Abstract
Weeds are among the greatest pests of agriculture, causing billions of dollars in crop losses each year. As crop field management practices have changed over the past 12 000 years, weeds have adapted in turn to evade human removal. This evolutionary change can be startlingly rapid, making weeds an appealing system to study evolutionary processes that occur over short periods of time. An understanding of how weeds originate and adapt is needed for successful management; however, relatively little emphasis has been placed on genetically characterizing these systems. Here, we review the current literature on agricultural weed origins and their mechanisms of adaptation. Where possible, we have included examples that have been genetically well characterized. Evidence for three possible, non-mutually exclusive weed origins (from wild species, crop-wild hybrids or directly from crops) is discussed with respect to what is known about the microevolutionary signatures that result from these processes. We also discuss what is known about the genetic basis of adaptive traits in weeds and the range of genetic mechanisms that are responsible. With a better understanding of genetic mechanisms underlying adaptation in weedy species, we can address the more general process of adaptive evolution and what can be expected as we continue to apply selective pressures in agroecosystems around the world.
Similar content being viewed by others
Introduction
Domesticated plants have long served as systems for the study of evolution and adaptation (Darwin, 1859). Crop domestication is an evolutionary process brought about by human interactions with plants and manipulation of their natural environment. When wild species are brought into cultivation, they are subjected to a suite of microevolutionary forces. Genetic drift, which randomly fixes alleles, is manifested through population bottlenecks imposed when a subset of individuals from wild populations is cultivated and propagated. Gene flow occurs via hybridization with wild relatives and among crop varieties, resulting in increased diversity across the genome. Strong artificial selection for domestication traits acts to fix alleles in the face of gene flow, and can result in selective sweeps in targeted genomic regions. The signatures of these microevolutionary processes can be sought in crop genomes, as selective sweeps and introgression are predicted to be limited to regions harboring genes involved in key traits, whereas drift and gene flow affect the entire genome (Doebley et al., 2006; Morrell et al., 2011).
As agriculture originated <12 000 years ago (Doebley et al., 2006), all evolutionary changes that take place in the agricultural context must have occurred within this recent time frame. This makes crop domestication an appealing and tractable system for studying the process of rapid evolutionary change. Moreover, because of their tremendous economic importance, many crops benefit from substantial research support, with community investment in genetic and genomic tools for applications in crop improvement. Multiple crop genomes have been sequenced to date, and a handful of these have been re-sequenced in several hundreds of individuals (for example, Lai et al., 2010; Lam et al., 2010). Comparative genomics is now being utilized to reveal similarities and differences in genome evolution during domestication (Morrell et al., 2011).
Although studies of crop domestication have revealed important insights into the genetic mechanisms underlying rapid evolutionary change under human-imposed selection, far less is known about the rapid evolution that can occur in the agricultural environment outside the boundaries of deliberate human selection. When weedy plants occupy this human-mediated environment, they can undergo rapid adaptive evolution to proliferate and escape eradication. Weeds are broadly defined as plants that are ‘growing where they are not desired’ or ‘plants out of place’ (Monaco et al., 2002). In this review we focus on weeds that invade and persist in agricultural fields, referred to herein as agricultural weeds. These weeds have evolved to survive in the artificial setting of crop fields, but they are not intentionally selected on by humans and are evolving through unintentional human-mediated selection (Warwick and Stewart, 2005); thus, agricultural weed evolution has occurred under the same forces that guide evolution in natural settings, including genetic drift, gene flow and natural selection.
Agricultural weeds are currently a leading cause of crop losses, resulting in ∼10% worldwide reduction in crop productivity (Oerke, 2006) and an estimated $33 billion annual cost to the United States alone (Pimentel et al., 2005). Whereas some of the traits that are adaptive in agricultural weeds are shared with those selectively favored in crops (such as herbicide and disease resistance), other weed-adaptive traits are similar to those favored in wild species (such as seed dispersal and dormancy). Therefore, agricultural weeds compose a unique evolutionary state, neither wild nor domesticated, that has developed in parallel to crop domestication. As study systems, agricultural weeds have many of the same advantages as crops in terms of the recent timing of evolutionary change, but they also offer the additional features of being subject to the forces of natural rather than artificial selection. Beyond their academic interest, understanding how weeds evolve is also clearly important for weed management and protection of the global food supply.
A complete understanding of the occurrence of agricultural weeds requires knowledge of two aspects of weed evolution. First, there is the phylogenetic origin of weed populations: from what wild and/or domesticated ancestors are agricultural weeds derived? Second, there is the origin of weed-adaptive traits per se: regardless of weed ancestry, what genetic mechanisms account for the traits that have enabled weeds to successfully invade the agricultural environment? As agricultural weeds are phylogenetically diverse, multiple evolutionary scenarios are likely to account for their appearance. Thousands of species are believed to function as agricultural weeds; about 250 are considered particularly problematic for agriculture worldwide, and these span multiple families and genera (Holm et al., 1977). Much as the current comparative mining of diverse crop genomes is helping us understand the common threads underlying the evolution of traits under domestication, comparative studies of weed evolution can address longstanding questions of how plant groups arise and adapt to novel environments.
There are two broad categories of agricultural weeds: those that are closely related to domesticated species, known as conspecific and congeneric weeds, and those that lack crop relatives. This difference has a key role in shaping weed origins and evolution. In particular, gene flow and introgression from crops is possible for weeds with crop relatives, and these forces influence the mechanisms by which weed-adaptive traits may evolve. For example, herbicide resistance may be acquired through crop-to-weed gene flow for weedy crop relatives, whereas this trait must evolve through de novo mutations or from pre-existing (standing) allelic variation in a weed without crop relatives. On the other hand, both classes of weed can potentially exchange genes with wild relatives, which provide an additional source of adaptive variation.
In this review, we survey the state of knowledge regarding the origin of agricultural weeds and their mechanisms of adaptation. We focus primarily on agricultural weed species for which genetic data have contributed to our understanding of their origins and mechanisms of adaptation to the agricultural environment. For more general reviews on agricultural weed evolution, we refer the reader to recent discussions by Ellstrand et al. (2010), Stewart et al. (2009), Neve et al. (2009), and Gressel (2005). Genetic resources are very unevenly distributed among weed taxa; the availability of genetic and genomic information for conspecific or congeneric crop species has often influenced which weed species get studied most. Nonetheless, the emergence of genomic tools that are applicable to non-model species offers nearly universal opportunities to advance the study of agricultural weeds, providing a wealth of study systems that can be used to understand the nature of rapid evolutionary change.
The origins of agricultural weeds
As the agricultural environment arose <12 000 years ago, plants that invade it represent organisms that have recently occupied new niches. Two potential mechanisms for weed colonization can be envisioned. First, plants that become agricultural weeds may be intrinsically capable of colonizing the agricultural environment. Plants within this group include generalists, with efficient dispersal mechanisms and phenotypically plastic genotypes (described by Baker, 1965), and plants that are specifically pre-adapted to growth and competition for resources in agricultural-like environments. Second, invasion of agricultural settings may be contingent on the process of adaptive evolution. The relative importance of plasticity/pre-adaptation vs adaptive evolution in the origins of weeds has been debated for many years in the weed science literature, and it may be that no single mechanism can account for all traits in any particular weed species (Baker, 1965; Clements et al., 2004). Additionally, changes that have occurred in cropping practices over time make it likely that ongoing adaptive evolution is a necessary feature of most successful agricultural weeds (Clements et al., 2004).
