Abstract
Purpose
Whole-exome sequencing (WES) has revolutionized Mendelian diagnostics, however, there is no consensus on the timing of data review in undiagnosed individuals and only preliminary data on the cost-effectiveness of this technology. We aimed to assess the utility of WES data reanalysis for diagnosis in Mendelian disorders and to analyze the cost-effectiveness of this technology compared with a traditional diagnostic pathway.
Methods
WES was applied to a cohort of 54 patients from 37 families with a variety of Mendelian disorders to identify the genetic etiology. Reanalysis was performed after 12 months with an improved WES diagnostic pipeline. A comparison was made between costs of a modeled WES pathway and a traditional diagnostic pathway in a cohort with intellectual disability (ID).
Results
Reanalysis of WES data at 12 months improved diagnostic success from 30 to 41% due to interim publication of disease genes, expanded phenotype data from referrer, and an improved bioinformatics pipeline. Cost analysis on the ID cohort showed average cost savings of US$586 (AU$782) for each additional diagnosis.
Conclusion
Early application of WES in Mendelian disorders is cost-effective and reanalysis of an undiagnosed individual at a 12-month time point increases total diagnoses by 11%.
Similar content being viewed by others
Introduction
Mendelian disorders have a major impact on families, occurring at a population frequency of at least 1%,1 for a worldwide incidence per year of 1.35 million births.2 These disorders require complex management and are clinically and genetically heterogeneous, making a molecular diagnosis challenging. This can lead to a lengthy diagnostic journey for patients who often remain undiagnosed. Next-generation sequencing technologies such as whole-exome sequencing (WES) have introduced powerful tools to identify the molecular etiology of Mendelian disorders,3 enabling improved diagnosis over traditional testing methodologies.
The early availability of WES reduces avoidable and invasive investigations and limits diagnostic odysseys. Significant cost savings have already been reported with appropriate use of next-generation sequencing4 when applied to Mendelian disorders.5,6,7,8 However, whether whole-genome sequencing (WGS) or WES is the most cost-effective genomic diagnostic technology remains unresolved. Despite the potential for WGS to improve copy-number variant detection, test cost has remained a barrier to the routine adoption of this technology over WES.9
Reanalysis of WES data could improve diagnostic rates in patients without an initial molecular etiology,10,11 however, there is no established timeframe for reanalysis. We therefore performed WES in a cohort of 54 patients from Australia with a variety of likely Mendelian disorders to assess both the clinical utility and reanalysis diagnostic potential of WES. The diagnostic success was improved following reanalysis of WES data 12 months after the initial assessment, and highlighted the contributing factors that will be important for future development of reanalysis strategies. Through a health economics comparison analysis of diagnostic costs between a WES strategy and a traditional diagnostic approach we demonstrate the potential cost savings of WES. We expect this will help to inform genomic diagnostic choices in the health-care setting.
Materials and methods
Cohort ascertainment
Patients were recruited through clinical genetics units in New South Wales (NSW) from July 2013 to July 2014. The selection criteria included a distinctive phenotype likely to have a monogenic etiology, a family structure consistent with Mendelian inheritance, availability of relevant family members for testing, the provision of consent for genomic testing, and no known genetic etiology. Prior diagnostic investigations had all been negative (examples in Supplementary Table S1 online) including chromosome microarray in patients with intellectual disability (ID). Fifty-four affected individuals, unaffected parents, or other affected relatives from 37 families underwent WES following informed consent. Study approval was granted by the ethics committee at the Prince of Wales Hospital Campus, Randwick (Human Research Ethics Committee ref. no. 13/094 and LNR/13/SCHN/112).
