Abstract
Purpose
The purpose of the study was to implement and prospectively evaluate the outcomes of a rapid genomic diagnosis program at two pediatric tertiary centers.
Methods
Rapid singleton whole-exome sequencing (rWES) was performed in acutely unwell pediatric patients with suspected monogenic disorders. Laboratory and clinical barriers to implementation were addressed through continuous multidisciplinary review of process parameters. Diagnostic and clinical utility and cost-effectiveness of rWES were assessed.
Results
Of 40 enrolled patients, 21 (52.5%) received a diagnosis, with median time to report of 16 days (range 9–109 days). A result was provided during the first hospital admission in 28 of 36 inpatients (78%). Clinical management changed in 12 of the 21 diagnosed patients (57%), including the provision of lifesaving treatment, avoidance of invasive biopsies, and palliative care guidance. The cost per diagnosis was AU$13,388 (US$10,453). Additional cost savings from avoidance of planned tests and procedures and reduced length of stay are estimated to be around AU$543,178 (US$424,101). The clear relative advantage of rWES, joint clinical and laboratory leadership, and the creation of a multidisciplinary “rapid team” were key to successful implementation.
Conclusion
Rapid genomic testing in acute pediatrics is not only feasible but also cost-effective, and has high diagnostic and clinical utility. It requires a whole-of-system approach for successful implementation.
Similar content being viewed by others

Introduction
Genetic conditions are the leading cause of death in infants and children in the developed world. Rare genetic disease diagnosis has been transformed by next-generation sequencing (NGS) technologies, including whole-exome sequencing (WES) and whole-genome sequencing (WGS), which are increasingly available as clinical tests. Despite common perceptions that rare genetic conditions are untreatable, early genomic diagnosis provides prognostic information, influences management decisions, and alters clinical outcomes.1 Studies limited to infants, primarily recruited from neonatal and pediatric intensive-care units (NICUs and PICUs),1,2,3,4,5,6 have consistently reported high diagnostic yields (30–73%), and NICUs have been highlighted as priority implementation areas for genomic medicine.7,8 Turnaround times for results appear to impact on clinical utility: results were clinically useful in 32.6% of diagnosed patients when WES was provided, with standard turnaround times (median 136 days),1 and clinically useful in 65–71% of diagnosed patients in rapid WGS (rWGS) and rapid WES (rWES) studies, where results were provided with median turnaround times of 12–23 days.3,5,6
The consistent delivery of timely and accurate results in the acute pediatric setting presents challenges for both clinical and laboratory services. The barriers to implementing genomic medicine in clinical practice are well documented,9 but there are few studies describing practical implementation experience.10,11,12 Reports of implementation experience in rapid genomics have focused primarily on the technical aspects of delivering results in less than 2 days as a proof of principle.7,13 Furthermore, while several studies have used modeling in retrospectively and prospectively ascertained cohorts to argue that genomic sequencing can reduce investigation costs in rare-disease diagnosis,14,15,16,17,18 an actual impact of standard or rapid genomic sequencing on health-care costs has not been demonstrated.
We developed and implemented an rWES diagnosis program at two tertiary pediatric hospitals using an accredited clinical test, with the aim of delivering rWES results in less than 21 days from study enrollment. We sought to prospectively evaluate patient-focused outcomes, assessing diagnostic and clinical utility and cost-effectiveness, as well as to conduct a formative process evaluation to identify and implement clinical and laboratory practices required to achieve rapid diagnostics.
Materials and methods
Study design and participants
Participants were recruited during clinical care by the clinical genetics services at the two tertiary pediatric hospitals in Melbourne, Australia: the Royal Children’s Hospital (RCH) and Monash Children’s Hospital (MCH), between April 2016 and September 2017. RCH has 250 inpatient beds, a 45-bed NICU, and a 28-bed PICU, whereas MCH has 230 inpatient beds and a 64-bed NICU. The patient-selection criteria are summarized in Table 1.
Potential participants were discussed by a panel of study investigators (Z.S., T.Y.T., A.Y., and S.M.W.) to determine eligibility, with a minimum of two clinical geneticists agreeing rWES was appropriate required for a patient to be enrolled. A core data set, including Human Phenotype Ontology (HPO) terms, was collected at enrollment and recorded using PhenoTips.19 All diagnostic investigations, including those planned, were collected from referring clinicians and from the medical records.