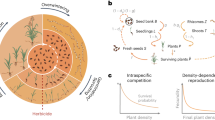
De Wet and Harlan (1975) proposed three routes to the origin of agricultural weeds. Figure 1 depicts these scenarios. First, wild colonizers that evolve or already possess traits specific for survival invade agricultural fields. These weeds may experience a demographic bottleneck in association with establishment in the field, resulting in reduced genome-wide neutral genetic diversity. However, gene flow can maintain high levels of genetic diversity if these weeds are in close proximity to populations occurring outside the agricultural setting. Second, hybridization between wild species and domesticated crops can occur, resulting in weeds with an admixed genomic makeup. This diversity can potentially fuel the evolution of adaptive weedy traits; the crop parent may also contribute traits that facilitate a pre-adaptation to crop fields. Weeds in this group may experience a genetic bottleneck if a small number of hybridization events are successful; however, continued hybridization with both wild and crop relatives after establishment could increase genetic diversity. Third, domesticated plants may escape cultivation and evolve into weed populations. Weeds directly descended from crops (‘de-domesticates’) will likely experience the strongest genetic bottleneck of all, due to the low level of diversity already present in crops compared to wild species. De-domesticated weeds may subsequently hybridize with crops and could hybridize with wild relatives, resulting in increases in genomic diversity.
Three alternate origins of agricultural weeds (a) wild colonizers, (b) wild and crop hybridization and (c) crop de-domesticates. Blue arrows represent reductions of diversity due to bottlenecks (genetic drift) and selection, with relative size indicating expected level of genetic diversity in each scenario. Orange stars represent new mutations and orange arrows represent gene flow through hybridization between groups, both representing increases in genetic diversity. The black X indicates a hybridization event between the crop and wild ancestors leading to the hybrid weed origin. Genes underlying weed adaptive traits come from standing variation, introgression and new mutations.
Although the last two categories of weed origins will result in weeds with close crop relatives, the first category does not eliminate the possibility of a weed having interfertile crop relatives. In practice, it is often difficult to separate out the effects of ancestry and subsequent introgression in weed genomes; thus, unequivocal assignment of one of these routes for the emergence of a given weedy species is not always possible. Additionally, few studies to date have addressed the demographic consequences of weed origins in specific weed species. Below, we discuss the available evidence for each of these routes of weed origins.
Wild to weed
Colonization of agricultural fields by wild plant species is as old as the practice of agriculture. Whereas selection for traits useful to humans was driving change in early crop species, wild plants were under selection to evade removal from crop fields. Wild plants that become weedy are often generalists, or are pre-adapted to infest agricultural fields. For example, witchweed (Striga hermonthica), a parasitic weed of cereal crops, has been shown to withstand a wide range of environmental conditions and can parasitize different hosts (Dawoud and Sauerborn, 1994; Ali et al., 2009). Based on AFLP (amplified fragment length polymorphism) markers, this genetically diverse outcrossing weed does not show genetic differentiation correlated with host specificity, but rather, it is correlated with geography (Welsh and Mohamed, 2011), suggesting that the weed is a generalist that can easily invade new crop hosts. Similarly, no genetic differentiation based on AFLP has been found between populations of broomrape (Orobanche foetida) infesting wild legumes and a population parasitizing cultivated vetch, suggesting a generalist ability to undergo host shifts onto cultivated species (Vaz Patto et al., 2008). Blackgrass (Alopecurus myosuroides), a non-parasitic weedy grass common in cereal fields worldwide, has high genetic diversity based on amplified fragment length polymorphisms within the populations, and little divergence between populations collected across Europe (Menchari et al., 2007). Local fields are likely connected by high levels of gene flow (Délye et al., 2010), but the lack of differentiation across large ranges suggests that this recently expanded weed did not require adaptive changes to colonize different cereal fields, although adaptation at a few loci cannot be ruled out. Interestingly, the above examples suggest that bottlenecks may not characterize weedy plant populations derived from generalists or from pre-adapted wild plants (Figure 1).
Adaptive evolution has been suggested to have a role in the origin of some wild-to-weed species, although the wild ancestors of weeds tend to be understudied, making conclusive inferences about the mechanisms of weed colonization difficult. One better-studied example of adaptive wild colonization involves the evolution of crop mimicry, the ability to escape human removal by physically resembling the crop in which weeds grow. Phenotypic variation in the progenitors of these weeds could be the source of some mimicry traits early in establishment; however, high levels of specialization have restricted many crop mimics to very specific agricultural settings (Barrett, 1983).
Barnyardgrass (Echinocloa oryzicola; Figure 2) is the best characterized crop mimic owing to its worldwide impact on rice yield (Smith, 1988), which makes it one of the costliest of all the agricultural weeds. Despite this impact, there is little genetic information about its evolutionary history, although recent studies have better resolved phylogenetic relationships within Echinochloa. Once thought to be conspecific with E. oryzicola, E. crus-galli (a separate hexaploid weed species) is now thought to have arisen through the hybridization of E. oryzicola (tetraploid) and an unknown diploid species (Aoki and Yamaguchi, 2008). E. crus-galli is also a successful weed of several crops but functions as a generalist, whereas E. oryzicola has adapted to mimic rice in morphology and phenology. Traits such as large seed size and poor seed dormancy prevent establishment outside of the rice field, suggesting that these traits are adaptive specifically in the agricultural setting (Barrett and Wilson, 1983). In addition, some biotypes possess traits that seem suited to particular farming practices, such as the ability to germinate under water (Smith, 1988). Barnyardgrass has congeneric crops including Indian and Japanese barnyard millet (E. frumentacea and E. esculenta, respectively) and a conspecific minor crop in China called Mosou barnyard millet, all of which could contribute alleles to the weed. However, ploidy differences between these groups suggest that gene flow from cultivars may not have a role in the weed’s evolution.
Some representative agricultural weeds. Clockwise from top left: barnyardgrass in a cultivated rice field (Katie Hyma), waterhemp infesting soybeans (Kate Waselkov), weedy rye infesting cultivated wheat (Jutta Burger), weedy sunflower in a maize field (Nolan Kane), morning glory (Michael Fethe), Conyza canadensis (horseweed) invading cultivated maize (Michael Fethe), black hull awned weedy rice in a rice field (Katie Hyma), California wild radish (Norman Ellstrand).
Another example of adaptive evolution enabling wild plants to colonize the agricultural environment is the evolution of herbicide resistance. This strategy is well illustrated by the weedy Amaranthus species, collectively known as pigweeds. Several Amaranthus crops are widely grown in Central and South America; however, these grain amaranths are genetically distinct from the weedy species, which are believed to have evolved from wild populations (Maughan et al., 2011). A striking example of recent weed invasion is that of waterhemp (A. tuberculatus) in corn and soybean fields (Figure 2) in the central United States over the past two decades (Steckel, 2007). Two unresolved hypotheses have been put forth for this phenomenon: (1) changes in weed management such as reduced tilling and reliance on post-emergence herbicides have favored rapid adaptive evolution by selection on standing genetic variation and/or de novo mutations (Costea et al., 2005), and (2) hybridization between waterhemp and other weedy Amaranthus species has resulted in the acquisition of genes allowing for successful invasion (Trucco et al., 2009). Weedy amaranths will likely continue to be problematic given their ability to rapidly evolve herbicide resistance (discussed below), produce an extraordinary number of seeds and compete for space and nutrients in the field. The evolution of herbicide resistance in wild-to-weed plants has also been reported for Conyza canadensis (Zheng et al., 2011; Figure 2), Lolium rigidum (Collavo and Sattin, 2012) and barnyardgrass (Hoagland et al., 2004; Talbert and Burgos, 2007), among others.
