Abstract
Purpose
Von Hippel–Lindau (VHL) disease is a rare hereditary cancer syndrome that reduces life expectancy. We aimed to construct a more valuable genotype–phenotype correlation based on alterations in VHL protein (pVHL).
Methods
VHL patients (n = 339) were recruited and grouped based on mutation types: HIF-α binding site missense (HM) mutations, non-HIF-α binding site missense (nHM) mutations, and truncating (TR) mutations. Age-related risks of VHL-associated tumors and patient survival were compared.
Results
Missense mutations conferred an increased risk of pheochromocytoma (HR = 1.854, p = 0.047) compared with truncating mutations. The risk of pheochromocytoma was lower in the HM group than in the nHM group (HR = 0.298, p = 0.003) but was similar between HM and TR groups (HR = 0.901, p = 0.810). Patients in the nHM group had a higher risk of pheochromocytoma (HR = 3.447, p < 0.001) and lower risks of central nervous system hemangioblastoma (CHB) (HR = 0.700, p = 0.045), renal cell carcinoma (HR = 0.610, p = 0.024), and pancreatic tumor (HR = 0.382, p < 0.001) than those in the combined HM and TR (HMTR) group. Moreover, nHM mutations were independently associated with better overall survival (HR = 0.345, p = 0.005) and CHB-specific survival (HR = 0.129, p = 0.005) than HMTR mutations.
Conclusion
The modified genotype–phenotype correlation links VHL gene mutation, substrate binding site, and phenotypic diversity (penetrance and survival), and provides more accurate information for genetic counseling and pathogenesis studies.
Similar content being viewed by others
Introduction
Von Hippel–Lindau (VHL) disease (OMIM 193300) is a rare autosomal dominant tumor syndrome resulting from mutations in the VHL tumor suppressor gene. The incidence of VHL mutation is approximately 1 in 36,000–53,000 births, with age-dependent high penetrance.1,2 Lesions associated with VHL disease include central nervous system hemangioblastoma (CHB), retinal angioma (RA), renal cell carcinoma (RCC), renal cyst, pancreatic tumor or cyst (PCT), pheochromocytoma (PHEO), endolymphatic sac tumor, and papillary cystadenoma of the epididymis or broad ligament.3,4,5,6 The presentations of VHL disease vary remarkably, and patients may develop symptoms from early childhood through adulthood.7,8,9,10 Phenotypically, VHL disease can be classified into four types: type 1 does not include PHEO, type 2A includes PHEO but not RCC, type 2B includes both PHEO and RCC, and type 2C is associated with PHEO as the sole manifestation.1
The VHL gene is located on chromosome 3p25.3 and was first identified in 1993 (ref.11). The encoded VHL protein (pVHL) forms a complex with elongation factor C and B (elongin C/B) named the VCB complex, which forms the VCB-CR complex along with cullin2 (CUL2) and RING finger protein (RBX1).12 This complex is critical for pVHL to function as an E3 ligase, an important enzyme in the ubiquitylation and proteasomal degradation of HIF-α and several other substrates. pVHL consists of one α-domain and one β-domain. The β-domain comprises several β-sheets and binds HIF-α via residues 65–117 (refs.5,13,14,15,16). The α-domain consists of three α-helices and binds elongin C.17 The pVHL–HIF pathway plays an important role in tumorigenesis in VHL disease. VHL germ-line mutations result in dysfunction of the E3 ligase and accumulation of HIF-α, leading to the upregulation of proangiogenic factors, such as vascular endothelial growth factor (VEGF) and platelet-derived growth factor β, which accelerate tumorigenesis.18
Genotype–phenotype correlations have been reported in many studies, providing valuable strategies for prophylactic surveillance and genetic counseling for presymptomatic individuals in families with VHL disease.19,20 The typical genotype–phenotype correlation is that VHL missense mutations are responsible for type 2 disease with a high risk of PHEO, and truncating mutations are responsible for type 1 disease with a low risk of PHEO.21 Deletion of all or part of the VHL gene as well as the nearby gene BRK1 (also known as HSPC300 and C3orf10) leads to RA and CHB with a low risk of RCC, sometimes called type 1B VHL disease.1,22,23 In clinical practice, a considerable portion of missense mutation carriers do not develop PHEO, implying that there must be difference in missense mutations among the patients. A structural analysis reported that VHL missense mutations in the elongin C binding domain were associated with a higher risk of PHEO.16 Another study including a small number of patients (n = 13) found that VHL missense mutations in the HIF-α binding site may predispose patients to age-related CHB risks.13 Moreover, in vitro studies showed that mutated pVHL involved in type 2C VHL disease retains the ability to degrade HIF-α degradation, suggesting that independent mechanisms are responsible for HIF-α degradation and the PHEO phenotype.24,25 In view of the fact that the binding of HIF-α to pVHL in the E3 ligase is the first step in mediating HIF-α degradation under normoxia conditions, we hypothesized that the genotype–phenotype correlation may be related to the alteration of HIF-α binding to pVHL due to VHL mutations.
