Abstract
Recent speciation events provide important insights into the understanding and conservation of Earth's biodiversity, representing recent adaptations to a changing environment and an important source of future evolutionary potential. However, the most frequently applied criterion for molecular-based speciation investigations, that of reciprocal monophyly of mitochondrial sequences, overlooks recent speciation events where insufficient time has passed for fixed molecular differences to develop between putative species. Two morphologically distinguishable forms of finless porpoise (genus Neophocaena) exist in sympatry in the strait of Taiwan, however the taxonomic relationship of these different forms is controversial. To test the hypothesis that the two forms represent different species, a study was conducted based on morphological characters and microsatellite and mitochondrial markers. The data suggest that the two forms are highly differentiated in terms of both morphology and genetic characteristics, despite being sympatric, and therefore represent different species as defined by the biological species concept. Moreover, the two forms appear to have been reproductively isolated since sharing a common ancestor prior to the last major glaciation event ∼18 000 years ago. However, this represents an insufficient amount of time for reciprocal monophyly to have developed, and thus previous studies based on this criterion have overlooked this speciation event and resulted in incorrect taxonomic classification of these forms.
Similar content being viewed by others
Introduction
Testing hypotheses of speciation or species-level taxonomy is one of the most fundamental tasks in population biology and is also one of the most controversial (Hey et al., 2003), with differing opinions on what species concepts are appropriate (Coyne and Orr, 2004), or even on whether these concepts are relevant at all (Hendry et al., 2000). Despite this controversy, it is necessary to work within an explicit scientific framework to address species-level issues, because incorrect taxonomic classification often has important implications for basic biological research and serious consequences for wildlife conservation (Daugherty et al., 1990).
Although most paradigms for testing species-level taxonomy are based on a similar idea of reproductive isolation (or nearly so) between the putative taxa, much of the controversy revolves around what data and criteria are indicative of reproductive isolation (Crandall et al., 2000; Hey et al., 2003; Coyne and Orr, 2004). The most frequently applied criterion for genetics-based speciation investigations is that of reciprocal monophyly, particularly of mitochondrial sequences (Moritz, 1994). However, this criterion assumes that divergence between the putative species has reached an equilibrium, which theoretically takes an average of 4Ne generations (assuming neutrality, Avise et al., 1984) and could require many more years than have passed since the hypothesized speciation event, particularly for species with large population sizes and/or long generation times. As a result, recent speciation events will not be detected using the typical criterion of reciprocal monophyly due to this lack of equilibrium and resulting lack of fixed molecular differences between recently derived species. If the goal of conserving biodiversity is to characterize and conserve evolutionary potential (Allendorf and Luikart, 2007), then studies based on reciprocal monophyly could be falling short of this goal not only by resulting in incorrect characterization of biodiversity, but perhaps more importantly by overlooking or discounting those taxa that have most recently evolved, and therefore represent the most recent adaptations to a changing environment and thus may be important sources of future evolutionary potential (Erwin, 1991).
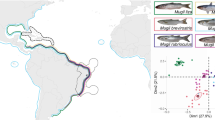
The taxonomy of finless porpoises (genus Neophocaena) is controversial, and is noted as having a ‘bewildering nomenclatural history’ (Rice, 1999). The currently accepted classification is of a single species with three subspecies that differ, for the most part, in distribution and morphology (Amano, 2002). Neophocaena phocaenoides phocaenoides is characterized by a wide area of tubercules on the dorsal surface and ranges from the Indian Ocean through the South China Sea; N. p. asiaeorientalis is characterized by a narrow tuberculed area on the dorsal ridge and represents those finless porpoises found in the Yangtze River; and N. p. sunameri is also characterized by a similar dorsal surface but is found in the East China Sea, Yellow Sea, Bohai and the waters of Korea and Japan (Kasuya, 1999; Rice, 1999; Amano, 2002; Figure 1). However, this prevailing classification was recently challenged by a study on cranial morphology that suggested the ‘wide’ (phocaenoides) and ‘narrow’ forms were differentiated enough to warrant separate species status, whereas the data did not support the further subdivision of the narrow form into the asiaeorientalis and sunameri subspecies (Jefferson, 2002; Jefferson and Hung, 2004).
Distribution and morphology of the two forms of finless porpoise. (a) A map of the range of the two forms and the area of sympatry. Note that the northern and southern limits of the area of sympatry are only rough estimates and need further assessment. (b) Images of the two forms, with the narrow ridge at the top and the wide form at the bottom. Photographs by JY Wang.
The biological species concept (BSC; Mayr, 1942) is arguably the most widely applied framework for testing species hypotheses (Avise, 2004). However, the primary difficulty in applying this concept is that it requires identifying whether reproductive isolation mechanisms, other than just spatial or geographic barriers to movement, have evolved between the putative species (Mayr, 1942). Testing this hypothesis often requires translocating individuals from one location to another, or using artificial settings to test reproductive compatibility (Rundle et al., 2000); however, these options are not feasible for many situations. The ranges of both forms of finless porpoise overlap in the Taiwan Strait and surrounding waters, which represents a large area of sympatry (Figure 1). This sympatric area provides a rare and ideal natural experiment for testing the status of the two forms of finless porpoises because it eliminates the possibility of spatial or geographic barriers to reproduction, leaving only the potential for biological isolation mechanisms to limit gene flow between the two forms.
Data from individuals in this area of sympatry were used to test the monotypic hypothesis of the finless porpoise under the theoretical framework of the biological species concept. The study was conducted in a ‘blind’ fashion, involving one research group conducting morphological analyses and another group conducting genetic analyses of the same individuals. Data were combined a posteriori and assessed for congruence between morphological and molecular characters to determine if the two forms are reproductively isolated despite being sympatric, and therefore represent different species as defined by the BSC.