Hybrid to weed
Hybridization of crops with wild relatives may result in genetic variants that have traits already adaptive in the agricultural field (those inherited from the crop parent) and traits that favor proliferation and persistence (those inherited from the wild parent), thereby creating a weedy plant. Hybridization can lead to establishment of crop-wild-weed complexes; the possibility of ongoing gene flow in these complexes can increase genome-wide genetic diversity, complicating predictions about the extent of bottlenecks and the major colonization strategies (that is, the roles of preadaptation or adaptive evolution). The expectation is that hybrid weeds may experience a population bottleneck during establishment, but may quickly recover if gene flow with wild and crop relatives persists (Figure 1). However, distinguishing ancient hybridization from ongoing gene flow is challenging, as is differentiating between genetic contributions from crop and wild species when both share ancestral polymorphisms. Examples of weeds with well-documented hybrid origins are described below.
The evolution of weedy sunflowers (Helianthus annuus) is well studied in several world regions. Sunflowers are native to North America, where the crop is often grown in proximity to wild Helianthus populations. Weedy sunflower strains in European fields most likely originated in seed plots in the US, where nearby populations of wild sunflower contributed pollen to cultivated sunflowers, producing weedy hybrids (Muller et al., 2011). These hybrid seeds were likely exported with cultivated seeds to Europe and established as weeds in the field. Genetic diversity is higher in European populations of weedy sunflower than in the populations of cultivated sunflower, but lower than the populations of US wild sunflower, indicating that the hybrid weeds underwent a small genetic bottleneck during establishment. Transcriptomics has revealed that weedy and cultivated sunflowers from Australia and Israel also show evidence of crop/weed introgression (Lai et al., 2012), indicating that hybridization is not uncommon in this group. Weedy sunflower populations in the United States (Figure 2) have been found to be more closely related to the locally occurring wild populations (Kane and Rieseberg, 2008), suggesting multiple direct wild-to-weed descent events, although ongoing gene flow between local wild and weedy populations cannot be completely ruled out. Regardless, the distinct origins of the United States vs old world weeds indicates that there are multiple routes to weediness in the sunflower group. Untangling the relative genomic input from wild and cultivated sunflower within the known hybrid weedy populations may shed light on which genes drive the emergence of weedy traits inherited from each parent in this group.
Cultivated, wild and weedy beets (Beta vulgaris) are phylogenetically complicated due to shared ancestry and hybridization. Weedy beets arose from hybridization between cultivated beets grown for seed production and wild inland beets, as indicated by microsatellite and chloroplast markers (Viard et al., 2002; Fénart et al., 2008). Seed beets are presumed to have been pollinated by wild beets and then planted in sugar beet fields. The weeds persist by undergoing self-fertilization (a crop trait), flowering the first year of cultivation (a wild trait), bolting early (a wild trait), and maintaining high levels of seed dormancy (a wild trait), reflecting a mixture of genetic input from wild and cultivated beets (Arnaud et al., 2010). Weedy beet populations do not show evidence of a recent genetic bottleneck, with similar levels of genetic diversity as wild beet populations (Fénart et al., 2008). However, weedy beet populations do show strong spatial structuring, likely due to seed dispersal by gravity and limited pollen flow among weedy populations (Arnaud et al., 2011).
There are several instances where weed origin is unclear, but where crop/weed hybridization currently occurs. These include two weedy sorghum species, johnsongrass (Sorghum halepense) and shattercane (S. bicolor). Johnsongrass (2n=40), a perennial grass that infests several crops and natural areas, can hybridize with cultivated sorghum (S. bicolor, 2n=20). However, it is not clear whether the weed’s origin was contingent on this hybridization event. Despite differences in ploidy, genetic introgression from the crop to the weed (through pollen) has been reported in weedy populations (Morrell et al., 2005). This introgression is thought to have contributed to the northward expansion of the weed in North America, providing the genetic variation necessary for a change in the weed’s phenology to adapt to colder winters (Warwick et al., 1986). Crop alleles have been reported in weed populations with no recent exposure to cultivated sorghum, indicating that introgressed alleles or weeds themselves have moved across large geographic distances.
Shattercane (S. bicolor, 2n=20) closely resembles the conspecific cultivated sorghum, although it differs from the crop in the ease with which it disperses seeds and its competitive ability, particularly in cornfields (Beckett et al., 1988). Shattercane is an annual plant whose distribution seems limited to regions of the world where sorghum is grown; its origin is unclear (Defelice, 2006), but is often described as a hybrid between cultivated sorghum and various wild relatives (Ejeta and Grenier, 2005). It is clear that shattercane can hybridize with grain sorghum, with introgression moving in the direction of crop to weed, potentially leading to the spread of alleles advantageous in agricultural environments (Ejeta and Grenier, 2005). There are several more weedy sorghum species, but the exact phylogeny has yet to be sorted out (Ejeta and Grenier, 2005), which impedes progress in understanding the origin and adaptation of these weedy species.
Crop to weed
Though suspected to be a common route to weed emergence, there are few verified examples of crops where volunteer strains have directly evolved into competitive weeds (but see Ellstrand et al., 2010). Weeds evolving directly from crops are evolutionarily intriguing, as they are expected to be more genetically depauperate than other weeds (Figure 1), and sources of adaptive traits should be limited to ancestral standing variation in the crop or new mutations. However, whether such weedy species can arise in the absence of any sort of genetic input from other wild groups is an open question. The best-known examples of crop origins of weedy plants include weedy rye (Secale cereale) and weedy rice (Oryza sativa), both discussed below. Although there are several other weedy species that have crop relatives, such as the sorghum relatives described above, data showing that the weed arose directly from the crop have not been produced. It is often difficult to distinguish hybrid from crop origins as shared alleles between weeds and crops or wild plants can be due to shared ancestry or introgression. To unravel these complicated phylogenies, genome-wide markers are needed for weed, crop and wild relatives. As these data become increasingly available, it is clear that untangling complex histories of adaptation and gene flow will require further improvement on our theoretical and simulation framework (Strasburg et al., 2012).
Weedy rye (Figure 2), which is particularly problematic in wheat fields, seems to have originated in cultivated rye fields in California (Burger et al., 2006). Volunteer rye has been reported as an agricultural weed in many locations; weedy rye differs from crop volunteers in that it has evolved several distinct traits, including seed shattering, smaller seeds and a delay in flowering time (Burger et al., 2007). Remarkably, these differences have evolved in only about 60 generations, the length of time rye has been cultivated in California. It is possible that flowering time differences evolved first, establishing a reproductive isolating barrier between weed and crop, and allowing for other differences to accumulate quickly in this group. Weedy rye does not show reduced diversity at allozyme and microsatellite markers when compared to its domesticated progenitors, indicating that it did not undergo a detectable genetic bottleneck in the populations studied (Burger et al., 2006). This finding stands in contrast to what might be expected for a recent weed origin from a crop species (Figure 1). An interesting feature of this system is that cultivated rye is itself thought to have originated through domestication of what was originally a weed of wheat and barley fields in Southwest Asia (Zohary, 1971). This implies that rye has evolved from wild to weed to crop to weed, a complex history indeed.