A phenotype-driven list of candidate genes for prioritized analysis was nominated for each participant by their clinician, as previously described.20 Information regarding the impact of rWES diagnosis on patient management was collected from the clinicians involved in each participant’s care.
Exome sequencing, data analysis and interpretation
Singleton rWES was performed at our National Association of Testing Authorities-accredited laboratory (VCGS Pathology), using SureSelect CRE exome capture (Agilent, Santa Clara, CA) on either HiSeq 4000 or NextSeq 500 sequencers (Illumina, San Diego, CA). The mean coverage obtained was 100 ×. Variants were characterized using the Melbourne Genomics Health Alliance shared bioinformatics pipeline, Cpipe21 version 2.0.1. Variant filtering and prioritization were performed as previously described.1,20 Only variants relevant to a participant’s phenotype were assessed with regard to pathogenicity. The data for each patient were reviewed by the referring clinical geneticist and a study clinical geneticist (Z.S.) in addition to the curating medical genomic scientist in order to prioritize variants for assessment. Criteria for classification were based on the principles outlined in the American College of Medical Genetics and Genomics standards for interpretation of sequence variants.22 Variant classifications were reviewed in a multidisciplinary team (MDT) meeting attended by clinical geneticists, other medical subspecialists, genetic counselors, medical genomic scientists and bioinformaticians. The quorum for an MDT meeting was the curating medical genomic scientist, one senior medical genomic scientist, and one clinical geneticist.
Cost-effectiveness
Cost data were collected as previously described.15,18 A clinical geneticist at each recruitment site (Z.S. and A.Y.) extracted all investigations, procedures, and assessments carried out for diagnostic purposes from the medical records. Costs of investigations and patient encounters were obtained from the hospital, state government, Medical Benefits Schedule, and from testing laboratories. The costs of individual investigations are presented in Supplementary Table S1 online. Average diagnostic costs per patient and cost per diagnosis were calculated. Bootstrapping using 500 replications was performed to estimate confidence intervals. The baseline cost of a day in an intensive-care unit (ICU) for a ventilated patient (AU$4,500) was used in calculations regarding savings arising from reduced length of stay following rapid diagnosis.
Costs were compared with those ascertained in our previously published study of an infant cohort undergoing the usual investigations and WES with standard turnaround times (sWES) in parallel.15 Specifically, we compared against the cost per diagnosis of usual care (including single-gene and multigene panel testing) as already published, and in addition calculated the cost per diagnosis of usual diagnostic care with sWES replacing all sequencing tests in that cohort.
Process evaluation
We conducted a formative evaluation to identify clinical and laboratory barriers to implementation. Information was collected on the following clinical service parameters as measured from the day of patient admission to tertiary pediatric hospital: time to clinical genetics referral, time to clinical genetics assessment, time to proposing patient for rWES, time to approval by patient approval panel, time to consent. These were collectively considered to be “time to ascertainment.” Information was also collected on laboratory service parameters as measured from the day on which sample and consent were made available to the laboratory: time to initiate sequencing, time to complete bioinformatics analysis, time to MDT review, and time to issue report. These were collectively considered to be “time to report.” Time to ascertainment and time to report were together taken as the “time to diagnosis.” Times are reported as calendar days.
The clinical and laboratory leads of the project (Z.S. and S.L.) reviewed the clinical and laboratory-service performance parameters following each case to identify delays and opportunities for improvement. Quarterly MDT meetings between the clinical geneticists, genetic counselors, specimen reception staff, and laboratory genomic scientists involved in the project were used to present patient outcomes, review service performance parameters, and discuss solutions to implementation barriers. Barriers and solutions were mapped to the relevant constructs of the Consolidated Framework for Implementation Research (CFIR). The CFIR is a conceptual framework that draws together multiple existing implementation models and enables systematic assessment of implementation barriers and facilitators across 39 constructs arranged in five domains (inner setting, outer setting, intervention characteristics, individuals’ characteristics, and process).23 The CFIR has been adopted by the Implementing Genomics in Practice Network to promote the comprehensive and shareable evaluation of implementation experiences in clinical genomics.24
Statistical analysis
Descriptive statistics were used to describe the demographic features, time course, and outcomes of the rWES and sWES cohorts. Dot plots were used to visualize the distribution of variables to determine whether to use parametric or nonparametric measures. Mean and standard deviation were determined for normally distributed variables. For non-normally distributed variables median and interquartile ranges were calculated. Power calculations were performed to compute power for the given sample size and values of the control-group and experimental-group proportions. Differences were investigated using two-sided t-tests for normally distributed variables, Wilcoxon-rank sum test for non-normally distributed variables, or chi-squared tests for categorical variables. P values are reported for results where P < 0.05. All analyses were performed using Stata 11.0 (Stata, College Station, TX).
