Abstract
Studies of the control and function of DNA methylation in Neurospora crassa have led to a greater understanding of heterochromatin formation. DNA methylation in Neurospora is dependent on trimethylation of histone H3 lysine 9 (H3K9me3) by the histone methyltransferase, DIM-5. The linkage between these two methyl marks is facilitated by heterochromatin protein 1 (HP1), which serves as an adapter protein. HP1 binds to the H3K9me3 and recruits the DNA methyltransferase, DIM-2. Although HP1 links H3K9me3 to DNA methylation, it also serves to recruit the DNA methylation modifier complex to the edges of heterochromatin regions, where it serves to limit the spreading of the heterochromatin by countering H3K9me3.
Similar content being viewed by others
Introduction
Constitutive heterochromatin, the densely staining fraction of the eukaryotic genome, has been historically regarded as an unimportant component of the genome. Compared with euchromatin, the fraction of the genome containing most of the genes, heterochromatin is regarded as a more ‘closed’ or ‘condensed’ chromatin state, which is refractory to DNA-based processes such as recombination and transcription. Recently, however, heterochromatin has attracted attention, in part because it has been implicated in human diseases, most notably cancer (Sharma et al., 2010). Moreover, there is increasing evidence that heterochromatin has roles in normal cells, for example promoting genome stability by preventing illegitimate recombination between repeats, serving to silence invasive elements and participating in centromere function (Grewal and Jia, 2007; Peng and Karpen, 2008; Rountree and Selker, 2009). Thus, there is considerable interest in how regions of heterochromatin are formed and maintained. Recent research implicates an intricate interplay of covalent modifications of histones and a variety of nuclear proteins that are positively or negatively influenced by these modifications. Covalent modifications of histones include methylation, acetylation, phosphorylation, and/or ubiquitination of various residues. In addition to histone modifications, in many eukaryotes chromatin can be modified by methylation of cytosines in the DNA. Modifications of histones and DNA should provide a virtually limitless combination of chromatin modifications that could be used for regulatory control. Unfortunately, our understanding of these modifications and their interplay is still in its infancy.
The use of simple model organisms, such as yeasts, filamentous fungi, flies, and Arabidopsis, has greatly aided our understanding of chromatin modifications and will continue to be invaluable in deciphering the chromatin modification ‘code’. However, some powerful model systems lack chromatin modifications that are present in higher eukaryotes, such as DNA methylation. Neurospora crassa is the simplest model organism to sport most of the chromatin modifications seen in higher eukaryotes and offers many advantages for their study. Some of the heterochromatin markers, such as DNA methylation, are not essential in Neurospora, facilitating dissection of the processes involved in the establishment and maintenance of heterochromatin.
DNA methylation and repeat-induced point mutation in Neurospora
Much of what has been learned about heterochromatin in Neurospora resulted from studies on DNA methylation. Approximately 1.5% of cytosines are methylated in the Neurospora genome (Selker and Stevens, 1987; Foss et al., 1993). None of this DNA methylation has been found in protein-coding genes; rather, nearly all is found within transposon relics found in the centromere regions, near the, telomeres and elsewhere. Essentially all of these sequences are relics of a genome defense system called repeat-induced point mutation (RIP) (Foss et al., 1993; Selker et al., 2003; Lewis et al., 2009). RIP apparently protects the genome from the assault of invasive DNAs (transposons, retrotransposons, viruses, and so on) (Rountree and Selker, 2009). RIP is a potent genome defense system, leaving duplicated sequences both mutated and methylated. The effectiveness of RIP is illustrated by the fact that no recognizable transposon is found within the wild-type Neurospora genome (Galagan et al., 2003). Early genetic studies showed that RIP senses duplicated sequences and subjects both sequences to numerous polarized transition mutations (G:C to A:T) in the stage of the life cycle between fertilization and nuclear fusion (Selker et al., 1987; Selker and Garrett, 1988). Up to ∼30% of the G:C pairs of duplicated sequences are mutated in a single passage through the sexual cycle (Cambareri et al., 1989; Cambareri et al., 1991). Although the mechanism of RIP remains to be elucidated, one of Neurospora's two putative DNA methyltransferases (DMT), RIP defective (RID), is essential for the process (Table 1) (Freitag et al., 2002). Sequences altered by RIP are typically methylated at the remaining cytosines and this methylation is not restricted to symmetrical sequences (for example, CpG), unlike the case of DNA methylation in most vertebrate cells (Selker and Stevens, 1985). Indeed, it appears that any cytosine residue in a methylated region can be modified and this methylation is typically dense but heterogeneous within a population of nuclei (Selker et al., 1993). Importantly, the DNA methylation does not depend on the sequence duplication that originally triggered RIP and is relatively independent of chromosomal position (Selker, 2004).
