Abstract
Purpose
We evaluated the American College of Medical Genetics and Genomics/Association for Molecular Pathology (ACMG/AMP) variant pathogenicity guidelines for internal consistency and compatibility with Bayesian statistical reasoning.
Methods
The ACMG/AMP criteria were translated into a naive Bayesian classifier, assuming four levels of evidence and exponentially scaled odds of pathogenicity. We tested this framework with a range of prior probabilities and odds of pathogenicity.
Results
We modeled the ACMG/AMP guidelines using biologically plausible assumptions. Most ACMG/AMP combining criteria were compatible. One ACMG/AMP likely pathogenic combination was mathematically equivalent to pathogenic and one ACMG/AMP pathogenic combination was actually likely pathogenic. We modeled combinations that include evidence for and against pathogenicity, showing that our approach scored some combinations as pathogenic or likely pathogenic that ACMG/AMP would designate as variant of uncertain significance (VUS).
Conclusion
By transforming the ACMG/AMP guidelines into a Bayesian framework, we provide a mathematical foundation for what was a qualitative heuristic. Only 2 of the 18 existing ACMG/AMP evidence combinations were mathematically inconsistent with the overall framework. Mixed combinations of pathogenic and benign evidence could yield a likely pathogenic, likely benign, or VUS result. This quantitative framework validates the approach adopted by the ACMG/AMP, provides opportunities to further refine evidence categories and combining rules, and supports efforts to automate components of variant pathogenicity assessments.
Similar content being viewed by others
Introduction
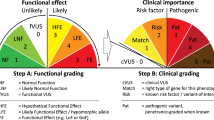
In 2008, Plon et al.1 published recommendations for sequence variant classification for seven cancer susceptibility genes. They coupled quantitative, probability-based thresholds for variant classification to a Bayesian approach for estimating probabilities of pathogenicity for variants of uncertain significance (VUS).1,2 Recently, Richards et al.3 (representing the American College of Genetics and Genomics and the Association for Molecular Pathology, ACMG/AMP) published guidelines for evaluating Mendelian disease gene variants. The ACMG/AMP guidelines codified multiple approaches to variant pathogenicity assessments in use by clinical genetic/genomic testing laboratories. The first stage of their process reduced this assessment into qualitatively distinct evidence types (e.g., functional, genetic, population, in silico.) and stratified the strength of evidence into categories (supporting, moderate, strong, very strong, and standalone). The second stage tallied evidence for or against pathogenicity using “combining criteria,” where various combinations would lead to semiquantitative categorical pathogenicity assessments (pathogenic, likely pathogenic, VUS, likely benign, or benign) similar to the five categories proposed by Plon et al.1
Juxtaposing these two achievements raises two questions. The first is whether the ACMG/AMP rules are internally consistent; the second, whether the systematic, qualitative, categorical ACMG/AMP combining criteria represent a Bayesian heuristic similar to that used for hereditary cancer variants. To address these questions, we analyzed the ACMG/AMP approach in a Bayesian framework.
Materials and methods
Bayesian reasoning starts with a prior probability (Prior_P), which is modified using conditional factors, expressed as probabilities, odds, or likelihoods, to raise or lower the Prior_P. Using Bayes rule to combine these factors results in a posterior probability (Post_P). We interpreted the evidence categories given in Tables 3 and 4 of Richards et al.3 as categorical conditional probabilities or odds of pathogenicity (OP), which could mathematically favor a pathogenic (odds >1) or benign (odds <1) interpretation. To test Bayesian compatibility of the ACMG/AMP combining criteria, we created an Excel spreadsheet that uses Bayes rule to calculate a Post_P from a Prior_P and the OP from four categories of pathogenic and two categories of benign data. Each of the rules for combining criteria to classify sequence variants described in Table 5 of Richards et al.3 was encoded (Supplementary Table S1 online). In principle, this approach could transform their qualitative Table 5 “combining criteria”3 into a formal, quantitative framework, based on several assumptions:
-
1
Each piece of evidence considered by ACMG/AMP was independent, which allows use of a naïve Bayesian classifier with multiple data types expressed individually as OP, the overall OP being obtained by multiplying the odds from each piece of evidence.
-
2
In encoding the combining criteria, we accepted all but one type of ACMG/AMP evidence, building a mathematical model that preserves the relative strengths of each type of evidence. We excluded BA1, “benign standalone,” because it is used as absolute evidence that a variant is benign, irrespective of other evidence, which is contrary to Bayesian reasoning. The BA1 filter is useful for excluding a variant from entering a Bayesian framework, and will be addressed separately by the ClinGen Sequence Variant Interpretation Working Group.
-
3
We transformed the adjectival ACMG/AMP descriptors (supporting, moderate, strong, or very strong) into four conditional probabilities—the odds of pathogenicity relationships. The strength of evidence relationships among the categories supporting

