Abstract
Eukaryotic genomes contain a large proportion of repetitive DNA sequences, mostly transposable elements (TEs) and tandem repeats. These repetitive sequences often colonize specific chromosomal (Y or W chromosomes, B chromosomes) or subchromosomal (telomeres, centromeres) niches. Sex chromosomes, especially non-recombining regions of the Y chromosome, are subject to different evolutionary forces compared with autosomes. In non-recombining regions of the Y chromosome repetitive DNA sequences are accumulated, representing a dominant and early process forming the Y chromosome, probably before genes start to degenerate. Here we review the occurrence and role of repetitive DNA in Y chromosome evolution in various species with a focus on dioecious plants. We also discuss the potential link between recombination and transposition in shaping genomes.
Similar content being viewed by others
Introduction
Transposable elements (TEs), both retrotransposons and DNA transposons, considered earlier as ‘junk DNA,’ are now viewed as major players in genome evolution (Kidwell and Lisch, 2001; Biemont and Vieira, 2006). Their distribution in genomes is variable: some are widespread, while others colonize specific chromosomal niches. Repeated sequence elements can be viewed as members of a community inhabiting the genome ecosystem (Leonardo and Nuzhdin, 2002; Brookfield, 2005).
The chromosomal distribution of TEs is often governed by the status of chromatin compaction and rates of recombination of particular genomic regions. In general, regions with suppressed or no recombination tend to accumulate repetitive DNA sequences (for review see Charlesworth et al., 1994; Gvozdev et al., 2005). Non-recombining parts of the Y chromosome have accumulated repetitive DNA, particularly in the mammalian (Erlandsson et al., 2000; Skaletsky et al., 2003) and Drosophila melanogaster (Pimpinelli et al., 1995) Y chromosomes, which are evolutionarily ancient, originating before 165 and 60 mya, respectively (Graves, 2005; Veyrunes et al., 2008). However, the process of accumulation of repetitive sequences is apparent even in evolutionarily young sex chromosomes, such as in Drosophila miranda (Steinemann and Steinemann, 1992), Silene latifolia (Hobza et al., 2006; Kejnovsky et al., 2006a) or Carica papaya (Liu et al., 2004). Repeat accumulation thus potentially represents an early process shaping the Y chromosome, even before the genes start to degenerate (Steinemann and Steinemann, 2005).
Sex chromosomes: special parts of genomes
The definition of a sex chromosome is not always clear-cut. It is generally applied to a pair of chromosomes on which the sex determination locus resides regardless of whether they are morphologically distinguishable (heteromorphic) and mostly non-recombining, or conversely, morphologically indistinguishable (homomorphic) and recombining along much of their length. In most species the males are the heterogametic (XY) sex, whereas the females are homogametic (XX). In some groups such as birds and lepidopterans, the reverse is true: females are heterogametic (ZW) and males are homogametic (ZZ). Here we will focus on the XY/XX system, but the same processes were observed in ZW/ZZ system as well (Traut and Marec, 1997; Ezaz et al., 2006; Tsuda et al., 2007).
The origin and evolution of sex chromosomes have interested evolutionary biologists for a long time. Although sex chromosomes evolve from a pair of autosomes (Ohno, 1967), over time they become different, both from each other and the autosomes, in gene content and structure. While sex chromosomes in most mammals are ancient, sex chromosomes in some fish, platypus, some insects and dioecious plants are evolutionarily young (for review see Vyskot and Hobza, 2004; Charlesworth et al., 2005). Despite the different ages of sex chromosomes in different taxonomic groups, they probably follow similar evolutionary trajectories with discrete identifiable stages.
The trajectory from autosomes to sex chromosomes may start with the emergence of a sex-determining gene with one allele that determines male individuals and the other female individuals. A two-loci model was suggested by Charlesworth et al. (2005) in which mutation in one gene on the proto-X chromosome results in male sterility and mutation in another gene on the proto-Y chromosome results in female sterility. Acquisition of a sex-determining gene(s) may be followed by the suppression of recombination in the vicinity of the gene(s). Later, other sex-determining genes that influence the development of a particular sex, or are antagonistic to the opposing sex, may accumulate around the sex-determining gene. Such sex-determining regions can, in some instances, translocate between chromosomes and create new sex chromosomes (Traut and Willhoeft, 1990; Willhoeft and Traut, 1990; Traut and Wollert, 1998). The initiating mechanisms of recombination suppression are not yet clear, though some models based on epigenetic silencing (Jablonka, 2004) or inversions (Lahn and Page, 1999) have been proposed. Zluvova et al. (2005) suggested that the inversion on the Y chromosome of Silene latifolia is a consequence of recombination arrest as opposed to its cause. In addition, non-recombining regions may expand through the accumulation of repetitive DNA sequences (Charlesworth, 1991), which often form heterochromatin. Ultimately the processes, because of a lack of recombination in meiosis, may lead to a lower expression of the Y-linked genes and eventually to their degeneration into pseudogenes (Bachtrog, 2006; Marais et al., 2008). Once the Y chromosome has become a genetic desert, the balance between mechanisms that expand the chromosome (for example, transposition) and those that cause it to contract (for example, deletions), will govern the long-term fate of the chromosome. At this stage, large regions of the Y chromosome without genes are dispensable and can be lost. It is not clear yet whether the processes of gene degeneration, TEs accumulation and expansions on one hand and contractions on the other hand are stepwise or are occurring simultaneously.
However, the lifetime of an old Y chromosome is often prolonged by the addition of segments transferred from autosomes (Graves, 2005). Acquisition of new genes from autosomes mediated by retrotransposition has been shown in humans (Lahn and Page, 1999), and a similar duplicative transfer has also been shown in the young Y chromosomes of Silene latifolia (Matsunaga et al., 2003). A final possibility is that sex is determined not by the specific Y-linked gene, but only by the ratio of X chromosome to autosomes (X/A ratio). Then the Y chromosome either remains as a genetic entity or could be lost entirely. A new autosomal pair can then be chosen to become a new pair of sex chromosomes and the cyclic process can continue. The persistence of the Y chromosome indicates that it can repeatedly arise de novo, for example, by the fusion between an autosome and an X chromosome followed by the fixation of the neo-X and the neo-Y chromosomes as was shown in grasshopper Podisma pedestris (Westerman and Hewitt, 1985; Veltsos et al., 2008).