In this study, we reanalyzed the genotype–phenotype correlation by investigating age-related penetrance of VHL-associated tumors and survival in different patient groups with missense mutations in the HIF-α binding site (HM group), missense mutations not in the HIF-α binding site (nHM group), and truncating mutations (TR group). The results refined the genotype–phenotype correlation in VHL disease and provide more accurate information for genetic consultation and pathogenesis studies.
Materials and methods
Medical ethics
This project was approved by the Medical Ethics Committee of Peking University First Hospital. Informed consent was obtained from patients or legal guardians.
Patient ascertainment and assessment
In this study, we included all the patients with VHL disease diagnosed at Peking University First Hospital prior to 1 May 2017. A total of 339 patients from 128 unrelated families were enrolled for analysis. The lifetime from birth to death or to the end of follow-up was used for survival analysis. The median included time was 37.5 years/person (range 1–75 years) with a total of 13,102 person-years. Individuals were diagnosed with VHL disease if they carried a VHL germ-line mutation or fulfilled the clinical criteria for VHL disease, as determined by medical records and interviews with the patients or family members.3,26,27 A VHL mutation was confirmed in at least one patient in a family. Follow-up was carried out from the time of first clinical presentation for symptomatic patients or the time of genetic diagnosis of VHL disease for asymptomatic carriers until death or 1 May 2017. Medical records, disease history, and imaging data for all affected patients and asymptomatic carriers were reviewed to determine age at first diagnosis of the five major VHL lesions: CHB, RA, RCC, PCT, and PHEO.
Examination of VHL mutations
VHL mutations were examined using peripheral blood samples from all members of the families except those who refused the test or died. The three exons and their flanking intronic sequences were amplified by polymerase chain reaction and sequenced. Large deletions were detected by multiplex ligation-dependent probe amplification (MLPA, P016-C2 kit, MRC-Holland, Amsterdam, The Netherlands) and confirmed by real-time quantitative polymerase chain reaction. The primers and conditions for amplification were described in our previous publication.28 Patients with missense mutations leading to a single amino acid change in pVHL were divided into the HM group (mutations in residues 65–117) and the nHM group.13 Patients with nonsense mutations predicted to cause truncated proteins, small indels resulting in frameshift, splice-site mutations, and large deletions were categorized into the TR group.
Statistical analysis
Age-related penetrance of the five major VHL lesions and survival analysis were calculated using Kaplan–Meier plot and log-rank analysis. The Cox regression model was used to assess the effect of genotype on risk of the five major VHL lesions and on hazard of death in different groups. A p value less than 0.05 was considered to be statistically significant. For multiple comparisons, Bonferroni’s correction was used to reduce the type I error rate. Data were analyzed using SPSS version 22.0 software (IBM-SPSS, Chicago, IL).