Treating Neurospora with the histone deacetylase inhibitor, Trichostatin-A, affects DNA methylation (Selker, 1998). Although this observation was the first indication that DNA methylation is closely linked to histone modifications in this organism, genetic studies have been most important in defining a sequence of chromatin events that lead to DNA methylation. Two mutants d efective i n DNA m ethylation (dim), dim-2 and dim-5, completely abolish DNA methylation in vegetative tissue were most informative (Foss et al., 1993; Tamaru and Selker, 2001). Unlike DNA methylation patterns in animals and plants, which are established by multiple DMTs in different pathways, one DMT, encoded by the dim-2 gene, is responsible for all DNA methylation found in vegetative tissue of Neurospora (Table 1) (Kouzminova and Selker, 2001). Although the role of dim-2 in DNA methylation was obvious, the role of dim-5, which encodes a histone methyltransferase, was less so, but provided a concrete link between histone modification and DNA methylation (Table 1) (Tamaru and Selker, 2001). The identification of these and other dim mutants, coupled with proteomic, bioinformatics, and high-throughput approaches have served to unlock the pathway that leads to heterochromatin formation in Neurospora.
Characteristics of heterochromatin in Neurospora
Heterochromatin is typically enriched for hypoacetylated histones, methylated H3K9, heterochromatin protein 1 (HP1) and DNA methylation (Bhaumik et al., 2007). To achieve a more thorough understanding of the relationship between such heterochromatin markers in Neurospora, Lewis et al. (2009) used a ChIP-chip approach to generate a high-resolution map of these heterochromatin markers along N. crassa Linkage Group VII. This analysis demonstrated that DNA methylation, H3K9me3, and HP1 are nearly perfectly co-localized at 44 discrete heterochromatic domains on this chromosome and showed that they do not overlap with H3K4me2, a mark of euchromatin (Lewis et al., 2009). All 44 of these domains appear to be relics of RIP, consistent with earlier studies showing that virtually all DNA methylation is associated with RIP’d sequences in Neurospora (Selker et al., 2003; Lewis et al., 2009).
Establishment of heterochromatin domains
In a number of organisms, most notably Schizosaccharomyces pombe (Cam and Grewal, 2004; Allshire and Selker, 2007), RNAi-dependent mechanisms have been implicated in the establishment of heterochromatin. In contrast, known elements of Neurospora's RNAi machinery (two dicer proteins, three RNA-dependent RNA polymerases, and two argonaut proteins) are not required for the initiation or maintenance of heterochromatin (Freitag et al., 2004a).
Methylation is at the core of heterochromatin formation in Neurospora as both dim-2 and dim-5 encode methyltransferases, the former for DNA methylation and the latter for H3K9 methylation. We will use a bottom up approach, starting with the DNA sequence characteristics that trigger heterochromatin formation, to discuss what has been learned and what remains to be discovered about how heterochromatin domains are established and maintained in this relatively simple organism.
RIP creates the signal
The majority of sequences altered by RIP trigger de novo methylation in vegetative tissue (Singer et al., 1995). This was first shown by removing the existing methylation, either by treatment with the methylation inhibitor 5-azacytidine or by molecular cloning followed by transformation back into Neurospora (Selker et al., 1987; Singer et al., 1995). More recently, the distribution of DNA methylation was analyzed on all of Linkage Group VII after passage through the DNA methylation mutant, dim-2, followed by re-introduction back into a DNA methylation-proficient background. Strikingly, DNA methylation was reestablished at all of the 44 heterochromatic regions identified on this chromosome (Lewis et al., 2009). As pointed out above, all DNA methylation in vegetative cells is dependent on DIM-2; RID is not required to establish DNA methylation in vegetative tissue (Freitag et al., 2002). Notably, although the majority of RIP’d sequences can trigger de novo DNA methylation in vegetative tissue, there are some examples of RIP’d sequences that cannot. These sequences, which are relatively lightly RIP’d, provided the first evidence of maintenance methylation in Neurospora (Singer et al., 1995). It is attractive to suppose that DNA methylation was established by RID during the sexual phase and was then maintained through vegetative growth by DIM-2.
What are the characteristics of a RIP’d sequence that cause it to trigger de novo DNA methylation? Although any cytosine in a duplicated sequence can be mutated, RIP has sequence preference for C residues preceding A's (that is, the dinucleotide CpA) (Margolin et al., 1998). This produces a recognizable signature, namely the resulting high density of TpA dinucleotides, and correspondingly low density of CpA dinucleotides (and the complementary GpT dinucleotides), rendering identification of RIP’d sequences straightforward (Margolin et al., 1998). Studies of the cis-acting signals of RIP’d sequences concluded that the higher A:T richness and TpA density are not essential features of the DNA methylation signal but are attributes of additive, positive signals that promote de novo DNA methylation (Miao et al., 2000). Further refinement of the de novo methylation signal revealed that both A and T residues are required on both strands and suggested that multiples of the sequence (TAAA)n or (TTAA)n constitute the most potent methylation signals (Tamaru and Selker, 2003). In addition, weak signals could by strengthened by extending their length, whereas the presence of G:C base pairs suppressed DNA methylation with Cs 5′ of ApT dinucleotides being the most inhibitory (Tamaru and Selker, 2003).