Weedy rice (Figure 2) is an obligate weed of rice worldwide, but populations in different regions of the world are thought to have independent evolutionary origins. Weedy rice in the United States has at least two distinct origins, resulting in two genetically distinct biotypes (Londo and Schaal, 2007; Reagon et al., 2010). Sequence data suggest that the two biotypes have descended from different cultivated Asian rice varieties (Reagon et al., 2010). These de-domestication events likely occurred after the non-shattering seed phenotype was fixed in domesticated Asian rice (discussed below), but before rice cultivation was introduced to the United States (Thurber et al., 2010). Coalescent and Bayesian simulations indicate that the timing of the de-domestication events differs for the two biotypes; however, both exhibit the genetic effects of strong population bottlenecks (Reagon et al., 2010), as expected for weeds descended from crops (Figure 1). Despite the fact that both cultivated and weedy rice are predominantly self-fertilizing, hybridization occasionally occurs, potentially allowing for adaptive traits such as herbicide resistance to be passed to the weed and between the two biotypes (Shivrain et al., 2009; Reagon et al., 2010, 2011). The independent evolution of two weed biotypes in the United States offers a promising opportunity to study the genetics of parallel adaptive evolution.
Beyond the United States, genetic and morphological data for weedy rice populations in some other parts of the world also suggest a de-domesticated origin, including weed strains in China (Cao et al., 2006), Japan (Akasaka et al., 2009) and Bhutan (Ishikawa et al., 2005). There is also evidence that not all weedy rice populations are necessarily descended directly from cultivated ancestors. In regions of the world where the wild progenitor of cultivated rice (O. rufipogon) persists, there is potential for wild rice to hybridize with weedy and cultivated rice (Majumder et al., 1997). Genomic data from cultivated, weedy and wild rice in these regions will be useful for determining the origin and level of gene flow between these groups.
The genetic basis of weed adaptation
From an evolutionary perspective, agricultural weeds capture our attention precisely because of their intrinsic capacity for rapid adaptation. Whether weedy phenotypes represent pre-adaptations or traits acquired during the colonization process, all agricultural weeds must possess traits that permit them to thrive in this recently created environment. Adaptive traits enabling agricultural weeds to survive and thrive vary from species to species, but many are familiar to us as weed-diagnostic traits that favor competitive ability and persistence in the agricultural setting. This set of traits, or the ‘agricultural weed syndrome’, includes rapid growth, high nutrient use efficiency, seed dormancy, efficient seed dispersal, crop mimicry and herbicide resistance. Longstanding questions about adaptation particularly applicable to agricultural weeds include the following: (1) What are the relative roles of new mutations, standing variation and introgression in the evolution of weed adaptive traits? (2) At the molecular level, are particular mutation types most often responsible for weedy adaptation? (3) What are the genes that underlie convergent evolution of weediness in different weed species or in different populations of one species? (4) In the arms race with humans, selective pressures have changed through weed history; can we detect the signatures of old vs new selection in weed genomes?
Understanding weed adaptation is currently hindered by our incomplete knowledge of the genetic basis of many weedy traits. Even in the few cases where the genetic basis of a weedy trait is well understood, genetic information from both wild and crop relatives is needed to distinguish ancestral (standing) genetic variation from adaptations evolving through de novo mutation in weed populations. Given both the economic consequences of weedy traits and the evolutionary appeal of the system, multiple efforts are underway to dissect the genetic basis of weed adaptation. Below, we discuss our current understanding of adaptive trait evolution in agricultural weeds.
Origins of adaptive traits
The origin of mutations underlying adaptive traits in organisms has generally been ascribed to one of the three possibilities: standing variation, introgression or new mutation. These are all possible sources of adaptive traits in agricultural weeds regardless of the weed’s primary phylogenetic origin. Characterization of the genetic provenance of weedy traits is perhaps most advanced in weedy rice, due to the genomic resources available. Genetic evidence, most of which comes from US weedy rice, points to multiple sources of weed-adaptive traits. We discuss examples of standing variation, new mutation and introgression contributing to weedy rice traits below, followed by examples from other weed species.
Proanthocyanidin pigmentation, resulting in dark reddish-brown color of the pericarp (bran), is one of the most obvious phenotypic characteristics of weedy rice (and the source of the weed’s common name, red rice). This trait is also characteristic of wild rice (O. rufipogon) and was selected against during crop improvement, leading to most modern domesticated rice cultivars lacking pericarp pigmentation, although some traditional landraces retain the ancestral trait. Loss of function mutations at the Rc gene, which encodes a regulatory protein in proanthocyanidin synthesis, are responsible for this transition from red to white pericarps in most domesticated rice (Sweeney et al., 2006). Clustering of weed Rc alleles with crop alleles of various red-pigmented varieties, both in US and Asian weeds (Gross et al., 2010; Gu et al., 2011), suggests that weeds have inherited the red pericarp phenotype from standing variation. Pericarp pigmentation likely has a direct role in seed dormancy (Gu et al., 2011), an important adaptive trait for weediness.
The origin of weed alleles is less clear for the shattering phenotype (seed dispersal), a trait that is characteristic of wild and weedy rice and was strongly selected against during domestication. The US, weedy rice strains freely disperse their seeds, whereas their putative Asian domesticated progenitors have reduced shattering (Thurber et al., 2010). Studies of sh4, a major gene implicated in loss of shattering during rice domestication, have revealed that weeds carry the derived reduced-function alleles that are fixed in the crop (Thurber et al., 2010). Thus, inheritance of an ancestral functional wild allele is not responsible for shattering in US weedy rice, an observation supported by developmental difference in the shattering phenotype between weeds and wild rice (Thurber et al., 2011). Instead, it appears that the US weeds have re-evolved shattering through sh4 mutational suppression involving an unknown genetic mechanism. Seed shattering is also characteristic of weedy rice that evolved from a cultivar in Japan, which also shares the sh4 domestication allele with cultivars (Akasaka et al., 2011). As genes other than sh4 are involved in the origin of shattering in US and Japanese weeds, novel mutations remain the most likely source for this adaptive trait. However, it is not possible to completely rule out standing variation in the diverse crop ancestor; although cultivated rice does not generally shatter, the range of variation in the crop is large (Thurber et al., 2010), and low-frequency shattering crop variants could exist.
Another likely example of novel mutations giving rise to adaptive weed phenotypes involves the evolution of response to novel temperature cues to avoid untimely germination in Chinese weedy rice, rather than reversion to the ancestral dormancy behavior typical of wild rice (Xia et al., 2011). Specifically, weedy rice collected from northern latitudes was found to germinate at a lower temperature after cold treatment than did weedy rice collected from more southern latitudes, whereas cultivated rice from all sampled regions germinated at the same temperature. Therefore, critical temperature cues for seed germination in Chinese weedy rice appear to correlate with local temperature and latitudinal gradients across the range (Xia et al., 2011). These traits are diverged from rice varieties cultivated in the region, which are thought to be the progenitors of the locally occurring weeds based on simple sequence repeat markers (Cao et al., 2006). As of yet, the gene(s) underlying this potentially adaptive trait are not known.