Forces different from those working on the Y chromosome or autosomes form the X chromosome. The X chromosome accumulates both recessive genes advantageous for males (masculinizing the X) and dominant genes advantageous for females (feminizing the X). In mammals, for example, genes primarily expressed in the brain and muscle or involved in sperm production, are enriched on the X chromosome (Hurst, 2001). These genes are often transferred onto the X chromosome by retrotransposition. In mammals, it was shown that retrotransposition both on and off the X chromosome is more frequent than between autosomes (Emerson et al., 2004; Khil et al., 2005) in which gene movement from the X chromosome is probably caused by meiotic sex chromosome inactivation (MSCI), during which sex chromosomes are condensed and transcription is repressed. Gene movement from the X chromosome then results in copies of the originally X-linked genes on autosomes.
It is not yet clear whether the accumulation of retrotransposing sequences on the X chromosome is a result of more efficient selection against this process in autosomes or preferential targeting to the X chromosome. Retrotranspositions to the X chromosome may also be a consequence of LINE element activity (Myers et al., 2002). However, the presence of recent elements (∼2 mya) on the X chromosome supports the latter argument (Myers et al., 2002). L1 elements show a two-fold higher enrichment on the human X chromosome and are clustered at the X inactivation centre. Perhaps the L1 elements serve as a DNA signal to propagate and stabilize X chromosome inactivation (Bailey et al., 2000; Lyon, 2000).
Processes acting in regions of reduced recombination
There are three models describing the population genetics of TEs in regions of reduced recombination (for review see Nuzhdin, 1999; Hua-Van et al., 2005). First, the ‘deleterious insertion model’ is based on the higher elimination of TEs from high-gene density regions (Charlesworth, 1991). Second, the ‘ectopic recombination model’ explains the higher abundance of TEs in low recombining regions by the reduced frequency of their removal by ectopic recombination (Langley et al., 1988). Third, the ‘deleterious transposition model’ is based on the deleterious effects of transposition, for example, the formation of chromosome breaks (Brookfield, 1991).
Centromeres are regions with lower recombination frequency (Beadle, 1932; Clarke and Carbon, 1980) and often exhibit interspersion of satellite sequences and TEs, as in cereals (Zhong et al., 2002), Arabidopsis (Copenhaver et al., 1999), insects (Sun et al., 2003) and fungi (Cambareri et al., 1998). There are indications of the evolutionary link between the centromere structure and TE's activity: centromeric satellite repeats may arise from DNA transposons (Kapitonov and Jurka, 1999). There are also similarities between the pogo-like superfamily of DNA transposons and the centromeric CENP-B protein (Smith and Riggs, 1996). CENP-B may have DNA nicking activity analogous to transposase, and it could promote the evolution of satellite arrays by stimulating homologous recombination (Kipling and Warburton, 1997). It is interesting that the CENP-B satellite is absent on the Y chromosome in a number of mammalian species from the only centromere in the genome that does not have a pairing partner in meiosis (Kipling and Warburton, 1997). Similarly, centromeric satellites STAR-C and STAR-Y are underrepresented only in centromeres of the Y chromosome in Silene latifolia (Hobza et al., 2007; Cermak et al., 2008).
The targeting of transposons can be achieved by the targeting domain of integrase, which interacts with specific DNA--protein complexes such as telomeric heterochromatin (Xie et al., 2001). Transposons can also recognize specific DNA sequence (Jurka, 1997) or DNA secondary structure, such as kinked DNAs (Jurka et al., 1998). Integration is also influenced by the epigenetic state of the target region. Many studies have been done on yeast showing targeting of telomeric or subtelomeric repeats as well as rDNA loci (Xie et al., 2001; Zhu et al., 2003). Some TEs integrate into silent regions while others into actively transcribed regions (Kirchner et al., 1995).
DNA-based elements and retrotransposons differ in mechanisms of their amplification. Their dynamics and distribution in genomes are different but the reasons why are not clear (Hua-Van et al., 2005) but it could be influenced by recombination. In Drosophila there is a negative correlation between abundance of DNA transposons and recombination rate (Rizzon et al., 2002). In contrast, in C. elegans, DNA transposons but not retrotransposons are located preferentially in regions with a high recombination rate (Duret et al., 2000). The contrasting relationships between TEs and recombination in these two species can be explained by differences in meiotic pairing and recombination mechanisms (Rizzon et al., 2002). The genome might be more accessible for transposon insertions in regions of intense recombination. It remains unclear whether recombination promotes transposon insertions or, vice versa (Duret et al., 2000), although the latter explanation is supported by the finding that MuDR transposase increases the frequency of meiotic crossovers in the vicinity of Mu insertions (Yandeau–Nelson et al., 2005). The relationship between transposons and recombination (Downs and Jackson, 1999; Kapitonov and Jurka, 2005) is likely to have significant evolutionary implications, especially as many DNA transposons have an affinity to insert into genes (Bureau and Wessler, 1992; Bureau et al., 1996).
Together, recombination processes and the activity of TEs, lead to a high turnover of elements in eukaryotic genomes, particularly in plants (Gaut et al., 2007). Ectopic recombination (unequal recombination, that is, recombination between homologous elements located in non-homologous positions) and TEs can cause large rearrangements. Ectopic recombination has been suggested to be the most important mechanism of genome size reduction and may provide a partial ‘return ticket from genomic obesity’ (Vicient et al., 1999; Petrov et al., 2003). The evidence that deletions are mediated by recombination is the presence of ‘solo long terminal repeats (LTRs)’ that remain in the genome after recombination between two LTRs deletes the internal regions (Vitte and Panaud, 2003). It is possible that this reduction took place on the small mammalian Y chromosome during its evolution. If ectopic recombination between retrotransposons played an important role in size reduction, we can predict a higher frequency of ‘solo LTRs’ on the human Y chromosome compared with other chromosomes.
Another recombination-related process is gene conversion, which can homogenize elements of a repeat family resulting in concerted evolution (Liao, 2003). Gene conversion most frequently occurs within, rather than between, chromosomes as indicated by the higher intrachromosomal similarity of TEs and tandem repeats (Hood et al., 2005; Krzywinski et al., 2005). Similarly, in white campion (Silene latifolia), TEs are more homogenous on the Y chromosome than on the X chromosome and autosomes (Kejnovsky et al., 2007).