Materials and methods
Sample collection
Analyzed specimens represent finless porpoises that were killed incidentally in fishing nets or were found washed ashore between 1993 and 2005. A total of 56 specimens were analyzed; 38 from the known area of sympatry within and around the Taiwan Strait, and 18 from Hong Kong (northern South China Sea), which is south of the area of sympatry and where, despite the analysis of over 100 carcasses, only the wide form has been found (T Jefferson and S Hung, personal communication). Samples from the area of sympatry include 15 from southern China, ranging from Xiamen to Dongshan Island of Fujian province; 17 from the Matsu Islands and 6 from western Taiwan and the eastern Taiwan Strait. Genetic analyses were conducted on 49 of the specimens (33 from the area of sympatry and 16 from Hong Kong), with the DNA from the remaining 7 being too degraded for reliable genetic analysis. Morphological measurements were conducted on 33 specimens, 31 from the area of sympatry and 2 from Hong Kong. The remaining specimens from Hong Kong could not be not rigorously assessed because the appropriate measurements were not recorded during the time of necropsy. However, this area is outside the range of sympatry and is only within the range of the wide form. Additionally, it was possible to confirm that all Hong Kong specimens were indeed representative of the wide form through qualitative assessment of photodocumentation of the sampled individuals. The analyses of the two data sets (based on morphological and genetic characters) were conducted independently by two research teams in a ‘blind’ fashion, where each team was blind to the results of the other until the data were combined a posteriori.
Morphological analyses
As suggested by its common name, the finless porpoise lacks a dorsal fin, but instead has a ridge that runs longitudinally along the middle of the back. Two conspicuous characteristics of the dorsal aspect differ between the ‘narrow’ and ‘wide’ forms (Gao and Zhou, 1995; Jefferson and Hung, 2004). The first is the width of the tuberculed area of the dorsal surface; the second is the number of longitudinal rows of tubercles that are distributed throughout the ridge. The function of these tubercles is unknown, but they possess an abundance of nerve endings, suggesting that they serve some sensory function (Kasuya, 1999). Previous studies have shown that these characteristics allow for reliable distinction between the two forms (Pilleri and Gihr, 1975; Wang, 1992). Morphological data were obtained during post-mortem examination and were based on two measurements of this dorsal ridge: (1) the greatest width of the tuberculed areas (to the nearest mm) and (2) the number of longitudinal rows of tubercules (counted at the point of the greatest width).
Molecular methods
Tissue (muscle, skin or heart) samples for genetic analyses were collected from each specimen and stored in a 20% dimethyl sulfoxide solution saturated with NaCl (Seutin et al., 1991) prior to analyses. Approximately 40 mg from each sample was used for subsequent extraction procedures. For skin samples, the tissue was frozen with liquid nitrogen, ground to a fine powder and transferred to a tube with 500 μl of lysis buffer (4 M urea, 0.2 M NaCl, 0.5% n-lauroyl sarcosine, 10 mM 1,2-cyclohexanediaminetetraacetic acid, 100 mM Tris-HCl, pH 8.0). For muscle samples, ∼40 mg of finely minced tissue was placed directly in 500 μl of lysis buffer. Samples were rotated in the lysis buffer at room temperature for⩾5 days, after which time they were subjected to three aliquots of proteinase K, each at a concentration of 2 U of proteinase K per milligram of tissue. The addition of proteinase K was as follows: after adding the first aliquot, samples were rotated at room temperature overnight; after adding the second aliquot the samples were placed in a 65 °C waterbath for 1 h, then transferred to a 37 °C incubator for 1 h; after adding the third aliquot, the samples were rotated at room temperature overnight. Approximately 250 μl of the tissue/lysis buffer solution was subsequently extracted using Qiagen DNeasy Tissue Extraction Kits (Qiagen Inc., Mississauga, Ontario, Canada). DNA quantity was estimated using PicoGreen (Singer et al., 1997), and DNA quality was examined by electrophoresis of 20 ng of DNA through 1.5% agarose gels stained with SYBR Green I (Cambrex, Rockland, ME, USA).
Samples were genotyped at nine microsatellite loci using the multiplex PCR protocol described in Supplementary Table 1. A 345 bp portion of the mitochondrial DNA control region was amplified using the primers t-PRO and Primer-2 from Yoshida et al. (2001), using the same PCR conditions as for the microsatellite loci, and an annealing temperature of 55 °C. After amplification, primers and unincorporated dNTPs were degraded using EXOSAP-IT (Dugan et al., 2002), and products were sequenced using the DYEnamic dye terminator kit (GE Healthcare, Piscataway, NJ, USA). Products were size-separated and visualized on a MegaBACE 1000 (GE Healthcare).
Statistical analysis (microsatellite data)
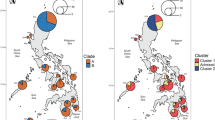
Estimation of the number of populations represented by the samples, and assignment of individuals to each population were performed using the Bayesian-based approach implemented in the program structure v.2 (Pritchard et al., 2000) that allows the number of populations and the membership of individuals within those populations to be estimated based solely on genetic data, without an a priori assignment of individuals into groups. The program was run with 500 000 MCMC steps as the burn-in time and 2 000 000 steps with recorded results, allowing for admixture and a correlation of allele frequencies between populations. The analyses were run allowing the sampled individuals to represent from one to five populations (K=1–5), and four iterations of the analyses were performed for each K. We tested for up to five populations to allow for unexpected subdivision over that of the two groups hypothesized to be present within the data set. Similarity in results across iterations was used to assess if the program had been run for enough steps. The average probability of the four runs for each K was taken as the probability for that K.