Introgression has been implicated as a source of adaptive weed traits in numerous cases, but genetic evidence (that is, identifying the introgressed allele that underlies a weedy trait) is often sparse. A subset of US weedy rice has been shown to possess alleles of the growth-related gene, SD1, introgressed from local rice cultivars, which are not implicated in the phylogenetic origin of the US weeds. These weeds are shorter, slower growing and earlier flowering than weeds that lack the introgressed allele (Reagon et al., 2011). In this case, the evidence for introgression is clear, but there is no proof yet that this phenotype is adaptive. It is possible that short stature and earlier flowering could be traits that are adaptive for crop mimicry, allowing weeds to escape eradication. Following the introduction of herbicide-resistant rice to US agriculture in the last decade, the emergence of some herbicide-resistant weed strains similarly appears to be linked to crop-weed hybridization (Shivrain et al., 2009).
For weeds that are products of hybridization events, the role of introgression in weed-adaptive phenotypes is often implicitly assumed. Identification of genes underlying adaptive traits in weeds can potentially reveal whether crop or wild alleles are contributing more to weed adaptation, and the extent to which genomic background and novel gene combinations drive weed adaptation. A possible example of this last category occurs in California wild radish (Figure 2), a weedy product of cultivated radish (Raphanus sativus) and the wild species jointed charlock (R. raphanistrum) (Hegde et al., 2006). California wild radish is notable in that it behaves as both an agricultural weed and occupant of natural areas. It displays higher fitness than both of its progenitors as determined by average fruit weight, rate of survival and reproductive output when grown in common gardens (Ridley and Ellstrand, 2009). These traits were likely acquired from the hybridization event itself; however, as the weeds are obligate outcrossers, heterosis, rather than particular wild or crop alleles, could also provide important contributions to weed fitness (Ridley and Ellstrand, 2009). This has been further supported by the construction of artificial radish hybrids which show elevated fitness in common garden experiments (Campbell et al., 2006).
For plant species that are not crop relatives, the emergence of a weedy life history in crop fields may or may not be due to recent de novo adaptive mutations. However, standing variation is believed to be the more likely source of adaptive traits, implying that these wild plants are pre-adapted to agricultural weediness. Unfortunately, there is usually too little information about the status of such agricultural weeds in their native (nonagricultural) environments to allow the determination of which mechanism is most prevalent. Some examples of each of these mechanisms are described below.
Standing variation seems to be involved in the success of Orobanche and Striga parasitic weeds, as described above. In addition, species that are naturally adapted to disturbed habitats may be pre-adapted to the agricultural environment. This form of pre-adaptation may be the case for pigweed amaranths, including waterhemp (A. tuberculatus) and Palmer amaranth (A. palmeri). On the other hand, the fact that amaranths have become major agricultural weeds only following the widespread use of herbicides suggests that the invasion of crop fields has occurred at least in part through recent adaptive evolution, in this case, through evolution of resistance against multiple classes of herbicide. Standing variation or pre-adaptation seems to be responsible for herbicide-resistant weedy ryegrass (Lolium rigidum) in fields that were never exposed to herbicides (Neve and Powles, 2005). This is also the case for morning glory (Ipomoea purpurea) populations that exhibit both tolerance and resistance in fields that were never exposed to glyphosate (Baucom and Mauricio, 2010). Standing variation has been implicated in some wild-to-weed plants that have crop relatives. For example, in a microsatellite study of the US weedy sunflowers, six ‘outlier’ loci that may have experienced selective sweeps were shared among populations (Kane and Rieseberg, 2008). As each of these populations is thought to have evolved independently from local wild populations, and there is no loss of diversity in weed populations, this suggests that genes necessary for adaptation in agricultural systems exist in the wild (Kane and Rieseberg, 2008).
In contrast to the above examples, the rapid evolution of herbicide resistance in weeds such as barnyardgrass (E. crus-galli and E. oryzicola) may largely be due to novel, adaptive mutations. Barnyardgrass populations from different world regions have evolved resistance to at least nine different mode of action herbicides via independent mutations (Hoagland et al., 2004; Talbert and Burgos, 2007). The occurrence of these independent mutations as well as the resistance to multiple herbicides within some single populations (Malik et al., 2010) suggests that these traits come from new mutations. Herbicide resistance has likely evolved de novo in other weeds due to the fitness cost associated with resistance when herbicide selection is not present. Examples include waterhemp (Duff et al., 2009) and foxtail grass (Darmency et al., 2011).
Mutation types
Although inferring trait origins is sometimes possible without knowledge of the exact gene underlying the trait, this is not true for assessment of mutation types and genetic convergence. Unfortunately, we do not yet have an extensive list of weed-adaptive genes that we can use to assess evolutionary patterns. Much debate in evolutionary biology has been framed around which types of mutations are more likely to underlie phenotypic and, in particular, adaptive change (for example, Hoekstra and Coyne, 2007; Stern and Orgogozo, 2008). Arguments have been put forth for structural (that is, changing protein sequence) and regulatory mutations (that is, changing gene expression). Parallel discussions on the types of genes involved in crop domestication have suggested that morphological and developmental changes are more likely to involve changes in transcriptional regulation, whereas biochemical and other metabolic changes are more likely to involve loss of function mutations at individual structural genes (Doebley et al., 2006). We suspect the same may be true for genes underlying weed-adaptive traits; however exceptions likely abound, as seen in herbicide resistance mutations discussed below. Additional discussion concerning mutation types has centered on the extent to which convergent evolution in different groups occurs through similar or different genetic mechanisms.
The most studied weed-adaptation genes are those that confer herbicide resistance, as gene identification is facilitated by knowledge of the metabolic targets of many herbicides. Typically, these genes are involved in metabolic pathways that are essential for plant function. Therefore, instead of loss of function mutations, we find mutations that alter the gene (so that the herbicide cannot bind) but maintain function. The mutation types identified so far that result in herbicide resistance in weeds are mostly structural, and include amino acid substitutions, copy number changes and deletions within the gene.
The most common mutations leading to herbicide resistance are amino acid substitutions in the target of the herbicide. One example is the herbicide that targets acetolactate synthase (ALS), also called acetohydroxyacid synthase, an enzyme involved in amino acid biosynthesis. Resistance to ALS herbicides is widespread, with 125 known resistant weedy species to date (Heap, 2012). Twenty-two different amino acid substitutions at seven sites in the ALS gene have been shown to cause resistance across multiple species. Similar findings have been found for herbicides that target acetyl-coenzyme A carboxylase, an enzyme needed for lipid metabolism (reviewed in Powles and Yu, 2010).
Palmer amaranth (A. palmeri) populations in the US have evolved resistance to several different mode of action herbicides, including glyphosate (Gaines et al., 2010; Heap, 2012). Glyphosate inhibits the enzyme 5-enolpyruvyl-shikimate-3-phosphate synthase (EPSPS). Resistance has been attributed to transposon-mediated gene amplification of EPSPS, which increases EPSPS expression, and presumably provides an excess of the enzyme, which cannot be targeted by the herbicide (Gaines et al., 2010). Between 30 and 50 copies of EPSPS are needed to survive glyphosate rates between 0.5 and 1 kg per ha (Gaines et al., 2011). Resistance first occurred in 2005 in Georgia and has rapidly spread to 11 other states (Heap, 2012). It is not clear whether resistance evolved in the years since glyphosate has been applied or stems from standing variation for gene copy number in the weed population.
