Abstract
Purpose
We systematically reviewed the published literature on test failure rates for the sequencing of cell-free DNA (cfDNA) in maternal plasma to identify Down syndrome.
Methods
We searched peer-reviewed English publications with diagnostic results on all pregnancies that provided test failure rates. Data on the odds of failure in Down syndrome and euploid pregnancies and the impact of repeat testing were extracted. Random-effects modeling was then used to identify moderators that could explain variability.
Results
Thirty articles satisfied the inclusion criteria for overall failure rates. Study location (Western and Asian with initial testing, and Western with repeat testing) were significant moderators with failure rates of 3.3, 0.6, and 1.2%, respectively (P = 0.001). The odds ratio for Down syndrome in successful versus failed tests was 0.98 (95% confidence interval: 0.62–1.55, I2 = 0%). Repeat testing from 14 large clinical cohort studies found that 83% (range: 52–100%) of failures were repeated, with 79% (range: 46–97%) being successful.
Conclusion
Lower failure rates in Asian studies may be related to not routinely measuring the fetal fraction and to fewer obese women. Repeat cfDNA testing is effective in providing reliable results after initial failures. Protocols for primary cfDNA screening should focus on Down syndrome, with less common and more structurally abnormal trisomy 18 and 13 pregnancies treated as adjuncts.
Similar content being viewed by others
Introduction
The aim of this study was to determine the overall cell-free DNA (cfDNA) test failure rates for plasma samples provided by women undergoing cfDNA as the primary prenatal screening test for Down syndrome in a general pregnancy population. First, we examined test failure rates among women with subsequently diagnosed euploid and Down syndrome pregnancies, which factors might help explain the variability in those rates, and whether failures occur at a similar rate in Down syndrome and euploid pregnancies. Second, we analyzed the impact on the testing success of a repeat cfDNA test using duplicate or subsequent plasma samples. Finally, these data were combined with data on the prevalences of the common autosomal aneuploidies to compare the rates of failure of alternative clinical protocols compared with follow-up cfDNA testing.
Test failures are uncommon when performing prenatal serum-based screening, but may occur due to a dropped sample or long time in transit at high temperatures. Ultrasound measurements of the nuchal translucency may also fail, but in the hands of experts with adequate time, failures only occur in a small percentage of women and are often associated with maternal obesity. Failure to meet test criteria (e.g., a sample drawn too early or too late, or a nuchal translucency request in the second trimester) would not be considered a test failure, but rather an inappropriate or inadequate sample. In general, serum- and ultrasound-based screening test results can be categorized into those that are screen-positive and those that are screen-negative. The rare failures are simply repeated or re-assigned (e.g., a request for a full integrated test but with a failed nuchal translucency measurement may be run instead as a serum integrated test). Also, there is no evidence that a failed serum or ultrasound test is a risk factor for chromosome abnormalities. These characteristics have led the prenatal screening performance for Down syndrome to be characterized only by the detection rate (sensitivity, that is, the proportion of all cases that have a positive screen) and the false-positive rate (1 – specificity, that is, the proportion of all controls with a positive screen).
Sequencing of cfDNA in the maternal circulation is different. Although some failures occur for the same reasons, others are due to failures in the complex methodology (e.g., poor polymerase chain reaction conditions, contamination, or too few aligned reads) or to sample characteristics that make the cfDNA testing inaccurate or inappropriate (e.g., low fetal fraction, or low amount of fetal or placenta DNA). Previous summaries of the literature have documented the reported failure rates, but have not fully explored the possible reasons for the wide range of rates. Some have reported initial failure rates of 5% or higher, while others found an increased risk of aneuploidies associated with failed samples compared to those with successful testing. In general, laboratories offering cfDNA screening tests attempt to simultaneously minimize the test failure rate, the false-positive rate, and false negatives. Unfortunately, most reports of clinical validity have used only data on those pregnancies with reported test results, rather than among all who have requested testing.
The aim of this systematic review is to summarize the literature on cfDNA test failure rates, Down syndrome (trisomy 21) risk among failures, and the impact of repeat testing so that practicing obstetrical care providers, genetic counselors, pregnant couples, professional organizations, and others have the most reliable information possible for decision-making and developing practice guidelines. The clinical setting for these analyses is cfDNA sequencing of maternal plasma as a primary screening test for Down syndrome in a general US pregnancy population. Test failures in high-risk pregnancies are somewhat less problematic because those women have likely already been informed by genetic professionals and been offered an invasive procedure and diagnostic testing. Factors that could impact failure rates include gestational age at testing, maternal weight or body mass index, whether fetal fraction is available, testing methodology, the number of aligned reads and depth of sequencing, and whether women with a test failure were subject to repeat testing. Repeat cfDNA testing is defined in this report as testing a new blood sample after the initial failure is identified. It does not include repeat sequencing of existing library material. Repeat testing might utilize a duplicate sample (a second tube of blood collected at the same time as the first sample), a subsequent sample (a new blood draw several days after the initial draw), or both.
Materials and methods
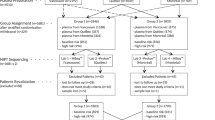
Three meta-analyses on the screening performance of cfDNA and common aneuploidies have been highly visible.1,2,3 The most recent of these3 was used as the baseline search target in a PubMed “related articles” search. Those results were limited to article types (clinical studies, clinical trials, meta-analyses, systematic reviews, or reviews) published between 1 January 2015 and 25 September 2017 and using English language. This time frame was chosen because systematic reviews performed earlier than 2015 would include only a few studies that were likely to be included in later reviews. This search identified 82 articles, of which five systematic reviews were identified, including the search target.2,3,4,5,6 Searching these reference lists identified one additional study in the relevant time period7 that was included and another8 that was not included (identifying only four studies all captured elsewhere). Three other systematic reviews published before 2015 were also identified.1,9,10 Only one review from 2014 was included.1 The other two9,10 were not, due again to the small number of referenced studies that were all identified by later reviews. The identification of potentially relevant literature was taken from these seven published systematic reviews and meta-analyses of cfDNA testing for Down syndrome and other related disorders (Figure 1).1,2,3,4,5,6,7 Studies included in these seven reports were then subject to seven specific inclusion criteria (Supplementary Table S1 online). Additional information was collected for each study that met these pre-specified criteria (Supplementary Table S1).