The program SPAGeDi (Hardy and Vekemans, 2002) was then used to obtain estimates of classical measures of differentiation (FST and RST) between the clusters identified with structure. Of perhaps more value than either of these estimates themselves is the relationship between the two. FST and RST provide information on different aspects of gene flow and also have different characteristics. Specifically, FST tends to have lower variance than RST and is based on frequency differences only; whereas RST considers the evolutionary relationships between alleles, but tends to have a higher variance than FST (Balloux and Lugon-Moulin, 2002). Thus, the lower variance in FST makes it more attractive for estimating differentiation on shorter time scales and/or when the migration rate is substantially larger than the mutation rate. However, RST is more appropriate when differentiation is over longer time scales and/or when the mutation rate is higher than the migration rate, leading to different allelic distributions between populations (Hardy et al., 2003). Thus, by comparing estimates of FST and RST it is possible to assess the relative roles of migration and mutation in driving observed patterns of differentiation. Tests for the significance of mutations on estimates of differentiation were performed using the randomization test implemented in SPAGeDi.
Statistical analysis (mitochondrial data)
Mitochondrial DNA control region sequences were aligned with ClustalX (Thompson et al., 1994) using a range of gap opening and extension penalties and compared by eye to establish the optimal alignment. The sequences were very similar, and all alignments were the same under the tested conditions. Estimates of variability for the control region were obtained using Arlequin v.3.1 (Excoffier et al., 2005). A minimum-spanning tree was generated based on the mutational differences between sequences. The most appropriate model of molecular evolution, and associated estimates of the transition/transversion ratio and α value for the γ distribution, were obtained from ModelGenerator (Keane et al., 2006). Population differentiation of the mtDNA sequences from individuals assigned to each group (based on the microsatellite data) was estimated using the analysis of molecular variance approach described in Excoffier et al. (1992) as implemented in the program Arlequin. The significance of the resulting estimates of FST and φST was tested using 1000 permutations.
Assessing contemporary and/or historical gene flow
The program BaysAss (Wilson and Rannala, 2003) was used to test for evidence of contemporary gene flow between the groups identified with structure. BaysAss uses multilocus genotype data and a Bayesian framework to estimate recent migration rates (within two generations) between populations. The program was run for 3 000 000 MCMC steps, with 1 000 000 steps as the burn-in and a sampling frequency of 2000. The δ values for allele frequencies, migration rates, and inbreeding values were set to 0.10. To provide a context for interpretation of these results, the migration rates obtained for the finless porpoise data were compared to those obtained by running simulated data, based on our null hypothesis of no gene flow (see below), through BaysAss with the same parameters.
To assess historical differentiation and gene flow between the two forms five microsatellite loci with the highest polymorphic information content, other than IGF1, were used together with the isolation with migration program (IM, Nielsen and Wakely, 2001; Hey and Nielsen, 2004) (note: the IGF1 locus is known to be linked to an insulin growth factor gene (Kirkpatrick, 1992), and therefore cannot be used for coalescent-based studies as it is influenced by selection, rather than just drift as is assumed in the coalescent). The program estimated six parameters, all of which were scaled by the neutral mutation rate μ. The parameters were (1) effective size of population 1, q1=4N1μ; (2) effective size of population 2, q2=4N2μ; (3) historical effective population size, qA=4NAμ; (4) time since common ancestor, t=tμ; (5) migration rate from population 1 to population 2 (going into the past), m1=m1/μ and (6) migration rate from population 2 to population 1, m2=m2/μ. For the analyses, a step-wise mutation model was used. The program was run under three different sets of conditions, and for multiple iterations under each set, to ensure that the MCMC steps had reached a stable distribution and that a similar result was obtained with each run. The first set of conditions were run for three iterations and consisted of a burn-in time of 500 000 MCMC steps; a run time of 5 000 000 MCMC steps; 20 chains with a geometric heating mode, a β value of 1.0 for the highest numbered chain and a scaling value of 0.9 for the degree of nonlinearity in the decline of β from chain 0 to chain 19; 100 chain-swapping attempts per step; a maximum m1 and m2 of 10; maximum q1, q2 and qA values of 10, 10 and 15, respectively; and maximum t of 15. For the second set of conditions, the number of chains was increased to 50, and the number of chain-swapping attempts per step was increased to 400. Two iterations were run under these conditions. One iteration was then run with the following changes: a burn-in time of 300 000 MCMC steps; a run time of 3 000 000 MCMC steps; 50 chains with a geometric heating mode, a β value of 1.0 for the highest numbered chain and a scaling value of 0.8; and 1000 chain-swapping attempts per step. These conditions were selected because previous trials showed that these conditions allowed for sufficient mixing between chains, and high updating rates for each parameter. Specifically, they resulted in effective sample size estimates greater than 50 for all parameters, and all mixing rates between successive chains to be greater than 70%. Finally, to assess if these conditions represented the appropriate run time and conditions to reach stationarity and convergence of these parameters, the program was run once for a much longer time using the following revised conditions: a burn-in time of 1 000 000 steps; a run time of 10 000 000 steps; 20 chains with a geometric heating mode, a β value of 0.8 for the highest numbered chain and a scaling value of 0.8 for the degree of nonlinearity in the decline of β from chain 0 to chain 19; and 100 chain-swapping attempts per step.
When using multiple loci, the overall neutral mutation rate represented in the parameters is the geometric mean of all of the mutation rates for the individual loci (Hey et al., 2004). Although a range of mutation rates have been reported for microsatellite loci, an increasing amount of data suggests an overall average mutation rate of 5 × 10−4 mutations per generation (Estoup and Angers, 1998). Therefore, we assumed an average neutral mutation rate for the microsatellite loci of 5 × 10−4.