Ethics approval
The study was part of the Melbourne Genomics Health Alliance project (http://www.melbournegenomics.org.au) and received Human Research Ethics Committee approval (13/MH/326). Parents provided written informed consent after genetic counseling regarding the testing.
Results
Participant demographics and indications for testing
A total of 46 patients were considered for rWES, 40 of whom were enrolled in the study. One patient was excluded owing to a prior completed sequencing-based test, and another was enrolled in an alternative genomic project. One patient was judged to have a low likelihood of a monogenic condition, and another did not meet complexity criteria. All but one of the families who were offered rWES accepted testing, although during pretest counseling several families expressed unwillingness to accept that their child could have a genetic condition (G.R.B., A.J., Y.P., J.E., personal observations). The one family who declined testing recontacted our team within a week to request rWES. Two approved patients died before sequencing started. Their samples were excluded from the rWES study and were processed using standard turnaround times.
The characteristics of enrolled participants and indications for testing are summarized in Table 2, and are compared with data from our previously published sWES infant cohort.1 Thirty-three patients (83% of the cohort) were recruited from RCH and the remainder from MCH. The median age at enrollment was 28 days (range 3 days to 4 years).
Diagnostic performance of rWES
rWES resulted in a molecular diagnosis of 23 conditions in 21 patients, with a diagnostic rate of 52.5% (Supplementary Table S2). All pathogenic and likely pathogenic variants identified in this study have been deposited in ClinVar with supporting evidence for pathogenicity (https://www.ncbi.nlm.nih.gov/clinvar/; submission ID: SUB3209813). Novel missense variants concordant with phenotype were identified in an additional three patients but currently do not meet American College of Medical Genetics and Genomics pathogenicity criteria (Supplementary Table S3).
rWES outperformed biochemical tests in two patients. It returned a diagnostic result of AADC deficiency (MIM 608643) in one patient in 14 days, enabling treatment to be started, while the cerebrospinal fluid neurotransmitter result sent to an interstate laboratory became available after 10 weeks. In another patient, a diagnosis of ECHS1 deficiency (MIM 616277) was made in 15 days, while the typical diagnostic pattern on urine metabolic screening was obscured in the acute setting by ketosis and acidosis and became apparent only once the patient was stable and the rWES diagnosis prompted retesting.
Impact on clinical management
rWES led to change in management in 12 diagnosed participants (Supplementary Table S2). This included the provision of lifesaving treatment in one patient diagnosed with riboflavin transporter defect, who at the time of diagnosis was ventilation-dependent via a tracheostomy. rWES was initiated on day 46 of admission to PICU, with a result available after 21 days. The patient was treated with riboflavin and discharged from the hospital after 179 days.
Nine patients (23%) died in the course of the study: three from the undiagnosed group and six from the diagnosed group. Five died before results were available and four after results were available. The result from rWES was instrumental in management being redirected toward palliation in the acute-care setting in two patients. One baby was relinquished for adoption, with the rWES diagnosis of a syndromic condition contributing to the parental decision.
In six patients, uninformative rWES, sometimes combined with improvement in clinical course, contributed to the diagnosis of a non-monogenic condition. Two of these patients avoided invasive tissue biopsies (lung and liver), which were scheduled but postponed owing to the availability of rWES (Supplementary Table S3).
Cost-effectiveness
The cost of providing rWES was AU$3,959 per patient. The total cost of diagnostic assessments and investigations, including rWES, in this cohort, was AU$281,143 and the cost per diagnosed patient was AU$13,388 (Table 3).