WES and bioinformatics analysis
DNA was extracted from EDTA blood. WES was performed using a Nextera extended exome kit or a NimbleGen (Madison, WI) VCRome rapid capture expanded exome kit, with libraries analyzed on Illumina (San Diego, CA, USA) HiSeq2500 instruments. Reads were aligned to the Human Reference Sequence hs37d5 using the Burrows–Wheeler Aligner (BWA-MEM; v0.7.12-r1039), followed by indel realignment and base quality score recalibration. Single-nucleotide and short insertion/deletion variants were identified using the GATK HaplotypeCaller (v3.3). Variant quality was assessed using variant quality score recalibration, then annotated using the Ensembl Variant Effect Predictor (http://www.ensembl.org/info/docs/tools/vep/index.html) including data from the 1000 Genomes Project (http://www.internationalgenome.org/), the Exome Variant Server database (ESP6500SI-V2; http://evs.gs.washington.edu/EVS/), the Exome Aggregation Consortium (http://exac.broadinstitute.org/), the Combined Annotation Dependent Depletion (CADD) database,12 OMIM (http://omim.org), and Orphanet (http://www.orpha.net/). Gender and familial relationship quality control steps were performed including the calculation of the coefficient of inbreeding and XY genotyping using the KING and PLINK software packages.
Annotated variants were converted into a searchable SQL databases using the GEnome MINIng (GEMINI) software.13 Interpretation of variants was limited to those annotated by Variant Effect Predictor, which in practice are the coding exons and their flanking 10 bases within introns.
Variant filtering and prioritization
Variants were filtered and prioritized by a clinical geneticist according to pedigree structure; however, all possible Mendelian inheritance patterns were assessed for each family. Variants were discarded if at a minor allele frequency of greater than 1% in population databases or in-house WES controls, or predicted to have a low impact on protein function by the GEMINI software. When assessing rare heterozygous variants for a de novo model, initial analysis was limited to those variants unique to the proband(s). Candidate variants were further assessed for pathogenicity using in silico prediction tools (SIFT, PolyPhen2, PROVEAN, CADD).12,14 Homozygous regions were identified in consanguineous families using HomozygosityMapper15 from either single-nucleotide polymorphism array data or WES VCF files. Variants were reviewed for their quality and underlying genomic architecture by uploading BAM files into the Integrative Genomics Viewer.16
Variants were reviewed by genetic pathologists utilizing the American College of Medical Genetics and Genomics guidelines17 and considered pathogenic based on adequate phenotype–genotype correlation in genes with published gene–disease evidence that included functional data and probands from separate families. Pathogenic variants identified by WES were frequently discussed by a multidisciplinary team and validated by Sanger sequencing on stored patient and parental samples.
12-month variant reanalysis
Repeat analysis in undiagnosed families was undertaken at 12 months following initial analysis. This timeframe was selected to take into account the clinical diagnostic resources required for reanalysis of a cohort of this size and was also based on a typical clinical genetics review cycle. Bioinformatics pipeline alterations included GEMINI and GATK (2.8 to 3.3) software updates, an increase in joint calling batch size, and development of an in-house variant-filtering platform, Seave, as a graphical user interface for GEMINI to streamline variant filtering and monthly updated gene lists from OMIM and Orphanet.
Economic analysis
Fourteen patients with ID (mixed syndromic (S-ID) and nonsyndromic (NS-ID)), for whom medical records were available, were selected to compare the diagnostic cost-effectiveness of a genomic pathway over the traditional diagnostic pathway. The proportion of diagnosed families was equivalent to that in the overall ID cohort before reanalysis (36 vs. 33% respectively). A clinical geneticist extracted diagnostic information from patient records. Costs for diagnostic encounters and procedures were determined by estimating staff time and using local salary data for 2016 from the NSW Health Department,18 alongside procedural and investigation costs listed in the Australian Medicare Benefits Schedule.19 Single-gene Sanger sequencing, deletion/duplication studies, and biochemical test costs were sought from referral laboratories. WES and WGS costs were obtained from local laboratories (Supplementary Table S1 online).
Total costs for the traditional pathway were calculated and compared with two alternative genomic diagnostic pathways (counterfactual arms): (i) WES available at initial contact with the clinical genetics service, and (ii) WES available at time of initial patient presentation with clinical symptoms that would warrant genomic testing. Genomic diagnostic pathway costs were calculated to include the WES costs, molecular pathology reporting, and data storage. Costs from the traditional pathway were removed if investigations, such as single-gene sequencing, or procedures could have been avoided if WES had been available. Some investigations or procedures were removed if timing of patient diagnosis meant these would have avoided, such as invasive lumbar punctures for diagnostic evaluation.