Recognition of the signal
As no single sequence motif serves as the ‘de novo DNA methylation signal’ in RIP’d sequences, it seems likely the factor(s) that are involved in recognizing RIP’d sequences show relatively indiscriminate recognition of the sequence alterations produced by RIP. A protein activity dubbed methyl/RIP-binding protein 1 has been detected by gel mobility shift experiments, which shows preference for RIP’d sequences, relative to its nonRIP’d counterpart; however, this activity has the strongest affinity for RIP’d sequences that are also methylated (Selker et al., 2002; Kothe and Selker, unpublished). Considering that the A:T-hook motif of proteins (for example, high mobility group proteins) is known to bind in the minor groove of A:T-rich sequences (Reeves, 2001), A:T-hook proteins are candidates for recognition of RIP’d sequences. Supporting this hypothesis, distamycin A, a chemical analog of the A:T-hook motif, was found to interfere with DNA methylation in Neurospora (Tamaru and Selker, 2003). Neurospora has at least 30 genes encoding proteins with an A:T-hook domain. Study of one of these A:T-hook motif proteins, led to the discovery of a putative histone demethylase (DNA Methylation Modifier (DMM-1); see below) that does have a role in DNA methylation, but did not turn out to be the elusive RIP recognition factor (Honda et al., 2010).
Histone H3K9 methylation marks the way
Neurospora's dim-5 encodes a histone methyltransferase that is responsible for the tri-methylation of the lysine 9 residue on histone H3 (H3K9me3) (Table 1) (Tamaru and Selker, 2001). Neurospora strains lacking functional DIM-5 show a complete loss of DNA methylation, indicating that H3K9me3 is essential for DNA methylation (Figure 1). Although the situation appears more complicated in higher organisms, there is evidence that methylation of lysine 9 of histone H3 (H3K9me) also has roles in DNA methylation at some sites in both plants and animals (Jackson et al., 2002; Lehnertz et al., 2003).
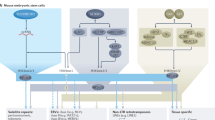
Model for de novo heterochromatin formation of RIP’d sequences. Mutations by RIP typically create an AT-rich cis-acting signal for de novo heterochromatin formation. In the upper section, DNA is shown wrapped around green nucleosomes with a region of RIP’d DNA represented in red and flanking unRIP’d DNA in black. The double arrow and associated question mark (?) represent the recognition of the RIP’d DNA by one or more unknown factors important for the initial step(s) in the establishment of heterochromatin. In the middle section, a sequence of events is depicted on a representative nucleosome from the RIP’d region. The cell cycle regulated H3S10p (yellow lollipops) is a posttranslational modification that is inhibitory to both the action of the DIM-5 complex (red oval with unidentified members of the complex represented by the ‘?’) and to the binding of the chromo-domain protein HP1 (green oval). Removal of H3S10p by the phosphatase, PP1 (orange oval), allows the DIM-5 complex to trimethylate H3K9 (red triple-headed lollipops) and initiate heterochromatin formation. HP1 binds to the H3K9me3 through its chromo-domain and recruits the DNA methyltransferase DIM-2 (blue oval) through a direct interaction of its chromo-shadow domain with two PXVXL-related motifs in DIM-2. DIM-2 methylates any cytosine residue (white circles) within the associated RIP’d DNA. In the bottom section, the fully modified RIP’d region is depicted in a more condensed state with bound HP1 concentrated at the flanks of the region where it recruits the DMM complex (light blue oval). The DMM complex prevents the heterochromatin from spreading into neighboring unRIP’d DNA by inhibiting H3K9me3.
As mentioned earlier, H3K9me3 shows extensive co-localization with DNA methylation at the 44 heterochromatin domains along Neurospora Linkage Group VII (Lewis et al., 2009). In the absence of DNA methylation, in a dim-2 mutant, H3K9me3 remained at these domains, confirming that DIM-5 is upstream of dim-2. How DIM-5 is targeted to RIP’d sequences is unclear, but presumably their is a factor or histone modification that is responsible for recruiting DIM-5. As pointed out above, unlike the situation in S. pombe, where the DIM-5 homologue, Clr4, is targeted by the RNA-induced transcriptional silencing complex (Zhang et al., 2008), the RNAi pathway does not appear to be involved in establishing or maintaining heterochromatin in Neurospora (Freitag et al., 2004a). Furthermore, unlike DIM-5, the H3K9 histone methyltransferases in S. pombe, Drosophila, and mammals contain a chromo-domain that binds to H3K9me3, which apparently helps to maintain and spread this modification in these systems (Zhang et al., 2008).
