Abstract
Transcription, translation and subsequent protein modification represent the transfer of genetic information from the archival copy of DNA to the short-lived messenger RNA, usually with subsequent production of protein. Although all cells in an organism contain essentially the same DNA, cell types and functions differ because of qualitative and quantitative differences in their gene expression. Thus, control of gene expression is at the heart of differentiation and development. Epigenetic processes, including DNA methylation, histone modification and various RNA-mediated processes, are thought to influence gene expression chiefly at the level of transcription; however, other steps in the process (for example, translation) may also be regulated epigenetically. The following paper will outline the role epigenetics is believed to have in influencing gene expression.
Similar content being viewed by others
Epigenetics and gene expression
Transcription, translation and subsequent protein modification represent the transfer of genetic information from the archival copy of DNA to short-lived messenger RNA, usually with subsequent production of protein. Although all cells in an organism contain essentially the same DNA, cell types and functions differ because of qualitative and quantitative differences in their gene expression, and control of gene expression is therefore at the heart of differentiation and development. The patterns of gene expression that characterize differentiated cells are established during development and are maintained as the cells divided by mitosis. Thus, in addition to inheriting genetic information, cells inherit information that is not encoded in the nucleotide sequence of DNA, and this has been termed epigenetic information. Epigenetics has been defined as ‘the study of mitotically (and potentially meiotically) heritable alterations in gene expression that are not caused by changes in DNA sequence’ (Waterland, 2006). However, some definitions of epigenetics are broader than this and do not necessarily encompass the requirement for heritability. For example, the US National Institutes of Health (2009) in their recent epigenomics initiative state that ‘epigenetics refers to both heritable changes in gene activity and expression (in the progeny of cells or of individuals) and also stable, long-term, alterations in the transcriptional potential of a cell that are not necessarily heritable’. Regardless of the exact definition, the epigenetic processes that stably alter gene expression patterns (and/or transmit the alterations at cell division) are thought to include: (1) cytosine methylation; (2) post-translational modification of histone proteins and remodelling of chromatin; and (3) RNA-based mechanisms.
Gene expression is a complex process involving numerous steps (Alberts et al., 2008) and, although a detailed account of gene expression is beyond the scope of this review, we will briefly summarize the major stages to place the topic of the review, how epigenetics interacts with gene expression, in context. The initial step in gene expression is the transcription of the DNA molecule into an exact RNA copy. To initiate transcription, RNA polymerase binds to a particular region of the DNA (the promoter) and starts to make a strand of mRNA complementary to one of the DNA strands. Post-transcriptional processing is critical: a methylated guanosine ‘cap’ is added to the 5′end of the transcribed RNA whereas splicing of the mRNA occurs via a step-wise series of cleavage and ligation events that remove intron sequences and bring exons together in an appropriate manner. Following splicing, the 3′ terminus of the mRNA is cleaved and a string of adenosine residues, known as a polyA tail, is added in preparation for mRNA transport from the nucleus to the cytoplasm. At this stage, the mRNA is ready to engage with ribosomes for translation. In translation, polypeptides are synthesised in a sequential stepwise fashion from N terminus to C terminus via three distinct steps—initiation, elongation and termination. After initiation, the genetic code is read in triplets of nucleotides (codons) specified by the mRNA and the specified amino acids are assembled during the elongation process and attached via a peptidyl transferase reaction, resulting in the formation of a peptide bond and the elongation of the peptide chain. Termination of translation occurs when one of the termination codons (UAG, UAA and UGA) signals the release of a completed polypeptide chain. The ribosome then disengages from the mRNA and the ribosomal subunits dissociate, ready to start the cycle again. The generated protein may undergo several post-translational modifications before it is used in its dedicated role.
The above description is a somewhat simplistic outline of gene expression but in reality it is far from simple. Numerous regulatory steps maintain the integrity of the process and work with external factors to control each stage. Epigenetic processes, including DNA methylation and histone modification, are thought to influence gene expression chiefly at the level of transcription; however, other steps in the process (for example splicing and translation) may also be regulated epigenetically. The following paper will outline the role epigenetics is believed to have in influencing gene expression.
DNA methylation and epigenetic regulation of gene expression
Methylation of the 5′-position of cytosine residues is a reversible covalent modification of DNA, resulting in production of 5-methyl-cytosine (Newell-Price et al., 2000) and approximately 3% of cytosines in human DNA are methylated (Nafee et al., 2008). In mammals, cytosine methylation is restricted to those located 5′ to a guanosine (commonly annotated as CpGs, where the intervening ‘p’ represents the phosphodiester bond linking cytosine- and guanosine-containing nucleotides) (Razin and Cedar, 1991; Weber and Schübeler, 2007). These methyl groups allow normal hydrogen bonding (Figure 1) and project into the major groove of DNA, changing the biophysical characteristics of the DNA. They are purported to have two effects: they inhibit the recognition of DNA by some proteins while they facilitate the binding of other proteins to the DNA (Prokhortchouk and Defossez, 2008). In general, DNA methylation is associated with gene repression (Miranda and Jones, 2007; Weber and Schübeler, 2007). As DNA methylation patterns can be maintained following DNA replication and mitosis (see DNA methyltransferases section below), this epigenetic modification is also associated with inheritance of the repressed state.
With four nucleotides (A, T, C and G), DNA contains 16 dinucleotide-pair possibilities. The CpG pairing occurs at a lower than expected frequency throughout most of the genome but at a higher than expected frequency in regions referred to as CpG islands (Gardiner-Garden and Frommer, 1987). CpG islands themselves are unevenly distributed throughout the genome and were initially thought to be concentrated in promoter regions of genes (Doerfler, 1981; Baylin, 2005; Miranda and Jones, 2007; Shen et al., 2007). More recent work (Illingworth et al., 2008), however, suggests that about half of the CpG islands in the genome are not associated with annotated promoters, but are located within genes (intragenically) or in intergenic locations. It is suggested that the latter CpG islands may mark the transcription start sites of non-coding RNAs (Illingworth et al., 2008). Either way, CpG islands are thought to have a major role in control of gene expression.