The average cost per patient, average cost per diagnosis, and incremental cost per additional diagnosis for the WES pathway compared with the traditional pathway were estimated for the initial analysis and 12-month reanalysis, the latter including rephenotyping costs. The uncertainty associated with these estimates was calculated using a bootstrapping methodology to create 1,000 replicated data sets by drawing a random sample of the 14 records, 1,000 times with replacement. The outcomes were then estimated for each replicated data set by generating 1,000 estimates and a distribution. The 95% confidence intervals (CIs) were calculated as uncertainty intervals for each outcome based on these distributions using the percentile method. Results of bootstrapped simulations are presented as scatterplots on cost-effectiveness planes where each point represents the result of each simulation (Figure 2). Analyses were performed in Microsoft Excel and SAS version 9.4.
The cost-effectiveness of WGS over WES was analyzed for a number of hypothetical scenarios with a range of diagnostic rates and WGS costs. WGS costs were calculated similarly to WES, with removal of investigations or procedures from the traditional pathway costs that would have been avoided if WGS were available. It was assumed that WGS and WES were available at initial patient presentation with features warranting genomic testing and that all patients diagnosed by WES would have been diagnosed by WGS. The Monte Carlo simulation method was used to randomize which of the undiagnosed patients after WES would be attributed a diagnosis with application of WGS. The simulation was iterated 1,000 times and 95% uncertainty intervals were estimated based on these data sets.
Results
Cohort description and overall results
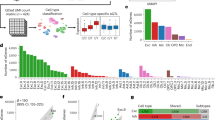
A diagnosis was made on initial WES analysis in 11/37 families (30%). Probands had a variety of disorders, the majority with S-ID (49%; 18 families), and the remainder skeletal (13%; 5), hematological (11%; 4), NS-ID (8%; 3), visual (8%; 3), neurological (5%; 2), metabolic (3%; 1), or a syndromal disorder (3%; 1 (Noonan syndrome no ID)). Families included probands (59%; 22 families) with no family history or multiple affected individuals (41%, 15) who were either sibling pairs or intergenerational affected individuals. Sixty-eight percent of referrals were from a pediatric age group. Analysis of families with a single proband was either performed as a trio study (35%; 13 families) or a singleton study (24%; 9 families). In families with multiple affected individuals, usually two affected individuals underwent WES, or in families with proposed X-linked inheritance, two affected male siblings and their mothers who had some similar but less marked features. Greater than 85% of target bases were covered more than 20 times in 70% of patients, with no association between sequencing quality and patient diagnosis (P = 0.16, Welch two-sample t-test).
The diagnostic utility was maximized in trio studies with 46% solved (six families) compared with 22% of singleton referrals (two families). For de novo analysis, trio studies had a marked reduction in the number of filtered variants for review compared with singleton studies. Importantly, filtering using genomic variants from in-house databases, achieved a significant reduction in variants requiring analysis (Figure 1) due both to variants not captured by global population databases and potentially due to laboratory artifacts. Families referred with multiple affected individuals also had a lower diagnostic rate (20% from three families).
Singleton WES led to an altered clinical diagnosis from a metabolic disorder to a syndromic form of ID (patient 11, Table 1). This patient had multisystem disease and developmental delay with a working diagnosis of a mitochondrial disorder, supported by a low complex IV level on respiratory chain enzymology from muscle biopsy. A pathogenic variant in HRAS previously reported in a patient with an attenuated form of Costello syndrome was identified. This clinical phenotype was less recognizable than that observed in other individuals with Costello syndrome and would not have been considered without the WES result.20 No additional pathogenic variants were identified in mitochondrial disease–related genes.
Reanalysis at a 12-month time point boosts diagnoses
Reanalysis of WES data at 12 months in undiagnosed individuals identified new diagnoses in four families, increasing the overall diagnostic rate from 30 to 41%. Two diagnoses were due to interim publications of new disease–gene associations including PPP2R5D linked to ID and overgrowth21 and TANGO2 linked to episodic metabolic crises and neurodegeneration22,23 (patients 8–10, Table 1). One additional large family with autosomal dominant retinitis pigmentosa had a new finding of a splice-site variant in a retinitis pigmentosa–related gene, PRPF31 (c.527+3A>G; patient 19). Although initial pathogenicity classification was of uncertain significance, consistent segregation studies and additional evidence relating to protein expression resulted in reclassification to likely pathogenic, with planned future pathogenicity review. This variant was absent in the initial WES analysis VCF file, but present at 12-month reanalysis. This was attributed to bioinformatics pipeline improvements, specifically a higher variant detection sensitivity with increased batch size for joint calling.
