Abstract
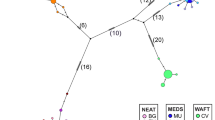
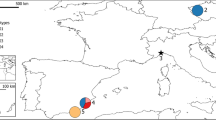
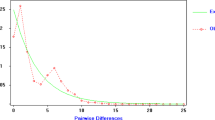
We determined the nucleotide sequence of the first 287 base pairs in the mitochondrial control region from 74 narwhals, Monodon monoceros, collected in the North-west Atlantic. We detected four polymorphic sites that defined five haplotypes, two of which were found in single specimens. The same DNA sequence was characterized in an additional 353 specimens by digestion with two restriction endonucleases. In this manner each specimen could be assigned to one of the three most common haplotypes. The nucleotide diversity for the total sample (as well as the sequenced subset) was estimated as 0.0017 and pairwise genetic distances between haplotypes ranged from 0.0035-0.0070. The low nucleotide diversity and the low average pair-wise genetic distance between haplotypes suggest a recent expansion in abundance from a small founding population. Despite the low degree of variation, frequencies of the common haplotypes differed markedly between areas. The results indicate isolation, even between geographically close areas, as well as fidelity to specific summer and autumn feeding grounds. Heterogeneity within a presumed single breeding ground suggests mixing of pods with different haplotypic composition.
Similar content being viewed by others
Article PDF
References
Amos, W, and Hoelzel, A R. 1991. Long-term preservation of whale skin for DNA analysis. Rep Int Whaling Comm Spec Issue, 13, 99–104.
Amos, B, Barrett, J, and Dover, G A. 1991. Breeding behaviour of pilot whales revealed by DNA fingerprinting. Heredity, 67, 49–55.
Amos, B, Schlötterer, C, and Tautz, D. 1993. Social structure of pilot whales revealed by analytical DNA profiling. Science, 260, 670–672.
Arnason, U, Gullberg, A, and Widegren, B. 1993. Cetacean mitochondrial DNA control region: Sequences of all extant baleen whales and two sperm whale species. Mol Biol Evol, 10, 960–970.
Baker, C S, Perry, A, Bannister, J L, Weinrich, M T, Abernethy, R B, and Calambodikis, J. et al. 1993. Abundant mitochondrial DNA variation and world-wide population structure in humpback whales. Proc Natl Acad Sci, USA, 90, 8239–8243.
Bérubé, M, and Palsbøll, P J. 1996. Identification of sex in Cetaceans by multiplexing with three ZFX and ZFY specific primers. Mol Ecol, 5, 283–287.
Dietz, R, and Heide-Jørgensen, M P. 1995. Movements and swimming speed of narwhals (Monodon monoceros) instrumented with satellite transmitters in Melville Bay, Northwest Greenland. Can J Zool, 73, 2106–2119.
Dietz, R, Heide-Jørgensen, M P, Born, E W, and Glahder, C M. 1994. Occurrence of narwhals, Monodon monoceros, and white whales, Delphinapterus leucas, in East Greenland. Meddelelser om Grønland, Bioscience, 39, 69–86.
Dizon, A E, Southern, S O, and Perrin, W F. 1991. Molecular analysis of mtDNA types in exploited populations of Spinner dolphins (Stenella longirostris). Rep Int Whaling Comm Spec Issue, 13, 183–202.
Duffield, D A, and Wells, R S. 1991. The combined application of chromosome, protein and molecular data for the investigation of social unit structure and dynamics in Tursiops truncatus. Rep Int Whaling Comm Spec Issue, 13, 155–170.
Gyllensten, U B, and Erlich, H A. 1988. Generation of single-stranded DNA by the polymerase chain reaction and its application to direct sequencing of the HLA-DQA locus. Proc Natl Acad Sci, USA, 85, 7652–7656.
Hay, K A, and Mansfield, A W. 1989. Narwhal Monodon monoceros Linnaeus, 1758. In: Ridgeway, H. and Harrison, R. (eds) River Dolphins and the Larger Toothed Whales Handbook of Marine Mammals, vol. 4, pp. 145–176. Academic Press, London.