In vertebrates, over 80% of CpG dinucleotides located outside of CpG islands are commonly methylated (Miranda and Jones, 2007; Nafee et al., 2008; Delcuve et al., 2009). In contrast, CpGs within CpG islands are generally not methylated or have relatively low levels of methylation (Miranda and Jones, 2007; Nafee et al., 2008; Delcuve et al., 2009). In keeping with this, approximately half of all transcribed genes have CpG islands within their coding regions, and these are reported to include all genes that are widely expressed and less than half of those that are expressed in a tissue-specific manner (Nafee et al., 2008).
DNA methyltransferases (DNMTs)
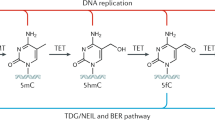
Cells have the ability to both methylate and demethylate DNA and this in turn is reported to influence specific gene expression (Wolfe, 1998; Ashraf and Ip, 1998; Kim et al., 2009). DNA methyltransferases (DNMTs) are the family of enzymes responsible for DNA methylation (Nafee et al., 2008; Kim et al., 2009; Delcuve et al., 2009) (Figure 2). To date, four DNMTs have been identified in mammals: DNMT1, DNMT2, DNMT3a and DNMT3b (Weber and Schübeler, 2007). DNMT1 maintains DNA methylation during replication by copying the methylation pattern of the parent DNA strand onto the newly synthesized strand (Newell-Price et al., 2000; Kim et al., 2009). DNMT3a and DNMT3b are responsible for de novo DNA methylation, targeting unmethylated CpG dinucleotides (Newell-Price et al., 2000; Wang et al., 2005; Suzuki et al., 2006; Hervouet et al., 2009), as well as working with DNMT1 to ensure propagation of methylation patterns during DNA replication (Weber and Schübeler, 2007). DNMT2 has reportedly only weak DNA methylation ability in vitro and appears to be involved in methylation of RNA (Goll et al., 2006). In terms of demethylation, relatively slow ‘passive’ DNA demethylation may occur if methylated CpGs fail to be propagated following DNA replication. However, more rapid ‘active’ demethylation also occurs, although the exact molecular mechanisms are not yet fully elucidated (Doerfler, 1981; Razin and Cedar, 1991; Kim et al., 2009). Plants use 5-methylcytosine glycosylases and the base excision repair pathway to remove excess cytosine methylation whereas, in mammals, active demethylation has been proposed to operate via several very different mechanisms, including deactivation of the aforementioned DNMTs (Doerfler, 1981; Razin and Cedar, 1991; Kim et al., 2009).
If methylation is involved in controlling gene expression, then genes that vary in their methylation status should show measurable and quantitative variations in their expression (Bird, 1984) and, furthermore, gene expression should be measurably altered by the methylation and demethylation of specific CpGs within specific genes. There are many examples of this, some of which are outlined in Table 1. In one case, Fuso et al. (2009) reported that the Presenilin 1 (PSEN1) 5′-flanking region has a site-specific methylation pattern that changes in response to metabolic stimuli, and that overexpression of this gene correlates with DNA demethylation. They showed that an induced B vitamin deficiency in mice resulted in DNA demethylation (and hyperhomocysteinemia), and caused PSEN1 overexpression. Furthermore, introduction of a methylating agent, S-adenosyl methionine (SAM), reversed both the demethylation and the overexpression of PSEN1. Similarly, Fang et al. (2003) examined the effect of the polyphenol, epigallocatechin-3-gallate (EGCG, a component of tea), on DNA methylation status of an oesophageal cancer cell line (KYSE 510) and noted a dose-dependent inhibitory effect of EGCG on DNMT activity. The reduction in DNMT activity resulted in CpG demethylation and reactivation of several methylation-silenced genes: p16INK4a, retinoic acid receptor β (RARβ), O6-methylguanine methyltransferase (MGMT), and human mutL homologue 1 (hMLH1). These examples show that cells do have the ability to methylate and demethylate specific genes and thus control the expression of these genes (with addition of methyl groups associated with gene silencing and removal of methyl groups associated with gene expression).
DNMT interaction with transcription factors
How does cytosine methylation influence gene expression? DNA methylation is suggested to lead to transcriptional silencing via multiple mechanisms. One mechanism involves DNMTs specifically interacting with transcription factors, resulting in site-specific methylation at promoter regions (Hervouet et al., 2009). This site-specific methylation is subsequently responsible for the assembly at these locations of proteins that recognise methylated DNA. These assemblies then directly influence further action of the transcriptional machinery (Kass et al., 1997; Wade, 2001) or cause alterations in chromatin structure, which in turn affect normal gene expression mechanisms (Prokhortchouk and Defossez, 2008).
The direct interaction of DNMTs with transcription factors (which interact with DNA at specific sites) and the resulting site-specific DNA methylation at promoters are purported to have an important role in gene regulation (Hervouet et al., 2009). Although limited in nature, data exist to show specific interactions between various transcription factors and DNMTs. Di Croce et al. (2002) provided the initial evidence indicating that DNMT3a was recruited to the RARβ2 promoter by the oncogenic transcription factor, PML–RAR, leading to promoter methylation and silencing of the RARβ2 gene. Brenner et al. (2005) and Wang et al. (2005) reported similar gene-specific transcriptional silencing, with the observation of suppression of P21 expression via Myc-targeted methylation of the P21 promoter. Within this pathway, p53 appears to recruit DNMT1, stimulating DNMT1-mediated DNA methylation and resulting in the repression of p21 expression. Hervouet et al. (2009) found that 79 known transcription factors interact directly with various DNMTs acting as potential DNA ‘anchors’ for the DNMTs, thus aiding in the site-specific methylation of promoter regions (Hervouet et al., 2009). Examples identified included the interaction of DNMT1 with Sp1 and Stat3; of DNMT3a with v myc, c-myc, ATF2 and ATF4; and of DNMT3b with Stat1, v-myc, Sp1, ATF2 and ATF4.
Thus, the dual ability of some transcription factors to bind to DNA via specific recognition sequences, and also to interact with DNMTs, may promote widespread site-specific DNA methylation at promoter regions. Once such site-specific methylation occurs, recruitment of methyl-binding proteins, as outlined below, may result in further effects on transcriptional activity and chromatin structure.
