 8.
Lynch M, Walsh B . (1998) Genetics and Analysis of Quantitative Traits. Sinauer Associates: Sunderland, MA, USA.
Mallona I, Weiss J, Egea-Cortines M . (2011). pcrEfficiency: a web tool for PCR amplification efficiency prediction. BMC Bioinformatics 12: 404.
Murakami M, Ichisaka T, Maeda M, Oshiro N, Hara K, Edenhofer F et al. (2004). mTOR is essential for growth and proliferation in early mouse embryos and embryonic stem cells. Mol Cell Biol 24: 6710–6718.
Oleksiak MF, Churchill GA, Crawford DL . (2002). Variation in gene expression within and among natural populations. Nat Genet 32: 261–266.
Olsvik PA, Lie KK, Jordal A-EO, Nilsen TO, Hordvik I . (2005). Evaluation of potential reference genes in real-time RT-PCR studies of Atlantic salmon. BMC Mol Biol 6: 21.
Palstra AP, Beltran S, Burgerhout E, Brittijn SA, Magnoni LJ, Henkel CV et al. (2013). Deep RNA sequencing of the skeletal muscle transcriptome in swimming fish. PLoS One 8: e53171.
Parrish DL, Behnke RJ, Gephard SR, McCormick SD, Reeves GH . (1998). Why aren’t there more Atlantic salmon (Salmo salar? Can J Fish Aquat Sci 55: 281–287.
Pfaffl MW . (2001). A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29: e45.
Price AL, Helgason A, Thorleifsson G, McCarroll SA, Kong A, Stefansson K . (2011). Single-tissue and cross-tissue heritability of gene expression via identity-by-descent in related or unrelated individuals. PLoS Genet 7: e1001317.
Ruijter JM, Ramakers C, Hoogaars WMH, Karlen Y, Bakker O, van den Hoff MJB et al. (2009). Amplification efficiency: linking baseline and bias in the analysis of quantitative PCR data. Nucleic Acids Res 37: e45.
Schlichting CD, Smith H . (2002). Phenotypic plasticity: linking molecular mechanisms with evolutionary outcomes. Evol Ecol 16: 189–211.
Schulte PM . (2004). Changes in gene expression as biochemical adaptations to environmental change: a tribute to Peter Hochachka. Comp Biochem Physiol B Biochem Mol Biol 139: 519–529.
Stewart TJ, Schaner T . (2002). Lake Ontario salmonid introductions 1970 to 1999: stocking, fishery and fish community influences. In: Lake Ontario Fish Communities and Fisheries: 2001 Annual Report of the Lake Ontario Management Unit. Queen’s Printer for Ontario: Picton, ON, Canada.
Storey JD, Madeoy J, Strout JL, Wurfel M, Ronald J, Akey JM . (2007). Gene-expression variation within and among human populations. Am J Hum Genet 80: 502–509.
Tedeschi JN, Kennington WJ, Tomkins JL, Berry O, Whiting S, Meekan MG et al. (2016). Heritable variation in heat shock gene expression: a potential mechanism for adaptation to thermal stress in embryos of sea turtles. Proc R Soc B 283: 20152320.
Torstensen Be, Nanton Da, Olsvik Pa, Sundvold H, Stubhaug I . (2009). Gene expression of fatty acid-binding proteins, fatty acid transport proteins (cd36 and FATP) and β-oxidation-related genes in Atlantic salmon (Salmo salar L.) fed fish oil or vegetable oil. Aquac Nutr 15: 440–451.
Videvall E, Sletvold N, Hagenblad J, Ågren J, Hansson B . (2016). Strong maternal effects on gene expression in Arabidopsis lyrata hybrids. Mol Biol Evol 33: 984–994.
Wheeler HE, Shah KP, Brenner J, Garcia T, Aquino-Michaels K et alConsortium Gte. (2016). Survey of the heritability and sparse architecture of gene expression traits across human tissues. PLoS Genet 12: e1006423.
Whitehead A, Crawford DL . (2005). Variation in tissue-specific gene expression among natural populations. Genome Biol 6: R13.
Whitehead A, Triant DA, Champlin D, Nacci D . (2010). Comparative transcriptomics implicates mechanisms of evolved pollution tolerance in a killifish population. Mol Ecol 19: 5186–5203.
Wolf JB, Wade MJ . (2009). What are maternal effects (and what are they not)? Philos Trans R Soc Lond B Biol Sci 364: 1107–1115.
Acknowledgements
We thank C Wilson, W Sloan, S Ferguson and B Lewis for their help and support at the Ontario Ministry of Natural Resources and Forestry (OMNRF) Codrington Research Facility. We thank C Black and A Smith for their assistance for sample collection. We thank R Hepburn for his help with the running of OpenArray chips. This work was supported by Natural Sciences and Engineering Research Council (NSERC) of Canada funding to DDH (Discovery and Strategic Project Grant) and TEP (Strategic Project Grant).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Supplementary information
Rights and permissions
About this article
Cite this article
He, X., Houde, A., Pitcher, T. et al. Genetic architecture of gene transcription in two Atlantic salmon (Salmo salar) populations. Heredity 119, 117–124 (2017). https://doi.org/10.1038/hdy.2017.24
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2017.24
This article is cited by
-
Heritability of resistance-related gene expression traits and their correlation with body size of clam Meretrix petechialis
Journal of Oceanology and Limnology (2020)