Simulations
Although the programs BayesAss and IM provide estimates of contemporary and historical migration rates, respectively (among other things), other methods are necessary for testing specific hypotheses regarding these estimates. Specifically, the hypothesis of two species is reliant on testing for gene flow occurring since the time of divergence. Thus, simulations were used to obtain the expected distribution of migration rate estimates (from BayesAss), or differentiation (quantified as FST and RST) under the null hypothesis (that of no migration since isolation). Simulations were conducted using the program EASYPOP v.2 (Balloux, 2001). Random mating populations with equal numbers of diploid males and females genotyped at nine unlinked microsatellite markers were simulated using a mutation rate of 5 × 10−4 per locus per generation, which is reasonable for microsatellite loci (Estoup and Angers, 1998). A mixed model of mutation was used with 92% of mutations representing single-step mutations and 8% of mutations resulting in any one of 50 possible allelic states at random (KAM). This mutation scheme is generally consistent with the data currently available on mutations in microsatellite regions (Di Rienzo et al., 1994; Renwick et al., 2001). The initial (ancestral) effective population size estimate obtained from IM was started out with ‘minimum’ diversity and allowed to reach equilibrium over 20 000 generations. The ancestral population was then split into three populations, two representing the extant effective population size estimates of the two forms obtained from IM, and the third representing a ‘dummy’ population that just contained the remaining individuals necessary to obtain the ancestral population size. The simulations were conducted with no migration between the populations after the hypothesized split. A total of 100 iterations were conducted, and these resulting populations were sampled and analyzed as described in the text.
Results and discussion
Specimens
Out of the 56 specimens analyzed, data necessary to conduct the analyses were obtained from 52, with the remaining four representing specimens for which only mitochondrial sequences were obtained, and thus there was no genotype or measurement data to use for group assignment. It is noteworthy that these four specimens were from Hong Kong, which is outside the area of sympatry and therefore the sample size from the critical specimens within the area of sympatry was not reduced.
Morphological characters
Measurements of the greatest width of the tuberculed area and the number of longitudinal rows of tubercles differentiated individuals into one of two clusters, which had distinct and clearly non-overlapping distributions (Figure 2a). All specimens could be unambiguously assigned to one of the two clusters, which represented the two different forms. For the ‘wide’ form, the greatest width of the tuberculed area varied from 4.0 to 10.0 cm and there were between 10 and 18 rows of tubercules, whereas for the ‘narrow’ form the greatest width of the tuberculed area was between 0.3 and 0.7 cm and there were only between 3 and 5 rows of tubercules. In addition to the clear and unequivocal visual differentiation between the characteristics of the two forms, this differentiation is also clear based on statistical analysis of the two clusters. The measurements for the greatest width of the dorsal ridge did not deviate from a normal distribution (Kolmogorov–Smirnov goodness of fit tests, both P-values>0.4), and a t-test detected a significant difference between the two forms (P<0.0001). The number of rows of tubercules did deviate from a normal distribution (P=0.0427 for the narrow-ridge specimens), and a significant difference of these values between the narrow- and wide-ridge forms was also detected (Wilcoxon rank test, P<0.0001).
Graphs showing the differentiation between the two forms. (a) Plotting individuals based on the two measured morphological characters (width of the tuberculed area and number of longitudinal rows of tubercles across the widest point of the tuberculed area), and (b) plotting individuals based on their probability of having ancestry in one of the two identified populations based on the estimates obtained from the program structure.
No specimens exhibiting intermediate morphology were found in this or previous studies. Thus, similar to the previous study based on skeletal characteristics (Jefferson, 2002), the external morphological data on its own suggest strong differentiation and reproductive isolation between the two forms.
Genetic characters
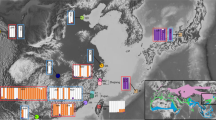
The analyses based on structure clearly indicated that the specimens came from two distinct clusters, and each individual was clearly and unambiguously assigned to one of the two groups (Figure 2b; Table 1). The differentiation between these groups was high, showing a significant difference in the distribution of alleles (Exact test, Raymond and Rousset, 1995, P<0.001). The FST estimate for the two populations was 0.26. The RST estimate (0.48) was almost twice as high, and was significantly higher than expected if mutation was not influencing estimates of differentiation (randomization test, P=0.006). The implication is that the migration rate must be notably smaller than the mutation rate and that sufficient time has passed for different alleles to accumulate in the different groups, indicating that the differentiation detected is of an evolutionary, rather than just a recent, time scale. Supporting this idea, it was found that the majority (40 of 69, 58%) of the detected alleles were confined to one of the two groups, and that the two forms have markedly different allele distributions at several loci (Figure 3).
Six variable sites in the sequenced portion of the mitochondrial control region were identified that resulted in seven haplotypes. Assessment of the sequences in relation to the two groups identified in the microsatellite analysis indicated that all but one haplotype were confined to one of the two groups. The shared haplotype (haplotype 6) represented the most frequent haplotype in the study (with a frequency of 0.78), and appeared to be the ancestral haplotype based on its central position in the star-like phylogeny of haplotype relationships (Figure 4). The estimated transition/transversion ratio was 22.08, and the α estimate for the γ distribution of mutation rate heterogeneity among sites was 0.02. On the basis of Akaike information criteria, and Bayesian information criteria, the HKY model of molecular evolution seemed most appropriate for downstream analyses. However, the HKY model is not an option in Arlequin, and therefore the Tamura–Nei model of evolution, which was ranked second to the HKY model in ModelGenerator, was used in the calculation of φST. The divergence between the groups was high, with FST and φST estimates (Excoffier et al., 1992) of 0.56 and 0.65, respectively (both P-values <0.001).
Minimum-spanning tree for each haplotype within the data set. Each hatch mark represents a transitional mutation and each rectangle represents a transversion. The presence of an unsampled haplotype was inferred, and is indicated by a box with a question mark. Haplotypes shaded gray were found only in sampled individuals of the narrow form, and those shaded black were found only in the wide form. Haplotype 6 was the only shared haplotype identified, having a frequency of 0.83 in the wide form and was found in one individual of the narrow form.
Combining morphological and genetic data
Combining the morphological and genetic data showed complete congruence between the methods. Both approaches clearly and independently indicated that the analyzed individuals represent two distinct groups, and the assignment of individuals into each group was also completely consistent (for example, the two groups identified in the genetic analyses represented the two identified morphotypes). Of the 52 specimens, 16 were found to represent the narrow ‘asiaeorientalis’ form, and 36 represented the wide ‘phocaenoides’ form. Thus, the combined data clearly indicated that the two morphotypes represented two distinct groups, with strong genetic differentiation between them. Moreover, the genetic data show that the differentiation is of an evolutionary time scale, suggesting that the two forms have been reproductively isolated, or nearly so, for a long period of time despite being sympatric. Thus, the data suggest that the two forms represent different species as defined by the BSC.