Protoxin resistance in waterhemp (A. tuberculatus) occurs from a very unusual codon deletion in PPX2L (Patzoldt et al., 2006). This gene encodes PPO, an enzyme involved in chlorophyll production. The deletion may have occurred multiple times independently in this group, based on polymorphism in the alleles with the deletion (Lee et al., 2008). Resistance to protoxin is uncommon as most plants contain two gene targets, PPX1 and PPX2, localized to the chloroplast and mitochondria respectively, resulting in two sites of action for this herbicide. PPX2L seems to replace the function of the two gene copies, with resistant plants lacking a second gene copy. Susceptible waterhemp plants, on the other hand, have a second PPX2 copy (Patzoldt et al., 2006; Lee et al., 2008). Therefore, resistance to protoxin may be a result of a loss of a gene copy and a codon deletion in the remaining copy.
Herbicide resistance in weeds is a good example of convergent evolution where both similar and different mutations have been implicated in the convergence of the resistance phenotype among groups or species. For example, ALS is a gene repeatedly targeted in the evolution of ALS-herbicide resistance; however multiple mutations at several sites within the gene have been implicated in different weed species, as mentioned above. Conversely, in resistance to triazine herbicides that inhibit photosynthesis, which occurs in 43 different genera worldwide to date (Heap, 2012), the same amino acid substitution is responsible for independent evolution of resistance in all known cases (Gronwald, 1997). Given that there seems to be a common suite of traits that are adaptive for weediness in several species, such as seed shattering, rapid growth and seed dormancy, more opportunities for the study of the genetic basis of weed convergence exist.
Weed evolutionary genetics: the path ahead
The impact of agricultural weeds on humans is immense. Beyond the economic costs caused by crop losses, once weeds are established in agricultural fields, their evolutionary dynamics begin to affect crops. In order to control weeds, humans breed crops and make genetic modifications that allow for the use of herbicides or other methods of control. Weeds then adapt to the changing practices, forcing humans to make further modifications, perpetuating the cycle. Crop−weed interactions offer a view of contemporary rapid evolution, a great example of the arms race between species (Neve et al., 2009). Weeds can also be exploited as crops in regions where nutrients and water limit agricultural potential. For example, the flax mimic, Camelina sativa or false flax, has been cultivated as an oilseed crop since the Bronze age with recent renewed interest as a biofuel crop (Hutcheon et al., 2010). Whether false flax was a weed that was brought into cultivation or a crop that became weedy is not clear, but traits such as drought tolerance and low nutrient input make it both a competitive weed and an easy crop (Putnam et al., 1993).
The combination of agricultural importance and unique evolutionary dynamics make agricultural crops a system ripe for the development as a model for studying adaptation and evolution. As genomic tools become more affordable and accessible even for species that are not laboratory model systems, investing in the development of genetic resources for weeds promises to yield great returns at the level of basic science and practical applications. Such calls for genetic and genomic resource development for weed species have already been made (Stewart et al., 2009), and data are starting to accumulate for some weedy species (for example, Anderson et al., 2007; Lee et al., 2009; Parasitic Plant Genome Project at http://ppgp.huck.psu.edu). However, few weedy species have been well characterized genetically. Weedy rice likely is the best genetically characterized weed, but this is due to characterization of its important crop relative. As we move into the genomics era, new tools to study the genetic basis for adaptive evolution such as whole genome sequencing, genome−wide association studies, nested association mapping and many more will allow for a better understanding of the world’s most problematic weeds. Ralph Waldo Emerson wrote that a ‘weed is a plant whose virtues have not yet been discovered’; we suggest that a weed’s greatest virtue is its ability to adapt.
Data archiving
There were no data to deposit.
References
Akasaka M, Konishi S, Izawa T, Ushiki J (2011). Histological and genetic characteristics associated with the seed-shattering habit of weedy rice (Oryza sativa L.) from Okayama, Japan. Breed Sci 61: 168–173.
Akasaka M, Ushiki J, Iwata H, Ishikawa R, Ishii T (2009). Genetic relationships and diversity of weedy rice (Oryza sativa L.) and cultivated rice varieties in Okayama Prefecture, Japan. Breed Sci 59: 401–409.
Ali RAMA, El-Hussein AA, Mohamed KI, Babiker AGT (2009). Specificity and genetic relatedness among Striga hermonthica strains in Sudan. Int J Life Sci 3: 1159–1166.
Anderson JV, Horvath DP, Chao WS, Foley ME, Hernandez AG, Thimmapuram J et al (2007). Characterization of an EST database for the perennial weed leafy spurge: an important resource for weed biology research. Weed Sci 55: 193–203.
Aoki D, Yamaguchi H (2008). Genetic relationship between Echinochloa crus-galli and Echinochloa oryzicola accessions inferred from internal transcribed spacer and chloroplast DNA sequences. Weed Biol Manag 8: 233–242.
Arnaud J-F, Cuguen J, Fénart S (2011). Metapopulation structure and fine-scaled genetic structuring in crop-wild hybrid weed beets. Heredity 107: 395–404.
Arnaud J-F, Fénart S, Cordellier M, Cuguen J (2010). Populations of weedy crop-wild hybrid beets show contrasting variation in mating system and population genetic structure. Evol Appl 3: 305–318.
Baker HG (1965). Characteristics and modes of origin of weeds. In: Baker HG, Stebbins GL, (eds). The Genetics of Colonizing Species. Academic Press: New York. pp 147–172.
Barrett SCH (1983). Crop mimicry in weeds. Econ Bot 37: 255–282.
Barrett SCH, Wilson BF (1983). Colonizing ability in the Echinochloa crus-galli complex (barnyard grass). II. Seed biology. Can J Bot 61: 556–562.
Baucom RS, Mauricio R (2010). Defence against the herbicide RoundUp predates its widespread use. Evol Ecol Res 12: 131–141.
Beckett TH, Stoller EW, Wax LM (1988). Interference of four annual weeds in corn (Zea mays). Weed Sci 36: 764–769.
Burger JC, Holt JM, Ellstrand NC (2007). Rapid phenotypic divergence of feral rye from domesticated cereal rye. Weed Sci 55: 204–211.
Burger JC, Lee S, Ellstrand NC (2006). Origin and genetic structure of feral rye in the western United States. Mol Ecol 15: 2527–2539.
Campbell LG, Snow AA, Ridley CE (2006). Weed evolution after crop gene introgression: greater survival and fecundity of hybrids in a new environment. Ecol Lett 9: 1198–1209.
Cao Q, Lu B-R, Xia H, Rong J, Sala F, Spada A et al (2006). Genetic diversity and origin of weedy rice (Oryza sativa f. spontanea) populations found in North-eastern China revealed by simple sequence repeat (SSR) markers. Ann Bot 98: 1241–1252.
Clements DR, DiTommaso A, Jordan N, Booth BD, Cardina J, Doohan D et al (2004). Adaptability of plants invading North American cropland. Agric Ecosyst Environ 104: 379–398.
Collavo A, Sattin M (2012). Resistance to glyphosate in Lolium rigidum selected in Italian perennial crops: bioevaluation, management and molecular bases of target-site resistance. Weed Res 52: 16–24.
Costea M, Weaver SE, Tardif FJ (2005). The biology of invasive alien plants in Canada. 3. Amaranthus tuberculatus (Moq.) Sauer var. rudis (Sauer) Costea & Tardif. Can J Plant Sci 85: 507–522.