Results
VHL gene mutations and clinical characteristics of the 339 Chinese patients with VHL disease
A total of 65 types of intragenic VHL mutations were identified within the 128 unrelated families, including 33 missense mutations (162 patients from 56 families) and 32 truncating mutations (177 patients from 72 families). Of the missense mutations, 56.1% were located in the HIF-α binding site, 35.8% were located in the elongin C binding site, and 8.1% were located in other sites (Supplementary Table S1 online). CHB (62.2%) was the most common manifestation in the patients, followed by PCT (46.0%), RCC (43.6%), RA (21.2%), and PHEO (13.5%), similar to our previous report.29
Genotype–phenotype correlations
Comparison of age-related tumor risks between missense mutation and truncating mutation carriers
As described in previous studies, missense mutation carriers showed an increased risk of PHEO (HR = 1.854, 95% confidence interval (CI) 1.009–3.406, p = 0.047), and a decreased risk of CHB (HR = 0.740, 95% CI 0.564–0.971, p = 0.030), RCC (HR = 0.577, 95% CI 0.414–0.804, p = 0.001), and PCT (HR = 0.608, 95% CI 0.441–0.838, p = 0.002) compared with those with truncating mutations. We further stratified the missense mutations as HIF-α binding site missense (HM) mutations and non-HIF-α binding site missense (nHM) mutations, and found that obvious phenotypic differences remained between nHM mutation carriers and TR mutation carriers; the nHM group had a higher risk of PHEO (HR = 3.023, 95% CI 1.584–5.769, p = 0.001) and lower risks of CHB (HR = 0.642, 95% CI 0.446–0.924, p = 0.017), RCC (HR = 0.545, 95% CI 0.353–0.840, p = 0.006), and PCT (HR = 0.431, 95% CI 0.273–0.683, p < 0.001). By contrast, risks of most tumors were similar between the HM group and the TR group, with the exception that the HM group had a lower risk of RCC (HR = 0.608, 95% CI 0.405–0.912, p = 0.016). Moreover, of the missense mutation carriers, the HM group had conferred a lower risk of PHEO (HR = 0.298, 95% CI 0.133–0.607, p = 0.003) and a higher risk of PCT (HR = 1.823, 95% CI 1.093–3.041, p = 0.021) than the nHM group (Supplementary Table S2).
Comparison of age-related risks for VHL-related tumors between patients with nHM mutations and patients with HM or TR mutations as one group (HMTR)
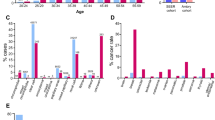
Due to the similarity of tumor risks in the TR and HM groups, we combined patients with HM or TR mutations into one group (HMTR) and compared this group with patients with nHM mutations. As expected, the nHM group was characterized by significantly higher age-related penetrance of PHEO (p < 0.001) and lower penetrance of CHB (p = 0.030), RCC (p = 0.033), and PCT (p < 0.001) compared with the HMTR group (Figure 1). Accordingly, multivariate Cox regression analysis demonstrated that nHM mutations were independently related to a higher age-related risk of PHEO (HR = 3.447, 95% CI 1.906–6.236, p < 0.001) and a lower risk of CHB (HR = 0.700, 95% CI 0.493–0.993, p = 0.045), RCC (HR = 0.610, 95% CI 0.398–0.936, p = 0.024), and PCT (HR = 0.382, 95% CI 0.243–0.600, p < 0.001) compared with the HMTR group (Table 1).
Comparison of age-related risks in patients with nHM and patients with HM mutations or TR (HMTR) mutations. (a) Overall, (b) CHB, (c) RA, (d) RCC, (e) PCT, and (f) PHEO. CHB, central nervous system hemangioblastoma; HM, HIF-α binding site missense mutation; HMTR, missense mutation located in the HIF-α binding site region or truncating mutation; nHM, missense mutation not located in the HIF-α binding site region; PCT, pancreatic cyst or tumor; PHEO, pheochromocytoma; RA, retinal angioma; RCC, renal cell carcinoma; TR, truncating mutation.
Relationship between clinical type and nHM, HM, and TR mutation types
Patients affected by PHEO before death or the end of follow-up were defined as type 2 patients, and the remaining patients were defined as type 1 patients. Patients with nHM mutations were more likely to develop type 2 disease than the HM (χ 2 = 13.02, p < 0.001) and TR (χ 2 = 18.81, p < 0.001) groups, while HM mutation carriers and TR mutation carriers showed similar phenotypic distributions (χ 2 = 0.005, p = 0.946, Supplementary Table S3). Similarly, patients in the combined HM and TR group (HMTR group) were more likely to develop type 1 disease than patients in the nHM group (χ 2 = 23.229, p < 0.001, Supplementary Table S4).