Following reanalysis of unsolved cases, 54% of trio studies were diagnosed (1 additional family), 44% of singletons (2 additional families), and 27% for multiple affected individuals (1 additional family). This represented an additional 15% diagnosed cases (4 of 26 families) at the 12-month time point. The average time to diagnosis from initial presentation was 12 years 8 months (Table 1). Sharing of candidate genes in matchmaker exchange did not result in additional diagnoses.
Cost-effectiveness analysis
The analysis of diagnostic costs and the cost-effectiveness of the counterfactual arms using WES over the traditional pathway utilized cost data from the subcohort of 14 patients with ID. Four patients (29%; 4 of 14 individuals) were diagnosed using the genomic diagnostic pathway with WES at the initial analysis and two additional patients were diagnosed when the variants were reanalyzed at the 12-month time point.
WES was more expensive if available at initial clinical genetics contact compared with the traditional diagnostic pathway (US$6,918 vs. $6,742). However, if WES had been available at the time of initial patient presentation the average cost was lower than the traditional pathway at $6,574 (Table 2; Supplementary Table S2 online). The estimated incremental cost per additional diagnosis with WES available at initial clinical genetics contact would have been $618 (95% CI: –$2,431; $17,439), contrasting with a cost saving of $586 (95%CI: –$3,769; $16,144) per additional diagnosis if WES had been available at the time of initial patient presentation (Table 2). Variant reanalysis costs spread across the entire cohort gave an additional average per patient cost of $134 (95% CI: $81; $184). An incremental cost per additional diagnosis was estimated at a cost saving of $77 (95%CI: –$2,990; $7,334) if WES was available at initial patient presentation.
The cost-effectiveness planes (Figure 2) demonstrate that WES, if available at the time of initial patient presentation with clinical symptoms, would be dominant (i.e., lower cost with a higher number of diagnoses) compared with the traditional diagnostic pathway for 55% of 1,000 bootstrapped simulations. However, if WES was available at initial contact with the clinical genetic service, it would be dominant for only 38% of 1,000 bootstrapped simulations. Thus, WES, if available at the time of initial patient presentation with clinical symptoms, is more likely to be cost saving.
Hypothetical comparison of WGS with WES
Because the diagnostic yield of WGS in ID is likely greater than WES,24,25 we were interested in assessing what additional costs would be required to use WGS over WES to achieve potential additional diagnoses. In the absence of data on WGS diagnostic rates in large cohorts with ID or neurodevelopmental disorders, we modeled WGS costs at various diagnostic rates and decreasing WGS price to identify at what point WGS would become more cost-effective than WES (Figure 3). At current costs for WGS and a 50% diagnostic rate, the incremental cost per additional diagnosis using WGS over WES was estimated at US$11,553 (95% CI: $3,325; $24,650; Figure 3, Supplementary Table S2 online).
Keeping a 50% diagnostic rate, if WGS costs were reduced by 17.9% (95% uncertainty range: 12.8%; 23.6%) there would be no statistically significant difference between the costs of diagnosis using WES and WGS. If the costs of WGS were reduced further, e.g., by 25%, our simulation suggests that there would be cost savings of $4,661 (95% CI: $689; $12,776) per additional diagnosis over WES. (Supplementary Table S2 online details further cost comparisons)
Discussion
We have demonstrated an improved diagnostic success in Mendelian disorders with reanalysis of WES data 12 months after original analysis. On initial analysis, additional diagnoses from WES over the traditional pathway were made in 30%, similar to other studies with unbiased clinical ascertainment,26,27,28,29 with 41% at reanalysis. This equates to new diagnoses at reanalysis in 15% of the unsolved group, which is similar to a previous report,11 highlighting the importance of genomic data reinterrogation. With time new disease genes are published, variants may be reclassified, and expanded phenotypic information becomes available. This, combined with improvements to bioinformatics pipelines and filtering strategies, contributes to an increased diagnostic yield, reinforcing the need for reanalysis.