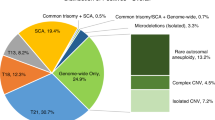
The current review was limited to Down syndrome as this was the focus of more studies, and there are more available data due to its higher prevalence and relevance. Using the 2015 US distribution of maternal ages,11 the age-associated risk of the common autosomal trisomies, their fetal loss rates during pregnancy,12 and survival to 1 year of age (93, 6, and 9% for Down syndrome, trisomy 18, and trisomy 13, respectively),13,14 the occurrence of Down syndrome among these common autosomal trisomies increases from 72% in the first trimester to 84% at term and reaches 99% 1 year after delivery. Focusing on Down syndrome is a reasonable first step in examining cfDNA test failures. It would be inappropriate to combine failure rates from one study that included only euploid and Down syndrome pregnancies with another study that also included trisomy 18, trisomy 13, sex chromosome aneuploidies, and select microdeletions. Screening for these additional disorders will be associated with higher failure rates. In addition, the presence of highly specific second-trimester ultrasound findings (e.g., heart defects, spina bifida, rocker bottom feet, and clenched fists) is far more common for trisomies 18 and 13 than for Down syndrome. This impacts the options for further testing should a test failure for one of these less common trisomies occur.
Once a study met the inclusion criteria, the additional information was extracted and added to an Excel spreadsheet (Microsoft, Redmond, WA), then independently verified by E.M.K. For important missing data (e.g., the number of Down syndrome and euploid failures, removing failures from other trisomies), the corresponding author was contacted via email for additional information. Insufficient or inappropriate samples (e.g., too little plasma or samples from twin pregnancies) were not considered to be test failures. Some results were estimated. For example, if the year of testing was not provided, it was assumed to be 1 year before the year of publication. The proportion of samples collected after 20 weeks’ gestation was estimated from figures or the reported mean and SD, interquartile range, or range. If the total, rather than the minimum number of aligned reads was reported, the proportion aligned was assumed to be 50%. We did not reassess the quality of individual studies as they had already been assessed and reported by the authors of the included meta-analyses.
Meta-analyses were performed using Comprehensive Meta-Analysis software (BioStat, Englewood, NJ). A random-effects model was chosen for the analysis of failure rates as it is likely that some of the categorical moderators (e.g., repeat testing) are associated with differing effect sizes. Default settings were used for the mixed-effects modeling. When computing an odds ratio where one cell was 0 (e.g., the number of test failures in Down syndrome pregnancies), 0.5 was added to all cells. This adjustment was also used when computing failure rates when the number of observed failures was zero. Significance was two-tailed at the P = 0.10 level for the examination of moderator variables and at the P = 0.05 level for all other comparisons. No adjustments were made for multiple comparisons. When relevant, the potential for publication bias was assessed using Duval and Tweedie’s trim and fill method (Comprehensive Meta-Analysis). In our report, the term “euploid” is used as the comparator. When possible, other aneuploidies were removed so that only Down syndrome and euploid pregnancies remained. However, in some studies, a few trisomy 18/13 pregnancies (labeled “non–Down syndrome” in the original studies) were included.
Results
Down syndrome cfDNA screening failure rates
A total of 7 structured reviews and 90 separate reports containing relevant information were identified by the search strategy (Figure 1). Of the 90 reports, 15 were published in a foreign language (most in Chinese). Of the remaining 75 reports, 30 satisfied the inclusion criteria (Table 1).15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44 The most common reason for exclusion was incomplete diagnostic testing follow-up (Supplementary Table S2). Additional comments and new information for the 30 included studies are contained in Supplementary Table S3. Twenty-two of these studied Western European populations,15,16,17,18,19,20,21,24,26,29,32,33,34,35,36,37,39,40,41,42,43,44 while eight studied Asian populations.22,23,25,27,28,30,31,38 Sixteen utilized massively parallel shotgun sequencing on an Illumina (San Diego, CA) platform, and six others used massively parallel shotgun sequencing on a semiconductor platform (Life Technologies, Grand Island, NY). Five reports utilized chromosome-specific sequencing (Ariosa Diagnostics, San Jose, CA) and the three others relied on identifying single-nucleotide polymorphisms (Natera, San Carlos, CA). Eighteen studies reported measuring fetal fraction on all samples, six reported this only for male fetuses, five did not measure or did not report fetal fraction, and one did not provide sufficient detail.31 Twenty-six studies provided test results for only the initial sample,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40 while four included results reported after repeat testing of all initial test failures.41,42,43,44
All 30 studies (26 with initial test results and 4 with repeat test results) provided sufficient detail to compute the overall test failure rate, and all but one30 provided data to compute failure rates among Down syndrome and euploid pregnancies. The 25 studies reporting complete initial results included 1,020 Down syndrome pregnancies (1,003 with successful testing and 17 with test failures), along with 9,868 euploid pregnancies (9,580 successful tests and 288 test failures). In the 4 studies reporting results after repeat testing, there were 444 Down syndrome (443 successful tests and 1 test failure) and 5,931 euploid pregnancies (5,850 successful tests and 81 test failures). All studies utilized designs that resulted in a relatively low ratio of euploid to Down syndrome pregnancies. In the 26 studies reporting initial sample testing, the average ratio of euploid to Down syndrome was 9.7:1 (range: 1.4:1–54:1) while in the 4 studies that utilized repeat sample testing, the average was 13.4:1 (range: 6.9:1–27.8:1).