The other important process acting in non-recombining regions is gene degeneration (Charlesworth, 1991; Charlesworth and Charlesworth, 2000). Degeneration could be a consequence of TE accumulation (Steinemann and Steinemann, 2005; Marais et al., 2008). The random inactivation model (Bachtrog, 2006) suggests that the process of gene inactivation is triggered by the disruption of promoter regions by TE insertion. TE insertions can lead to an epigenetic phenomenon, or global changes in chromatin status (heterochromatization). In D. miranda the random inactivation model best explains neo-Y chromosome inactivation. Recent results by Zhang et al. (2008) on the papaya show that in MSY (male-specific region of the Y chromosome), but not in corresponding X chromosomal region, there are multiple knob-like heterochromatin structures. Such knobs are usually composed of satellite DNA and transposons. Moreover, in the papaya they show that the knob regions are extensively methylated. Heterochromatinization of the region, which may have originally served as a defense against TEs, probably started, or at least accelerated, the degeneration of the MSY in the papaya (Zhang et al., 2008).
Bachtrog et al. (2008) showed that in the neo-Y chromosome of D. miranda there are 20 times as many TE insertions compared with the neo-X chromosome. They also show that the fraction of TE insertions is similar in functional and non-functional genes. These data suggest the role of TE insertions adjacent to the genes in gene inactivation on the neo-Y. It is still not clear whether TEs affect the vicinity of genes in terms of epigenetic status by chromatin modification in the vicinity of TE insertion or by producing antisense transcripts of adjacent genes and subsequent inactivation by RNAi (Slotkin and Martienssen, 2007).
Accumulation of repetitive DNA in plant sex chromosomes
The main stages of the sex chromosome evolution: the establishment of the sex-determining region, local suppression of recombination, accumulation of repeats, degeneration of genes, and shrinkage by deletions: have been shown in several plant species. In contrast to mammals, most dioecious plants do not possess heteromorphic sex chromosomes (Gorelick, 2005). However about one-tenth of angiosperm species have separate sexes (Yampolsky and Yampolsky, 1922), and of these only a few species are known to have heteromorphic sex chromosomes (Vyskot and Hobza, 2004). In plants with heteromorphic sex chromosomes, the Y chromosome is usually larger than the X chromosome: the reverse of the situation in mammals. Perhaps the larger Y in plants is caused by the accumulation of repeats; indeed, plant genomes in general show high levels of genome plasticity compared with mammals. There are several plant model species that enable the study of different stages of sex chromosome evolution.
The sex in the squirting cucumber (Ecballium elaterium) is determined by one locus localized on a structurally and functionally undifferentiated pair of autosomes. The Y chromosome of the asparagus (Asparagus officinalis) is probably evolutionarily young, as the YY genotype is viable and only a small amount of the Y chromosome contains constitutive heterochromatin (Loptien, 1979). Sex chromosomes of the kiwi fruit (Actinidia chinensis) are at a similar stage (Harvey et al., 1997). In the papaya (Carica papaya), the MSY is short: about 8–9 Mbp (Yu et al., 2008), covering about 20% of the Y chromosome. The MSY region has lost many genes, and the YY genotype is lethal. Despite the short period of time for the divergence of the X and Y, the MSY region of the Y chromosome is already enriched for local segmental duplications, TEs and plastid DNA insertions compared with its X partner (Liu et al., 2004; Yu et al., 2007).
Diversification of sex chromosomes can lead to heteromorphic sex chromosomes that are found in mammals, Drosophila, and some dioecious plants: liverwort (Marchantia polymorpha), white campion (Silene latifolia), sorrel (Rumex acetosa), hop (Humulus lupulus) and hemp (Cannabis sativa). In hop, the Y chromosome is the smallest chromosome in the genome (Winge, 1929); however, there are no data concerning the specific structure of sex chromosomes in this species. Hemp shows male-specific accumulation of LINE-like retrotransposons (Sakamoto et al., 2000), as well as a high abundance of MADC3 and MADC4 (male-associated DNA sequences in C. sativa), encoding gag/pol polyproteins of copia-like retrotransposons (Sakamoto et al., 2005). The Y chromosome of liverwort has specific repeat sequences that contain multiplicated genes. These data suggest that the Y chromosome-amplified protein-coding genes evolved in concert with specific repeat sequences (Okada et al., 2001; Ishizaki et al., 2002).
In the white campion, the Y chromosome is the largest chromosome in the karyotype and is mostly non-recombining with the X chromosome. The homologous genes characterized on the X and Y chromosomes of S. latifolia have not diverged much, yet the YY genotype is not viable, indicating a significant erosion of crucial genes on the Y chromosome. It is interesting that the Y chromosome of S. latifolia is euchromatic except at subtelomeric regions, which contain clusters of tandem repeats (reviewed below).
Sorrel is unique among plants because it has XY1Y2 males and XX females (Kihara and Ono, 1923). Sex is determined by the ratio of X chromosomes to autosomes. It is also the only example of a plant possessing largely heterochromatic Y chromosomes (Pazourkova, 1964) condensed at interphase (Ruiz Rejon et al., 1994; Lengerova and Vyskot, 2001). Sorrel Y chromosomes contain two types of satellite DNA called RAE180 and RAYS (Shibata et al., 1999, 2000). RAYSI and RAYSIII are specific for both Y chromosomes whereas RAYSII is specific for the Y1 chromosome (Mariotti et al., 2009). RAE180 is a major component of the Y1 chromosome, but is also located on the Y2 and at a single autosomal locus. RAYSI is restricted only to the Y chromosomes in R. acetosa and also occurs in several related species with multiple sex chromosome systems (Shibata et al., 1999, 2000; Navajas–Perez et al., 2005; Cunado et al., 2007).
Silene latifolia: a model dioecious plant with heteromorphic sex chromosomes
Silene latifolia (2n=24 white campion) has the most studied heteromorphic sex chromosomes in plants. The first indirect evidence that X and Y chromosomes differed in S. latifolia, not only in size but also in DNA composition, was differential labeling of X and Y chromosomes by fluorescent in situ hybridization (FISH) using microdissected sex chromosome-derived probes. The hybridization conditions of the experiments (low amount of probe and short hybridization time) showed that chromosomes probably differ in their repetitive DNA structure (Hobza et al., 2004). In later experiments, several bacterial artificial chromosome (BAC) clones that originally served as anonymous cytogenetic markers (Lengerova et al., 2004) were sequenced and new repetitive DNA sequences from the Y chromosomes were characterized. It was also shown that chloroplast DNA accumulates preferentially on Y chromosome (Kejnovsky et al., 2006a).