Assessing gene flow
Despite the high levels of genetic divergence between the two forms, there were shared alleles at both the mitochondrial and nuclear markers. Shared alleles between putative groups (particularly of mitochondrial haplotypes) are often taken at face value as evidence for contemporary gene flow, and interpreted as rejecting the hypothesis of reproductive isolation and evolutionary distinctiveness of the taxonomic units of interest (Moritz, 1994). However, if speciation is relatively recent, then alleles will be shared between the putative species in the absence of gene flow solely due to their common ancestry. In these cases, the criteria typically used will falsely reject the hypothesis of reproductive isolation, and therefore not detect recent speciation events until sufficient time has passed for complete lineage sorting to occur.
To determine if the shared alleles between the two forms of finless porpoise were due to a low rate of migration or common ancestry, the coalescent-based program IM was used. The multiple iterations of these analyses provided similar results (except for t, see below), suggesting that the conditions used were appropriate for this data set and provided good estimation of the parameters (Table 2). Additionally, the graphs of the likelihood distributions for one of the iterations are presented in Supplementary Figure 1 and show that under the conditions used the range of values tested is appropriate for these data, and to show that in all cases the posterior distributions are either complete or are ‘flat’, meaning that they have leveled out after having a peak elsewhere in the distribution.
Estimates of the ancestral effective population size (qA), and both extant effective population sizes (q1 and q2) were 8150, 6200 and 650, respectively (Table 2). The estimated time since divergence was 456 generations; however there was wide variation around this estimate between runs, suggesting that the estimates of t are unreliable under the current conditions (Table 2). The estimated migration rate in one direction was zero, and although the other migration rate estimate was 0.26 migrants per generation (Table 2), the likelihood of this migration rate was not vastly higher than the likelihood of no migration (average log-likelihood ratio= −1.06).
Although a statistical nonzero migration rate estimate was obtained, the next step involved testing if the hypothesis of no gene flow could actually be rejected. Likelihood ratio tests are not appropriate for the data generated by IM (Nielsen and Wakely, 2001), and the ideal method would be to conduct simulations-based estimates of qA, q1, q2 and t, and use these data to obtain a distribution of expected migration rate estimates from IM under the scenario of no migration, sensu Nielsen and Wakely (2001). However, analyzing a reasonable number simulated data sets in this manner would require a prohibitive amount of computational time. As an alternative, simulations were still conducted based on the null hypotheses, but were used to generate expected distributions of FST and RST between the two forms (instead of migration estimates from IM), which could then be compared to the observed values.
The initial simulations were based on the effective population size (qA, q1 and q2) and time since divergence (t) estimates from IM, with complete reproductive isolation since divergence. A total of 100 simulations were conducted, based on individuals genotyped at nine microsatellite loci, and for each simulation the resulting populations were sampled in the same manner as in the actual populations (for example, the same numbers of individuals were sampled from each simulated population as were sampled from the actual populations). Thus, our sampling regime was also incorporated into the simulations. For each simulation, FST and RST estimates were obtained using the program SPAGeDi (Hardy and Vekemans, 2002).
For both FST and RST, the observed values were much larger than were found in any of the iterations of the simulated conditions (Figure 5a). These data not only reject the hypothesis of measurable gene flow between the putative species within the past 456 generations (t estimate from IM, P<0.01), but also suggest that the two groups have been isolated for a much longer time period. Although multiple estimates of t may be consistent with the observed data, we hypothesized that the two forms diverged during the last glaciation, and thus focused the remaining analyses on testing this hypothesis.
Observed estimates of differentiation (FST and RST) versus those expected under two hypothesized scenarios. (a) Expected differentiation based on no migration since a time of divergence (t) of 456 generations ago (estimate of t obtained from IM). (b) Expected differentiation if the two forms diverged during the last ice age (∼18 000 years or 2570 generations ago) and have been reproductively isolated since. In all cases the height of the ‘observed’ bar is not to scale for the y axis, but rather just represents a visualization of where the observed value falls in relation to the expected values. Expected values for each scenario are based on 100 simulations conducted as outlined in the text.
During the last glacial maximum (∼17 000–18 000 years ago), lower sea levels resulted in the emergence of a land bridge that connected Taiwan to mainland China, eliminating the Taiwan Strait and thus separating the northern and southern coastal waters of China (Calder, 1983; Voris, 2000). We postulate that this event divided the ancestral population of finless porpoises and resulted in the two present species of finless porpoises diverging in allopatry. By about 11 000 years before present, the Taiwan Strait reappeared with rising sea levels due to glacial retreat (Voris, 2000) and secondary contact between the two new species was established. To test this hypothesis we conducted 100 simulations in the same manner as before (for example, with no migration since divergence), except under this scenario the time since divergence used was 18 000 years; finless porpoises reach sexual maturity between 4 and 9 years (Amano, 2002; Jefferson and Hung, 2004), and we used a conservative estimate of 7 years for the generation time; therefore, this represents a time since divergence of ∼2570 generations. The estimates of effective population size were kept the same as for the previous simulations. Under this scenario the observed estimates of FST and RST (and significantly, the ratio between the two) were precisely the same as the mode of the distribution of expected values under the scenario of complete reproductive isolation since the last ice age (Figure 5b). Thus, the data reject the hypothesis of measurable gene flow within the past 456 generations, and are exactly what would be expected if the two forms have been reproductively isolated since the last ice age.
Using the smallest estimate of Ne (650) obtained from IM, complete lineage sorting (at least of mitochondrial sequences) between the putative species should occur after approximately 4Ne=2600 generations. Thus, despite the large number of years that have passed since the hypothesized speciation event, the long generation time of finless porpoises indicates that the time since divergence (∼2570 generations) has not been quite long enough to expect reciprocal monophyly between the two forms. These data demonstrate that the criterion of reciprocal monophyly is inappropriate for detecting ‘recent’ speciation events.