All 30 data sets were analyzed for the overall failure rate, stratified by each of the nine potential moderator variables listed in Table 1 (Supplementary Figure S1). The results (labeled analysis 1) are summarized in Table 2 and ordered by increasing P value. Only three were significant: study location (P = 0.020), repeat testing (P = 0.022), and fetal fraction (P = 0.042). Using a priori reasoning, studies that repeated testing on a duplicate or subsequent sample should be separated from those reporting results only on initial sample testing. The study location was also highly significant, with the studies from Asian countries having lower failure rates than those from Western countries (0.6 versus 2.4%, respectively). Of the eight Asian studies, four did not measure fetal fraction, two measured it only in males, one measured it in all pregnancies, and one did not report it. This is significantly different from the 22 Western studies, of which only one did not measure fetal fraction, while four measured it only in males, and the remaining 17 measured it in all pregnancies (categories of no measurement/male only versus all pregnancies, Fisher’s exact test P = 0.0053). None of the studies in Asian populations reported the maternal weight or body mass index, but two previous prenatal screening reports from China reported average weights of 52 kg45 and 60 kg.46 Six studies in the Western population reported average or median weights between 65 and 70 kg16,20,29,36,42,43 and another two reported average or median body mass indexes of 2341 and 27.17 This systematic difference would be expected to result in fewer instances of low fetal fraction in obese women in the Asian populations and therefore an expected lower test failure rate. For these reasons, the eight studies from Asian countries formed a separate group.
The 18 studies15,16,17,18,19,20,21,24,26,29,32,33,34,35,36,37,39,40 from Western populations that reported initial failures were then reanalyzed by the moderator variables (analysis 2 in Table 2, and Supplementary Figure S2). None of those examined (i.e., volume of plasma, fetal fraction measurement, or test methodology) was helpful in explaining the remaining variability. The three final groups were then reanalyzed (“final” analysis in Table 2). This stratification resulted in a significant reduction in the variability (P = 0.001) and final estimates (0.6, 3.3, and 1.2%, respectively) of failure rates. The corresponding Forest plot is shown in Figure 2.
Forest plot of the overall cfDNA test failure rate in 30 included studies, stratified into three subgroups. Dotted vertical lines indicate the 95% confidence interval (CI) for each group. The study weighting within each group is indicated by the size of the circle (small to large: <5, 5–9, 10–19, and ≥20%).
Risk of Down syndrome in successful and failed cfDNA screening tests
Eighteen studies15,16,17,18,20,26,28,29,31,32,33,34,35,36,40,41,42,43,44 stratified the failed samples into Down syndrome and euploid pregnancies. The remaining 12 reported no failures and were, therefore, uninformative. The results were examined using a fixed-effects model as there is no expectation that the ratio of Down syndrome to euploid failures should be dependent on the overall failure rate. Figure 3 shows the results, with a consensus odds ratio of 0.98 (95% confidence interval (CI): 0.62–1.55; range: 0.14–5.53), indicating that test failures in Down syndrome are no more, or less, likely than in euploid pregnancies. One of the 18 studies34 had an odds ratio that was significantly higher than the consensus (P = 0.014), but the overall heterogeneity was considered low (Cochran’s Q = 14.9, P = 0.60, I2 = 0%). A random-effects model resulted in nearly identical results (data not shown). An examination of potential publication bias indicated a slightly higher odds ratio of 1.13 (95% CI: 0.73–1.76; Duval and Tweedie’s trim and fill), but this was not significantly different from the original estimate.
Forest plot of the odds ratio (OR) for the failure rate in Down syndrome versus euploid pregnancies from 18 included studies. Two studies42,44 reported results after repeat testing of a second plasma sample. The study weighting is indicated by the size of the circle (small to large: <5, 5–9, 10–19, and ≥20%). CI, confidence interval; *P = 0.014; **P = 0.94.
Repeat cfDNA testing after an initial Down syndrome test failure
The strategy of repeat testing of a duplicate sample when the initial sample failed for Down syndrome screening was utilized by four studies.41,42,43,44 In three of these,41,42,44 it was possible to obtain the initial failure rates as well. The initial and repeat failure rates and the percentage reduction for these studies were 4.0, 0.4, and 93%, 5.2, 0.8, and 85%, and 9.6, 2.9, and 70%, respectively. These results may overestimate the effect of repeat testing, as this strategy allows for a conservative call on the initial samples with the knowledge that more information will be generated before making a final clinical interpretation. The design for these three studies ensured that all women with a failed initial test had a second sample suitable for testing.
An independent estimate of the reduction in failed tests can be gleaned from large prospective cohort studies that reported initial cfDNA test failure rates and documented the impact of repeat testing. Using the same seven meta-analyses as a guide, we searched the 44 excluded reports for additional cohort studies that might have reported such data and included at least 1,000 pregnancies. The search was supplemented with more recent publications and totaled 26 studies (Supplementary Table S4). Of these 26 studies, 12 provided only the initial failure rate and were excluded from further analyses. The remaining 14 studies2,47,48,49,50,51,52,53,54,55,56,57,58,59 reported the number of initial failures, the number of those retested and the number of retested samples yielding informative results for Down syndrome screening. Among the 304,462 initial cfDNA tests performed (Supplementary Table S5), there were 4,322 failures for an overall rate of 1.3% (95% CI: 0.8–2.2%; range: 0.1–5.6%). Among these women, 4,050 repeat samples were obtained (83%, 95% CI: 74–90%; range 52–100%). Of those repeated, 3,659 were successful (79%, 95% CI: 62–89%; range: 46–97%). Although there was considerable heterogeneity among these rates, which was probably due to clinical practice, removing each study and recomputing these rates showed that no one study dominated the final estimate. Among the four studies reporting pregnancy outcomes,47,51,53,55 there were 179 successful repeat tests (176 euploid pregnancies that screened negative, one false-positive euploid, one true-positive trisomy 18, and one true-positive trisomy 13).
Also of interest is the risk of Down syndrome among those pregnancies that failed cfDNA testing twice (initial and duplicate samples). These data are available from the four studies41,42,43,44 that routinely repeated all initial failures (Table 1, bottom four rows). Among the 82 “double failures” reported, there were one Down syndrome and 81 euploid pregnancies. Compared with the risk of Down syndrome in women with successful testing (including repeated results) from the same four studies, the odds ratio was 0.57 (95% CI: 0.16–2.03), but this reduction in Down syndrome risk was not significantly different from the consensus odds ratio of 0.98.