Relationship between survival and nHM, HM, and TR mutation types
The median survival of missense mutation carriers was 6 years longer than that of truncating mutation carriers (median survival 66 y vs. 60 y), but the difference was not statistically significant (p = 0.056, Figure 2a). Likewise, patients with nHM mutations had a significantly better survival than those with HM or TR mutations (median survival 72 y vs. 62 y vs. 60 y, p = 0.013) (Figure 2b). After combining the HM and TR groups into the HMTR group, the median survival of patients with nHM mutations was 12 years longer than that of HMTR mutation carriers (72 y vs. 60 y, p = 0.003) (Figure 2c). In addition, multivariate Cox regression analysis showed that an nHM mutation was an independent protective factor for survival of patients with VHL disease (HR = 0.345, 95% CI 0.165–0.724, p = 0.005) (Table 2).
Comparison of survival and CHB-specific survival in patients with different mutation groups. (a) Comparison of survival in patients with missense mutations and truncating mutations. (b) Comparison of survival in patients with nHM, HM, and TR mutations. (c) Comparison of survival in patients with nHM mutations and HM or TR (HMTR) mutations. (d) Comparison of CHB-specific survival in patients with missense mutations and with truncating mutations. (e) Comparison of CHB-specific survival in patients with nHM, HM, and TR mutations. (f) Comparison of CHB-specific survival in patients with nHM mutations and HM or TR (HMTR) mutations. CHB, central nervous system hemangioblastoma; HM, HIF-α binding site missense mutation; HMTR, missense mutation located in the HIF-α binding site region or truncating mutation; nHM, missense mutation not located in the HIF-α binding site region; TR, truncating mutation.
Relationship between CHB and RCC specific survival and nHM, HM, and TR mutation types
As CHB and RCC were the most common causes of death for patients with VHL disease, we also assessed CHB- and RCC-specific survival in different mutation groups. For CHB-specific survival, we found no difference between missense mutation carriers and truncating mutation carriers (p = 0.388) (Figure 2d). However, patients with nHM mutations had longer CHB-specific survival than patients in the HM group, the TR group (p = 0.002) (Figure 2e), and the combined HMTR group (p < 0.001) (Figure 2f). Multivariate Cox regression analysis confirmed that an nHM mutation was independently related to a better CHB-specific survival in patients with VHL disease (HR = 0.129, 95% CI 0.031–0.532, p = 0.005) (Table 2). For RCC-specific survival, no difference was observed between patients with different types of mutations, probably due to the small number of RCC-specific deaths in this cohort (Supplementary Figure S1).
Discussion
The typical genotype–phenotype correlation divides patients with VHL disease into two subtypes based on the existence of PHEO and links type 1 disease to truncating mutations and type 2 disease to missense mutations. In this study, we found that HM mutations resulted in similar risks of VHL-related PHEO, CHB, PCT, and RA as truncating mutations, suggesting that HM mutations should also be associated with type 1 VHL disease. Thus, we propose the modified genotype–phenotype correlation described in Table 3; VHL gene truncating mutations and HM mutations are related to a lower risk of PHEO and higher risks of CHB and PCT, while nHM mutations are related to a higher risk of PHEO. Meanwhile, patients with TR mutations are more likely to develop RCC than those with HM mutations. In terms of survival, the typical genotype–phenotype correlation fails to predict prognosis based on patient genotype. This modified correlation, however, indicates that the median survival of patients with nHM mutations is 12 years longer than that of patients with HM and TR mutations. Therefore, the new genotype–phenotype correlation seems more reasonable and provides more information for clinical practice.
Since the genotype–phenotype correlation in VHL disease was first described, many researchers have attempted to update it. Cascon et al.22 found that among patients with type 1 VHL disease, those with a contiguous deletion of VHL and the nearby gene BRK1 usually developed hemangioblastoma but had an obvious low risk of RCC, which was designated the type 1B subtype. Inactivation of both VHL and BRK1 in RCC may lead to a growth disadvantage for cancer cells. In another study, Ong et al.21 subdivided missense mutations based on whether the amino acid substitution involved a surface residue or was buried within the protein core and demonstrated that surface missense mutations were related to a higher risk of PHEO. Similar to our study, they discriminated missense mutations at the protein level and emphasized alterations of protein structure, while we directly evaluated alterations of the critical sites for HIF-α binding, the first step for HIF-α ubiquitylation and degradation. However, overlap may exist between the two categorization methods due to the similar mechanisms underlying VHL disease phenotypes.