It could be argued, based on the assumption that the Ne of mtDNA is one-quarter that of nuclear markers (Halliburton, 2004), that enough time has passed to expect complete lineage sorting for the mitochondrial sequences. However, it has been shown that this assumption is rarely valid in biologically realistic situations. For example, the presence of even extremely subtle polygyny and/or female philopatry reducing female dispersal relative to that of males can result in the Ne of mtDNA being markedly higher than that of nuclear markers (Chesser and Baker, 1996; Hoelzer, 1997). Strong female philopatry and male polygyny are two characteristics that are widespread throughout cetacean species (Mann et al., 2000), and are indeed common to many mammalian species. Therefore, in this case it does not seem reasonable to assume that the mitochondrial Ne is markedly less than the Ne estimated from nuclear markers, or that enough time has passed to expect complete lineage sorting of the mitochondrial sequences. Confirmation of this can be seen in estimates of θ for the ancestral population (θA), population 1 (θ1), and population 2 (θ2) obtained from IM based on the mitochondrial versus nuclear loci. Mean estimates based on the mitochondrial data are θA=20.37, θ1=13.95 and θ2=1.7; whereas for the nuclear loci these estimates are θA=16.3, θ1=12.4 and θ2=1.3. The mitochondrial estimates are already scaled for mtDNA, and thus these estimates are directly comparable to the diploid nuclear marker estimates. Note that in all cases the mtDNA estimates are higher than those based on the nuclear loci. The differences in Ne estimates will be even higher because Ne=θ/4μ, where μ is the mutation rate per generation. Mutation rates in the control region are typically much lower than those of microsatellites (Li, 1997), which would result in much larger estimates of Ne based on the mitochondrial data than on the nuclear data.
The tests of contemporary gene flow (from BaysAss) also resulted in nonzero migration rate estimates (m), with m estimates of 0.0406 (95% CI: 0.00179–0.124) and 0.0116 (95% CI: 0.000301–0.0443) from Pop1 to Pop2, and Pop2 to Pop1, respectively. However, BaysAss is known to overestimate migration rates when the true rate is zero (Faubet et al., 2007). Thus, to provide context for the interpretation of these results, the simulated data were used. Specifically, the 100 sets of sampled genotypes simulated under our hypothesis of reproductive isolation since the last ice age were run through BayesAss to obtain the expected migration rate estimates for these data if the true migration rate is zero. As with the data on FST and RST, the observed migration rate estimates, while nonzero, fall well within the range of expected mean migration rate estimates from BayesAss with this data set if these two forms have been reproductively isolated since the last ice age (P<0.38 for m estimate from Pop1 to Pop2; P<0.58 for m estimate from Pop2 to Pop1). Thus, the data on both historical and contemporary gene flow are consistent with the hypothesis that the two forms have been reproductively isolated since the last ice age.
It is noteworthy that although the number of specimens analyzed was limited, the study design and analytical approaches were conducted in such a way that the analyses and resulting conclusions should be robust despite this potential limitation. The study design was based on maximizing the number of specimens analyzed from the area of sympatry, as these are the individuals necessary to test the hypothesis of reproductive isolation in the most direct manner, and focusing on just these samples (even if the sample size is small) maximizes the probability of detecting gene flow if it is occurring. On the analytical side, both methods of estimating gene flow (IM and BayesAss) have been shown to provide reliable results when sample sizes are small if a decent number of loci are used (Wilson and Rannala, 2003; Won and Hey, 2005). Indeed, one assumption when conducting analyses based on the coalescent (such as with IM) is that the sample size is much smaller than the effective size of the population (Wakeley and Takahashi, 2003), and it is therefore desirable to analyze a relatively small number of samples. Additionally, the results of all genetic analyses were put into context based on simulations of the hypothesized scenarios. The simulation processes included sampling the simulated populations in the same manner as we sampled the actual populations (for example, the same number of individuals were sampled from each simulated population as were sampled from the two forms of finless porpoises), and therefore sample size was integrated into all analytical procedures and subsequent interpretations. Combined, this study design and analytical approach allows for robust testing of this taxonomic hypothesis, although more details of this system will be of course obtainable with a larger data set in the future.
Taxonomic implications/recommendations
Combining all of the information suggests that the two forms of finless porpoise represent distinct groups that have been reproductively isolated from one another (despite currently living in sympatry) since sharing a common ancestor ∼18 000 years ago, and therefore represent different biological species. It is noteworthy that the two forms would also qualify as separate species had we chosen other criteria to test this hypothesis. For example, the genetic data reject both recent and historical genetic exchangeability, and the morphological (and previous skeletal) data reject ecological exchangeability. Thus, the two forms would also qualify as different species under the criteria of Crandall et al. (2000).
We recommend that the two species be referred to as Neophocaena phocaenoides (G. Cuvier, 1829) and N. asiaeorientalis as suggested previously (Jefferson, 2002). To facilitate clearer communication of the two species, we propose the following common names: Indo-Pacific finless porpoise for N. phocaenoides, and narrow-ridged finless porpoise for N. asiaeorientalis. Population structure within these two species, as well as the relationship between these forms and a third recognized morphotype that inhabits the coastal waters of Japan, requires further study.