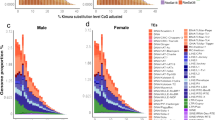
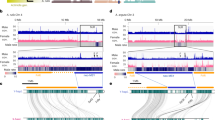
Tandem repeats in S. latifolia were described by Buzek et al. (1997), Matsunaga et al. (1999), Garrido-Ramos et al. (1999), Sykorova et al. (2001), Kazama et al. (2003, 2006) and Kazama and Matsunaga (2008). The subtelomeric tandem repeat X-43.1 (Buzek et al., 1997) similarly as Ssp15 (Sykorova et al., 2001) are located not only on the sex chromosomes but also on the autosomes. The role of tandem repeat amplification in Y chromosome evolution was shown by the characterization of the TRAYC element (tandem repeat accumulated on the Y chromosome) (Hobza et al., 2006) and STAR-Y (S ilene tandem repeat amplified on the Y chromosome) (Hobza et al., 2007). Both sequences are located in the centromeric regions of some autosomes and are highly abundant on the Y chromosome. The data suggest a centromeric origin of TRAYC and STAR-Y sequences, which then accumulated on the Y chromosome in S. latifolia. Sequence analysis also showed that microsatellites are overrepresented on the Y chromosome (Kubat et al., 2008). Comparisons of microsatellite distributions between R. acetosa and S. latifolia chromosomes show that some motifs (for example, CAA or TAA) are strongly accumulated in non-recombining regions of the Y chromosome in both species (Figure 1), whereas other motifs have different patterns: (GC)15 is abundant on the Y chromosomes in S. latifolia and absent in R. acetosa. Perhaps some simple DNA repeats regularly expand on Y chromosomes as speculated Kubat et al. (2008). This may be because of their DNA conformational properties or dependent on other circumstances (for example, the stage of evolution of the sex chromosomes, co-transposition with some retroelements).
Chromosomal distribution of labeled microsatellites—containing oligonucleotides (CAA)10 in Silene latifolia (a) and Rumex acetosa (d), microsatellites (TAA)10 in S. latifolia (b) and R. acetosa (e), and microsatellites (GC)15 in S. latifolia (c) and R. acetosa (f). Chromosomes were counterstained with DAPI (blue). Microsatellite probes were directly labeled with Cy3 during synthesis (red signals). The tandem repeat X-43.1, which is present at most subtelomeres of S. latifolia but on the Y chromosome only on its q-arm, was labeled with SpectrumGreen-conjugated nucleotides (green signals in a–c). In R. acetosa, Y1 and Y2 chromosomes were specifically labeled by RAYS1 probe (green in e, f). The X and Y chromosomes are indicated. Bar=10 μm.
Matsunaga et al. (2002) and Obara et al. (2002) studied the retrotransposons in S. latifolia. Later, Pritham et al. (2003) isolated a transcriptionally active DNA transposon residing on the Y chromosome. The first active MITE elements in S. latifolia were described by Bergero et al. (2008). A comprehensive systematic study of repetitive DNA in S. latifolia showed that Copia retroelements are the most abundant DNA element on the Y chromosome (Cermak et al., 2008). Cermak et al. (2008) also showed the enigmatic distribution of the Ogre-like retroelement, which colonizes only recombining parts of the genome of S. latifolia (is absent in non-recombining parts of the Y chromosome). This distribution of Ogre-like elements could be explained either by their spreading in connection with the recombination machinery or by their activity only in females, for example, because of some inhibition mechanisms (for example, RNAi) present only in males. The different evolutionary pressures acting on non-recombining regions influence TE distribution and affect sex chromosome evolution. We speculate that for some TEs recombination is an essential condition for transposition (Cermak et al., 2008). We can also speculate that different repetitive elements occupy the non-recombining part of the Y chromosome in different evolutionary steps of X--Y diversification.
Figure 2 represents a comprehensive summary of repetitive DNA distribution on sex chromosomes in S. latifolia. Most TEs are distributed uniformly along both the X and Y chromosomes. Two exceptions are Retand elements, which are localized at subtelomeres (Kejnovsky et al., 2006b) and Ogre-like elements, which are present on whole X chromosome but restricted to the PAR region of the Y chromosome (Cermak et al., 2008). Tandem repeats colonize the centromeres (STAR-C) and subtelomeres (X-43.1) of X chromosome, whereas in the Y chromosome STAR-C and STAR-Y are located in the middle of both arms and X-43.1 is at the subtelomere of the q-arm. Telomere-like sequences are present also in centromeres of the X and Y chromosomes (Uchida et al., 2002). It is evident that the Y chromosome has a different composition and localization of repetitive DNA compared with X chromosome and autosomes.
The presence of sex chromosomes and their tendency to accumulate repetitive DNA gives this dioecious species evolutionary potential different from what one might expect in the hermaphroditic species (Meagher and Costich, 2008). The content of repetitive DNA may have a role in phenotypic features (Meagher and Vassiliadis, 2005; Biemont, 2008). For example, a negative correlation between DNA content and flower and leaf size has been shown in four Silene species (Meagher and Costich, 2004).
Conclusions and prospects
Despite much theoretical and experimental progress, our understanding of sex chromosome evolution is more diagrammatic than dynamic. The contribution of many processes such as accumulation of repetitive DNA, degeneration of genes, additions of genes, small or large deletions, inversions, or heterochromatinization, is not clear. Neither is the sequence or timing of these events (for example, was accumulation followed by degeneration or vice versa, Steinemann and Steinemann, 2005) nor the interactions of these processes are well understood (for example, insertions of TE can cause degeneration of genes, Marais et al., 2008).
We are also far from understanding the dominant processes working on the Y chromosome such as the preferential accumulation of repetitive DNA in regions with suppressed recombination. Accumulation may be explained by reduced selective pressure against insertions. However, the targeted insertion of repeats onto the Y chromosome may also play a role. TE-rich regions on the Y chromosomes could be converted into heterochromatin and, as a result, can attract more TEs because of targeted insertion into heterochromatin regions (Zhu et al., 2003).