DNA methylation and methyl-binding proteins
Various DNA methyl-binding proteins (MBPs) exist, and are grouped into similar ‘families’ according to their structural similarity. One family shares a related DNA-binding domain (methylated DNA-binding domain, MBD) and the MBD family includes the proteins MBD1, MBD2, MBD3, MBD4 and MECP2. MBD1-3 proteins are transcriptional repressors that act through various mechanisms, resulting in the recruitment of co-repressors and histone deacetylases (Wade, 2001). Recruitment of histone deacetylases results in a distinct compaction of DNA, leading to the characteristic remodelling of chromatin (Wade, 2001). MBD4 is a thymidine glycosylase repair enzyme. It is not associated with transcriptional inactivation and is likely to have a role in limiting the mutagenicitiy of methylcytosine (Newell-Price et al., 2000; Wade, 2001). MECP2 is probably the best characterized of the MBD family. It binds methylated CpG via its MBD domain and exerts repressive effects on transcription over distances of several hundred base pairs via its second functional domain, a transcriptional repression domain (Razin, 1998; Newell-Price et al., 2000). This repressor domain recruits the co-repressor, Sin3 complex, which contains histone deacetylase 1 and 2, or other co-repressor complexes (Delcuve et al., 2009). Alternatively MECP2 can alter chromatin compaction by binding to linker DNA and nucleosomes, resulting in a physical barrier to the transcriptional machinery.
The second family of MBPs contains a common zinc finger domain and consists of the proteins Kaiso, ZBTB4 and ZBTB38 (Prokhortchouk and Defossez, 2008). The nucleo/cytoplasmic distribution of this family of proteins is variable and is said to respond to intracellular signalling, including the Wnt pathway (Prokhortchouk and Defossez, 2008). A recent paper by Iioka et al. (2009) suggests that Kaiso can regulate transcriptional activity via modulating histone deacetylase 1 (HDAC1) and β-catenin complex formation, and interacting with transcriptional factors such as LEF1 and its homologs. The third family of methyl DNA-binding proteins contains UHRF1 and UHRF2 (also known as ICBP90 and NIRF, respectively), which recognize and bind semi-methylated DNA through their SET- and RING finger-associated domains (SRA proteins) (Newell-Price et al., 2000; Prokhortchouk and Defossez, 2008). Binding of SRA proteins to methylated DNA directs DNMT1 to these sites, resulting in further alteration of DNA methylation and additional recruitment of other MBPs and their associated activities.
Thus, methyl-binding proteins react to the methylation status of DNA at specific sites, often associated with promoters of genes. These methyl-binding proteins appear to exert their effect by recruiting additional enzymes, such as histone deacetylases, which, as described in the following sections, also have important roles in epigenetic control of gene expression.
Histones and epigenetic regulation of gene expression
In eukaryotic cells, DNA and histone proteins form chromatin, and it is in this context that transcription takes place. The basic unit of chromatin is the nucleosome, and consists of an octamer of two molecules of each of the four canonical histone molecules (H2A, H2B, H3 and H4), around which is wrapped 147 bp of DNA (Alberts et al., 2008). Another type of histone (linker histone, H1) binds to the DNA between the nucleosomes. Histones help package DNA so that it can be contained in the nucleus but more recently their involvement in regulating gene expression has been shown.
The core histones are highly conserved basic proteins with globular domains around which the DNA is wrapped with relatively unstructured flexible ‘tails’ that protrude from the nucleosome. The tails are subject to a variety of post-translational modifications (PTMs) (Allis et al., 2007; Berger, 2007; Kouzarides, 2007), the best characterized of which are small covalent modifications: methylation, acetylation and phosphorylation. Other modifications include ubiquitination, sumoylation, ADP ribosylation and deimination, and the non-covalent proline isomerization that occurs in histone H3. Lysine residues can accept one, two or three methyl residues, while arginine can be mono- or dimethylated. Because histones contain high concentrations of these basic amino acids, the potential for complexity is obvious. In addition, in certain situations some of the core histones may be replaced by less abundant variant histones; variants of H2A and H3 (but not of H2B and H4) are known (Henikoff and Smith, 2007).
Most histone PTMs are dynamic and are regulated by families of enzymes that promote or reverse the modifications (Allis et al., 2007; Berger, 2007; Kouzarides, 2007). For example, histone acetyltransferases (HATs) and histone deacetyl transferases (HDACs) add and remove acetylation. Histone methyltransferases add methyl groups to arginine (protein arginine methyltransferases, PRMTs) and lysine (histone lysine methyltransferases, HKMTs) residues; arginine methylation is reversed by deiminases, which convert the side chain to citrulline, whereas two types of lysine demethylases have recently been identified. Histone kinases phosphorylate serine and threonine residues and phosphates are removed by various phosphatases. The factors regulating the modification enzymes and the methods by which they are targeted to specific loci are areas of intense investigation.
In addition to covalent modification of histones, chromatin structure is also controlled by families of enzymes that use the energy associated with ATP hydrolysis to effect changes in nucleosome arrangement or composition. These chromatin remodelling complexes belong to two families, the SNF2H or ISWI family and the Brahma or SWI/SNF family (Allis et al., 2007). The SNF2H/ISWI complexes act by mobilizing nucleosomes along the DNA, whereas the Brahma/SWI/SNF complexes transiently alter the structure of the nucleosomes, thus exposing the DNA-histone contacts. Some of the remodelling complexes promote the replacement of conventional core histones with variant forms, thus acting as ‘exchanger complexes’ (Allis et al., 2007). The chromatin remodelling complexes act in concert with the enzymes that covalently modify histones to facilitate transcription or alternatively may act to maintain repressed chromatin.
Histone modifications and gene expression
Although the literature detailing the effects of histone modifications on transcription is complex and constantly expanding, three general principles are thought to be involved (Kouzarides and Berger, 2007):
-
i)
PTMs directly affect the structure of chromatin, regulating its higher order conformation and thus acting in cis to regulate transcription;
-
ii)
PTMs disrupt the binding of proteins that associate with chromatin (trans effect);
-
iii)
PTMs attract certain effector proteins to the chromatin (trans effect).
Early studies indicated that histone acetylation positively correlates with transcription (as opposed to low levels of acetylation associated with transcriptionally silent chromatin). It is now thought that DNA-bound activators of transcription recruit HATs to acetylate nucleosomal histones, whereas transcription repressors recruit HDACs to deacetylate histones (Kouzarides and Berger, 2007). Many coactivators and corepressors possess HAT or HDAC activity respectively, or associate with such enzymes, and the enzymatic activity is necessary for their effects on transcription. The enzymes are often part of larger complexes that have additional functions to histone modification, such as recruiting TATA-binding protein.