HP1 recruits DIM-2
Once H3K9me3 is established, how does this lead to DNA methylation by DIM-2? Proteins with a chromo-domain are known to bind to methylated lysine residues in histones. Most notably, HP1, first described in Drosophila, has been shown to bind to methylated H3K9 (Bannister et al., 2001; Lachner et al., 2001). HP1 has been implicated in the spreading and maintenance of H3K9 methylation (Hall et al., 2002; Schotta et al., 2002). Neurospora has several chromo-domain proteins, one of which also has a chromo-shadow domain, characteristic of HP1 homologues. In contrast to the results in fission yeast and flies, inactivation of the hpo gene, which encodes the HP1 homologue of Neurospora, showed very little alteration in H3K9me3 (Lewis et al., 2009). However, hpo mutants show a complete loss of DNA methylation in vegetative tissue, supporting the idea that this protein is required for reading the H3K9me3 mark and relaying this information to the DNA methylation machinery (Table 1) (Freitag et al., 2004b). Consistent with this, HP1 was found to generally co-localize with H3K9me3 along Linkage Group V11 in Neurospora (Lewis et al., 2009). In addition, HP1 was found localized to discrete heterochromatic foci in nuclei and this localization was lost when H3K9me3 was eliminated by mutation of dim-5 (Freitag et al., 2004b). More recent work showed that the chromo-shadow domain of HP1 interacts directly with the two PXVXL-related domains of DIM-2 (Table 1) (Honda and Selker, 2008). Thus, HP1 provides the link between H3K9me3 established by DIM-5 and DNA methylation established by DIM-2 (Figure 1). Although these studies of HP1 in Neurospora have provided a direct link between HP1 and DNA methyaltion, HP1 proteins have also been shown to interact and/or colocalize with DMTs and methyl-binding proteins in mammals (Bachman et al., 2001; Fujita et al., 2003; Fuks et al., 2003; Reese et al., 2003). On the other hand, the Arabidopsis HP1 homologue, TFL2/LHP1, is dispensable for DNA methylation (Malagnac et al., 2002).
Containing heterochromatin domains: role of a putative histone demethylase complex
Although silencing of transposons by DNA methylation may be beneficial to an organism, presumably it is important to prevent the DNA methylation from spreading into, and potentially silencing, neighboring genes (Cambareri et al., 1996; Slotkin and Martienssen, 2007). Recent findings suggest that Neurospora uses a protein complex that prevents spreading of DNA methylation and the underlying H3K9me3. The protein DMM-1 (DNA Methylation Modulator; Honda et al., 2010) is the central player. DMM-1 is a 1458 amino-acid protein with three A:T-hook motifs near its C-terminus, making it a candidate to bind to the AT-rich RIP’d sequences, a JmjC domain near its N-terminus, and two cysteine-rich domains (Table 1). Interestingly, JmjC domains are frequently implicated in histone demethylation (Klose et al., 2006). A null mutation of dmm-1 resulted in severe growth defects and a disruption of normal DNA methylation patterns; many RIP’d transposons showed spreading of DNA methylation into neighboring sequences. Honda et al. (2010) hypothesized that the growth defects in dmm-1 mutants result from the inappropriate silencing of critical genes because of the spreading of DNA methylation from RIP’d transposons. Consistent with this hypothesis, the dmm-1 growth defects were complemented when DNA methylation was eliminated in a dim-2 background. Furthermore, the spreading of methylation into a neighboring gene (NCU02414) in a dmm-1 mutant resulted in reduced gene expression that was relieved in a dmm-1 dim-2 double mutant.
A second DMM protein, DMM-2, was found to co-immunoprecipitate with DMM-1 (Honda et al., 2010). DMM-2 is a 1133 amino-acid protein containing a fungal-specific Zn(II)2Cys6 DNA-binding domain (Table 1). Although dmm-2 strains show DNA methylation defects congruous with dmm-1 strains, the hypermethylation in dmm-2 strains is less extreme. Consistent with this observation, dmm-2 strains do not have the severe growth defects seen in dmm-1 strains. Presumably, DNA methylation spreads into and silences important endogenous genes in dmm-1 strains whereas the less pronounced hypermethylation defect of dmm-2 strains results in less aberrant silencing and thus less growth defects.
Although histone demethylase activity has not been detected in vitro for DMM-1, it is attractive to speculate that the JmjC domain of DMM-1 has histone demethylase activity that prevents the spreading of DNA methylation by removing H3K9me3 deposited by DIM-5 (Figure 1). Such a role would be consistent with the observed localization of DMM-1 at the edges of methylated regions (Honda et al., 2010). Interestingly, this localization is dependent on both the presence of H3K9me3 and on interaction with HP1, the location of which (Table 1; Figure 1) is found preferentially at the edges of methylated regions (Lewis et al., 2009). Thus, HP1 not only serves as an adapter to promote DNA methylation by recruiting the DMT, DIM-2, but also to recruit the DMM-1/2 complex that prevents the spreading of DNA methylation (Figure 1). Somewhat similar anti-silencing mechanisms that use a JmjC domain protein have been identified in Arabidopsis and fission yeast (Zofall and Grewal, 2006; Saze et al., 2008; Miura et al., 2009).