Application of these results to the US general pregnancy population
The setting specified for this review was primary cfDNA screening for Down syndrome in a general pregnancy population. While the data presented so far have focused on Down syndrome, the failure rates for trisomy 18 and trisomy 13 are known to be higher than those in euploid pregnancies.34,60,61 Using the 2015 US maternal age distribution and recent age-associated risks (Supplementary Table S6), the first-trimester prevalences of Down syndrome, trisomy 18 and trisomy 13 are 1:340, 1:1,130 and 1:3,510, respectively. In a general pregnancy population cohort of 100,000 women in the first trimester of pregnancy, this corresponds to 294 Down syndrome, 88 trisomy 18, and 28 trisomy 13 pregnancies, along with 99,590 euploid pregnancies. Given a 3.3% expected initial failure rate for the euploid and Down syndrome pregnancies (Table 2 and Figure 2), 3,286 euploid and 9.7 Down syndrome samples would fail. If the failure rate in trisomy 18 and 13 were twice as high (6.6%), another 5.8 and 1.8 failures would occur, respectively. The overall failure rate would be 3.303%, as euploid failures dominate the overall rate. The risk of any of the common trisomies among these failures would be about 0.53% (1:190) compared with the risk in the overall population of 0.41% (1:243). Were the analysis to be shifted to term, the failed cases would be 3,292 euploid, 6.6 Down syndrome, 1.6 trisomy 18, and 1.0 trisomy 13. The risk of delivering a trisomic fetus among the group with a failed initial test at term would be 0.28% (1:360) compared with the risk in the entire population at term of 0.24% (1:420). If diagnostic testing were to be accepted by the 3,301 women with test failures (3.301%), it would identify about 9 pregnancies associated with term deliveries of infants with common trisomies, 6 or 7 of which would have Down syndrome. If the procedure-related loss was 1 in 400, there would be an expected 8 unaffected pregnancies lost—approximately a 1:1 ratio of trisomies detected to euploid procedure-related losses.
Discussion
This systematic review of cfDNA test failures focuses on Down syndrome screening in the general pregnancy population. Initial test failure rates were found to be highly dependent on the geographic location of the study (Figure 2), with lower rates in Asian (0.6%) compared with Western (3.3%) populations. This is probably due to two factors: the lower proportion of Asian women who are obese compared with Western women (obesity is associated with lower fetal fractions), and the fact that few Asian studies (two of seven; 29%) measured fetal fraction in all pregnancies while most Western studies did (13 of 18; 72%). Test failure rates were also significantly different among Western studies between women with repeat testing (1.2%) and those with testing of only an initial sample (3.3%). Our review also confirmed, on a much larger sample set, that the risk of Down syndrome associated with test failures is no different from the risk of Down syndrome associated with successful testing (odds ratio = 0.98). We did not quantify the risk for trisomy 18 and 13 in test failures. Finally, the review quantified the impact of repeat testing that might be expected in practice in two ways. The initial failure rates in three studies41,42,44 routinely testing a duplicate sample as part of the protocol were reduced by 93, 85, and 70%, respectively. In a series of 18 cohort studies, 83% of women with an initial failure submitted a subsequent sample and 79% of those tests returned a usable result. This suggests that two-thirds of initial failed tests resolve upon repeat testing of a subsequent sample.
One review of failure rates62 concluded that laboratories using the massively parallel shotgun sequencing method have the lowest (average: 1.6%), followed by the chromosome-specific sequencing method (3.6%), while those using the single-nucleotide polymorphism method have the highest (6.4%). Unlike the current study, this review did not attempt to distinguish repeat from initial testing or to account for the expected higher failure rates when disorders in addition to Down syndrome were included. Although failure rates due to fetal fraction are probably similar regardless of the disorders included, there are additional reasons for failed interpretations for the other autosomal and sex chromosome aneuploidies. In their subsequent analysis, these authors modeled the impact of test failures on the detection and false-positive rates as well as positive predictive values in a general pregnancy population. However, they assumed that all women with initial test failures would require invasive procedures and diagnostic testing. Another study modeled the impact of test failures on the detection and false-positive rates in a general pregnancy population.63 Their model employed separate rates for those failing due to low fetal fraction versus technical failures and found that the Down syndrome detection rate dropped by 1–2% with little impact on the false-positive rate. However, that study also did not consider the impact of repeat testing or subsequent targeted second-trimester ultrasound. Another study addressed duplicate versus subsequent sampling.64 As might be expected, collecting and testing a subsequent sample was more effective in resolving test failures (56%) than testing a duplicate sample (39%). This is probably due to the added within-woman variability associated with a later sample draw. However, these results are complicated to interpret, as duplicate samples are routinely collected and tested by this laboratory and are available for nearly every pregnant woman. Thus, a direct comparison with the summary of studies provided earlier cannot be made (Supplementary Table S5).
The American College of Obstetricians and Gynecologists65 recommends that women whose initial cfDNA test fails (e.g., the results are not reported, indeterminate, uninterpretable, or no-call) receive genetic counseling and be offered comprehensive ultrasound evaluation and diagnostic testing because of an increased risk of aneuploidy.34,61 In one of these references,61 there were 488 test failures that included 13 chromosome abnormalities (2.7%). The odds ratio for these abnormalities in failed versus successful testing was reported to be 8.7. However, only 6 of the 13 abnormalities were targets for current prenatal screening tests (three Down syndrome, one trisomy 18, and two trisomy 13 pregnancies). The other abnormalities were triploidy (four), trisomy 13 mosaic (one), del 11p (WAGR syndrome), and a structural abnormality in chromosome 1. Restricting the analysis to only the common autosomal trisomies, the revised odds ratio is 4.0; that is, half the original, but still statistically significant. For comparison with the current study, the odds ratio for test failures in Down syndrome versus euploid pregnancies is 2.62 (95% CI: 0.84–8.87, P = 0.12).61 If this study had been included in the current review (Figure 3), the odds ratio would have been the third highest of 19 studies. The second report34 enrolled 15% of women before 10 weeks and these early samples were responsible for 40% of all test failures. Most clinical laboratories do not accept samples at 9 weeks or earlier. As shown in Figure 3, the odds ratio for Down syndrome in this report34 was the highest of all 18 included studies, and the only one significantly higher than the consensus odds ratio of 0.98. These two studies34,61 are not representative of the literature regarding Down syndrome risk in failed cfDNA tests.