Three patients avoided invasive tissue biopsies as a result of rWES being available. The combined cost of these biopsies is estimated to be AU$2,428.43. In another patient, orders for Prader-Willi methylation studies (AU$250) and nerve conduction studies (AU$500) were canceled following an rWES diagnosis. The patient treated for riboflavin transporter defect is estimated to have avoided an additional 115 days in PICU because the rWES result was provided in 21 days as opposed to the 136 days it would have taken for an sWES result. Based on a cost of AU$4,500 per day for a ventilated patient in a PICU, the total cost savings for this patient can be estimated to be around AU$517, 500. For one of the patients, whose care was redirected toward palliation, the rWES result became available at the same time as a diagnostic anatomical pathology result, and therefore the rWES result is not considered to have altered the length of stay in an ICU. However, in the second patient, the diagnosis of a neurodegenerative condition prompted earlier redirection of care to palliation. Consensus medical opinion from the treating team is that the length of stay in the ICU was reduced by at least 5 days (AU$22,500). The total cost savings in this cohort are therefore estimated to be around AU$543,178.
Barriers to rWES implementation and solutions implemented
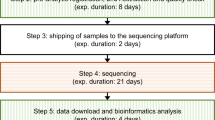
A number of clinical and laboratory barriers to rWES implementation were identified throughout the study. These, and the solutions implemented, are presented in Table 4 and temporally illustrated in relation to the test turnaround times in Figure 1. Key strategies included utilizing the relative advantage of rWES to engage team members in changing work practices; appointing formal implementation leaders to clearly define the project goals and identify problems and solutions using processes of continuous reflection and evaluation, resulting in the creation of a dedicated network (“rapid team”) with its own dedicated communication strategy and standard operating procedures. These strategies delivered efficiency gains without an increase in test cost. Costs were increased through the allocation of additional resources to obtain further reductions in time to report—in particular, the dynamic use of HiSeq and NextSeq instruments to enable more flexible batching of samples, with 21 of 40 samples (52%) sequenced on NextSeq. Several parts of the program benefited from increased flexibility on the part of clinical and laboratory staff to deal with single urgent cases at short notice, as well as willingness to change working patterns, for example, by increasing the deployment of genetic counselors in the inpatient setting. Successful implementation was also dependent on staff being able to adapt established working patterns, for example, by defining quorum requirements for patient approval and multidisciplinary variant review meetings to enable rapid single-case processing while maintaining quality. Demonstrating the relative advantage of rWES in terms of clinical utility and cost-effectiveness to referrers and clinical geneticists was an important factor in achieving reductions in time to ascertainment.
A timeline for the study is provided on the X-axis, demonstrating the increase in throughput over time, and the principal interventions implemented to reduce time to report in response to continuous case review are depicted above. MDT, multidisciplinary team meeting; rWES, rapid singleton whole-exome sequencing.
Impact on clinician time
Referrals for inpatient genetics consultation at our principal recruitment center (RCH) have nearly doubled during the course of the study, from 150 per year in 2014 to 273 in 2016; a further increase is observed in 2017, with 150 inpatients referred in the first 6 months alone. The average number of genetic counselor–inpatient contacts per rWES family was 1.8 (range 0–4 contacts), and overall duration 68 min (range 15–150 min). The average number of clinical geneticist–inpatient contacts per rWES family was 4 (range 1–6 contacts), with an estimated 30 min spent with families who did not receive a diagnosis and 90 min with those who received a diagnosis.
Discussion
We have demonstrated that rapid genomic-sequencing results have the power to deliver actual reductions in cost per diagnosis by altering the diagnostic pathway. At AU$13,388 per diagnosis, a diagnosis by rWES costs half that of diagnosis by usual care in our previously reported infant cohort (AU$27,050.36).15 It is also comparable to the cost per diagnosis of sWES (AU$10,843.60) in that cohort, despite the higher cost of rWES and the selection of high-complexity patients. The increased cost of rWES was offset by reductions in the number of outpatient subspecialist assessments, imaging and anatomical pathology tests for diagnostic purposes..
The majority of results were delivered during the child’s initial tertiary hospital presentation, affording the opportunity to influence early management in 57% of diagnosed patients, delivering more personalized care. Given the high cost of ICU admission, rapid genomic sequencing results have the power to produce large savings by altering lengths of stay, and the overall cost savings from rWES, AU$543,178, far exceeded the cost of providing rWES in the entire cohort (AU$157,960). It is likely that further reductions in both time to ascertainment and time to result will deliver additional cost savings.