Cell cycle propagation of heterochromatin
Proper propagation of chromatin domains during passage through the cell cycle is essential to insure that regulatory controls are maintained. Interplay of histone modifications is critical for this propagation. Notably, phosphorylation of the serine 10 residue of histone H3 (H3S10p) is required for the faithful propagation of heterochromatin through the cell cycle. Increased H3S10p is required in fission yeast during S phase to drive the transcription of repeats in heterochromatin regions to generate small RNAs that nucleate heterochromatin formation by recruiting the RNA-induced transcriptional silencing and ClrC complexes (Chen et al., 2008; Kloc et al., 2008; Zhang et al., 2008). Such an RNAi-based mechanism may not be necessary for heterochromatin in Neurospora, however (Freitag et al., 2004a). The H3S10 residue is known to be phosphorylated by a number of kinases in response to environmental signals to activate genes, or in response to internal signals that initiate chromosome condensation and segregation during mitosis and meiosis (Prigent and Dimitrov, 2003; Nowak and Corces, 2004; Johansen and Johansen, 2006). Phosphorylation of H3S10 occurs at the onset of mitosis, disrupting the binding of HP1 to methylated H3K9 at pericentric heterochromatin (Fischle et al., 2005; Hirota et al., 2005; Chen et al., 2008; Kloc et al., 2008). Re-association of HP1 cannot occur until H3S10p is dephosphorylated (Fischle et al., 2005; Hirota et al., 2005; Chen et al., 2008). Structural studies have shown that DIM-5 directly interacts with S10 of H3, the residue next to its site of action, H3K9, and that this interaction is critical for its activity (Zhang et al., 2003; Rathert et al., 2008; Adhvaryu et al., unpublished results). In vitro assays with H3 peptide substrates have shown that phosphorylation of S10 inhibits the activity of mammalian SUV39H1, fission yeast Clr4, and Neurospora DIM-5 (Rea et al., 2000; Nakayama et al., 2001; Adhvaryu et al., unpublished results). Consistent with the idea that H3S10p is inhibitory to DIM-5 binding and activity, a partial loss-of-function mutant defective in the gene encoding the H3S10 protein phosphatase, PP1, results in growth and developmental defects as well as deficits of H3K9me3 and DNA methylation (Adhvaryu and Selker, 2008). Hence, removal of the H3S10p modification by PP1 during the cell cycle is required to allow DIM-5 and HP1 binding, resulting in the propagation of heterochromatin (Figure 1).
Function of heterochromatin
As mentioned above, virtually all heterochromatin domains in Neurospora appear to be based on sequences altered by RIP (Lewis et al., 2009); the only currently known exception being the tandemly arranged rRNA repeats. Most RIP’d sequences are relics of transposons and it is presumably in the best interest of an organism to protect its genome by silencing these invasive elements (Galagan et al., 2003). The mutations that occur by RIP would seem to be an effective barrier against these transposons. However, as a result of these transition mutations, expression of RIP’d sequences could lead to the production of dominant negative proteins, or RNAs with deleterious effects. Thus, relics of RIP may be packaged into heterochromatin to silence their expression.
Removal of DNA methylation, either with the drug 5-azacytidine or with a dim-2 mutation, can lead to activation of transcription (Irelan and Selker, 1997; Rountree and Selker, 1997). Furthermore, DNA methylation appears to exert its suppressive effect downstream of transcription initiation in Neurospora (Rountree and Selker, 1997). These Neurospora studies showed that once DNA methylation is removed, expression is restored suggesting that DNA methylation is the critical modification that ‘locks down’ the heterochromatin, making it refractory to transcription. Interestingly, although H3K9 methylation is largely upstream of DNA methylation, H3K9me3 was lost from a RIP’d allele of the Neurospora am gene (amRIP8) in a dim-2 mutant (Tamaru et al., 2003). This amRIP8 allele is known to have a functional promoter (Singer et al., 1995; Rountree and Selker, 1997). In contrast, H3K9me3 was not lost from any of the natural heterochromatic regions along Linkage Group VII in a dim-2 mutant (Lewis et al., 2009). Lewis et al. (2009) suggested that the extensive RIP of the heterochromatic sequences on Linkage Group VII probably inactivated promoter sequences. Thus, in the absence of DNA methylation, transcriptional activity may be capable of overcoming and preventing the formation of heterochromatin.
In animals, DNA methylation induced transcriptional repression has been attributed to a series of methylated DNA-binding proteins that complex with chromatin modifiers to establish, or at least perpetuate, a silenced chromatin state (Fuks, 2005; Sasai and Defossez, 2009). A number of methylated DNA-binding proteins have been identified in plants based on homology to the mammalian methyl-binding domain motif, but Neurospora lacks proteins with homology to higher eukaryotic methylated DNA-binding proteins (Zemach and Grafi, 2007). However, as mentioned above, a methylated DNA-binding activity (methyl/RIP-binding protein 1) has been detected that has the strongest affinity for RIP’d methylated sequences (Selker et al., 2002). The role of this factor in the establishment and perpetuation of heterochromatin requires further work.