The pathogenesis of VHL-related PHEO has been reported to be different from that of other highly vascular tumors (RCC, CHB, and RA). In CHB and RCC, upregulated HIF-α expression and consequent overexpression of VEGF and other HIF-related genes are the main causative factors of tumor progression.1,5 Thus, VHL gene mutations disrupting HIF-α binding (HM and TR) can increase the risk of age-related CHB and RCC, consistent with our results. In PHEO, HIF-α (especially HIF-2α) dysregulation results in overexpression of HIF-inducible genes, including glucose transporter 1 (GLUT1), VEGF, EPO, and many others, which play important roles in tumor development.30 Recently, new somatic and germ-line mutations in EPAS1 (HIF2A) have been found to be the cause of a syndrome consisting of multiple and recurrent PHEOs and paragangliomas and polycythemia.31,32 However, the development of PHEO seems to be HIF-independent. In patients with a typical type 2C phenotype, mutant pVHL is still able to degrade HIF, and it has been hypothesized that mutations related to PHEO may induce a gain of function through an intact but altered pVHL.24 In addition, Forman et al.16 reported that mutations in the elongin C binding domain of pVHL were associated with PHEO. These binding sites are implicated in the p53-mediated apoptosis of sympathetic neuronal precursor cells, which can then go on to form PHEO.16,33 In our study, most of the nHM mutations were located in the elongin C binding domain, and the high risk of PHEO in this group validates the above hypothesis.
Gallou et al.34 found that mutations located in missense cluster regions (MCR-1, residues 74–90; MCR-2, residues 130–136) were associated with an increased risk of RCC in French patients with VHL disease. However, Ong et al.21 and our previous work29 found no difference in RCC risk between patients with missense mutations within the MCR regions and those with missense mutations outside of the MCR regions. In this study, patients with HM mutations had similar phenotypes as those with TR mutations, except for a lower risk of RCC. This indicates the existence of other mechanisms in the pathogenesis of RCC besides the dysregulated VHL–HIF pathway. We also found that HM mutations conferred an increased risk of PCT. It is hypothesized that the pathogenesis of PCT is also HIF-dependent. Hammel et al.35 found that the hyperproduction of VEGF may favor pancreatic cyst formation in patients with VHL disease, which supports our results. Thus, HM mutations resulting in HIF-α upregulation and the consequent high VEGF expression are important steps in the development of PCT.
We found that patients with nHM mutation have a much better survival than those with HMTR mutation. This can be explained by the higher risk in HMTR group for CHB, which is the leading cause of death in VHL patients.36 In addition, among patients affected by CHB, different tumor locations may be related to different prognosis because tumors in the medulla grow faster compared with those in the cerebellum.37 We analyzed survival data of the 194 VHL patients with clear information of tumor location in our cohort, and found that individuals with medulla tumors had an obvious worse survival than those with tumors in other locations (p < 0.001, Supplementary Figure S2). Further results showed that patients in the nHM group had a lower incidence of medullary hemangioblastomas than those in the HMTR group (5.9% vs. 20.6%, Supplementary Table S5), but the difference was not statistically significant (p = 0.112). Therefore, we hypothesize that patients with non-HIF-α binding site missense mutation had a lower risk for CHB and lower incidence of medullary hemangioblastomas, which leads to better survival. However, considering that only 34 patients in the nHM group were affected by central nervous system hemangioblastoma (two of them were medullary hemangioblastomas) and the p value (Supplementary Table S5) was not significant, it needs further confirmation.
Although we describe a more reasonable genotype–phenotype correlation for VHL disease, more work needs to be done to improve it. The typical genotype–phenotype correlation subdivides type 2 disease into groups 2A, 2B, and 2C, and we failed to distinguish these groups with specific missense mutations. Further structural and functional studies will help construct a more accurate genotype–phenotype correlation. In type 1 disease, contiguous deletion of VHL and the nearby gene BRK1 is reported to be related to type 1B disease, with an obviously low risk of RCC. We could not subgroup patients with TR mutations for further analysis because we only had three patients with a contiguous VHL and BRK1 deletion.
In this study, which examined a large cohort of Chinese patients with VHL disease, we describe a modified genotype–phenotype correlation according to alterations of the HIF-α binding site in pVHL. This correlation allows us to predict tumor risks and patient survival based on the mutation type they have. The phenotypic differences among mutation subtypes will facilitate studies of the pathogenesis of VHL-related tumors.