Other groups have highlighted the value of rapid genomic results in redirecting care toward palliation.5 However, we found that the majority of such decisions in our cohort were made on clinical grounds, often before a result was available. Australia has some of the highest reported rates of withdrawal and withholding of medical treatment in the pediatric setting.25 Australian ICU clinicians may be accustomed to counseling families about palliation in the absence of a secure diagnosis and hence generally did not defer palliation discussions until rWES results were available.
Many of the barriers to implementation of genomic medicine have been well described; they include insufficient sequencing, bioinformatics, data interpretation, and workforce capacity and capability, as well as lack of evidence for effectiveness and lack of funding.9 Rapid genomic sequencing not only magnifies existing bottlenecks and barriers but poses additional laboratory and clinical challenges through the introduction of time pressure in systems that are still evolving and lack absorptive capacity.26 Using an implementation framework, CFIR, allowed the systematic and comprehensive identification of clinical and laboratory barriers throughout the study. This practical approach provided an opportunity to identify and implement a broad range of real-time solutions and strategies to optimize effectiveness of implementation in a timely manner. Overcoming obstacles to deliver an rWES service in a clinical setting required change in the work patterns of all health professionals involved, including ICU clinicians, clinical geneticists, clinical genetics trainees, genetic counselors, specimen reception staff, bioinformaticians, and laboratory genomic scientists. This was facilitated by the formation of a joint clinical and laboratory implementation leadership. The clear relative advantage of rWES in terms of clinical utility and cost-effectiveness proved to be a powerful engagement tool with other subspecialist groups. Finally, successful implementation in our setting required the creation of a defined multidisciplinary network (rapid team), which developed standard operating procedures, established dedicated communication channels, and engaged in continuous reflection and evaluation, creating a virtuous cycle through precision medicine, implementation science, and a learning health-care system.27
From a laboratory perspective, our experience demonstrates that 10-fold improvements in WES turnaround times can be achieved through efficiency gains, and supplementary use of smaller-capacity sequencers. The use of such sequencers to alleviate batching issues has been highlighted by other groups attempting rapid genomics5 and largely accounted for doubling the per-sample cost to AU$3,959 in our setting. The median time to result of 16 days is comparable to that achieved by other groups utilizing singleton5 or trio WGS,6 or trio WES3 of 12–23 days. Decreasing time to result beyond the best (9 days) achieved here would require instituting 24-hour, 7-days-per-week sample processing and analysis, and true “on-demand” sequencing independent of batching considerations, as well as deploying specialized infrastructure, such as dedicated rapid analysis bioinformatics pipelines.13 While our study utilized singleton WES to contain costs, we consider trio sequencing to be necessary for rapid turnaround results, to allow the immediate and final classification of a number of recessive and de novo variants and potentially increase diagnostic yield.28 Similarly, although we used sequential single-nucleotide polymorphism microarray and rWES to contain costs, there are patients in whom the pretest probability of chromosomal abnormality is low and testing can proceed in parallel. Ultimately, WGS may be the preferred testing modality for rapid genomics owing to its ability to provide both copy-number and single-nucleotide variation data, despite the difference in cost, which is currently considerable.
From a clinical perspective, timely diagnosis relies not only on the ability to process rWES samples in the laboratory but also on timely and appropriate patient selection and test initiation. Clinical genetics is typically perceived as a nonacute outpatient medical specialty. Although most clinical genetics services provide inpatient and out-of-hours consultation, there has traditionally been little expectation for urgent clinical genetics opinion or test results. We have prioritized access to genomic testing for patients early in the diagnostic trajectory as a first-tier sequencing test since 2014,1,18 and specifically prioritized inpatients in ICUs. We have observed a doubling in the number of referrals to our inpatient service, particularly since the introduction of rWES, and an increase in the number of patient/family contacts by clinical geneticists and genetic counselors occurring in the inpatient setting. As test turnaround times continue to decrease, we can anticipate greater demand for out-of-hours genetic services. Resourcing the increase in demand for inpatient clinical genetics consultations and upskilling ICU clinicians in appropriate test initiation and consent would both be important ingredients for the success of rapid genomic-sequencing programs.
From a genetic counseling perspective, the acute setting reduces or eliminates counselors’ opportunity for preparatory information-gathering regarding family medical and psychosocial history. Families are frequently highly distressed by their child being acutely unwell, while being simultaneously faced with complex decisions regarding diagnostic investigations, immediate treatment, long-term management, and/or palliation. Concerns have also been raised about the potential for genomic testing in the newborn period to alter family dynamics.29 Traditionally, we have deferred complex genetic testing in this patient group to the ambulatory setting following discharge or death, which may have provided parents valuable processing time. Despite these concerns, we found a very high rate of acceptance of genomic testing by families. Factors influencing uptake of genomic testing are likely to be complex and would benefit from further study, particularly across different health-care settings and implementation models.