Acetylation of lysine residues neutralizes their positively charged side chains, reducing the strength of the binding of histone tails to negatively charged DNA, ‘opening’ the chromatin structure and facilitating transcription and/or exposing DNA-binding sites. Many of the HDACs that remove acetyl groups are found in large multisubunit complexes, components of which target the complexes to genes, leading to transcriptional repression. Phosphorylation of histone H3 (serine 10) is also associated with transcription. The increased negative charge that phosphorylation confers on the histone may alter the structure of the nucleosome (model (i) above), whereas phosphorylation may also serve to dislodge proteins bound to chromatin or to attract proteins that enhance transcription (models ii and iii).
Of the various methylated residues that have been identified, several have been highly characterized. These include the lysine (K) residues, K4, K9, K27, K36 and K79 of histone H3, and K20 of histone H4, and arginine (R) residues R2, R17 and R26 of H3, and R3 of H4 (Kouzarides and Berger, 2007). The consequences of methylation can be either positive or negative with respect to transcriptional activity, depending on the position of the modified residue within the histone tail. Thus, H3 (K4, K36 and K79) are associated with activation, whereas H3 (K9, K27) and H4K20 are associated with repression (Kouzarides and Berger, 2007).
Histone deimination involves the conversion of arginine to citrulline by removal of an imino group and is catalysed by peptidyl arginine deiminases (PADIs, also known as PADs; the first enzyme identified was PAD4) (Cuthbert et al., 2004; Wang et al., 2004). If the arginine is monomethylated, deimination effectively results in antagonism of arginine methylation, as the methyl group is removed with the imino residue. The action of PAD4 has been best studied in the context of estrogen regulation of pS2 transcription, where PAD4 represses transcription. A cycle of methylation of arginine residues, followed by deimination by PAD4 and accumulation of citrullinated histones, has been shown in the cyclic on and off regulation of PS2 transcription. This repression involves coordinated action with the histone deacetylase, HDAC1 (Denis et al., 2009).
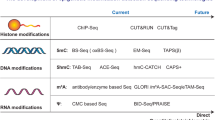
Chromatin immunoprecipitation (ChIP) using modification-specific antibodies has revolutionised the study of histone PTMs and, when combined with microarray analysis (ChIP on CHIP) or sequencing (ChIP-Seq) of immunoprecipitated DNA, has allowed for analysis of modifications on a global genome scale (Mikkelsen et al., 2007; Spivakov and Fisher, 2007). Such studies have shown that certain modifications are consistently associated with actively transcribed or repressed genes. For example H3K4me3 is associated with the promoters of actively transcribed genes and H3K36me3 is enriched in the body of such genes, specifically in exons. In contrast, H3K9me3 is enriched over the promoter and body of repressed genes. The consistency of these findings in species as diverse as yeast, human and mouse has allowed the use of histone modification ‘signatures’ to search for novel transcribed genes (Ozsolak et al., 2008; Won et al., 2008; Guttman et al., 2009; Heintzman et al., 2007, 2009). In these studies, previously non-annotated protein-coding genes, as well as numerous non-coding RNAs that are likely to be functional, were identified.
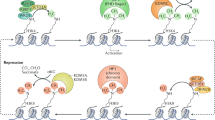
In summary, the covalent modification status of histone proteins, together with nucleosome composition and arrangement comprises an epigenetic layer of information that facilitates or inhibits gene expression (Figure 3).
Open and closed chromatin configurations are influenced by post-translational histone modifications. In the upper panel, DNA is wrapped around histones that possess activating modifications (green circles and blue triangles). In the lower panel, DNA is wrapped around histones with repressing modifications (red circles and orange triangles). The bent arrow indicates a transcription start site; this is more accessible to RNA polymerase in the open chromatin configuration.
RNA-based mechanisms and epigenetic regulation of gene expression
RNA-based mechanisms of epigenetic regulation are less well understood than mechanisms based on DNA methylation and histones. RNA involvement in regulating monoallelic expression of imprinted genes (see Barlow article, this issue) and in X-chromosome inactivation (see Wutz article, this issue) is well established, but recent studies have implicated RNA-based mechanisms in more widespread epigenetic regulation (Bernstein and Allis, 2005; Costa, 2008; Mattick et al., 2009).
Non-coding infrastructural RNAs have been known for a long time (Alberts et al., 2008) and include tRNAs, rRNAs, small nuclear RNAs (snRNAs) and small nucleolar RNAs (snoRNAs). These are involved in translation and splicing, and function by sequence-specific recognition of RNA substrates and also in catalysis. In addition to their infrastructural roles, some of these may also have regulatory roles; for example U1 snRNA is involved in regulating the activity of transcription initiation by RNA polymerase II through interaction with the transcription initiation factor TFIIH (Kwek et al., 2002). More recent studies indicate that the majority of the genome is transcribed into RNA transcripts, most of which do not code for protein (Kapranov et al., 2007; Amaral and Mattick, 2008 and references therein). These non-coding RNAs (ncRNAs) range from very short molecules to extremely large transcripts and are usually classified based on length, subcellular location, orientation with respect to the nearest protein-coding gene and/or function (if known). A widely used classification divides ncRNAs into small (less than 200 nt, and typically much shorter) and long (greater than 200 nt and often much longer) species.