One of the key challenges of delivering next-generation sequencing results to patients is the interpretation of variant pathogenicity,30 particularly for novel variation. Having both expert clinical genomic staff and an in-house database of curated variants are crucial for the effective identification of pathogenic variants and the removal of laboratory-specific artifacts. The reduced diagnostic yield in singleton compared with trio analyses (22–44 vs. 46–54%) was similar to those reported in other studies31 reinforcing that trio genomic testing is more successful. Variant interpretation for heterozygous calls in a singleton referral remains challenging due to high variant numbers requiring assessment without parental alleles. Although cost saving for reagents, singleton analyses come at an opportunity cost given a three- to fourfold increase in analysis and reporting times (unpublished data).
The importance of phenotype to guide WES analysis
Our experience is that the addition of a multidisciplinary team forum for variant assessment assisted in facilitating more accurate diagnoses and improved patient management. Multidisciplinary team staff included clinical geneticists, genetic counselors, genetic pathologists, scientists, and genomicists who had performed the analyses. Accurate and comprehensive phenotype information was essential for correct and timely diagnosis.
For some referrals, limited phenotypic information inadequately informed the interpretation of filtered WES variants. An example was a singleton patient with phenotypic information limited to vitamin B12 deficiency (patient 2, Table 1). A single variant of interest was identified in CUBN, associated with autosomal recessive vitamin B12 deficiency. A copy-number variant on the second allele was not found. At 12-month reanalysis the WES data search was broadened and a heterozygous pathogenic variant was identified in RPS26 associated with Diamond–Blackfan anemia. Clinical record review identified phenotype information unavailable at referral that Diamond–Blackfan anemia had been raised previously as a diagnosis due to continued lack of response to vitamin B12 treatment and a mildly raised red cell adenosine deaminase enzyme level.
Two brothers with ID and progressive spasticity had a homozygous predicted pathogenic variant in TANGO2 (patients 9 and 10, Table 1) identified. Although an initial assessment of the phenotype did not show the expected metabolic features associated with TANGO2 mutations, further clinical follow-up identified episodic weight-bearing difficulties, ataxia, and hypothyroidism. Consultation with international TANGO2 disease experts confirmed a consistent clinical presentation. This was an important new diagnosis to guide management 10 months later when one of the brothers presented with life-threatening rhabdomyolysis requiring intensive care admission.
Genomic diagnoses enable optimized clinical management
Genomic technologies not only provide diagnoses, but similar to the TANGO2 family, there are now many reports describing alteration to patient management.32,33,34,35 Some diagnoses relate to treatable disorders that can be ameliorated with early therapy,35 surveillance, or specialist assessment. Therefore the early use of genomic technologies in undiagnosed Mendelian disorders should now be considered best practice. Whether there is alteration to management, the impact of having a diagnosis and identification of the molecular etiology should not be underestimated. Not least of these is the impact on reproductive management including preimplantation genetic diagnosis for conditions with high recurrence or the identification of a de novo pathogenic variant.
This is further highlighted by the patient whose alteration of diagnosis from a pure mitochondrial disorder to Costello syndrome shifted the focus from mitochondrial therapy to surveillance and management for cardiac and cancer-related complications known to occur in Costello syndrome. Interestingly, the overlap between RASopathies and mitochondrial disorders has been previously recognized,36 which may be a feature of mitochondrial dysfunction in these patients due primarily to the Ras-MAPK mutation or contribution of a separate mitochondrial genome variant.
Cost savings when WES applied early
This study has demonstrated that the use of WES at the time of initial patient presentation in a cohort of individuals with ID results in incremental cost savings of approximately $586 for each additional diagnosis compared with the traditional diagnostic pathway through the avoidance of unnecessary diagnostic management. Given these diagnostic cost savings are in an Australian context, greater cost savings may be anticipated in countries with fewer constraints on diagnostic testing such as the United States.37 Further downstream savings resulting from early diagnoses through WES would be expected such as the cost of altered clinical management, additional life span, and quality of life gained, as well as the ability of families to make informed reproductive decisions. Some families with high recurrence risks may decide not to have another child or may choose options such as preimplantation genetic diagnosis to avoid recurrence. The high management cost for people with ID38 would not be required for those families making such reproductive choices. While no incidental findings were reported in this study, these could increase costs incurred through additional management, although this may be outweighed by cost savings from earlier diagnoses of treatable disease.