, moderate

, strong

, and very strong

defined by the ACMG/AMP guidelines3 were scaled exponentially such that

,

, etc. The “evidence of benign impact” categories were assigned reciprocal OP to the corresponding pathogenicity categories. If N is the number of criteria with a given strength of evidence category in a classification rule (detailed in Richards et al.3 Table 5), with categories named as subscripted above, the OP attributable to each of the pathogenic data combining rules given in Table 5 is expressible as a specific example of equation (1):

The corresponding equation for the benign data combining rules is expressible as a specific example of equation (2):

-
4
The ACMG/AMP defined the term “likely pathogenic” to mean >90% certainty of a variant being disease-causing, but below a higher “pathogenic” threshold. In Bayesian terms, these translate to a Post_P of >0.90 for likely pathogenic and, if compatible with the definitions established by Plon et al.,1 a higher pathogenic threshold at Post_P >99% certainty of pathogenicity. These probability thresholds, plus the extent to which different combining rules do (or do not) arrive at equivalent Post_Ps, provide criteria for judging the internal consistency of the qualitative rules and their compatibility with a Bayesian framework.
-
5
Obtaining numeric results for purposes of illustration requires that we either select a Prior_P or test a range of Prior_Ps. For simplicity, we have provided a set of calculations using a Prior_P of 0.10. This Prior_P is reasonable for a panel testing scenario where it is likely that the laboratory would encounter on the order of 10 variants in a panel of biologically relevant susceptibility genes and perhaps one of them is the actual pathogenic variant. It is also the approximate empirically measured Prior_P for the combination of missense substitutions, in-frame indels, and proximal splice junction variants in BRCA1 and BRCA2.2,4,5 Finally, the structure of the ACMG/AMP criteria include an implicit lower constraint of 0.10 for a Prior_P, as they specify that if none of the criteria are met, the variant is a VUS, and thus the prior must be between 0.10 and 0.90, which is the Post_P range for a VUS. There are many other potential Prior_Ps for different scenarios if the underlying assumptions of the ACMG/AMP framework were discarded. For the present analysis, as described above, we chose to accept the ACMG/AMP assumptions, but future work should be undertaken to explore whether other approaches that violate those assumptions may yield improved results.
We then tested the ACMG/AMP guidelines in this quantitative framework, under the hypothesis that we could identify an algorithmically plausible value of the exponent X and then biologically plausible combinations of numerical values of (i) the strength of the very strong evidence

(which, combined with the exponent X, determines the strength of the strong, moderate, and supporting evidence) and (ii) the Prior_P that would yield Post_Ps similar to those defined by the ACMG/AMP. Specifically, (i) the Post_Ps of the likely pathogenic combinations would be between 0.90 and 0.99, (ii) the Post_Ps of the likely benign combinations would be between 0.001 and <0.10, (iii) the pathogenic combinations would have Post_Ps >0.99, and (iv) the one benign category has Post_P <0.001. In fact, there was a uniquely optimal value for the exponent X, under which many combinations of

and Prior_P meet these criteria for a large majority of the combining rules. In Table 1 we calculate several post_Ps using X = 2.0, a Prior_P of 0.10, and

=350 (other combinations were calculated; data not shown).
Code availability
Exploration of the Bayesian compatibility of the ACMG/AMP combining criteria that we describe here can be entirely replicated using the Supplementary Table S1.
Results
The classical formulation of Bayes rule is specified in equation (3):

For the purposes of this analysis, we defined

as the probability of pathogenicity (or prior probability (Prior_P),

as the probability of the evidence for pathogenicity,

as the probability of pathogenicity given the evidence (or posterior probability Post_P), and

as the probability of the evidence, given that the variant is pathogenic. This equation can be rearranged to accommodate odds instead of probabilities, and simplified in equation (4):