Small non-coding RNAs
Small ncRNAs are generally derived from larger RNA precursor molecules, by cleavage with RNAse III-family enzymes (typically Drosha and Dicer) and include microRNAs (miRNAs), short interfering RNAs (siRNAs), PIWI-interacting RNAs (piRNAs), and repeat-associated RNAs (rasiRNAs), in addition to other less well-characterized species. These RNAs have been described in recent excellent reviews (Farazi et al., 2008; Amaral and Mattick, 2008; Carthew and Sontheimer, 2009; Malone and Hannon, 2009) and some are briefly addressed below.
miRNAs (∼22 nt long) are derived from imperfect hairpin structures present in long ncRNA precursors or introns (of coding or non-coding genes), and are processed in two consecutive cleavage steps by Drosha and Dicer. Mature miRNAs base-pair with target mRNAs to inhibit translation (if they pair with the target RNA imperfectly) or direct mRNA degradation via the RISC complex (if they pair perfectly with their target). Recently miRNA regulation of de novo DNA methylation was demonstrated in mouse embryo stem cells (Benetti et al., 2008; Sikkonen et al., 2008).
siRNAs are similar in size to miRNAs (∼21 nt long) but differ in that they are derived from double-stranded RNA precursors that are processed by Dicer. They usually base-pair with perfect matches to their target mRNAs and direct them for degradation; however, they may also repress translation, if they base-pair with less complementarity. siRNAs also participate in transcriptional gene silencing, particularly for silencing transposable elements, and this function is well-characterized in plants, where it involves the siRNA guiding DNA methylation to genomic regions homologous to the siRNA sequence. In Saccharomyces pombe and probably in animals, siRNA-directed transcriptional gene silencing involves recruitment of histone methyltransferases and generation of heterochromatin.
piRNAs are 28–33 nt in length and associate with PIWI-family proteins in male germ cells and in oocytes. They are apparently processed from single-stranded precursors by a Dicer-independent (but poorly characterized) pathway. The rasiRNAs of Drosophila and the ‘21U’ RNAs of Caenorhabditis elegans appear to correspond to piRNAs. These RNAs are involved in control of transposable element activity in the germ lines of Drosophila, Caenorhabditis elegans, fish and mammals and are essential for germline viability. Maternal piRNAs deposited in Drosophila oocytes alter the phenotype of progeny in a heritable manner and thus may act as vectors of epigenetic inheritance (Malone and Hannon, 2009).
Long non-coding RNAs
Long ncRNAs (lncRNAs) are usually defined as being greater than 200 nt (Ponting et al., 2009). This is not a great definition but, in the absence of evidence of functionality for most lncRNAs, it is widely used. An initial concern was that lncRNAs represent ‘loose’ or ‘noisy’ transcription (Mattick, 2005; Struhl, 2007; Ponting et al., 2009) and/or artefacts of experimental procedures (e.g., contamination with genomic DNA, fragments of pre-mRNA etc). However it is now generally accepted that there is a large amount of such transcripts in most eukaryotes. Ponting et al. (2009) categorize lncRNAs in five categories: (1) sense or (2) anti-sense, when overlapping one or more exons of another transcript on the same or opposite strand respectively; (3) bidirectional, when expression of the lncRNA and of a neighbouring coding transcript on the opposite strand are initiated in close genomic proximity; (4) intronic, when it is derived wholly from within an intron of a second transcript; or (5) intergenic when it lies within the genomic interval between two genes.
As a group, the function of lncRNAs is not well understood. Some may well represent transcriptional noise or experimental artefacts, and others may serve as precursors of short RNAs. However, in many cases it seems that long ncRNAs regulate gene expression in their own right, either because the actual transcript is functional or because of the act of their transcription per se. Evidence supporting the contention that many lncRNAs are functional is that they exhibit tissue-specificity, are regulated during development, localize to specific cellular compartments, are associated with human disease and/or show evidence of evolutionary selection (Wilusz et al., 2009). Wilusz et al. (2009) describe the various ways in which lncRNAs can have regulatory effects. In some cases, the simple act of transcription of a lncRNA can increase or decrease transcription from a downstream promoter, either by altering RNA Polymerase II recruitment or by altering chromatin configuration. In other cases, hybridization of an antisense transcript with a sense transcript may result in altered splicing of the sense transcript, or could lead to the generation of endogenous siRNAs following Dicer-mediated processing. lncRNAs may interact with protein partners, modulating the activity or localization of these proteins within the cell, or the RNA may be processed to give rise to various types of small regulatory RNAs. Table 2 lists some examples of functional lncRNAs.
The inability to ascribe functions to many lncRNAs may reflect the inadequacy or insensitivity of current experimental methods of detecting function. As an initial approach to systematic high-throughput screening for function, Willingham et al. (2005) used 12 cell-based assays and an siRNA approach to probe the function of 512 lncRNAs that showed a high degree of sequence conservation between mouse and human (average size ∼2 kb). In this relatively small screen, they identified eight functional lncRNAs. Interestingly, these turned out to be involved in pathways that were already intensively studied and in which a role of lncRNAs had not been previously suspected (hedgehog signalling, nuclear trafficking). This study suggests that the potential involvement of lncRNAs in regulatory processes is currently underestimated.
Interaction of lncRNAs with chromatin-modifying complexes
An emerging theme in the study of lncRNAs is their interaction with chromatin modifying enzymes. HOTAIR, a lncRNA transcribed from the HOXC cluster, binds to the polycomb repressive complex PRC2 and targets this to the HOXD cluster where several genes are repressed (Rinn et al., 2007). (PRC2 possesses methyltransferase activity and trimethylates H3K27). In a similar manner, Zhao et al. (2008) reported that the lncRNA Xist, which plays an important role in X-chromosome inactivation in female mammal cells, as well as a shorter internal transcript, RepA, interact with PRC2 complexes (specifically with the Ezh2 subunit) and recruit them to the inactive X-chromosome where they subsequently serve to trimethylate H3K27. Interestingly, the lncRNA Tsix, a known Xist antagonist, also interacts with PRC2 and it is suggested that Tsix could block X-chromosome inactivation by titrating away PRC2 (Zhao et al., 2008).
Two lncRNAs transcribed at imprinted loci have also been shown to interact with chromatin-modifying complexes. Airn is a transcript that is produced from a promoter in intron 2 of the paternally-derived Igf2r allele. The Airn transcript, or the act of its transcription, is required for monoallelic expression of Igf2r and of two other protein-coding genes in the Igf2r cluster, Slc22a2 and Slc22a3 (Sleutels et al., 2002). Nagano et al. (2008) reported that Airn accumulates at the paternally-derived Slc22a3 promoter in placenta. They further showed that Airn interacts with G9a, a histone H3K9 methyltransferase, and that this interaction is required for recruitment of G9a to the paternally-derived Slc22a3 promoter where allele-specific methylation of H3K9 and Slc22a3 repression occurs. Interestingly, silencing of Igf2r by Airn appears to occur by a different mechanism, and does not involve accumulation of Airn and G9a at the Igf2r promoter (Nagano et al., 2008). Kcnq1ot1 is generated from a promoter in intron 10 of the paternally-derived Kcnq1 gene and is linked to the silencing of 8–10 protein coding genes at this imprinted locus. Pandey et al. (2008) have shown that Kcnq1ot1 interacts with PRC2 and with G9a in a lineage-specific manner (interaction occurs in placenta but not in fetal liver) and these associations correlate with lineage-specific differences in repressive chromatin modifications at the locus.