Darmency H, Picard JC, Wang T (2011). Fitness costs linked to dinitroaniline resistance mutation in Setaria. Heredity 107: 80–86.
Darwin CR (1859) On the Origin of Species. John Murray: London.
Dawoud DA, Sauerborn J (1994). Impact of drought stress and temperature on the parasitic weeds Striga hermonthica and Alectra vogelii in their early growth stages. Exp Agr 30: 249–257.
De Wet JMJ, Harlan JR (1975). Weeds and domesticates: evolution in the man-made habitat. Econ Bot 29: 99–107.
Defelice MS (2006). Shattercane, Sorghum bicolor (L.) Moench Ssp. Drummondii (nees ex steud.) de wet ex davidse—black sheep of the family. Weed Technol 20: 1076–1083.
Délye C, Clément JAJ, Pernin F, Chauvel B, Le Corre V (2010). High gene flow promotes the genetic homogeneity of arable weed populations at the landscape level. Basic Appl Ecol 11: 504–512.
Doebley JF, Gaut BS, Smith BD (2006). The molecular genetics of crop domestication. Cell 127: 1309–1321.
Duff MG, Al-Khatib K, Peterson DE (2009). Relative competitiveness of protoporphyrinogen oxidase-resistant common waterhemp (Amaranthus rudis). Weed Sci 57: 169–174.
Ejeta G, Grenier C (2005). Sorghum and its weedy hybrids. In: Gressel J, (ed.). Crop Ferality and Volunteerism. CRC Press: Boca Raton. pp 123–135.
Ellstrand NC, Heredia SM, Leak-Garcia JA, Heraty JM, Burger JC, Yao L et al (2010). Crops gone wild: evolution of weeds and invasives from domesticated ancestors. Evol Appl 3: 494–504.
Fénart S, Arnaud J-F, De Cauwer I, Cuguen J (2008). Nuclear and cytoplasmic genetic diversity in weed beet and sugar beet accessions compared to wild relatives: new insights into the genetic relationships within the Beta vulgaris complex species. Theor Appl Genet 116: 1063–1077.
Gaines T, Shaner DL, Ward SM, Leach JE, Preston C, Westra P (2011). Mechanism of resistance of evolved glyphosate-resistant Palmer amaranth (Amaranthus palmeri). J Agric Food Chem 59: 5886–5889.
Gaines T, Zhang W, Wang D, Bukun B, Chisholm ST, Shaner DL et al (2010). Gene amplification confers glyphosate resistance in Amaranthus palmeri. Proc Natl Acad Sci USA 107: 1029–1034.
Gressel JB (2005) Crop Ferality and Volunteerism. CRC Press: Boca Raton.
Gronwald JW (1997). Resistance to PS II inhibitor herbicides. In: De Prado R, Jorrin J, Garcia-Torres L, (eds). Weed and Crop Resistance to Herbicides. Kluwer: Dordrecht. pp 53–59.
Gross BL, Reagon M, Hsu S-C, Caicedo AL, Jia Y, Olsen KM (2010). Seeing red: the origin of grain pigmentation in US weedy rice. Mol Ecol 19: 3380–3393.
Gu X-Y, Foley ME, Horvath DP, Anderson JV, Feng J, Zhang L et al (2011). Association between seed dormancy and pericarp color is controlled by a pleiotropic gene that regulates abscisic acid and flavonoid synthesis in weedy red rice. Genetics 189: 1515–1524.
Heap I (2012). International survey of herbicide resistant weeds. http://www.weedscience.org/In.asp (Accessed May 25, 2012).
Hegde SG, Nason JD, Clegg JM, Ellstrand NC (2006). The evolution of California’s wild radish has resulted in the extinction of its progenitors. Evolution 60: 1187–1197.
Hoagland RE, Norsworthy JK, Carey F, Talbert RE (2004). Metabolically based resistance to the herbicide propanil in Echinochloa species. Weed Sci 52: 475–486.
Hoekstra HE, Coyne JA (2007). The locus of evolution: evo devo and the genetics of adaptation. Evolution 61: 995–1016.
Holm LG, Plucknett DL, Pancho JV, Herbeger JP (1977) World’s Worst Weeds. Distribution and Biology. University of Hawaii: Honolulu.
Hutcheon C, Ditt RF, Beilstein M, Comai L, Schroeder J, Goldstein E et al (2010). Polyploid genome of Camelina sativa revealed by isolation of fatty acid synthesis genes. BMC Plant Biol 10: 233.
Ishikawa R, Toki N, Imai K, Sato YI, Yamagishi H, Shimamoto Y et al (2005). Origin of weedy rice grown in Bhutan and the force of genetic diversity. Genet Resour Crop Evol 52: 395–403.
Kane NC, Rieseberg LH (2008). Genetics and evolution of weedy Helianthus annuus populations: adaptation of an agricultural weed. Mol Ecol 17: 384–394.
Lai J, Li R, Xu X, Jin W, Xu M, Zhao H et al (2010). Genome-wide patterns of genetic variation among elite maize inbred lines. Nat Genet 42: 1027–1030.
Lai Z, Kane NC, Kozik A, Hodgins K, Dlugosch KM, Barker MS et al (2012). Genomics of Compositae weeds: EST libraries, microarrays, and evidence of introgression. Am J Bot 99: 209–218.
Lam H-M, Xu X, Liu X, Chen W, Yang G, Wong F-L et al (2010). Resequencing of 31 wild and cultivated soybean genomes identifies patterns of genetic diversity and selection. Nat Genet 42: 1053–1059.
Lee RM, Hager AG, Tranel PJ (2008). Prevalence of a novel resistance mechanism to PPO-inhibiting herbicides in waterhemp (Amaranthus tuberculatus). Weed Sci 56: 371–375.
Lee RM, Thimmapuram J, Thinglum KA, Gong G, Hernandez AG, Wright CL et al (2009). Sampling the waterhemp (Amaranthus tuberculatus) genome using pyrosequencing technology. Weed Sci 57: 463–469.
Londo JP, Schaal B (2007). Origins and population genetics of weedy red rice in the USA. Mol Ecol 16: 4523–4535.
Majumder ND, Ram T, Sharma AC (1997). Cytological and morphological variation in hybrid swarms and introgressed population of interspecific hybrids (Oryza rufipogon Griff. Oryza sativa L.) and its impact on evolution of intermediate types. Euphytica 94: 295–302.
Malik MS, Burgos NR, Talbert RE (2010). Confirmation and control of propanil-resistant and quinclorac-resistant barnyardgrass (Echinochloa crus-galli) in rice. Weed Technol 24: 226–233.
Maughan P, Smith S, Fairbanks D, Jellen E (2011). Development, characterization, and linkage mapping of single nucleotide polymorphisms in the grain amaranths (Amaranthus sp.). Plant Genome J 4: 92–101.
Menchari Y, Délye C, Le Corre V (2007). Genetic variation and population structure in black-grass (Alopecurus myosuroides Huds.), a successful, herbicide-resistant, annual grass weed of winter cereal fields. Mol Ecol 16: 3161–3172.
Monaco TJ, Weller SC, Ashton FM (2002) Weed Science: Principles and Practices. John Wiley and Sons: New York.
Morrell PL, Buckler ES, Ross-Ibarra J (2011). Crop genomics: advances and applications. Nat Rev Genet 13: 85–96.