Current studies suggest that while the cost of WGS is higher than WES, it has a higher diagnostic yield.24,39 At this time, whether WGS is a cost-effective option compared with WES depends upon the value of obtaining additional diagnoses, particularly because the initial outlay may be offset by downstream cost savings, including recurrence avoidance of diagnoses with high service provision costs such as ID.37 Further, health providers commonly place increased importance on the upfront costs for diagnostic testing and may consider a small increase in diagnostic rate to be of secondary importance when selecting testing methodologies. Increased WGS diagnostic rates result from more consistent exon coverage and the ability to analyze copy-number variation and the mitochondrial genome.39 A fall in the cost of WGS would improve the cost-effectiveness of this technology, highlighting the importance of prospective cost-effectiveness studies in clinical cohorts applying WGS in undiagnosed WES cohorts.
Two-thirds of this cohort was ascertained from a pediatric age group, enriched for complex medical conditions. Individuals with Mendelian disorders with allelic and phenotypic heterogeneity have been shown to benefit from a genomic approach to diagnosis. The cost savings demonstrated in this study for ID, representing the most common, heterogeneous, and diagnostically difficult group, confirm genomic testing is the diagnostic methodology of choice from an economic perspective. A de novo etiology was identified in the majority of families (11/15, 73%), with clear implications for low recurrence risks.
The average time to diagnosis of 12 years and 8 months reflects the length of the diagnostic odyssey faced by many families who have a child affected by an unknown Mendelian disorder. A diagnosis at presentation through genomic testing, particularly in the context of a confirmed de novo etiology, highlights the importance of adequate clinical resourcing to restore reproductive confidence at a time when family expansion is still practical. The increased diagnostic yield, potential for alteration in diagnosis and disease-specific management, and the cost savings demonstrated in ID also support the need to access and resource genomic testing early in the diagnostic process.
Consideration of the unsolved cohort
Despite WES data reanalysis, 60% of this cohort remained undiagnosed. We expect annual reanalysis of WES data will continue to provide additional diagnoses with an ongoing increased yield of at least 10% per annum if gene discovery continues at the present rate, which may be facilitated by global databases matching candidate genes with new phenotypes. There was a surprisingly low diagnostic yield in families referred with multiple affected individuals, which may be a reflection of the large number of families with male-limited ID (33% of multiple affected families) where etiology could relate to noncoding region variation or other mechanisms missed by WES. The application of WGS may increase diagnoses through improved gene coverage, structural variation calling, and the identification of novel mutation mechanisms in noncoding regions.40 The ongoing refinement of DNA capture systems, including greater read depth to improve mosaic mutation detection, and bioinformatics pipelines will also be of value to increase diagnostic rates.
Conclusion
In a 12-month reanalysis of WES data in Mendelian disorders, diagnostic yields increased from 30 to 41%. The main contributing factors were accurate phenotyping through close clinical and laboratory collaboration, inclusion of an in-house variant database, and, most importantly, the identification of new disease genes.
We recommend that in patients with genetically heterogeneous or suspected but undiagnosed Mendelian disorders, genomic testing should be used as a first-line genetic investigation with early involvement of clinical genetics services. Early diagnoses provide cost savings to health systems and these should be leveraged to increase genomic resources for reanalysis after 12 months or when instigated by referrers.
References
World Health Organization. Genes and human disease. http://www.who.int/genomics/public/geneticdiseases/en/index2.html. Accessed 22 March 2015.
Central Intelligence Agency. The World Factbook. 2016. https://www.cia.gov/library/publications/the-world-factbook/geos/xx.html; Accessed 22 March 2018.
Rabbani B, Tekin M & Mahdieh N. The promise of whole-exome sequencing in medical genetics. J Hum Genet 2014;59:5–15.
Shashi V, McConkie-Rosell A, Rosell B et al. The utility of the traditional medical genetics diagnostic evaluation in the context of next-generation sequencing for undiagnosed genetic disorders. Genet Med 2014;16:176–182.
Bamshad MJ, Ng SB, Bigham AW et al. Exome sequencing as a tool for Mendelian disease gene discovery. Nat Rev Genet 2011;12:745–755.
Gilissen C, Hoischen A, Brunner HG & Veltman JA. Unlocking Mendelian disease using exome sequencing. Genome 2011;11:64.