References
Gossage L, Eisen T, Maher ER. VHL, the story of a tumour suppressor gene. Nat Rev Cancer 2015;15:55–64.
Maher ER, Neumann HP, Richard S. von Hippel-Lindau disease: a clinical and scientific review. Eur J Hum Genet 2011;19:617–623.
Chou A, Toon C, Pickett J, Gill AJ. von Hippel-Lindau syndrome. Front Horm Res 2013;41:30–49.
Lonser RR, Glenn GM, Walther M et al. von Hippel-Lindau disease. Lancet 2003;361:2059–2067.
Nordstrom-O’Brien M, van der Luijt RB, van Rooijen E et al. Genetic analysis of von Hippel-Lindau disease. Hum Mutat 2010;31:521–537.
Filling-Katz MR, Choyke PL, Oldfield E et al. Central nervous system involvement in von Hippel-Lindau disease. Neurology 1991;41:41–46.
Maher ER, Yates JR, Harries R et al. Clinical features and natural history of von Hippel-Lindau disease. Q J Med 1990;77:1151–1163.
Webster AR, Richards FM, MacRonald FE, Moore AT, Maher ER. An analysis of phenotypic variation in the familial cancer syndrome von Hippel-Lindau disease: evidence for modifier effects. Am J Hum Genet 1998;63:1025–1035.
Ning XH, Zhang N, Li T et al. Telomere shortening is associated with genetic anticipation in Chinese von Hippel-Lindau disease families. Cancer Res 2014;74:3802–3809.
Wang JY, Peng SH, Ning XH et al. Shorter telomere length increases age-related tumor risks in von Hippel-Lindau disease patients. Cancer Med 2017;6:2131–2141.
Latif F, Tory K, Gnarra J et al. Identification of the von Hippel-Lindau disease tumor suppressor gene. Science 1993;260:1317–1320.
Kibel A, Iliopoulos O, DeCaprio JA, Kaelin WG Jr. Binding of the von Hippel-Lindau tumor suppressor protein to elongin B and C. Science 1995;269:1444–1446.
Lee JS, Lee JH, Lee KE et al. Genotype-phenotype analysis of von Hippel-Lindau syndrome in Korean families: HIF-alpha binding site missense mutations elevate age-specific risk for CNS hemangioblastoma. BMC Med Genet 2016;17:48.
Hon WC, Wilson MI, Harlos K et al. Structural basis for the recognition of hydroxyproline in HIF-1 alpha by pVHL. Nature 2002;417:975–978.
Min JH, Yang H, Ivan M, Gertler F, Kaelin WG Jr, Pavletich NP. Structure of an HIF-1alpha -pVHL complex: hydroxyproline recognition in signaling. Science 2002;296:1886–1889.
Forman JR, Worth CL, Bickerton GR, Eisen TG, Blundell TL. Structural bioinformatics mutation analysis reveals genotype-phenotype correlations in von Hippel-Lindau disease and suggests molecular mechanisms of tumorigenesis. Proteins 2009;77:84–96.
Stebbins CE, Kaelin WG Jr, Pavletich NP. Structure of the VHL-ElonginC-ElonginB complex: implications for VHL tumor suppressor function. Science 1999;284:455–461.
Kim WY, Kaelin WG. Role of VHL gene mutation in human cancer. J Clin Oncol 2004;22:4991–5004.
Cybulski C, Krzystolik K, Murgia A et al. Germline mutations in the von Hippel-Lindau (VHL) gene in patients from Poland: disease presentation in patients with deletions of the entire VHL gene. J Med Genet 2002;39:E38.
Sgambati MT, Stolle C, Choyke PL et al. Mosaicism in von Hippel-Lindau disease: lessons from kindreds with germline mutations identified in offspring with mosaic parents. Am J Hum Genet 2000;66:84–91.
Ong KR, Woodward ER, Killick P, Lim C, Macdonald F, Maher ER. Genotype-phenotype correlations in von Hippel-Lindau disease. Hum Mutat 2007;28:143–149.
Cascon A, Escobar B, Montero-Conde C et al. Loss of the actin regulator HSPC300 results in clear cell renal cell carcinoma protection in von Hippel-Lindau patients. Hum Mutat 2007;28:613–621.