Conclusion and perspectives
Although our understanding of how heterochromatin domains are established and maintained in Neurospora has grown considerably, many questions remain to be answered regarding these processes, including but certainly not limited to: What factor(s) recognizes RIP’d sequences and recruit the DIM-5 complex to these sequences? Where in the process of heterochromatin formation is histone deacetylation critical? How does the DMM complex inhibit H3K9me3 at the boundaries of heterochromatin regions? What other histone modifications have a role in heterochromatin formation and/or maintenance? Why do hpo-1 and dim-5 mutants display growth defects, unlike dim-2? The answer to this latter question most likely has to do with the role of HP1 and DIM-5 in additional cellular processes, such as chromosomal segregation. DNA methylation can be regarded as the ‘final layer’ of heterochromatin, which though not essential for the cell, ‘locks’ down the heterochromatin to prevent transcription of invasive elements.
References
Adhvaryu KK, Selker EU (2008). Protein phosphatase PP1 is required for normal DNA methylation in Neurospora. Genes Dev 22: 3391–3396.
Allshire R, Selker EU (2007). Fungal models for epigenetic research: Schizosaccharomyces pombe and Neurospora crassa. Epigenetics In: Allis CD, Jenuwein T, Reinberg D (eds). Cold Spring Harbor Laboratory Press: Cold Spring Harbor, New York, USA, 101–125.
Bachman KE, Rountree MR, Baylin SB (2001). Dnmt3a and Dnmt3b are transcriptional repressors that exhibit unique localization properties to heterochromatin. J Biol Chem 276: 32282–32287.
Bannister AJ, Zegerman P, Partridge JF, Miska EA, Thomas JO, Allshire RC et al. (2001). Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410: 120–124.
Bhaumik SR, Smith E, Shilatifard A (2007). Covalent modifications of histones during development and disease pathogenesis. Nat Struct Mol Biol 14: 1008–1016.
Cam H, Grewal SI (2004). RNA interference and epigenetic control of heterochromatin assembly in fission yeast. Cold Spring Harb Symp Quant Biol 69: 419–427.
Cambareri EB, Foss HM, Rountree MR, Selker EU, Kinsey JA (1996). Epigenetic control of a transposon-inactivated gene in Neurospora is dependent on DNA methylation. Genetics 143: 137–146.
Cambareri EB, Jensen BC, Schabtach E, Selker EU (1989). Repeat-induced G-C to A-T mutations in Neurospora. Science 244: 1571–1575.
Cambareri EB, Singer MJ, Selker EU (1991). Recurrence of repeat-induced point mutation (RIP) in Neurospora crassa. Genetics 127: 699–710.
Chen ES, Zhang K, Nicolas E, Cam HP, Zofall M, Grewal SI (2008). Cell cycle control of centromeric repeat transcription and heterochromatin assembly. Nature 451: 734–737.
Fischle W, Tseng BS, Dormann HL, Ueberheide BM, Garcia BA, Shabanowitz J et al. (2005). Regulation of HP1-chromatin binding by histone H3 methylation and phosphorylation. Nature 438: 1116–1122. (e-pub ahead of print 12 October 2005).
Foss HM, Roberts CJ, Claeys KM, Selker EU (1993). Abnormal chromosome behavior in Neurospora mutants defective in DNA methylation. Science 262: 1737–1741.
Freitag M, Lee DW, Kothe GO, Pratt RJ, Aramayo R, Selker EU (2004a). DNA methylation is independent of RNA interference in Neurospora. Science 304: 1939.
Freitag M, Hickey PC, Khlafallah TK, Read ND, Selker EU (2004b). HP1 is essential for DNA methylation in Neurospora. Mol Cell 13: 427–434.
Freitag M, Williams RL, Kothe GO, Selker EU (2002). A cytosine methyltransferase homologue is essential for repeat-induced point mutation in Neurospora crassa. Proc Natl Acad Sci USA 99: 8802–8807.
Fujita N, Watanabe S, Ichimura T, Tsuruzoe S, Shinkai Y, Tachibana M et al. (2003). Methyl-CpG binding domain 1 (MBD1) interacts with the Suv39h1-HP1 heterochromatic complex for DNA methylation-based transcriptional repression. J Biol Chem 278: 24132–24138.
Fuks F (2005). DNA methylation and histone modifications: teaming up to silence genes. Curr Opin Genet Dev 15: 490–495.
Fuks F, Hurd PJ, Deplus R, Kouzarides T (2003). The DNA methyltransferases associate with HP1 and the SUV39H1 histone methyltransferase. Nucleic Acids Res 31: 2305–2312.
Galagan JE, Calvo SE, Borkovich KA, Selker EU, Read ND, Jaffe D et al. (2003). The genome sequence of the filamentous fungus Neurospora crassa. Nature 422: 859–868.
Grewal SI, Jia S (2007). Heterochromatin revisited. Nat Rev Genet 8: 35–46.
Hall IM, Shankaranarayana GD, Noma KI, Ayoub N, Cohen A, Grewal SI (2002). Establishment and maintenance of a heterochromatin domain. Science 297: 2232–2237.
Hirota T, Lipp JJ, Toh BH, Peters JM (2005). Histone H3 serine 10 phosphorylation by Aurora B causes HP1 dissociation from heterochromatin. Nature 438: 1176–1180.