Rabbani B, Mahdieh N, Hosomichi K, Nakaoka H & Inoue I. Next-generation sequencing: impact of exome sequencing in characterizing Mendelian disorders. J Hum Genet 2012;57:621–632.
Zhang X. Exome sequencing greatly expedites the progressive research of Mendelian diseases. Front Med 2014;8:42–57.
Jacob H, Abrams K, Bick D et al. Genomics in clinical practice: lessons from the front lines. Sci Transl Med 2013;5:194cm5.
Biesecker LG & Green RC. Diagnostic clinical genome and exome sequencing. N Engl J Med 2014;370:2418–2425.
Wenger AM, Guturu H, Bernstein JA & Bejerano G. Systematic reanalysis of clinical exome data yields additional diagnoses: implications for providers. Genet Med 2017;19:209–214.
Kircher M, Witten DM, Jain P, O’Roak BJ, Cooper GM & Shendure J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet 2014;46:310–315.
Paila U, Chapman BA, Kirchner R & Quinlan AR. GEMINI: integrative exploration of genetic variation and genome annotations. PLoS Comput Biol 2013;9:e1003153.
Choi Y & Chan AP. PROVEAN web server: a tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 2015;31:2745–2747.
Seelow D, Schuelke M, Hildebrandt F & Nurnberg P. Homozygosity Mapper—an interactive approach to homozygosity mapping. Nucleic Acids Res 2009;37:W593–W599.
Thorvaldsdottir H, Robinson JT & Mesirov JP. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 2013;14:178–192.
Richards S, Aziz N, Bale S et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015;17:405–423.
NSW Health. NSW Health Renumerations and Conditions. NSW Government Health. 2016. http://www.health.nsw.gov.au/careers/conditions/Pages/default.aspx; Accessed 22 March 2018.
Australian Government Department of Health. Medicare Benefits Schedule Book. April 2014. http://www.mbsonline.gov.au/internet/mbsonline/publishing.nsf/Content/Home; Accessed 22 March 2018.
Denayer E, Parret A, Chmara M et al. Mutation analysis in Costello syndrome: functional and structural characterization of the HRAS p.Lys117Arg mutation. Hum Mutat 2008;29:232–239.
Loveday C, Tatton-Brown K, Clarke M et al. Mutations in the PP2A regulatory subunit B family genes PPP2R5BPPP2R5C and PPP2R5D cause human overgrowth. Hum Mol Genet 2015;24:4775–4779.
Kremer LS, Distelmaier F, Alhaddad B et al. Bi-allelic truncating mutations in TANGO2 cause infancy-onset recurrent metabolic crises with encephalocardiomyopathy. Am J Hum Genet 2016;98:358–362.
Lalani SR, Liu P, Rosenfeld JA et al. Recurrent muscle weakness with rhabdomyolysis, metabolic crises, and cardiac arrhythmia due to bi-allelic TANGO2 mutations. Am J Hum Genet 2016;98:347–357.
Gilissen C, Hehir-Kwa JY, Thung DT et al. Genome sequencing identifies major causes of severe intellectual disability. Nature 2014;511:344–347.
Soden SE, Saunders CJ, Willig LK et al. Effectiveness of exome and genome sequencing guided by acuity of illness for diagnosis of neurodevelopmental disorders. Sci Transl Med 2014;6:265ra168.
Yang Y, Muzny DM, Reid JG et al. Clinical whole-exome sequencing for the diagnosis of Mendelian disorders. N Engl J Med 2013;369:1502–1511.
Farwell KD, Shahmirzadi L, El-Khechen D et al. Enhanced utility of family-centered diagnostic exome sequencing with inheritance model–based analysis: results from 500 unselected families with undiagnosed genetic conditions. Genet Med 2015;17:578–586.
Lazaridis KN, Schahl KA, Cousin MA et al. Outcome of whole exome sequencing for diagnostic odyssey cases of an individualized medicine clinic. Mayo Clin Proc 2016;91:297–307.
Retterer K, Juusola J, Cho MT et al. Clinical application of whole-exome sequencing across clinical indications. Genet Med 2016;18:696–704.