Conclusions
It is important that studies testing species boundaries clearly and explicitly present a testable hypothesis based on a species concept and its criteria (Sites and Crandall, 1997). However, in almost all earlier taxonomic investigations of finless porpoises, testable hypotheses and criteria for accepting species-level designations were lacking. Additionally, previous genetic investigations were based solely on mtDNA and used geography as a basis for a priori grouping of specimens (Yang et al., 2002), both methods of which are widely used in taxonomic investigations in general. This case study demonstrates that although mtDNA can be highly informative, it is of limited value (and can lead to incorrect conclusions) on its own in testing hypotheses of recent speciation events and α-level taxonomy, where lineage sorting is incomplete. In these situations, Mendelian-inherited nuclear loci may be more informative for assessing reproductive isolation. Nuclear markers also allow reproductive entities to be identified based on the genetic data solely (Pritchard et al., 2000), allowing for truly independent tests of congruence between genotypic and phenotypic characters. This approach also removes the necessity of grouping individuals based on a priori criteria, such as geography, which can clearly lead to erroneous taxonomic conclusions when cryptic species are sympatric. Lastly, the primary difficulty in identifying recent speciation events revolves around the interpretation of shared alleles. Historically, and in many current studies, the interpretation of shared alleles has been oversimplified and taken at face value as evidence of gene flow; however with incomplete lineage sorting alleles can be shared between putative species despite complete reproductive isolation solely due to common ancestry. As shown in this case study, recent analytical advances have made it possible to differentiate between these scenarios, greatly increasing both the power to detect recently derived species, and the resolution for analyses of evolutionary histories. Such power is of increasing importance given the growing concern for biodiversity conservation issues and recognition that sibling species and/or cryptic species in sympatry may be quite common, particularly in marine environments (Knowlton, 1993).
References
Allendorf FW, Luikart G (2007). Conservation and the Genetics of Populations. Blackwell: Malden.
Amano M (2002). Finless porpoise, Nephocaena phocaenoides. In: Perrin WF, Würsig B, Thewissen JGM (eds). Encyclopedia of Marine Mammals. Academic Press: San Diego, pp 432–435.
Avise JC (2004). Molecular Markers, Natural History, and Evolution. Sinauer Associates: Sunderland.
Avise JC, Neigel JE, Arnold J (1984). Demographic influences on mitochondrial DNA lineage survivorship in animal populations. J Mol Evol 20: 99–105.
Balloux F (2001). EASYPOP (Version 1.7): a computer program for population genetics simulation. J Hered 92: 301–302.
Balloux F, Lugon-Moulin N (2002). The estimation of population differentiation with microsatellite markers. Mol Ecol 11: 155–165.
Calder N (1983). Timescale: An Atlas of the Fourth Dimension. Viking Press: New York.
Chesser RK, Baker RJ (1996). Effective sizes and dynamics of uniparentally and diparentally inherited genes. Genetics 144: 1225–1235.
Coyne JA, Orr HA (2004). Speciation. Sinauer Associates: Sunderland.
Crandall KA, Bininda-Edmonds ORP, Mace GM, Wayne RK (2000). Considering evolutionary processes in conservation biology. Trends Ecol Evol 15: 290–295.
Daugherty CH, Cree A, Hay JM, Thompson MB (1990). Neglected taxonomy and continuing extinctions of tuatara (Sphenodon). Nature 347: 177–179.
Di Rienzo A, Peterson AC, Garza JC, Valdes AM, Slatkin M, Freimer NB (1994). Mutational processes of simple-sequence repeat loci in human populations. Proc Natl Acad Sci USA 91: 3166–3170.
Dugan KA, Lawrence HS, Hares DR, Fisher CL, Budowle B (2002). An improved method for post-PCR purification for mtDNA sequence analysis. J Forensic Sci 47: 811–818.
Erwin TL (1991). An evolutionary basis for conservation strategies. Science 253: 750–752.
Estoup A, Angers B (1998). Microsatellites and minisatellites for molecular ecology: theoretical and empirical considerations. In: Carvalho GR (ed). Advances in Molecular Ecology. IOS Press: Amsterdam, pp 55–86.
Excoffier L, Laval G, Schneider S (2005). Arlequin ver.3.0: an integrated software package for population genetics data analysis. Evol Bioinform Online 1: 47–50.
Excoffier L, Smouse P, Quattro J (1992). Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial restriction data. Genetics 131: 479–491.
Faubet P, Waples RS, Gaggiotti OE (2007). Evaluating the performance of a multilocus Bayesian method for the estimation of migration rates. Mol Ecol 16: 1149–1166.
Gao A, Zhou K (1995). Geographical variation of external measurements and three subspecies of Neophocaena phocaenoides in Chinese waters. Acta Theriologica Sinica 15: 81–92.
Halliburton R (2004). Introduction to Population Genetics. Pearson: Upper Saddle River.
Hardy OJ, Charbonnel N, Fréville H, Heuertz M (2003). Microsatellite allele sizes: a simple test to assess their significance on genetic differentiation. Genetics 163: 1467–1482.
Hardy OJ, Vekemans X (2002). SPAGeDi: a versatile computer program to analyze spatial genetic structure at the individual or population levels. Mol Ecol Notes 2: 618–620.
Hendry AP, Vamosi SM, Latham SJ, Heilbuth JC, Day T (2000). Questioning species realities. Conserv Genet 1: 67–76.
Hey J, Nielsen R (2004). Multilocus methods for estimating population sizes, migration rates and divergence time, with applications to the divergence of Drosophila pseudoobscura and D persimilis. Genetics 167: 747–760.
Hey J, Waples RS, Arnold ML, Butlin RK, Harrison RG (2003). Understanding and confronting species uncertainty in biology and conservation. Trends Ecol Evol 18: 597–603.
Hey J, Won Y-J, Sivasundar A, Nielsen R, Markert JA (2004). Using nuclear haplotypes with microsatellites to study gene flow between recently separated cichlid species. Mol Ecol 13: 909–919.
Hoelzer GA (1997). Inferring phylogenies from mtDNA variation: mitochondrial-gene trees versus nuclear-gene trees revisited. Evolution 51: 622–626.
Jefferson TA (2002). Preliminary analysis of geographic variation in cranial morphometrics of the finless porpoise (Neophocaena phocaenoides). Raffles Bull Zool (Supplement 10): 3–14.
Jefferson TA, Hung SK (2004). Neophocaena phocaenoides. Mammal Species 746: 1–12.