This study is limited by its small size. As the numbers of patients requiring rapid testing are low even in major tertiary pediatric centers, full evaluation of effectiveness and of implementation approaches would be facilitated by multicenter studies. In terms of health economic evaluation, the greatest difficulty is in accurately capturing the impact of rWES on clinical trajectory and length of stay; cohorts tend to be heterogeneous and include many patients with ultrarare conditions, with poorly understood natural history. Evaluation of rapid genomic testing would be enhanced by collection of additional parameters such as family experience, empowerment, and decisional regret.
Rapid genomic testing challenges traditional clinical and laboratory genetics service models and requires a whole-of-system approach for successful implementation. Our study extends the body of evidence indicating that the provision of rapid genomic results in acute pediatric care is feasible and has high diagnostic and clinical utility. Importantly, we demonstrate cost savings from a clinical rapid WES program. Barriers to implementation are likely to be common across health-care systems and we identify solutions for these that were successful in our setting. Developing the capability to reliably produce rapid genomic results for selected patients has applications outside of acute pediatric care, notably in the oncology and prenatal settings, and requires substantial investment to further optimize test performance and equity of access.
References
Stark Z, Tan TY, Chong B et al. A prospective evaluation of whole-exome sequencing as a first-tier molecular test in infants with suspected monogenic disorders. Genet Med 2016;18:1090–1096.
Daoud H, Luco SM, Li R et al. Next-generation sequencing for diagnosis of rare diseases in the neonatal intensive care unit. CMAJ 2016;188:E254–260.
Meng L, Pammi M, Saronwala A et al. Use of exome sequencing for infants in intensive care units: ascertainment of severe single-gene disorders and effect on medical management. JAMA Pediatr 2017: e173438.
Soden SE, Saunders CJ, Willig LK et al. Effectiveness of exome and genome sequencing guided by acuity of illness for diagnosis of neurodevelopmental disorders. Sci Transl Med 2014;6:265ra168.
van Diemen CC, Kerstjens-Frederikse WS, Bergman KA et al. Rapid targeted genomics in critically ill newborns. Pediatrics 2017;140:e20162854.
Willig LK, Petrikin JE, Smith LD et al. Whole-genome sequencing for identification of Mendelian disorders in critically ill infants: a retrospective analysis of diagnostic and clinical findings. Lancet Resp Med 2015;3:377–387.
Kingsmore SF, Petrikin J, Willig LK, Guest E. Emergency medical genomes: a breakthrough application of precision medicine. Genome Med 2015;7:82.
Petrikin JE, Willig LK, Smith LD, Kingsmore SF. Rapid whole genome sequencing and precision neonatology. Semin Perinatol 2015;39:623–631.
Manolio TA, Abramowicz M, Al-Mulla F et al. Global implementation of genomic medicine: we are not alone. Sci Transl Med 2015;7:290ps13.
Gaff CL, M Winship I, Forrest SM et al. Preparing for genomic medicine: a real world demonstration of health system change. NPJ Genom Med 2017;2:31.
Manolio TA, Chisholm RL, Ozenberger B et al. Implementing genomic medicine in the clinic: the future is here. Genet Med 2013;15:258–267.
Roberts MC, Kennedy AE, Chambers DA, Khoury MJ. The current state of implementation science in genomic medicine: opportunities for improvement. Genet Med 2017;19:858–863.
Miller NA, Farrow EG, Gibson M et al. A 26-hour system of highly sensitive whole genome sequencing for emergency management of genetic diseases. Genome Med 2015;7:100.
Monroe GR, Frederix GW, Savelberg SM et al. Effectiveness of whole-exome sequencing and costs of the traditional diagnostic trajectory in children with intellectual disability. Genet Med 2016;18:949–56.
Stark Z, Schofield D, Alam K et al. Prospective comparison of the cost-effectiveness of clinical whole-exome sequencing with that of usual care overwhelmingly supports early use and reimbursement. Genet Med 2017;19:867–874.