The common abnormalities that would be found more often in failed tests are trisomy 18, trisomy 13, and triploidy due to the clearly reduced average fetal fractions.60,66,67 However, these disorders are likely to be detectable through targeted ultrasound with highly specific second-trimester findings such as heart defects and limb abnormalities,68,69 and would not justify the need for offering invasive testing to all women with failed cfDNA tests. However, in settings where cfDNA testing is secondary to a positive screen for maternal age or serum screening, the offer of invasive testing would be appropriate.
The American College of Medical Genetics and Genomics70 recommends offering diagnostic testing to women with an initial cfDNA test failure. A repeat blood draw is not considered appropriate. This is based on the well-confirmed observation that the average fetal fraction increases only slightly between 10 and 20 weeks of gestation (0.1%71 to 0.44%64 per week). However, the authors do not consider differences between the initial and repeat fetal fraction estimates that are due to within-woman variability and between-run assay variability. Furthermore, this ignores that about half of all test failures are not related to fetal fraction.
Based on the information contained in this review, it is possible to construct evidence-based options for women experiencing an initial test failure as part of general pregnancy population cfDNA-based screening for Down syndrome and trisomies 18 and 13. The primary obstetrical care provider always has the option of immediately referring patients with a test failure to a specialist for follow-up. However, the following options could be considered relevant for primary care providers. (i) If available, offer a repeat cfDNA test from the same laboratory as soon as possible. Waiting longer is unlikely to improve the success rate given the short half-life of cfDNA. Consider the gestational age to ensure that this delay would not limit the woman’s reproductive options. For an individual, a successful repeat test may depend on her maternal weight and the initially reported fetal fraction. One commercial laboratory is now including a table on their reports with initial test failures providing repeat testing success rates stratified by maternal weight and initial fetal fraction, which could be of use in counseling. Sending a repeat sample to another laboratory will probably incur additional costs with no clear advantage. (ii) Offer conventional serum or ultrasound screening, such as a first-trimester combined test, especially if a first trimester nuchal translucency measurement has already been performed. The results could be available within a day or two. (iii) Offer targeted ultrasound in the early second trimester to look for highly specific structural anomalies and other findings indicative of a trisomy or other chromosome disorder (e.g., triploidy). If found, offer an invasive procedure and diagnostic testing. If there are no ultrasound findings, the risks for these common trisomies are reduced by 80% or more.68,69 (iv) Offer an invasive procedure or diagnostic test if definitive results are desired.
Using the results of this review, it is possible to model the impact of a repeat cfDNA test and targeted second-trimester ultrasound in the hypothetical group of 3,303 women with an initial first-trimester cfDNA test failure described earlier. Approximately 66% (83 × 79%; Supplementary Table S5) of the women with a euploid or Down syndrome pregnancy would have a repeat sample collected and tested successfully. The repeat test result would be highly accurate in the 2,155 euploid and 6 or 7 Down syndrome pregnancies. Repeat testing is likely to be less successful in trisomy 18 and 13.60,64,66,72,73 However, these conditions are also far less common, especially later in pregnancy. When coupled with a mid-second trimester targeted ultrasound, most of these cases would be identified, along with the occasional triploid pregnancy. The vast majority (95%) of the remaining euploid pregnancies would have a normal ultrasound with a corresponding reduction in risk, while one or two of the three to four remaining Down syndrome pregnancies might also have a detectable ultrasound abnormality. Thus, a follow-up protocol incorporating repeat cfDNA testing and targeted ultrasound would probably identify the vast majority of common trisomies while reducing the need for diagnostic testing to about 165 (5%) of the 3,303 women. This would result in reduced costs and, perhaps, a single procedure-related loss.
There are limitations to the data summarized in this report. Many of the included studies are several years old and test failure rates may have changed over time. We did not assess the quality of individual studies, as nearly all had already been assessed by one or more of the previous systematic reviews1,2,3,5,6,7 using the QUADAS-2 criteria.74 Although each of these reviews raised concerns about potential biases and applicability, especially with respect to patient selection and study flow and timing, only one study5 used these quality measures in its analyses (low-quality studies were removed). None of the reviews explored study quality with the detection rate, false-positive rate, or failure rate.
The new information in this systematic review helps clarify some of the possible reasons why failure rates differ so widely in the literature and in practice. Restricting analyses to only Down syndrome and euploid pregnancies (or some other common set of outcomes) is one factor, but further analyses reveal that there are important differences in both screening philosophy and population characteristics that may contribute to systematic differences in the failure rates reported from Asian and Western countries. Finally, this report provides more clarity on the impact of repeat testing after an initial test failure. Repeat testing is already incorporated into most clinical testing protocols. In addition, collecting and testing a subsequent sample is highly effective in providing reliable screening results to women with an initial failed test. The offer of repeat testing should consider the reasons for failure, the fetal fraction and maternal weight, and that the gestational age is sufficiently early so as to not limit reproductive choices.
References
Gil MM, Akolekar R, Quezada MS, Bregant B & Nicolaides KH. Analysis of cell-free DNA in maternal blood in screening for aneuploidies: meta-analysis. Fetal Diagn Ther 2014;35:156–173.
Gil MM, Quezada MS, Revello R, Akolekar R & Nicolaides KH. Analysis of cell-free DNA in maternal blood in screening for fetal aneuploidies: updated meta-analysis. Ultrasound Obstet Gynecol 2015;45:249–266.