Morrell PL, Williams-Coplin TD, Lattu AL, Bowers JE, Chandler JM, Paterson AH (2005). Crop-to-weed introgression has impacted allelic composition of johnsongrass populations with and without recent exposure to cultivated sorghum. Mol Ecol 14: 2143–2154.
Muller M-H, Latreille M, Tollon C (2011). The origin and evolution of a recent agricultural weed: population genetic diversity of weedy populations of sunflower (Helianthus annuus L.) in Spain and France. Evol Appl 4: 499–514.
Neve P, Powles S (2005). High survival frequencies at low herbicide use rates in populations of Lolium rigidum result in rapid evolution of herbicide resistance. Heredity 95: 485–492.
Neve P, Vila-Aiub M, Roux F (2009). Evolutionary-thinking in agricultural weed management. New Phytol 184: 783–793.
Oerke EC (2006). Crop losses to pests. J Agric Sci 144: 31.
Patzoldt WL, Hager AG, McCormick JS, Tranel PJ (2006). A codon deletion confers resistance to herbicides inhibiting protoporphyrinogen oxidase. Proc Natl Acad Sci USA 103: 12329–12334.
Pimentel D, Zuniga R, Morrison D (2005). Update on the environmental and economic costs associated with alien-invasive species in the United States. Ecol Econ 52: 273–288.
Powles SB, Yu Q (2010). Evolution in action: plants resistant to herbicides. Annu Rev Plant Biol 61: 317–347.
Putnam DH, Budin JT, Field LA, Breene WM (1993). Camelina: a promising low-input oilseed. In: Janick J, Simon JE, (eds). New Crops. Wiley: New York. pp 314–322.
Reagon M, Thurber CS, Gross BL, Olsen KM, Jia Y, Caicedo AL (2010). Genomic patterns of nucleotide diversity in divergent populations of US weedy rice. BMC Evol Biol 10: 180–196.
Reagon M, Thurber CS, Olsen KM, Jia Y, Caicedo AL (2011). The long and the short of it: SD1 polymorphism and the evolution of growth trait divergence in US weedy rice. Mol Ecol 20: 3743–3756.
Ridley CE, Ellstrand NC (2009). Evolution of enhanced reproduction in the hybrid-derived invasive, California wild radish (Raphanus sativus). Biol Invasions 11: 2251–2264.
Shivrain VK, Burgos NR, Sales MA, Mauromoustakos A, Gealy DR, Smith KL et al (2009). Factors affecting the outcrossing rate between ClearfieldTM rice and red rice (Oryza sativa). Weed Sci 57: 394–403.
Smith RJ (1988). Weed thresholds in southern US rice, Oryza sativa. Weed Technol 2: 232–241.
Steckel LE (2007). The dioecious Amaranthus spp.: here to stay. Weed Technol 21: 567–570.
Stern DL, Orgogozo V (2008). The loci of evolution: how predictable is genetic evolution? Evolution 62: 2155–2177.
Stewart CN, Tranel PJ, Horvath DP, Anderson JV, Rieseberg LH, Westwood JH et al (2009). Evolution of weediness and invasiveness: charting the course for weed genomics. Weed Sci 57: 451–462.
Strasburg JL, Sherman NA, Wright KM, Moyle LC, Willis JH, Rieseberg LH (2012). What can patterns of differentiation across plant genomes tell us about adaptation and speciation? Philos Trans R Soc Lond B Biol Sci 367: 364–373.
Sweeney MT, Thomson MJ, Pfeil BE, McCouch S (2006). Caught red-handed: Rc encodes a basic helix-loop-helix protein conditioning red pericarp in rice. Plant Cell 18: 283–294.
Talbert RE, Burgos NR (2007). History and management of herbicide-resistant barnyardgrass (Echinochloa crus-galli) in Arkansas rice. Weed Technol 21: 324–331.
Thurber CS, Hepler PK, Caicedo AL (2011). Timing is everything: early degradation of abscission layer is associated with increased seed shattering in US weedy rice. BMC Plant Biol 11: 14.
Thurber CS, Reagon M, Gross BL, Olsen KM, Jia Y, Caicedo AL (2010). Molecular evolution of shattering loci in US weedy rice. Mol Ecol 19: 3271–3284.
Trucco F, Tatum T, Rayburn AL, Tranel PJ (2009). Out of the swamp: unidirectional hybridization with weedy species may explain the prevalence of Amaranthus tuberculatus as a weed. New Phytol 184: 819–827.
Vaz Patto MC, Diaz-Ruiz R, Satovic Z, Roman B, Pujadas-Salva AJ, Rubiales DR (2008). Genetic diversity of Moroccan populations of Orobanche foetida: evolving from parasitising wild hosts to crop plants. Weed Res 48: 179–186.
Viard F, Bernard J, Desplanque B (2002). Populations of weedy crop-wild hybrid beets show contrasting variation in mating system and population genetic structure. Theor Appl Genet 104: 688–697.
Warwick SI, Phillips D, Andrews C (1986). Rhizome depth: the critical factor in winter survival of Sorghum halepense (L.) Pers. (johnson grass). Weed Res 26: 381–388.
Warwick SI, Stewart CN (2005). Crops come from wild plants—how domestication, transgenes, and linkage together shape ferality. In: Gressel JB, (ed.). Crop Ferality and Volunteerism. CRC Press: Boca Raton. pp 9–30.
Welsh AB, Mohamed KI (2011). Genetic diversity of Striga hermonthica populations in Ethiopia: evaluating the role of geography and host specificity in shaping population structure. Int J Plant Sci 172: 773–782.
Xia H-B, Xia H, Ellstrand NC, Yang C, Lu B-R (2011). Rapid evolutionary divergence and ecotypic diversification of germination behavior in weedy rice populations. New Phytol 191: 1119–1127.
Zheng D, Kruger GR, Singh S, Davis VM, Tranel PJ, Weller SC et al (2011). Cross-resistance of horseweed (Conyza canadensis) populations with three different ALS mutations. Pest Manag Sci 67: 1486–1492.
Zohary D (1971). Origin of south-west Asiatic cereals: wheats, barley, oats, and rye. In: Davis P, Harper P, Hedge I, (eds). Plant Life in South-West Asia. Royal Botanic Garden: Edinburgh. pp 235–263.
Acknowledgements
This study was funded in part by a grant from the US National Science Foundation Plant Genome Research Program (IOS-1032023) to ALC, KMO and Yulin Jia.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Vigueira, C., Olsen, K. & Caicedo, A. The red queen in the corn: agricultural weeds as models of rapid adaptive evolution. Heredity 110, 303–311 (2013). https://doi.org/10.1038/hdy.2012.104
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2012.104
Keywords
This article is cited by
-
Transcriptomic analysis provides insight into the genetic regulation of shade avoidance in Aegilops tauschii
BMC Plant Biology (2023)
-
Common evolutionary trajectory of short life-cycle in Brassicaceae ruderal weeds
Nature Communications (2023)
-
Weed genomics: yielding insights into the genetics of weedy traits for crop improvement
aBIOTECH (2023)
-
Sustainability, Efficiency, and Circularity of Weedy Rice Management Strategies
Circular Economy and Sustainability (2021)
-
Whole genomes and transcriptomes reveal adaptation and domestication of pistachio
Genome Biology (2019)