Hoelzel, A R, and Dover, G A. 1991. Genetic differentiation between sympatric killer whale populations. Heredity, 66, 191–195.
Hoelzel, A R, Halley, J, O'Brien, S J, Campagna, C, Arnbom, T, and Le Boeuf, B. et al. 1993. Elephant seal genetic variation and the use of simulation models to investigate historical population bottlenecks. J Hered, 84, 443–449.
Hudson, R R. 1992. Reply to Roff and Bentzen. Mol Biol Evol, 9, 969.
Hudson, R R, Boos, D D, and Kaplan, N L. 1992. A statistical test for detecting geographic subdivision. Mol Biol Evol, 9, 138–151.
IWC, 1992. Report of the sub-committee on small cetaceans. Rep Int Whaling Comm, 42, 178–234.
Koski, W R, and Davis, R A. 1994. Distribution and numbers of narwhals (Monodon monoceros) in Baffin Bay and Davis Strait. Meddelelser om Grønland, Bioscience, 39, 15–40.
Maniatis, T, Fritsch, E F, and Sambrook, J. 1982. Molecular cloning A Laboratory Manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY.
Nei, M. 1987. Molecular Evolutionary Genetics. Columbia University Press, New York.
Palsbøll, P J, Clapham, P J, Mattila, D K, Larsen, F, Sears, R, Siegismund, H R. et al. 1995. Distribution of mtDNA haplotypes in North Atlantic humpback whales: the influence of behaviour on population structure. Mar Ecol Progress Series, 116, 1–10.
Reeves, R R, and Heide-Jørgensen, M P. 1994. Commercial aspects of narwhal exploitation in Greenland, with emphasis on the exportation of tusk ivory. Meddelelser om Grønland, Bioscience, 39, 119–134.
Roff, D E, and Bentzen, P. 1992. Detecting geographic subdivision: A comment on a paper by Hudson et al. Mol Biol Evol, 9, 968.
Rosel, P E, Dizon, A E, and Heyning, J E. 1994. Genetic analysis of sympatric morphotypes of common dolphins (genus Delphinus). Mar Biol, 119, 159–167.
Saiki, R K, Gelfand, D H, Stoffel, S, Scharf, S J, Higuchi, R, and Horn, G T. et al. 1988. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science, 239, 487–491.
Sanger, F. 1981. Determination of nucleotide sequences in DNA. Science, 214, 1205–1210.
Schaeff, C, Kraus, S, Brown, M, Perkins, J, Payne, R, Gaskin, D, and Boag, P. 1991. Preliminary analysis of mitochondrial DNA variation within and between right whale species Eubalaena glacialis and Eubalaena australis. Rep Int Whaling Comm Spec Issue, 13, 155–170.
Siegstad, H, and Heide-Jørgensen, M P. 1994. Ice entrapments of narwhals (Monodon monoceros) and white whales (Delphinapterus leucas) in Greenland. Meddelelser om Grønland, Bioscience, 39, 119–134.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Palsbøll, P., Heide-Jørgensen, M. & Dietz, R. Population structure and seasonal movements of narwhals, Monodon monoceros, determined from mtDNA analysis. Heredity 78, 284–292 (1997). https://doi.org/10.1038/hdy.1997.43
Received:
Issue Date:
DOI: https://doi.org/10.1038/hdy.1997.43
Keywords
This article is cited by
-
The evolution of menopause in toothed whales
Nature (2024)
-
Long-term isolation at a low effective population size greatly reduced genetic diversity in Gulf of California fin whales
Scientific Reports (2019)
-
Analyses of ovarian activity reveal repeated evolution of post-reproductive lifespans in toothed whales
Scientific Reports (2018)
-
Genetic diversity of the Tibetan antelope (Pantholops hodgsonii) population of Ladakh, India, its relationship with other populations and conservation implications
BMC Research Notes (2016)
-
Genetics and fatty acids assist in deciphering narwhal (Monodon monoceros) social groupings
Polar Biology (2015)