There are still not many data that show how, or even whether, non-recombining regions of plant sex chromosomes degenerate (Marais et al., 2008). Different effects of TE insertion (either genetic or epigenetic) on degeneration of the Y chromosome may play different roles in plants and animals. It could be that degeneration of the Y chromosome is as an evolutionary phenomenon restricted to animals, which, unlike plants, do not undergo strong selection in haploid gametophytes (Armstrong and Filatov, 2008). On the other hand, genes in plants are usually present in multiple copies or as large gene families, a consequence of duplications and polyploidization events. The loss of one allele of a specific gene, or a decrease in expression level as a result of degeneration should therefore not present a dramatic genetic imbalance for plant species that have other functional copies. Duplicity of genes could even accelerate the degeneration of Y chromosomes in plants. Another consequence of the back-up of genetic information in multiplicated plant genomes is different evolution of dosage compensation in plants. There are still no data that dosage compensation in plant sex chromosomes has evolved. Determining which evolutionary patterns after the evolution of sex chromosomes are common both in plants and animals and which ones are restricted to only one of these kingdoms will depend on an increasing amount of data from emerging plant models.
References
Armstrong SJ, Filatov DA (2008). A cytogenetic view of sex chromosome evolution in plants. Cytogenet Genome Res 120: 241–246.
Bachtrog D (2006). Expression profile of a degenerating neo-Y chromosome in Drosophila. Curr Biology 16: 1694–1699.
Bachtrog D, Hom E, Wong KM, Maside X, de Jong P (2008). Genomic degradation of a young Y chromosome in Drosophila miranda. Genome Biol 9: R30.
Bailey JA, Carrel L, Chakravarti A, Eichler EE (2000). Molecular evidence for a relationship between LINE-1 elements and X chromosome inactivation: The Lyon repeat hypothesis. Proc Natl Acad Sci USA 97: 6634–6639.
Beadle GW (1932). A possible influence of the spindle fibre on crossing-over in Drosophila. Proc Natl Acad Sci USA 18: 160–165.
Bergero R, Forrest A, Charlesworth D (2008). Active miniature transposons from a plant genome and its nonrecombining Y chromosome. Genetics 178: 1085–1092.
Biemont C (2008). Within-species variation in genome size. Heredity 101: 297–298.
Biemont C, Vieira C (2006). Junk DNA as an evolutionary force. Nature 443: 521–524.
Brookfield JFY (1991). Models of repression of transposition in P-M hybrid dysgenesis by P cytotype and by zygotically encoded repressor proteins. Genetics 128: 471–486.
Brookfield JFY (2005). The ecology of the genome—mobile DNA elements and their hosts. Nature Rev Genet 6: 128–136.
Bureau TE, Ronald PC, Wessler SR (1996). A computer-based systematic survey reveals the predominance of small inverted-repeat elements in wild-type rice genes. Proc Natl Acad Sci USA 93: 8524–8529.
Bureau TE, Wessler SR (1992). Tourist: a large family of small inverted repeat elements frequently associated with maize genes. Plant Cell 4: 1283–1294.
Buzek J, Koutnikova H, Houben A, Riha K, Janousek B, Siroky K et al. (1997). Isolation and characterization of X chromosome-derived DNA sequences from a dioecious plant Melandrium album. Chromosome Res 5: 57–65.
Cambareri EB, Aisner R, Carbon J (1998). Structure of the chromosome VII centromeric region in Neurospora crassa: degenerate transposon and simple repeats. Mol Cell Biol 18: 5465–5477.
Cermak T, Kubat Z, Hobza R, Koblizkova A, Widmer A, Macas J et al. (2008). Survey of repetitive sequences in Silene latifolia with respect to their distribution on sex chromosomes. Chromosome Res 16: 961–976.
Charlesworth B (1991). The evolution of sex chromosomes. Science 251: 1030–1033.
Charlesworth B, Charlesworth D (2000). The degeneration of Y chromosomes. Phil Trans R Soc Lond B Biol Sci 355: 1563–1572.
Charlesworth B, Sniegowski P, Stephan W (1994). The evolutionary dynamics of repetitive DNA in eukaryotes. Nature 371: 215–220.
Charlesworth D, Charlesworth B, Marais G (2005). Steps in the evolution of heteromorphic sex chromosomes. Heredity 95: 118–128.
Clarke L, Carbon J (1980). Isolation of the centromere-linked CDC10 gene by complementation in yeast. Proc Natl Acad Sci USA 77: 2173–2177.
Copenhaver GP, Nickel K, Kuromori T, Benito MI, Kaul S, Lin X et al. (1999). Genetic definition and sequence analysis of Arabidopsis centromeres. Science 286: 2253–2256.
Cunado N, Navajas-Pereze R, de la Herran R, Ruiz Rejon C, Ruiz Rejon M, Santos JL et al. (2007). The evolution of sex chromosomes in the genus Rumex (Polygonaceae): identification of a new species with heteromorphic sex chromosomes. Chromosome Res 15: 825–832.
Downs AJ, Jackson SP (1999). Involvement of DNA end-binding protein Ku in Ty element retrotransposition. Mol Cell Biol 19: 6260–6268.
Duret L, Marais G, Biemont C (2000). Transposons but not retrotransposons are located preferentially in regions of high recombination rate in Caenorhabditis elegans. Genetics 156: 1661–1669.
Emerson JJ, Kaessmann H, Betrán E, Long M (2004). Extensive gene traffic on the mammalian X chromosome. Science 303: 537–540.
Erlandsson R, Wilson JF, Paeaebo S (2000). Sex chromosome transposable element accumulation and male-driven substitutional evolution in humans. Mol Biol Evol 17: 804–812.
Ezaz T, Stiglec R, Veyrunes F, Graves JAM (2006). Relationship between vertebrate ZW and XY sex chromosome systems. Curr Biol 16: R736–R743.
Garrido-Ramos MA, de la Herran R, Ruiz Rejon M, Ruiz Rejon C (1999). A subtelomeric satellite DNA family isolated from the genome of the dioecious plant Silene latifolia. Genome 42: 442–446.
Gaut BS, Wright SI, Rizzon C, Dvorak J, Anderson LK (2007). Recombination: an underappreciated factor in the evolution of plant genomes. Nature Rev Genet 8: 77–84.
Gorelick R (2005). Theory for why dioecious plants have equal length sex chromosomes. Am J Bot 92: 979–984.