Kassahn KS, Scott HS & Caramins MC. Integrating massively parallel sequencing into diagnostic workflows and managing the annotation and clinical interpretation challenge. Hum Mutat 2014;35:413–423.
Lee H, Deignan JL, Dorrani N et al. Clinical exome sequencing for genetic identification of rare Mendelian disorders. JAMA 2014;312:1880.
Choi M, Scholl UI, Ji W et al. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc Natl Acad Sci USA 2009;106:19096–19101.
Majewski J, Wang Z, Lopez I et al. A new ocular phenotype associated with an unexpected but known systemic disorder and mutation: novel use of genomic diagnostics and exome sequencing. J Med Genet 2011;48:593–596.
Zhan Z, Liao X, Du J et al. Exome sequencing released a case of X-linked adrenoleukodystrophy mimicking recessive hereditary spastic paraplegia. Eur J Med Genet 2013;56:375–378.
Sawyer SL, Hartley T, Dyment DA et al. Utility of whole-exome sequencing for those near the end of the diagnostic odyssey: time to address gaps in care: whole-exome sequencing for rare disease diagnosis. Clin Genet 2016;89:275–284.
Kleefstra T, Wortmann S, Rodenburg R et al. Mitochondrial dysfunction and organic aciduria in five patients carrying mutations in the Ras-MAPK pathway. Eur J Hum Genet 2011;19:138–144.
Stark Z, Schofield D, Alam K et al. Prospective comparison of the cost-effectiveness of clinical whole-exome sequencing with that of usual care overwhelmingly supports early use and reimbursement. Genet Med 2017;19:867–874.
Doran CM, Einfeld SL, Madden RH et al. How much does intellectual disability really cost? First estimates for Australia. J Intellect Dev Disabil 2012;37:42–49.
Belkadi A, Bolze A, Itan Y et al. Whole-genome sequencing is more powerful than whole-exome sequencing for detecting exome variants. Proc Natl Acad Sci USA 2015;112:5473–5478.
Eldomery MK, Coban-Akdemir Z, Harel T et al. Lessons learned from additional research analyses of unsolved clinical exome cases. Genome Med 2017;9:26.
Acknowledgments
Whole-exome sequencing was funded through a donation by the Kinghorn Foundation to the Garvan Institute of Medical Research. L.J.E. is supported through an Australian Government Research Training Program Scholarship. M.J.C. is supported by Cancer Institute NSW (13/ECF/1-46). T.R. was supported by the Australian National Health and Medical Research Council (APP1117394 Transforming the Diagnosis and Management of Severe Neurocognitive Disorders through Genomics; APP1045465 Gene Identification in Familial Orofacial Clefts by Next Generation Sequencing of Exomes and p63 regulatory elements and an International Biomedical Fellowship 512123 Identification of genes for intellectual disability and brain malformations). We thank the patients, their families, and physicians for their participation in this study. We also thank Jiang Tao, Paula Morris, and Kerith-Rae Dias from the Kinghorn Centre for Clinical Genomics for sequencing these patients.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
L.J.E. and M.E.D. are employees of Genome.One, a whole-owned subsidiary of the Garvan Institute for Medical Research. M.F.B., E.P.K, E.L., and C.W. are employees of Randwick Genetics, NSW Health Pathology East. The other authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Ewans, L.J., Schofield, D., Shrestha, R. et al. Whole-exome sequencing reanalysis at 12 months boosts diagnosis and is cost-effective when applied early in Mendelian disorders. Genet Med 20, 1564–1574 (2018). https://doi.org/10.1038/gim.2018.39
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2018.39
Keywords
This article is cited by
-
Rapid genomic sequencing for genetic disease diagnosis and therapy in intensive care units: a review
npj Genomic Medicine (2024)
-
Systematic reanalysis of genomic data by diagnostic laboratories: a scoping review of ethical, economic, legal and (psycho)social implications
European Journal of Human Genetics (2024)
-
Variants in mitochondrial disease genes are common causes of inherited peripheral neuropathies
Journal of Neurology (2024)
-
Semiautomated approach focused on new genomic information results in time and effort-efficient reannotation of negative exome data
Human Genetics (2024)
-
Next-generation sequencing and comprehensive data reassessment in 263 adult patients with neuromuscular disorders: insights into the gray zone of molecular diagnoses
Journal of Neurology (2024)