Honda S, Lewis ZA, Huarte M, Cho LY, David LL, Shi Y et al. (2010). The DMM complex prevents spreading of DNA methylation from transposons to nearby genes in Neurospora crassa. Genes Dev 24: 443–454.
Honda SH, Selker EU (2008). Direct interaction between DNA methylatransferase DIM-2 and HP1 is required for DNA methylation in Neurospora crassa. Mol Cell Biol 28: 6044–6055.
Irelan JT, Selker EU (1997). Cytosine methylation associated with repeat-induced point mutation causes epigenetic gene silencing in Neurospora crassa. Genetics 146: 509–523.
Jackson JP, Lindroth AM, Cao X, Jacobsen SE (2002). Control of CpNpG DNA methylation by the KRYPTONITE histone H3 methyltransferase. Nature 416: 556–560.
Johansen KM, Johansen J (2006). Regulation of chromatin structure by histone H3S10 phosphorylation. Chromosome Res 14: 393–404.
Kloc A, Zaratiegui M, Nora E, Martienssen R (2008). RNA interference guides histone modification during the S phase of chromosomal replication. Curr Biol 18: 490–495.
Klose RJ, Yamane K, Bae Y, Zhang D, Erdjument-Bromage H, Tempst P et al. (2006). The transcriptional repressor JHDM3A demethylates trimethyl histone H3 lysine 9 and lysine 36. Nature 442: 312–316.
Kouzminova EA, Selker EU (2001). Dim-2 encodes a DNA-methyltransferase responsible for all known cytosine methylation in Neurospora. EMBO J 20: 4309–4323.
Lachner M, O’Carroll D, Rea S, Mechtler K, Jenuwein T (2001). Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410: 116–120.
Lehnertz B, Ueda Y, Derijck AA, Braunschweig U, Perez-Burgos L, Kubicek S et al. (2003). Suv39h-mediated histone h3 lysine 9 methylation directs DNA methylation to major satellite repeats at pericentric heterochromatin. Curr Biol 13: 1192–1200.
Lewis ZA, Honda S, Khlafallah TK, Jeffress JK, Freitag M, Mohn F et al. (2009). Relics of repeat-induced point mutation direct heterochromatin formation in Neurospora crassa. Genome Res 19: 427–437.
Malagnac F, Bartee L, Bender J (2002). An Arabidopsis SET domain protein required for maintenance but not establishment of DNA methylation. EMBO J 21: 6842–6852.
Margolin BS, Garrett-Engele PW, Stevens JN, Yen-Fritz D, Garrett-Engele C, Metzenberg RL et al. (1998). A methylated Neurospora 5S rRNA pseudogene contains a transposable element inactivated by RIP. Genetics 149: 1787–1797.
Miao VP, Freitag M, Selker EU (2000). Short TpA-rich segments of the zeta-eta region induce DNA methylation in Neurospora crassa. J Mol Biol 300: 249–273.
Miura A, Nakamura M, Inagaki S, Kobayashi A, Saze H, Kakutani T (2009). An Arabidopsis jmjC domain protein protects transcribed genes from DNA methylation at CHG sites. EMBO J 28: 1078–1086.
Nakayama J, Rice JC, Strahl BD, Allis CD, Grewal SI (2001). Role of histone H3 lysine 9 methylation in epigenetic control of heterochromatin assembly. Science 292: 110–113.
Nowak SJ, Corces VG (2004). Phosphorylation of histone H3: a balancing act between chromosome condensation and transcriptional activation. Trends Genet 20: 214–220.
Peng JC, Karpen GH (2008). Epigenetic regulation of heterochromatic DNA stability. Curr Opin Genet Dev 18: 204–211.
Prigent C, Dimitrov S (2003). Phosphorylation of serine 10 in histone H3, what for? J Cell Sci 116: 3677–3685.
Rathert P, Zhang X, Freund C, Cheng X, Jeltsch A (2008). Analysis of the substrate specificity of the Dim-5 histone lysine methyltransferase using peptide arrays. Chem Biol 15: 5–11.
Rea S, Eisenhaber F, O’Carroll D, Strahl BD, Sun ZW, Schmid M et al. (2000). Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406: 593–599.
Reese BE, Bachman KE, Baylin SB, Rountree MR (2003). The methyl-CpG binding protein MBD1 interacts with the p150 subunit of chromatin assembly factor 1. Mol Cell Biol 23: 3226–3236.
Reeves R (2001). Molecular biology of HMGA proteins: hubs of nuclear function. Gene 277: 63–81.
Rountree MR, Selker EU (1997). DNA methylation inhibits elongation but not initiation of transcription in Neurospora crassa. Genes Dev 11: 2383–2395.
Rountree MR, Selker EU (2009). Genome defense: the neurospora paradigm. In: Ferguson-Smith AC, Greally JM, Martienssen RA (eds). Epigenomics. Springer: Dordrecht, The Netherlands, pp 321–341.
Sasai N, Defossez PA (2009). Many paths to one goal? The proteins that recognize methylated DNA in eukaryotes. Int J Dev Biol 53: 323–334.
Saze H, Shiraishi A, Miura A, Kakutani T (2008). Control of genic DNA methylation by a jmjC domain-containing protein in Arabidopsis thaliana. Science 319: 462–465.