Walsh M, Bell KM, Chong B et al. Diagnostic and cost utility of whole exome sequencing in peripheral neuropathy. Ann Clin Transl Neurol 2017;4:318–325.
Vissers LE, van Nimwegen KJ, Schieving JH et al. A clinical utility study of exome sequencing versus conventional genetic testing in pediatric neurology. Genet Med 2017;19:1055–1063.
Tan TY, Dillon OJ, Stark Z et al. Diagnostic impact and cost-effectiveness of whole-exome sequencing for ambulant children with suspected monogenic conditions. JAMA Pediatr 2017;171:855–862.
Girdea M, Dumitriu S, Fiume M et al. PhenoTips: patient phenotyping software for clinical and research use. Hum Mutat 2013;34:1057–1065.
Stark Z, Dashnow H, Lunke S et al. A clinically driven variant prioritization framework outperforms purely computational approaches for the diagnostic analysis of singleton WES data. Eur J Hum Genet 2017;25:1268–1272.
Sadedin SP, Dashnow H, James PA et al. Cpipe: a shared variant detection pipeline designed for diagnostic settings. Genome Med 2015;7:68.
Richards S, Aziz N, Bale S et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015;17:405–424.
Damschroder LJ, Aron DC, Keith RE, Kirsh SR, Alexander JA, Lowery JC. Fostering implementation of health services research findings into practice: a consolidated framework for advancing implementation science. Implement Sci 2009;4:50.
Orlando LA, Sperber NR, Voils C et al. Developing a common framework for evaluating the implementation of genomic medicine interventions in clinical care: the IGNITE Network’s Common Measures Working Group. Genet Med; e-pub ahead of print 14 September 2017.
Stark Z, Hynson J, Forrester M. Discussing withholding and withdrawing of life-sustaining medical treatment in paediatric inpatients: audit of current practice. J Paediatr Child Health 2008;44:399–403.
Cohen WM, Levinthal DA. Absorptive-capacity—a new perspective on learning and innovation. Admin Sci Quart 1990;35:128–152.
Chambers DA, Feero WG, Khoury MJ. Convergence of implementation science, precision medicine, and the learning health care system: a new model for biomedical research. JAMA 2016;315:1941–1942.
Farwell KD, Shahmirzadi L, El-Khechen D et al. Enhanced utility of family-centered diagnostic exome sequencing with inheritance model-based analysis: results from 500 unselected families with undiagnosed genetic conditions. Genet Med 2015;17:578–586.
Frankel LA, Pereira S, McGuire AL. Potential psychosocial risks of sequencing newborns. Pediatrics 2016;137(suppl 1):S24–29.
Acknowledgments
The study was funded by the founding organizations of the Melbourne Genomics Health Alliance and the State Government of Victoria (Department of Health and Human Services). The involvement of the Australian Genome Research Facility was supported by sponsorship from Bioplatforms Australia and the NCRIS program. We thank the patients and families for participating in this study. We are grateful to Ravi Savarirayan, David Amor, Martin Delatycki, Lilian Downie, Emma Krzesinski, Amanda Moody, David Tingay, Kevin Wheeler, Anastasia Pellicano, Leah Hickey, Ruth Armstrong, Trisha Prentice, and Julia Gunn for referring patients to the study; Amber Boys for cytogenetics support; Michael Tamayo and Audrey Chong for sample processing support; Chris Ieng for bioinformatics support; and Hamidul Huque for statistical support.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Stark, Z., Lunke, S., Brett, G.R. et al. Meeting the challenges of implementing rapid genomic testing in acute pediatric care. Genet Med 20, 1554–1563 (2018). https://doi.org/10.1038/gim.2018.37
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2018.37
Keywords
This article is cited by
-
Role of next generation sequencing in diagnosis and management of critically ill children with suspected monogenic disorder
European Journal of Human Genetics (2024)
-
Rapid genomic sequencing for genetic disease diagnosis and therapy in intensive care units: a review
npj Genomic Medicine (2024)
-
Rapid genomic testing in critically ill patients with genetic conditions: position statement by the Human Genetics Society of Australasia
European Journal of Human Genetics (2024)
-
What is the power of a genomic multidisciplinary team approach? A systematic review of implementation and sustainability
European Journal of Human Genetics (2024)
-
A model to implement genomic medicine in the neonatal intensive care unit
Journal of Perinatology (2023)