A recent study demonstrated that a large number of lncRNAs in several human cell types associate with complexes that add repressive chromatin marks and, using an siRNA approach, provided evidence that these associations are functional in a target-specific manner (Khalil et al., 2009). Interactions of lncRNAs with chromatin-modifying complexes may also serve to target activating modifications to specific loci. Thus, in Drosophila, three lncRNAs recruit the trithorax protein Ash1 to the Ultrabithorax locus maintaining Ubx transcription (Sanchez-Elsner et al., 2006) while Dinger et al. (2008) suggest that two lncRNAs (Evx1as and Hoxb5/6as) function in a similar way in mammals. Therefore, the varying expression patterns of lncRNAs in different cell types and their ability to interact specifically with chromatin-modifying complexes provides a plausible mechanism for establishing and maintaining the epigenetic landscapes characteristic of differentiated cells.
Conclusion
The establishment of stable patterns of gene expression is a pre-requisite of normal differentiation and is accomplished by the imposition of a layer of lineage-specific epigenetic information onto the genome. This information (the epigenome) thus distinguishes one cell type from another and also appears to comprise the molecular memory that is inherited by daughter cells at mitosis. The epigenome encompasses a number of molecular components, with cytosine methylation, histone proteins and their post-translational modification, chromatin remodeling complexes and various non-coding RNAs playing important roles. Most aspects of gene expression are influenced by epigenetic mechanisms, from relative accessibility of genes in the chromosomal landscape, through transcription and post-transcriptional RNA processing and stability, to translation. The variety of molecular ‘players’ identified so far, and the array of mechanisms involved, together with the interplay among them all, suggests that even apparently simple patterns of gene regulation may represent dynamic and complex operations.
Unlike the genome, which is essentially identical in all cells of a vertebrate and stable throughout the life-time of an individual, the epigenome differs from cell to cell and is plastic, changing with time and with exposure to the environment (Jirtle and Skinner, 2007; Szyf, 2009a). The epigenome appears to be particularly vulnerable to environmental influences during certain stages of development (cleavage, perinatal period, puberty) and alterations in gene expression patterns induced at these times may persist for long periods, influencing the phenotype of the adult. Such long-term changes in gene expression patterns represent an attractive molecular basis for the hypothesis that the origin of adult disease lies in environmental exposure events during an individual's pre- or post-natal development (Barker, 2007; Heijmans et al., 2008; Szyf, 2009b). For this reason, research into the epigenetic regulation of gene expression will continue unabated.
References
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2008). Molecular Biology of the Cell. 5th edn. Garland Science: New York.
Allis CD, Jenuwein T, Reinberg D (2007). Overview and concepts. In: Allis CD, Jenuwein T, Reinberg D (eds). Epigenetics. CSHL Press: New York. pp 23–61.
Amaral PP, Mattick JS (2008). Noncoding RNA in development. Mamm Genome 19: 454–492.
Ashraf SI, Ip YT (1998). Transcriptional control: repression by local chromatin medication. Curr Biol 8: R683–R686.
Barker DJ (2007). The origins of the developmental origins theory. J Intern Med 261: 412–417.
Baylin SB (2005). DNA methylation and gene silencing in cancer. Nat Clin Pract Oncol S 1: S4–11.
Benetti R, Gonzalo S, Jaco I, Muñoz P, Gonzalez S, Schoeftner S et al. (2008). A mammalian microRNA cluster controls DNA methylation and telomere recombination via Rbl2-dependent regulation of DNA methyltransferases. Nat Struct Mol Biol 15: 268–279.
Berger SL (2007). The complex language of chromatin regulation during transcription. Nature 447: 407–412.
Bernstein E, Allis CD (2005). RNA meets chromatin. Genes Dev 19: 1635–1655.
Bird AP (1984). DNA methylation versus gene expression. J Embryol Exp Morphol 83 (Suppl): 33–40.
Brenner C, Deplus R, Didelot C, Loriot A, Vire E, De Smet C (2005). Myc represses transcription through recruitment of DNA methyltransferases corepressor. EMBO J 24: 336–346.
Carthew RW, Sontheimer EJ (2009). Origins and mechanisms of miRNAs and siRNAs. Cell 136: 642–655.
Costa FF (2008). Non-coding RNAs, epigenetics and complexity. Gene 410: 9–17.
Cuthbert GL, Daujat S, Snowden AW, Erdjument-Bromage H, Hagiwara T, Yamada M et al. (2004). Histone deimination antagonizes arginine methylation. Cell 118: 545–553.
Delcuve GP, Rastegar M, Davie JR (2009). Epigenetic control. J Cell Physiol 219: 243–250.
Denis H, Deplus R, Putmans P, Yamada M, Metivier R, Fuks F (2009). Functional connection between deimination and deacetylation of histones. Mol Cell Biol 29: 4982–4993.
Di Croce L, Raker VA, Corsaro M, Fazi F, Fanelli M, Faretta M et al. (2002). Methyltransferase recruitment and DNA hypermethylation of target promoters by an oncogenic transcription factor. Science 295: 1079–1082.
Dinger ME, Amaral PP, Mercer TR, Pang KC, Bruce SJ, Gardiner BB et al. (2008). Long noncoding RNAs in mouse embryonic stem cell pluripotency and differentiation. Genome Res 18: 1433–1445.
Doerfler W (1981). DNA methylation—a regulatory signal in eukaryotic gene expression. J Gen Virol 57: 1–20.
Fang MZ, Wang Y, Ai N, Hou Z, Sun Y, Lu H et al. (2003). Tea polyphenol (-)-epigallocatechin-3-gallate inhibits DNA methyltransferase and reactivates methylation-silenced genes in cancer cell lines. Cancer Res 63: 7563–7570.
Farazi TA, Juranek SA, Tuschl T (2008). The growing catalog of small RNAs and their association with distincy Argonaute/Piwi family members. Development 135: 1201–1214.