Kasuya T (1999). Finless porpoise—Neophocaena phcaenoides (G. Cuvier, 1829). In: Ridgway SH, Harrison R (eds). Handbook of Marine Mammals. Academic Press: San Diego, Vol 6, pp 411–442.
Keane TM, Creevey CJ, Pentony MM, Naughton TJ, McInerney JO (2006). Assessment of methods for amino acid matrix selection and their use on empirical data shows that ad hoc assumptions for choice of matrix are not justified. BMC Evol Biol 6: 29.
Kirkpatrick BW (1992). Identification of a conserved microsatellite site in the porcine and bovine insulin-like growth factor-I gene 5′ flank. Anim Genet 23: 543–548.
Knowlton N (1993). Sibling species in the sea. Annu Rev Ecol Syst 24: 189–216.
Li W-H (1997). Molecular Evolution. Sinauer Associates: Sunderland.
Mann J, Connor RC, Tyack PL, Whitehead H (eds) (2000). Cetacean Societies: Field Studies of Dolphins and Whales. University of Chicago Press: Chicago.
Mayr E (1942). Systematics and the Origin of Species. Columbia University Press: New York.
Moritz C (1994). Defining ‘evolutionary significant units’ for conservation. Trends Ecol Evol 9: 373–375.
Nielsen R, Wakely J (2001). Distinguishing migration from isolation: a Markov chain Monte Carlo approach. Genetics 158: 885–896.
Pilleri G, Gihr M (1975). On the taxonomy and ecology of the finless black porpoise, Neophocaena (Cetacea, Delphinidae). Mammalia 39: 657–673.
Pritchard JK, Stephens M, Donnelly P (2000). Inference of population structure using multilocus genotype data. Genetics 155: 945–959.
Raymond M, Rousset F (1995). GENEPOP: population genetics software for exact tests and ecuminicism. J Hered 86: 248–249.
Renwick A, Davison L, Spratt H, King JP, Kimmel M (2001). DNA dinucleotide evolution in humans: fitting theory to facts. Genetics 159: 737–747.
Rice DW (1999). Marine Mammals of the World: Systematics and Distribution. Society for Marine Mammalogy Special Publication 4: Lawrence.
Rundle HD, Nagel L, Boughman JW, Schluter D (2000). Natural selection and parallel speciation in sympatric sticklebacks. Science 287: 306–308.
Seutin G, White BN, Boag PT (1991). Preservation of avian blood and tissue samples for DNA analyses. Can J Zool 69: 82–90.
Singer VL, Jones LJ, Sue ST, Haugland RP (1997). Characterization of PicoGreen reagent and development of a fluorescent-based solution assay for double-stranded DNA quantitation. Anal Biochem 249: 228–238.
Sites Jr JW, Crandall KA (1997). Testing species boundaries in biodiversity studies. Conserv Biol 11: 1289–1297.
Thompson JD, Higging DG, Gibson TJ (1994). CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22: 4673–4680.
Voris HK (2000). Maps of Pleistocene sea levels in southeast Asia: shorelines, river systems and time durations. J Biogeogr 27: 1153–1167.
Wakeley J, Takahashi T (2003). Gene genealogies when the sample size exceeds the effective size of the population. Mol Biol Evol 20: 208–213.
Wang P (1992). The morphological characters and the problem of subspecies identifications of the finless porpoise. Fish Sci 11: 4–9.
Wilson GA, Rannala B (2003). Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163: 1177–1191.
Won Y-J, Hey J (2005). Divergence population genetics of chimpanzees. Mol Biol Evol 22: 297–307.
Yang G, Ren W, Zhou K, Liu S, Ji G, Yan J et al. (2002). Population genetic structure of finless porpoises, Neophocaena phocaenoides, in Chinese waters, inferred from mitochondrial control region sequences. Mar Mammal Sci 18: 336–347.
Yoshida H, Yoshioka M, Shirakihara M, Chow S (2001). Population structure of finless porpoises (Neophocaena phocaenoides) in coastal waters of Japan based on mitochondrial DNA sequences. J Mammal 82: 123–130.
Acknowledgements
We thank all those who were involved in providing specimens for this study: ECM Parsons, Lien-Jiang County government of Taiwan, S Leatherwood, P Wang and L-S Chou. We also thank S Leatherwood and P Wang for encouraging JYW to study finless porpoises and TA Jefferson for stimulating discussions about finless porpoises, encouragement and support. We especially thank SK Hung, ER Secchi, G Abel, PE Rosel and BA McLeod for their support. Funding was provided by the Whale and Dolphin Conservation Society (UK), Ocean Park Conservation Foundation Hong Kong (OPCFHK), FormosaCetus Research and Conservation Group (Canada and Taiwan), and the National Sciences and Engineering Research Council of Canada (NSERC). The computational aspect of this work was made possible by the facilities of the Shared Hierarchical Academic Research Computing Network (SHARCNET: www.sharcnet.ca). We thank three anonymous reviewers whose comments greatly improved the paper.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Heredity website (http://www.nature.com/hdy)
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, J., Frasier, T., Yang, S. et al. Detecting recent speciation events: the case of the finless porpoise (genus Neophocaena). Heredity 101, 145–155 (2008). https://doi.org/10.1038/hdy.2008.40
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2008.40
Keywords
This article is cited by
-
Gapless genome assembly of East Asian finless porpoise
Scientific Data (2022)
-
Mitochondrial genomics reveals the evolutionary history of the porpoises (Phocoenidae) across the speciation continuum
Scientific Reports (2020)
-
Phylogenetic relationships between different raccoon dog (Nyctereutes procyonoides) populations based on four nuclear and Y genes
Genes & Genomics (2020)
-
Quaternary climate change drives allo-peripatric speciation and refugial divergence in the Dysosma versipellis-pleiantha complex from different forest types in China
Scientific Reports (2017)
-
Organization and characteristics of the major histocompatibility complex class II region in the Yangtze finless porpoise (Neophocaena asiaeorientalis asiaeorientalis)
Scientific Reports (2016)