Gil MM, Accurti V, Santacruz B, Plana MN & Nicolaides KH. Analysis of cell-free DNA in maternal blood in screening for aneuploidies: updated meta-analysis. Ultrasound Obstet Gynecol 2017;50:302–314.
Jin J, Yang J, Chen Y & Huang J. Systematic review and meta-analysis of non-invasive prenatal DNA testing for trisomy 21: implications for implementation in China. Prenat Diagn 2017;37:864–873.
Iwarsson E, Jacobsson B, Dagerhamn J, Davidson T, Bernabe E & Heibert Arnlind M. Analysis of cell-free fetal DNA in maternal blood for detection of trisomy 21, 18 and 13 in a general pregnant population and in a high risk population—a systematic review and meta-analysis. Acta Obstet Gynecol Scand 2017;96:7–18.
Mackie FL, Hemming K, Allen S, Morris RK & Kilby MD. The accuracy of cell-free fetal DNA-based non-invasive prenatal testing in singleton pregnancies: a systematic review and bivariate meta-analysis. BJOG 2017;124:32–46.
Taylor-Phillips S, Freeman K, Geppert J et al. Accuracy of non-invasive prenatal testing using cell-free DNA for detection of Down, Edwards and Patau syndromes: a systematic review and meta-analysis. BMJ Open 2016;6:e010002.
Yang H, Xu HB, Liu TT & He XL. Systematic review of noninvasive prenatal diagnosis for abnormal chromosome genetic diseases using free fetal DNA in maternal plasma. Genet Mol Res 2015;14:10603–10608.
Mersy E, Smits LJ, van Winden LA et al. Noninvasive detection of fetal trisomy 21: systematic review and report of quality and outcomes of diagnostic accuracy studies performed between 1997 and 2012. Hum Reprod Update 2013;19:318–329.
Verweij EJ, van den Oever JM, de Boer MA, Boon EM & Oepkes D. Diagnostic accuracy of noninvasive detection of fetal trisomy 21 in maternal blood: a systematic review. Fetal Diagn Ther 2012;31:81–86.
Centers for Disease Control and Prevention. CDC WONDER https://wonder.cdc.gov/controller/datarequest/D66. Accessed 11 September 2017.
Savva GM, Walker K & Morris JK. The maternal age-specific live birth prevalence of trisomies 13 and 18 compared to trisomy 21 (Down syndrome). Prenat Diagn 2010;30:57–64.
Kucik JE, Shin M, Siffel C et al. Trends in survival among children with Down syndrome in 10 regions of the United States. Pediatrics 2013;131:e27–e36.
Rasmussen SA, Wong LY, Yang Q, May KM & Friedman JM. Population-based analyses of mortality in trisomy 13 and trisomy 18. Pediatrics 2003;111:777–784.
Alberti A, Salomon LJ, Le Lorc’h M et al. Non-invasive prenatal testing for trisomy 21 based on analysis of cell-free fetal DNA circulating in the maternal plasma. Prenat Diagn 2015;35:471–476.
Ashoor G, Syngelaki A, Wagner M, Birdir C & Nicolaides KH. Chromosome-selective sequencing of maternal plasma cell-free DNA for first-trimester detection of trisomy 21 and trisomy 18. Am J Obstet Gynecol 2012;206:322.e1–5.
Bianchi DW, Platt LD, Goldberg JD, Abuhamad AZ, Sehnert AJ & Rava RP. Genome-wide fetal aneuploidy detection by maternal plasma DNA sequencing. Obstet Gynecol 2012;119:890–901.
Chiu RW, Akolekar R, Zheng YW et al. Non-invasive prenatal assessment of trisomy 21 by multiplexed maternal plasma DNA sequencing: large scale validity study. BMJ 2011;342:c7401.
Dheedene A, Sante T, De Smet M et al. Implementation of non-invasive prenatal testing by semiconductor sequencing in a genetic laboratory. Prenat Diagn 2016;36:699–707.
Ehrich M, Deciu C, Zwiefelhofer T et al. Noninvasive detection of fetal trisomy 21 by sequencing of DNA in maternal blood: a study in a clinical setting. Am J Obstet Gynecol 2011;204:205.e1–11.
Guex N, Iseli C, Syngelaki A et al. A robust second-generation genome-wide test for fetal aneuploidy based on shotgun sequencing cell-free DNA in maternal blood. Prenat Diagn 2013;33:707–710.
Jeon YJ, Zhou Y, Li Y et al. The feasibility study of non-invasive fetal trisomy 18 and 21 detection with semiconductor sequencing platform. PLoS ONE 2014;9:e110240.
Jiang F, Ren J, Chen F et al. Noninvasive Fetal Trisomy (NIFTY) test: an advanced noninvasive prenatal diagnosis methodology for fetal autosomal and seks chromosomal aneuploidies. BMC Med Genomics 2012;5:57.
Johansen P, Richter SR, Balslev-Harder M et al. Open source non-invasive prenatal testing platform and its performance in a public health laboratory. Prenat Diagn 2016;36:530–536.
Kim S, Jung H, Han SH et al. Comparison of two high-throughput semiconductor chip sequencing platforms in noninvasive prenatal testing for Down syndrome in early pregnancy. BMC Med Genomics 2016;9:22.
Koumbaris G, Kypri E, Tsangaras K et al. Cell-free DNA analysis of targeted genomic regions in maternal plasma for non-invasive prenatal testing of trisomy 21, trisomy 18, trisomy 13, and fetsekssex. Clin Chem 2016;62:848–855.
Lau TK, Chen F, Pan X et al. Noninvasive prenatal diagnosis of common fetal chromosomal aneuploidies by maternal plasma DNA sequencing. J Matern Fetal Neonatal Med 2012;25:1370–1374.
Lee MY, Cho DY, Won HS et al. Performance of Momguard, a new non-invasive prenatal testing protocol developed in Korea. Obstet Gynecol Sci 2015;58:340–345.