Feng J, Bi C, Clark BS, Mady R, Shah P, Kohtz JD (2006). The Evf-2 noncoding RNA is transcribed from the Dlx-5/6 ultraconserved regions and functions iias a Dlx-2 transcriptional coactivator. Genes Dev 20: 1470–1484.
Fujiki K, Kano F, Shiota K, Murata M (2009). Expression of the peroxisome proliferator activated receptor γ gene is repressed by DNA methylation in visceral adipose tissue of mouse models of diabetes. BMC Biol 7: 38.
Fuso A, Nicolia V, Pasqualato A, Fiorenza MT, Cavallaro RA, Scarpa S (2009). Changes in Presenilin 1 gene methylation pattern in diet induced B-vitamin deficiency. Neurobiol Aging 10: 1016–1028.
Gardiner-Garden M, Frommer M (1987). CpG islands in vertebrate genomes. J Mol Biol 196: 261–282.
Goll MG, Kirpekar F, Maggert KA, Yoder JA, Hsieh C-L, Zhang X (2006). Methylation of tRNA Asp by the DNA methyltransferase homolog Dnmt2. Science 311: 395–398.
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D et al. (2009). Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458: 223–227.
Heijmans BT, Tobi EW, Stein AD, Putter H, Blauw GJ, Susser ES et al. (2008). Persistent epigenetic differences associated with prenatal exposure to famine in humans. Proc Natl Acad Sci USA 105: 17046–17049.
Heintzman ND, Stuart RK, Hon G, Fu Y, Ching CW, Hawkins RD et al. (2007). Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat Genet 39: 311–318.
Heintzman ND, Hon GC, Hawkins RD, Kheradpour P, Stark A, Harp LF et al. (2009). Histone modifications at enhancers reflect global cell-type-specific gene expression. Nature 459: 108–112.
Henikoff S, Smith MM (2007). Histone variants and epigenetics. In: Allis CD, Jenuwein T, Reinberg D (eds). Epigenetics. CSHL Press: New York, pp 249–264.
Hervouet E, Vallette FM, Cartron PF (2009). Dnmt3/transcription factor interactions as crucial players in targeted DNA methylation. Epigenetics 4: 1–13.
Hirota K, Miyoshi T, Kugou K, Hoffman CS, Shibata T, Ohta K (2008). Stepwise chromatin remodelling by a cascade of transcription initiation of non-coding RNAs. Nature 456: 130–134.
Iioka H, Doerner SK, Tamai K (2009). Kaiso is a bimodal modulator for Wnt/beta-catenin signaling. FEBS Lett 583: 627–632.
Illingworth R, Kerr A, Desousa D, Jorgensen H, Ellis P, Stalker J et al. (2008). A novel CpG island set identifies tissue specific methylation at developmental gene loci. PLoS Biol 6: e22.
Jirtle RL, Skinner MK (2007). Environmental epigenomics and disease susceptibility. Nat Rev Genet 8: 253–262.
Kapranov P, Willingham AT, Gingeras TR (2007). Genome wide transcription and the implications for genomic organization. Nat Rev Genet 8: 413–423.
Kass SU, Pruss D, Wolffe AP (1997). How does DNA methylation repress transcription? Trends Genet 13: 444–449.
Khalil AM, Guttman M, Huarte M, Garber M, Raj A, rivea Morales D et al. (2009). Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Nat Acad Sci USA 106: 11667–11672.
Kim JK, Samaranayake M, Pradham S (2009). Epigenetic mechanisms in mammals. Cell Mol Life Sci 66: 596–612.
King-Batoon A, Leszczynska JM, Klein CB (2008). Modulation of gene methylation by genistein or lycopene in breast cancer cells. Environ Mol Mutagen 49: 36–45.
Kouzarides T (2007). Chromatin modifications and their function. Cell 128: 693–705.
Kouzarides T, Berger SL (2007). Chromatin modifications and their mechanism of action. In: Allis CD, Jenuwein T, Reinberg D (eds). Epigenetics. CSHL Press: New York. pp 191–209.
Kwek KY, Murphy S, Furger A, Thomas B, O'Gorma W, Kimura H et al. (2002). U1 snRNA associates with TFIIH and regulates transcriptional initiation. Nat Struct Biol 9: 800–805.
Malone CD, Hannon GJ (2009). Small RNAs as guardians of the genome. Cell 136: 656–668.
Martens JA, Laprade L, Wilson F (2004). Intergenic transcription is required to repress the Saccharomyces cerevisiae SER3 gene. Nature 429: 571–574.
Martianov I, Ramadass A, Barros AS, Chow N, Akoulitchev A (2007). Repression of the human dihydrofolate reductase gene by a non-coding interfering transcript. Nature 445: 666–670.
Mattick JS (2005). The functional genomics of noncoding RNA. Science 309: 1527–1528.
Mattick JS, Amarel PP, Dinger ME, Mercer TR, Mehler MF (2009). RNA regulation of epigenetic processes. BioEssays 31: 51–59.
Mikkelsen TS, Ku M, Jaffe DB, Isaac B, Lieberman E, Giannoukos G et al. (2007). Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448: 553–560.
Miranda TB, Jones PA (2007). DNA methylation: the nuts and bolts of repression. J Cell Physiol 213: 384–390.
Nafee TM, Farrell WE, Carroll WD, Fryer AA, Ismail KMK (2008). Epigenetic control of fetal gene expression. BJOG 115: 158–168.
Nagano T, Mitchell JA, Sanz LA, Pauler FM, Ferguson-Smith AC, Feil R et al. (2008). The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 322: 1717–1720.
Newell-Price J, Clark AJL, King P (2000). DNA methylation and silencing of gene expression. Trends Endocrinol Metab 11: 142–148.
Novikova SI, He F, Bai J, Cutrufello NJ, Lidow MS, Undieh AS (2008). Maternal cocaine administration in mice alters DNA methylation and gene expression in hippocampal neurons of neonatal and prepubertal offspring. PLoS One 3: e1919.
Ozsolak F, Poling LL, Wang Z, Liu XS, Roeder RG, Zhang X et al. (2008). Chromatin structure analyses identify miRNA promoters. Genes Dev 22: 3172–3183.
Pandey M, Shukla S, Gupta S (2009). Promoter demethylation and chromatin remodeling by green tea polyphenols leads to re-expression of GSTP1 in human prostate cancer cells. Int J Cancer 126: 2520–2533.