Graves JAM (2005). Recycling the Y chromosome. Science 307: 50–51.
Gvozdev VA, Kogan GL, Usakin LA (2005). The Y chromosome as a target for acquired and amplified genetic material in evolution. BioEssays 27: 1256–1262.
Harvey CF, Gill GP, Fraser LG, McNeilage MA (1997). Sex determination in Actinidia. 1. Sex-linked markers and progeny sex ratio in diploid A. chinensis. Sex Plant Reprod 10: 149–154.
Hobza R, Kejnovsky E, Vyskot B, Widmer A (2007). The role of chromosomal rearrangements in the evolution of Silene latifolia sex chromosomes. Mol Genet Genomics 278: 633–638.
Hobza R, Lengerova M, Cernohorska H, Rubes J, Vyskot B (2004). FAST-FISH with laser beam microdissected DOP-PCR probe distinguishes the sex chromosomes of Silene latifolia. Chromosome Res 12: 245–250.
Hobza R, Lengerova M, Svoboda J, Kubekova H, Kejnovsky E, Vyskot B (2006). An accumulation of tandem DNA repeats on the Y chromosome in Silene latifolia during early stages of sex chromosome evolution. Chromosoma 115: 376–382.
Hood ME, Katawczik M, Giraud T (2005). Repeat-induced point mutation and the population structure of transposable elements in Microbotryum violaceum. Genetics 170: 1081–1089.
Hua-Van A, Le Rouzic A, Maisonhaute C, Capy P (2005). Abundance, distribution and dynamics of retrotransposable elements and transposons: similarities and differences. Cytogenet Genome Res 110: 426–440.
Hurst LD (2001). Sex and the X. Nature 411: 149–150.
Ishizaki K, Shimizu-Ueda Y, Okada S, Yamamoto M, Fujisawa M, Yamato KT et al. (2002). Multicopy genes uniquely amplified in the Y chromosome-specific repeats of the liverwort Marchantia polymorpha. Nucleic Acids Res 30: 4675–4681.
Jablonka E (2004). The evolution of the peculiarities of mammalian sex chromosomes: an epigenetic view. BioEssays 26: 1327–1332.
Jurka J (1997). Sequence patterns indicate an enzymatic involvement in integration of mammalian retroposons. Proc Natl Acad Sci USA 94: 1872–1877.
Jurka J, Klonowski P, Trifonov EN (1998). Mammalian retroposons integrate at kinkable DNA sites. J Biomol Struct Dyn 15: 717–721.
Kapitonov VV, Jurka J (1999). Molecular paleontology of transposable elements from Arabidopsis thaliana. Genetica 107: 27–37.
Kapitonov VV, Jurka J (2005). RAG1 core and V(D)J recombination signal sequences were derived from Transib transposons. PLoS Biol 3: 998–1010.
Kazama Y, Matsunaga S (2008). The use of repetitive sequences in cytogenetic studies of plant sex chromosomes. Cytogenet Genet Res 120: 247–254.
Kazama Y, Sugiyama R, Matsunaga S, Shibata F, Uchida W, Hizume M et al. (2003). Organization of the KpnI family of chromosomal distal-end satellite DNAs in Silene latifolia. J Plant Res 116: 317–326.
Kazama Y, Sugiyama R, Suto Y, Uchida W, Kawano S (2006). The clustering of four subfamilies of satellite DNA at individual chromosome ends in Silene latifolia. Genome 49: 520–530.
Kejnovsky E, Hobza R, Kubat Z, Widmer A, Marais GAB, Vyskot B (2007). High intrachromosomal similarity of retrotransposon long terminal repeats: Evidence for homogenization by gene conversion on plant sex chromosomes? Gene 390: 92–97.
Kejnovsky E, Kubat Z, Hobza R, Lengerova M, Sato S, Tabata S et al. (2006a). Accumulation of chloroplast DNA sequences on the Y chromosome of Silene latifolia. Genetica 128: 167–175.
Kejnovsky E, Kubat Z, Macas J, Hobza R, Mracek J, Vyskot B (2006b). Retand: a novel family of gypsy-like retrotransposon harboring an amplified tandem repeat. Mol Genet Genomics 276: 254–263.
Khil PP, Oliver B, Camerini-Otero RD (2005). X for intersection: retrotransposition both on and off the X chromosome is more frequent. Trends Genet 21: 3–7.
Kidwell MG, Lisch D (2001). Perspective: transposable elements, parasitic DNA, and genome evolution. Evolution Int J Org Evolution 55: 1–24.
Kihara H, Ono T (1923). Cytological studies on Rumex L., I. Chromosomes of Rumex acetosa L. Bot Mag Tokyo 37: 84–90.
Kipling D, Warburton PE (1997). Centromeres, CENP-B and Tigger too. Trend Genet 13: 141–145.
Kirchner J, Connolly CM, Sandmeyer SB (1995). Requirement of RNA polymerase III transcription factors for in vitro position-specific integration of a retroviruslike element. Science 267: 1488–1491.
Krzywinski J, Sangare D, Besansky NJ (2005). Satellite DNA from the Y chromosome of the malaria vector Anopheles gambiae. Genetics 169: 185–196.
Kubat Z, Hobza R, Vyskot B, Kejnovsky E (2008). Microsatellite accumulation on the Y chromosome in Silene latifolia. Genome 51: 350–356.
Lahn BT, Page DC (1999). Retroposition of autosomal mRNA yielded testis-specific gene family on human Y chromosome. Nature Genet 21: 429–433.
Langley CH, Montgomery E, Hudson R, Kaplan N, Charlesworth B (1988). On the role of unequal exchange in the containment of transposable element copy number. Genet Res 52: 223–235.
Lengerova M, Kejnovsky E, Hobza R, Macas J, Grant SR, Vyskot B (2004). Multicolor FISH mapping of the dioecious model plant, Silene latifolia. Theor Appl Genet 108: 1193–1199.
Lengerova M, Vyskot B (2001). Sex chromatin and nucleolar analyses in Rumex acetosa L. Protoplasma 217: 147–153.
Leonardo TE, Nuzhdin SV (2002). Intracelullar battlegrounds: conflict and cooperation between transposabel elements. Genet Res Camb 80: 155–161.
Liao D (2003). Concerted evolution. In: Encyclopedia of the Human Genome. Macmillian Publishers Ltd, Nature Publishing Group: London. pp 1–6.