Lefkowitz RB, Tynan JA, Liu T et al. Clinical validation of a noninvasive prenatal test for genomewide detection of fetal copy number variants. Am J Obstet Gynecol 2016;215:227.e1–227.e16.
Liang D, Lv W, Wang H et al. Non-invasive prenatal testing of fetal whole chromosome aneuploidy by massively parallel sequencing. Prenat Diagn 2013;33:409–415.
Liao C, Yin AH, Peng CF et al. Noninvasive prenatal diagnosis of common aneuploidies by semiconductor sequencing. Proc Natl Acad Sci USA 2014;111:7415–7420.
Nicolaides KH, Syngelaki A, Gil M, Atanasova V & Markova D. Validation of targeted sequencing of single-nucleotide polymorphisms for non-invasive prenatal detection of aneuploidy of chromosomes 13, 18, 21, X, and Y. Prenat Diagn 2013;33:575–579.
Norton ME, Brar H, Weiss J et al. Non-Invasive Chromosomal Evaluation (NICE) study: results of a multicenter prospective cohort study for detection of fetal trisomy 21 and trisomy 18. Am J Obstet Gynecol 2012;207:137.e1–137.e8.
Pergament E, Cuckle H, Zimmermann B et al. Single-nucleotide polymorphism-based noninvasive prenatal screening in a high-risk and low-risk cohort. Obstet Gynecol 2014;124:210–218.
Persico N, Boito S, Ischia B et al. Cell-free DNA testing in the maternal blood in high-risk pregnancies after first-trimester combined screening. Prenat Diagn 2016;36:232–236.
Poon LC, Dumidrascu-Diris D, Francisco C, Fantasia I & Nicolaides KH. IONA test for first-trimester detection of trisomies 21, 18 and 13. Ultrasound Obstet Gynecol 2016;47:184–187.
Sehnert AJ, Rhees B, Comstock D et al. Optimal detection of fetal chromosomal abnormalities by massively parallel DNA sequencing of cell-free fetal DNA from maternal blood. Clin Chem 2011;57:1042–1049.
Shaw SW, Hsiao CH, Chen CY et al. Noninvasive prenatal testing for whole fetal chromosomal aneuploidies: a multicenter prospective cohort trial in Taiwan. Fetal Diagn Ther 2014;35:13–17.
Sparks AB, Struble CA, Wang ET, Song K & Oliphant A. Noninvasive prenatal detection and selective analysis of cell-free DNA obtained from maternal blood: evaluation for trisomy 21 and trisomy 18. Am J Obstet Gynecol 2012;206:319.e1–319.e9.
Stumm M, Entezami M, Haug K et al. Diagnostic accuracy of random massively parallel sequencing for non-invasive prenatal detection of common autosomal aneuploidies: a collaborative study in Europe. Prenat Diagn 2014;34:185–191.
Benachi A, Letourneau A, Kleinfinger P et al. Cell-free DNA analysis in maternal plasma in cases of fetal abnormalities detected on ultrasound examination. Obstet Gynecol 2015;125:1330–1337.
Palomaki GE, Kloza EM, Lambert-Messerlian GM et al. DNA sequencing of maternal plasma to detect Down syndrome: an international clinical validation study. Genet Med 2011;13:913–920.
Porreco RP, Garite TJ, Maurel K et al. Noninvasive prenatal screening for fetal trisomies 21, 18, 13 and the csekson sex chromosome aneuploidies from maternal blood using massively parallel genomic sequencing of DNA. Am J Obstet Gynecol 2014;211:365.e1–12.
Verweij EJ, Jacobsson B, van Scheltema PA et al. European Non-Invasive Trisomy Evaluation (EU-NITE) study: a multicenter prospective cohort study for non-invasive fetal trisomy 21 testing. Prenat Diagn 2013;33:996–1001.
Miao ZY, Liu X, Shi TK, Xu Y, Song QH & Tang SH. First trimester, second trimester, and integrated screening’for Down’s syndrome in China. J Med Screen 2012;19:68–71.
Song Y, Huang S, Zhou X et al. Non-invasive prenatal testing for fetal aneuploidies in the first trimester of pregnancy. Ultrasound Obstet Gynecol 2015;45:55–60.
Chitty LS, Wright D, Hill M et al. Uptake, outcomes, and costs of implementing non-invasive prenatal testing’for Down’s syndrome into NHS maternity care: prospective cohort study in eight diverse maternity units. BMJ 2016;354:i3426.
Eiben B, Krapp M, Borth H et al. Single nucleotide polymorphism-based analysis of cell-free fetal DNA in 3000 cases from Germany and Austria. Ultrasound Int Open 2015;1:E8–E11.
Flock A, Tu NC, Ruland A, Holzgreve W, Gembruch U & Geipel A. Non-invasive prenatal testing (NIPT: Europe’s first multicenter post-market clinical follow-up study validating the quality in clinical routine. Arch Gynecol Obstet 2017;296:923–928.
Lau TK, Cheung SW, Lo PS et al. Non-invasive prenatal testing for fetal chromosomal abnormalities by low-coverage whole-genome sequencing of maternal plasma DNA: review of 1982 consecutive cases in a single center. Ultrasound Obstet Gynecol 2014;43:254–264.
McLennan A, Palma-Dias R, da Silva Costa F, Meagher S, Nisbet DL & Scott F. Noninvasive prenatal testing in routine clinical practice—an audit of NIPT and combined first-trimester screening in an unselected Australian population. Aust NZ J Obstet Gynaecol 2016;56:22–28.
Oepkes D, Page-Christiaens GC, Bax CJ et al. Trial by Dutch laboratories for evaluation of non-invasive prenatal testing. Part I—clinical impact. Prenat Diagn 2016;36:1083–1090.
Palomaki GE, Kloza EM, O’Brien BM, Eklund EE & Lambert-Messerlian GM. The clinical utility of DNA-based screening for fetal aneuploidy by primary obstetrical care providers in the general pregnancy population. Genet Med 2017;19:778–786.