McNeill A, Rattenberry E, Barber R, Killick P, MacDonald F, Maher ER. Genotype-phenotype correlations in VHL exon deletions. Am J Med Genet A 2009;149A:2147–2151.
Hoffman MA, Ohh M, Yang H, Klco JM, Ivan M, Kaelin WG Jr. von Hippel-Lindau protein mutants linked to type 2C VHL disease preserve the ability to downregulate HIF. Hum Mol Genet 2001;10:1019–1027.
Clifford SC, Cockman ME, Smallwood AC et al. Contrasting effects on HIF-1alpha regulation by disease-causing pVHL mutations correlate with patterns of tumourigenesis in von Hippel-Lindau disease. Hum Mol Genet 2001;10:1029–1038.
Chittiboina P, Lonser RR. Von Hippel-Lindau disease. Handb Clin Neurol 2015;132:139–156.
Melmon KL, Rosen SW. Lindau’s disease. Review of the literature and study of a large kindred. Am J Med 1964;36:595–617.
Wu P, Zhang N, Wang X et al. Family history of von Hippel-Lindau disease was uncommon in Chinese patients: suggesting the higher frequency of de novo mutations in VHL gene in these patients. J Hum Genet 2012;57:238–243.
Peng S, Shepard MJ, Wang J et al. Genotype-phenotype correlations in Chinese von Hippel-Lindau disease patients. Oncotarget 2017;8:38456–38465.
Young RM, Simon MC. Untuning the tumor metabolic machine: HIF-alpha: pro- and antitumorigenic? Nat Med 2012;18:1024–1025.
Zhuang Z, Yang C, Lorenzo F et al. Somatic HIF2A gain-of-function mutations in paraganglioma with polycythemia. N Engl J Med 2012;367:922–930.
Taieb D, Yang C, Delenne B et al. First report of bilateral pheochromocytoma in the clinical spectrum of HIF2A-related polycythemia-paraganglioma syndrome. J Clin Endocrinol Metab 2013;98:E908–913.
Roe JS, Kim H, Lee SM, Kim ST, Cho EJ, Youn HD. p53 stabilization and transactivation by a von Hippel-Lindau protein. Mol Cell 2006;22:395–405.
Gallou C, Chauveau D, Richard S et al. Genotype-phenotype correlation in von Hippel-Lindau families with renal lesions. Hum Mutat 2004;24:215–224.
Hammel PR, Vilgrain V, Terris B et al. Pancreatic involvement in von Hippel-Lindau disease. The Groupe Francophone d’Etude de la Maladie de von Hippel-Lindau. Gastroenterology 2000;119:1087–1095.
Binderup ML, Jensen AM, Budtz-Jorgensen E, Bisgaard ML. Survival and causes of death in patients with von Hippel-Lindau disease. J Med Genet 2017;54:11–18.
Lonser RR, Butman JA, Huntoon K et al. Prospective natural history study of central nervous system hemangioblastomas in von Hippel-Lindau disease. J Neurosurg 2014;120:1055–1062.
Acknowledgments
This work was supported by the National Natural Science Foundation of China (grant 81572506), the Special Health Development Research Project of Capital (grant 2016-2-4074), and the Beijing Municipal Science and Technology Commission (grant Z151100003915126). We thank Ding-fang Bu, Medical Experiment Center, Peking University First Hospital, for his guidance regarding this study.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Disclosure
The authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Liu, SJ., Wang, JY., Peng, SH. et al. Genotype and phenotype correlation in von Hippel–Lindau disease based on alteration of the HIF-α binding site in VHL protein. Genet Med 20, 1266–1273 (2018). https://doi.org/10.1038/gim.2017.261
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.261
Keywords
This article is cited by
-
VHL Ser65 mutations enhance HIF2α signaling and promote epithelial-mesenchymal transition of renal cancer cells
Cell & Bioscience (2022)
-
Cardio-onco-metabolism: metabolic remodelling in cardiovascular disease and cancer
Nature Reviews Cardiology (2022)
-
Adipose-derived exosomes deliver miR-23a/b to regulate tumor growth in hepatocellular cancer by targeting the VHL/HIF axis
Journal of Physiology and Biochemistry (2019)