Pandey RR, Mondal T, Mohammad F, Enroth S, Redrup L, Komorowski J et al. (2008). Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol Cell 32: 232–246.
Ponting CP, Oliver PL, Reik W (2009). Evolution and functions of long noncoding RNAs. Cell 136: 629–641.
Prokhortchouk E, Defossez PA (2008). The cell biology of DNA methylation in mammals. Biochim Biophys Acta 1783: 2167–2173.
Qiang M, Denny A, Chen J, Ticku MK, Yan B, Henderson G (2010). The site specific demethylation in the 5-regulatory area of NMDA receptor 2B subunit gene associated with CIE-induced up-regulation of transcription. PLoS One 5: e8798.
Razin A (1998). CpG methylation, chromatic structure and gene silencing—a three-way connection. EMBO J 17: 4905–4908.
Razin A, Cedar H (1991). DNA methylation and gene expression. Microbiol Rev 55: 451–458.
Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA et al. (2007). Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129: 1311–1323.
Sanchez-Elsner T, Gou D, Kremmer E, Sauer F (2006). Noncoding RNAs of trithorax response elements recruit Drosophila Ash1 to Ultrabithorax. Science 311: 1118–1123.
Shen L, Kondo Y, Guo YI, Zhang J, Zhang LI, Ahmed S et al. (2007). Genome-wide profiling of DNA methylation reveals a class of normally methylated CpG island promoters. PLoS Genet 3: 2023–2036.
Sikkonen L, Hugenschmidt T, Berninger P, Gaidatzis D, Mohn F, Artus-Revel CG et al. (2008). MicroRNAs control de novo DNA methylation through regulation of transcriptional repressors in mouse embryonic stem cells. Nat Struct Mol Biol 15: 259–267.
Sleutels F, Zwart R, Barlow DP (2002). The non-coding Air RNA is required for silencing autosomal imprinted genes. Nature 415: 810–813.
Spivakov M, Fisher AG (2007). Epigenetic signatures of stem-cell identity. Nat Rev Genet 8: 263–271.
Struhl K (2007). Transcriptional noise and the fidelity of initiation by RNA polymerase II. Nat Struct Mol Biol 14: 103–105.
Suzuki M, Yamada T, Kihara-Negishi F, Sakurai T, Hara E, Tenen DG et al. (2006). Site-specific DNA methylation of a complex of PU.1 and Dnmt3a/b. Oncogene 25: 2477–2488.
Szyf M (2009a). The early life environment and the epigenome. Biochim Biophys Acta 1790: 878–885.
Szyf M (2009b). Early life, the epigenome and human health. Acta Paediatr 98: 1082–1084.
US National Institutes of Health, NIH Common Fund (2009). Epigenomics. http://nihroadmap.nih.gov/epigenomics/.
Wade PA (2001). Methyl CpG binding proteins: coupling chromatin architecture to gene regulation. Oncogene 20: 3166–3173.
Wang Y, Wysocka J, Sayegh J, Lee YH, Perlin JR, Leonelli L et al. (2004). Human PAD4 regulates histone arginine methylation levels via demethylimination. Science 306: 279–283.
Wang YA, Kamarova Y, Shen KC, Jiang Z, Hahn MJ, Wang Y et al. (2005). DNA methyltransferases-3a interacts with p53 and represses p53-mediated gene expression. Cancer Biol Ther 4: 1138–1143.
Wang X, Arai S, Song X, Reichart D, Du K, Pascual G et al. (2008). Induced ncRNAs allosterically modify RNA-binding proteins in cis to inhibit transcription. Nature 454: 126–130.
Waterland RA (2006). Epigenetic mechanisms and gastrointestinal development. J Pediatr 149: S137–S142.
Weber M, Schübeler D (2007). Genomic patterns of DNA methylation: targets and function of an epigenetic mark. Curr Opin Cell Biol 19: 273–280.
Willingham AT, Orth AP, Batalov S, Peters EC, Wen BG, Aza-Blanc P et al. (2005). A strategy for probing the function of noncoding RNAs finds a repressor of NFAT. Science 309: 1570–1573.
Wilusz JE, Sunwoo H, Spector DL (2009). Long noncoding RNAs: functional surprises from the RNA world. Genes Dev 23: 1494–1504.
Wolfe AP (1998). Packaging principle: how DNA methylation and histone acetylation control the transcriptional activity of chromatin. J Exp Zool 282: 239–244.
Won KJ, Chepelev I, Ren B, Wang W (2008). Prediction of regulatory elements in mammalian genomes using chromatin signatures. BMC Bioinformatics 9: 547.
Xiang N, Zhao R, Song G, Zhong W (2008). Selenite reactivates silenced genes by modifying DNA methylation and histones in prostate cancer cells. Carcinogenesis 29: 2175–2181.
Young TL, Matsuda T, Cepko CL (2005). The noncoding RNA taurine upregulated gene 1 is required for differentiation of the murine retina. Curr Biol 15: 501–512.
Zhao J, Sun BK, Erwin JA, Song J-J, Lee JT (2008). Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322: 750–756.
Acknowledgements
The authors acknowledge support from the Strategic Research Fund of UCD's College of Life Sciences and thank Shaun O'Boyle for figure preparation and Carlotta Sacchi for critical reading of the manuscript. CMN acknowledges funding from Science Foundation Ireland (RFP06/EEB007).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Gibney, E., Nolan, C. Epigenetics and gene expression. Heredity 105, 4–13 (2010). https://doi.org/10.1038/hdy.2010.54
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2010.54
Keywords
This article is cited by
-
Current and future therapeutic strategies for high-grade gliomas leveraging the interplay between epigenetic regulators and kinase signaling networks
Journal of Experimental & Clinical Cancer Research (2024)
-
Sex-specific epigenetics drive low GPER expression in gastrointestinal smooth muscles in type 2 diabetic mice
Scientific Reports (2024)
-
Intron-mediated enhancement of DIACYLGLYCEROL ACYLTRANSFERASE1 expression in energycane promotes a step change for lipid accumulation in vegetative tissues
Biotechnology for Biofuels and Bioproducts (2023)
-
Methionine consumption by cancer cells drives a progressive upregulation of PD-1 expression in CD4 T cells
Nature Communications (2023)
-
The maternal effects of dietary restriction on Dnmt expression and reproduction in two clones of Daphnia pulex
Heredity (2023)