Quezada MS, Gil MM, Francisco C, Orosz G & Nicolaides KH. Screening for trisomies 21, 18 and 13 by cell-free DNA analysis of maternal blood at 10–11 weeks’ gestation and the combined test at 11–13 weeks. Ultrasound Obstet Gynecol 2015;45:36–41.
Sago H & Sekizawa A Japan NIPT consortium. Nationwide demonstration project of next-generation sequencing of cell-free DNA in maternal plasma in Japan: 1-year experience. Prenat Diagn 2015;35:331–336.
Samura O, Sekizawa A, Suzumori N et al. Current status of non-invasive prenatal testing in Japan. J Obstet Gynaecol Res 2017;43:1245–1255.
Taneja PA, Snyder HL, de Feo E et al. Noninvasive prenatal testing in the general obstetric population: clinical performance and counseling considerations in over 85 000 cases. Prenat Diagn 2016;36:237–243.
Zhang H, Gao Y, Jiang F et al. Non-invasive prenatal testing for trisomies 21, 18 and 13: clinical experience from 146,958 pregnancies. Ultrasound Obstet Gynecol 2015;45:530–538.
Zhou Q, Pan L, Chen S et al. Clinical application of noninvasive prenatal testing for the detection of trisomies 21, 18, and 13: a hospital experience. Prenat Diagn 2014;34:1061–1065.
Revello R, Sarno L, Ispas A, Akolekar R & Nicolaides KH. Screening for trisomies by cell-free DNA testing of maternal blood: consequences of a failed result. Ultrasound Obstet Gynecol 2016;47:698–704.
Norton ME, Jacobsson B, Swamy GK et al. Cell-free DNA analysis for noninvasive examination of trisomy. N Engl J Med 2015;372:1589–1597.
Yaron Y. The implications of non-invasive prenatal testing failures: a review of an under-discussed phenomenon. Prenat Diagn 2016;36:391–396.
Grati FR & Kagan KO. Rate of no result in cell-free DNA testing and its influence on test performance metrics. Ultrasound Obstet Gynecol 2017;50:134–137.
Kinnings SL, Geis JA, Almasri E et al. Factors affecting levels of circulating cell-free fetal DNA in maternal plasma and their implications for noninvasive prenatal testing. Prenat Diagn 2015;35:816–822.
Committee on Practice Bulletins—Obstetrics, Committee on Genetics, Society for Maternal-Fetal Medicine. Practice Bulletin No. 163: Screening for Fetal Aneuploidy. Obstet Gynecol 2016;127:e123–e137.
Palomaki GE, Kloza EM, Lambert-Messerlian GM et al. Circulating cell free DNA testing: are some test failures informative? Prenat Diagn 2015;35:289–293.
Musci T, Schmid M, Song K, Wang E & Fergus K. Fetal aneuploidy screening results in maternal plasma samples redrawn due to insufficient fetal cfDNA in the initial sample. Prenat Diagn 2016;36(suppl 1):16.
Tongsong T, Sirichotiyakul S, Wanapirak C & Chanprapaph P. Sonographic features of trisomy 18 at midpregnancy. J Obstet Gynaecol Res 2002;28:245–250.
Watson WJ, Miller RC, Wax JR, Hansen WF, Yamamura Y & Polzin WJ. Sonographic detection of trisomy 13 in the first and second trimesters of pregnancy. J Ultrasound Med 2007;26:1209–1214.
Gregg AR, Skotko BG, Benkendorf JL et al. Noninvasive prenatal screening for fetal aneuploidy, 2016 update: a position statement of the American College of Medical Genetics and Genomics. Genet Med 2016;18:1056–1065.
Wang E, Batey A, Struble C, Musci T, Song K & Oliphant A. Gestational age and maternal weight effects on fetal cell-free DNA in maternal plasma. Prenat Diagn 2013;33:662–666.
Ashoor G, Syngelaki A, Poon LC, Rezende JC & Nicolaides KH. Fetal fraction in maternal plasma cell-free DNA at 11–13 weeks’ gestation: relation to maternal and fetal characteristics. Ultrasound Obstet Gynecol 2013;41:26–32.
Rava RP, Srinivasan A, Sehnert AJ & Bianchi DW. Circulating fetal cell-free DNA fractions differ in autosomal aneuploidies and monosomy X. Clin Chem 2014;60:243–250.
Whiting P, Rutjes AW, Reitsma JB, Bossuyt PM & Kleijnen J. The development of QUADAS: a tool for the quality assessment of studies of diagnostic accuracy included in systematic reviews. BMC Med Res Methodol 2003;3:25.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
No outside funding was received for this project. G.E.P. was, or is, principal investigator for three investigator-developed and industry-sponsored studies relating to cfDNA (NCT00877292, NCT01966991, and NCT03087357). E.M.K. received grant funding from all three studies. G.E.P.’s institution has received consulting fees and/or an honorarium from Ansh Labs, Roche USA, and Illumina.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Palomaki, G.E., Kloza, E.M. Prenatal cell-free DNA screening test failures: a systematic review of failure rates, risks of Down syndrome, and impact of repeat testing. Genet Med 20, 1312–1323 (2018). https://doi.org/10.1038/gim.2018.22
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2018.22
Keywords
This article is cited by
-
Whole genome non-invasive prenatal testing in prenatal screening algorithm: clinical experience from 12,700 pregnancies
BMC Pregnancy and Childbirth (2022)
-
The effect of hemolysis on quality control metrics for noninvasive prenatal testing
BMC Medical Genomics (2022)
-
Antenatal screening for chromosomal abnormalities
Archives of Gynecology and Obstetrics (2022)
-
Prenatal screening for trisomy 21: a comparative performance and cost analysis of different screening strategies
BMC Pregnancy and Childbirth (2020)
-
Non-invasive prenatal testing of fetal chromosomal aneuploidies: validation and clinical performance of the veracity test
Molecular Cytogenetics (2019)