Liu Z, Moore PH, Ma H, Ackerman CM, Ragiba M, Yu Q et al. (2004). A primitive Y chromosome in papaya marks incipient sex chromosome evolution. Nature 427: 348–352.
Loptien H (1979). Identification of the sex chromosome pair in asparagus (Asparagus officinalis L.). Zeitschrift fur Pflanzenzuchtung 82: 162–173.
Lyon MF (2000). LINE-1 elements and X chromosome inactivation: a function for ‘junk’ DNA? Proc Natl Acad Sci USA 97: 6248–6249.
Marais GAB, Nicolas M, Bergero R, Chambrier P, Kejnovsky E, Moneger F et al. (2008). Evidence for degeneration of the Y chromosome in the dioecious plant Silene latifolia. Current Biology 18: 1–5.
Mariotti B, Manzano S, Kejnovsky E, Vyskot B, Jamilena M (2009). Accumulation of Y-specific satellite DNAs during the evolution of Rumex acetosa sex chromosomes. Molec Genet Genomics (in press).
Matsunaga S, Isono E, Kejnovsky E, Vyskot B, Dolezel J, Kawano S et al. (2003). Duplicative transfer of a MADS box gene to a plant Y chromosome. Mol Biol Evol 20: 1062–1069.
Matsunaga S, Kawano S, Michimoto T, Higashiyama T, Nakao S, Sakai A et al. (1999). Semi-automatic laser beam microdissection of the Y chromosome and analysis of Y chromosome DNA in a dioecious plant, Silene latifolia. Plant Cell Physiol 40: 60–68.
Matsunaga S, Yagisawa F, Yamamoto M, Uchida W, Nakao S, Kawano S (2002). LTR retrotransposons in the dioecious plant Silene latifolia. Genome 45: 745–751.
Meagher TR, Costich DE (2004). Junk' DNA and long-term phenotypic evolution in Silene section Elisanthe (Caryophyllaceae). Proc R Soc Lond B 271: S493–S497.
Meagher TR, Costich DE (2008). Junk' DNA and phenotypic evolution in Silene section Siphnomorpha. Genet Res Camb 90: 111–118.
Meagher TR, Vassiliadis C (2005). Phenotypic impacts of repetitive DNA in flowering plants. New Phytol 168: 71–80.
Myers JS, Vincent BJ, Udall H, Watkins WS, Morrish TA, Kilroy GE et al. (2002). A comprehensive analysis of recently integrated human Ta L1 elements. Am J Hum Genet 71: 312–326.
Navajas-Perez R, Schwarzacher T, de la Heran R, Ruiz Rejon C, Ruiz Rejon M, Garrido-Ramos MA (2005). The origin and evolution of the variability in a Y-linked specific satellite-DNA of Rumex acetosa and its relatives. Gene 368: 61–71.
Nuzhdin SV (1999). Sure facts, speculations, and open questions about the evolution of transposable element copy number. Genetica 107: 129–137.
Obara M, Matsunaga S, Nakao S, Kawano S (2002). A plant Y chromosome –STS marker encoding a degenerate retrotransposon. Genes Genet Syst 77: 393–398.
Ohno S (1967). Sex Chromosomes and Sex Linked Genes. Springer-Verlag: New York.
Okada S, Sone T, Fujisawa M, Nakayama S, Takenaka M, Ishizaki K et al. (2001). The Y chromosome in the liverwort Marchantia polymorpha has accumulated unique repeat sequences harboring a male-specific gene. Proc Natl Acad Sci USA 98: 9454–9459.
Pazourkova Z (1964). Sex chromatin in Rumex acetosa L. Preslia 36: 422–424.
Petrov DA, Aminetzach YT, Davis JC, Bensasson D, Hirsh AE (2003). Size matters? Non-LTR retrotransposable elements and ectopic recombination in Drosophila. Mol Biol Evol 20: 880–892.
Pimpinelli S, Berloco M, Fanti L, Dimitri P, Bonaccorsi S et al. (1995). Transposable elements are stable structural components of Drosophila melanogatser heterochromatin. Proc Natl Acad Sci USA 92: 3804–3808.
Pritham EJ, Zhang YH, Feschotte C, Kesseli RV (2003). An Ac-like transposable element family with transriptionally active Y-linked copies in the white campion, Silene latifolia. Genetics 165: 799–807.
Rizzon C, Marais G, Gouy M, Biemont C (2002). Recombination rate and the distribution of transposable elements in the Drosophila melanogaster genome. Genome Res 12: 400–407.
Ruiz Rejon C, Jamilena M, Garrido Ramos M, Parker JS, Ruiz Rejon M (1994). Cytogenetic and molecular analysis of the multiple sex chromosome system of Rumex acetosa. Heredity 72: 209–215.
Sakamoto K, Abe T, Matsuyama T, Yoshida S, Ohmido N, Fukui K et al. (2005). RAPD markers encoding retrotransposable elements are linked to the male sex in Cannabis sativa L. Genome 48: 931–936.
Sakamoto K, Ohmido N, Fukui K, Kamada H, Satoh S (2000). Site-specific accumulation of a LINE-like retrotransposon in a sex chromosome of the dioecious plant Cannabis sativa. Plant Mol Biol 44: 723–732.
Shibata F, Hizume M, Kuroki Y (1999). Chromosome painting of Y chromosomes and isolation of a Y chromosome-specific repetitive sequence in the dioecious plant Rumex acetosa. Chromosoma 108: 266–270.
Shibata F, Hizume M, Kuroki Y (2000). Differentiation and the polymorphic nature of the Y chromosome revealed by repetitive sequences in the dioecious plant, Rumex acetosa. Chromosome Res 8: 229–236.
Skaletsky H, Kuroda-Kawaguchi T, Minx PJ, Cordum HS, Hillier L, Brown LG et al. (2003). The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature 423: 825–837.
Slotkin RK, Martienssen R (2007). Transposable elements and the epigenetic regulation of the genome. Nature Rev Genet 8: 272–285.
Smith AFA, Riggs AD (1996). Tiggers and other DNA transposon fossils in the human genome. Proc Natl Acad Sci USA 93: 1443–1448.
Steinemann M, Steinemann S (1992). Degenerating Y chromosome of Drosophila miranda: a trap for retrotransposons. Proc Natl Acad Sci USA 89: 7591–7595.