Schotta G, Ebert A, Krauss V, Fischer A, Hoffmann J, Rea S et al. (2002). Central role of Drosophila SU(VAR)3-9 in histone H3-K9 methylation and heterochromatic gene silencing. EMBO J 21: 1121–1131.
Selker EU (1998). Trichostatin A causes selective loss of DNA methylation in Neurospora. Proc Natl Acad Sci USA 95: 9430–9435.
Selker EU (2004). Genome defense and DNA methylation in neurospora. Cold Spring Harb Symp Quant Biol 69: 119–124.
Selker EU, Cambareri EB, Jensen BC, Haack KR (1987). Rearrangement of duplicated DNA in specialized cells of Neurospora. Cell 51: 741–752.
Selker EU, Freitag M, Kothe GO, Margolin BS, Rountree MR, Allis CD et al. (2002). Induction and maintenance of nonsymmetrical DNA methylation in Neurospora. Proc Natl Acad Sci USA 99: 16485–16490.
Selker EU, Fritz DY, Singer MJ (1993). Dense non-symmetrical DNA methylation resulting from repeat-induced point mutation (RIP) in Neurospora. Science 262: 1724–1728.
Selker EU, Garrett PW (1988). DNA sequence duplications trigger gene inactivation in Neurospora crassa. Proc Natl Acad Sci USA 85: 6870–6874.
Selker EU, Jensen BC, Richardson GA (1987). A portable signal causing faithful DNA methylation de novo in Neurospora crassa. Science 238: 48–53.
Selker EU, Stevens JN (1985). DNA methylation at asymmetric sites is associated with numerous transition mutations. Proc Natl Acad Sci USA 82: 8114–8118.
Selker EU, Stevens JN (1987). Signal for DNA methylation associated with tandem duplication in Neurospora crassa. Mol Cell Biol 7: 1032–1038.
Selker EU, Tountas NA, Cross SH, Margolin BS, Murphy JG, Bird AP et al. (2003). The methylated component of the Neurospora crassa genome. Nature 422: 893–897.
Sharma S, Kelly TK, Jones PA (2010). Epigenetics in cancer. Carcinogenesis 31: 27–36.
Singer MJ, Marcotte BA, Selker EU (1995). DNA methylation associated with repeat-induced point mutation in Neurospora crassa. Mol Cell Biol 15: 5586–5597.
Slotkin RK, Martienssen R (2007). Transposable elements and the epigenetic regulation of the genome. Nat Rev Genet 8: 272–285.
Tamaru H, Selker EU (2001). A histone H3 methyltransferase controls DNA methylation in Neurospora crassa. Nature 414: 277–283.
Tamaru H, Selker EU (2003). Synthesis of Signals for De Novo DNA Methylation in Neurospora crassa. Mol Cell Biol 23: 2379–2394.
Tamaru H, Zhang X, McMillen D, Singh PB, Nakayama J, Grewal SI et al. (2003). Trimethylated lysine 9 of histone H3 is a mark for DNA methylation in Neurospora crassa. Nat Genet 34: 75–79.
Zemach A, Grafi G (2007). Methyl-CpG-binding domain proteins in plants: interpreters of DNA methylation. Trends Plant Sci 12: 80–85.
Zhang K, Mosch K, Fischle W, Grewal SI (2008). Roles of the Clr4 methyltransferase complex in nucleation, spreading and maintenance of heterochromatin. Nat Struct Mol Biol 15: 381–388.
Zhang X, Yang Z, Khan SI, Horton JR, Tamaru H, Selker EU et al. (2003). Structural basis for the product specificity of histone lysine methyltransferases. Mol Cell 12: 177–185.
Zofall M, Grewal SI (2006). Swi6/HP1 recruits a JmjC domain protein to facilitate transcription of heterochromatic repeats. Mol Cell 22: 681–692.
Acknowledgements
We thank Zachary Lewis, Shinji Honda, and Keyur Adhvaryu for helpful discussions. This work was supported by NIH Grant GM025690-22 to EUS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Rountree, M., Selker, E. DNA methylation and the formation of heterochromatin in Neurospora crassa. Heredity 105, 38–44 (2010). https://doi.org/10.1038/hdy.2010.44
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2010.44
Keywords
This article is cited by
-
Structural and functional specificity of H3K36 methylation
Epigenetics & Chromatin (2022)
-
Nuclear organization and regulation of the differentiated state
Cellular and Molecular Life Sciences (2021)
-
DNA methyltransferase implicated in the recovery of conidiation, through successive plant passages, in phenotypically degenerated Metarhizium
Applied Microbiology and Biotechnology (2020)
-
Sodium valproate and 5-aza-2′-deoxycytidine differentially modulate DNA demethylation in G1 phase-arrested and proliferative HeLa cells
Scientific Reports (2019)
-
The transcription factor PRO44 and the histone chaperone ASF1 regulate distinct aspects of multicellular development in the filamentous fungus Sordaria macrospora
BMC Genetics (2018)