Steinemann S, Steinemann M (2005). Y chromosomes: born to be destroyed. BioEssays 27: 1076–1083.
Sun X, Le HD, Wahlstorm JM, Karpen GH (2003). Sequence analysis of a functional Drosophila centromere. Genome Res 13: 182–194.
Sykorova E, Fajkus J, Mikako I, Fukui K (2001). Transition between two forms of heterochromatin at plant subtelomeres. Chromosome Res 9: 309–323.
Traut W, Marec F (1997). Sex chromosome differentiation in some species of Lepidoptera. Chromosome Res 5: 283–291.
Traut W, Willhoeft U (1990). A jumping sex determining factor in the fly Megaselia scalaris. Chromosoma 99: 407–412.
Traut W, Wollert B (1998). An X/Y DNA segment from an early stage of sex chromosome differentiation in the fly Megaselia scalaris. Genome 41: 289–294.
Tsuda Y, Nishida-Umehara C, Ishijima J, Yamada K, matsuda Y (2007). Comparison of the Z and W sex chromosomal architectures in elegant crested tinamou (Eudromia elegans) and ostrich (Struthio camelus) and the process of sex chromosome differentiation in paleognathous birds. Chromosoma 116: 159–173.
Uchida W, Matsunaga S, Sugiyama R, Shibata F, Kazama Y, Miyazawa Y et al. (2002). Distribution of interstitial telomere-like repeats and their adjacent sequenced in a dioecious plant, Silene latifolia. Chromosoma 111: 313–320.
Veltsos P, Keller I, Nichols RA (2008). The inexorable spread of a newly arisen neo-Y chromosome. PLoS Genetics 4: 1–9.
Veyrunes F, Waters PD, Miethke P, Rens W, McMilla D, Alsop AE et al. (2008). Bird-like sex chromosomes of platypus imply recent origin of mammal sex chromosomes. Genome Res 18: 1–9.
Vicient CM, Suoniemi A, Anamthawat-Jonsson K, Tanskane J, Beharav A, Nevo E et al. (1999). Retrotransposon BARE-1 and its role in genome evolution in the genus Hordeum. Plant Cell 11: 1769–1784.
Vitte C, Panaud O (2003). Formation of solo-LTRs through unequal homologous recombination counterbalances amplification of LTR retrotransposons in rice Oryza sativa L. Mol Biol Evol 20: 528–540.
Vyskot B, Hobza R (2004). Gender in plants: sex chromosomes are emerging from the fog. Trends Genet 20: 432–438.
Westerman M, Hewitt GM (1985). Chromosome banding in Podisma pedestris. Heredity 55: 157–161.
Willhoeft U, Traut W (1990). Molecular differentiation of the homorphic sex chromosomes in Megaselia scalaris (Diptera) detected by random DNA probes. Chromosoma 99: 237–242.
Winge O (1929). On the nature of sex chromosomes in Humulus. Hereditas 12: 53–63.
Yampolsky C, Yampolsky H (1922). Distribution of the sex forms in the phanerogamic flora. Bibliography of Genetics 3: 1–62.
Yandeau-Nelson, Zhou Q, Yao H, Xu X, Nikolau BJ, Schnable PS (2005). MuDR transposase increases the frequency of meiotic crossovers in the vicinity of a Mu insertion in the maize a1 gene. Genetics 169: 917–929.
Yu Q, Hou S, Feltus FA, Jones MR, Murray JE, Veatch O et al. (2008). Low X/Y divergence in four pairs of papaya sex-linked genes. Plant J 53: 124–132.
Yu Q, Shaobin H, Hobza R, Feltus FA, Wang X, Jin W et al. (2007). Chromosomal location and gene paucity of the male specific region on papaya Y chromosome. MGG 278: 177–185.
Xie WW, Gai X, Zhu Y, Zappulla DC, Sternglanz R, Voytas D (2001). Targeting of the yeast Ty5 retrotransposon to silent chromatin is mediated by interactions between integrase and Sir4p. Mol Cell Biol 21: 6606–6614.
Zhang W, Wang X, Yu Q, Ming R, Jiang J (2008). DNA methylation and heterochromatinization in the male-specific region of the primitive Y chromosome of papaya. Genome Res 18: 1938–1943.
Zhong CX, Marshall JB, Topp C, Mroczek R, Kato A, Nagaki K et al. (2002). Centromeric retroelements and satellites interact with maize kinetochore protein CENH3. Plant Cell 14: 2825–2836.
Zhu Y, Dai J, Fuerst PG, Voytas DF (2003). Controlling integration specificity of a yeast retrotransposon. Proc Natl Acad Sci USA 100: 5891–5895.
Zluvova J, Janousek B, Negrutiu I, Vyskot B (2005). Comparison of the X and Y chromosome organization in Silene latifolia. Genetics 170: 1431–1434.
Acknowledgements
The authors thank Professor Andrew Leitch and Dr Jiri Macas for their comments on the manuscript and Bc. Julia Svoboda for revision of the manuscript. This research was supported by the LC06004 grant from Ministry of Education and grants no. AV0Z50040507 and AV0Z50040702 from the Academy of Sciences of the Czech Republic.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kejnovsky, E., Hobza, R., Cermak, T. et al. The role of repetitive DNA in structure and evolution of sex chromosomes in plants. Heredity 102, 533–541 (2009). https://doi.org/10.1038/hdy.2009.17
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2009.17
Keywords
This article is cited by
-
Accumulation of retrotransposons contributes to W chromosome differentiation in the willow beauty Peribatodes rhomboidaria (Lepidoptera: Geometridae)
Scientific Reports (2023)
-
In silico analysis of steviol glycoside biosynthesis related genes of Helianthus annuus shows its potential for sweetener production
Genetic Resources and Crop Evolution (2023)
-
Characterization of repetitive DNA on the genome of the marsh rat Holochilus nanus (Cricetidae: Sigmodontinae)
Molecular Genetics and Genomics (2023)
-
Genetic insights: mapping sex-specific loci in Siamese cobra (Naja kaouthia) sheds light on the putative sex determining region
Genes & Genomics (2023)
-
GISH painting of the Y chromosomes suggests advanced phases of sex chromosome evolution in three dioecious Cannabaceae species (Humulus lupulus, H. japonicus, and Cannabis sativa)
Protoplasma (2023)