The spreadsheet that we created to explore Bayesian compatibility of the ACMG/AMP combining criteria was programmed to enable a simple grid search, or parameter sweep, that could be used to either optimize or find plausible values of key variables. In this spreadsheet, the Prior_P and Odds_Path for the “very strong” category

were independently searchable variables. The four categories of pathogenic data included in the spreadsheet were used to model the ACMG/AMP pathogenic evidence categories “supporting,” “moderate,” “strong,” and “very strong.” Each of the ACMG/AMP combining rules was modeled in the spreadsheet.
An obvious constraint on the relative strength of the pathogenic evidence categories is that their OP had to ascend from supporting to very strong. In a naïve Bayesian calculation, the OPs from all of the observational evidence are multiplied together to get an overall OP. Here, this multiplication process implies that if an ACMG/AMP combining rule included two evidence criteria from the same category, then the OP from that category would equal the OP of a single evidence squared. That element of Bayesian reasoning led to the hypothesis that, if the ACMG/AMP combining criteria are Bayes-compatible, then the most natural way to model the relative strength of the evidence categories would be to treat the ordered series of categories as an exponential series with a uniform exponential step from one category to the next. Hence, the relative strength of the ordered evidence categories “supporting,” “moderate,” and “strong” were linked to “very strong” through a single exponent, X, which was a third variable that could be optimized by a grid search.
An initial observation from the grid search was that when the exponent X was set to exactly 2.0, 7 pairs of pathogenic combining rules had identical overall OP and identical Post_Ps, as did 10 pairs of likely pathogenic combining rules; no other value we tested resulted in more than 1 pair of rules with identical Post_Ps. This result was independent to Prior_P and the

. Summing across the exponents of the odds path equations presented in Table 1 (or Supplementary Table S1) from the ACMG/AMP guidelines shows that the matching pairs of combined OP and Post_Ps is a feature that emerges from the repeating structure of the ACMG/AMP combining rules and the arithmetic used to encode them in the exponent of the OP equation.
With the value of the exponent X set at 2.0, we were left with two critical variables to explore, the prior probability (Prior_P) and the combined odds of pathogenicity (OP), which is the totality of the various types of evidence from Table 3 of ACMG/AMP. We tested a range of OP for the O PVst category, as all other levels of evidence strength can be derived from this value via assumption 3. We explored a range of values for

and Prior_P to test whether the ACMG/AMP heuristic could be modeled as a Bayesian formulation. We judged these trials by determining if the calculations could yield a value for Post_P that was internally consistent with the ACMG/AMP rules. We first set out to find a minimum O PVSt . The

of 81 was a unique minimum bound that could simultaneously meet the likely pathogenic and likely benign Post_P thresholds, but only if the Prior_P = 0.25 (Figure 1). This is because OP of 81 are the exact odds required to convert a Prior_P of 0.10 to a Post_P of 0.90 using Bayes’ rule, and the ratio of OP between the likely pathogenic rules (ii, iii, iv, v, and vi) and the likely benign rule (ii) was exactly

.
Permissible solutions to likely pathogenic (LP) and likely benign (LB) equations. X–Y combinations of prior probability of pathogenicity (Prior_P) and odds very strong (O Vst ) lying to the right of the blue (dashed) curve satisfy combining rules likely pathogenic (ii–vi). X–Y combinations of Prior_P and OVst lying to the left of the red (solid) curve satisfy the likely benign combining rule (ii). Values between the two curves and above their intersection at (Prior_P=0.25, O Vst =81) simultaneously meet LP and LB criteria. Values outside of the two curves and below their intersection at (Prior_P=0.25, O Vst =81) meet neither LP nor LB criteria. The black triangle marks the minimum simultaneous solution of LP and LB at Prior_P=0.25, O Vst =81. The black circle marks the solution of LP and LB at Prior_P=0.10, O Vst =350 illustrated in Table 1.
Equations representing each of the ACMG/AMP combining rules are presented in Table 1 and combinations of Prior_P and

that simultaneously satisfy the six rules likely pathogenic (ii, iii, iv, v, and vi) and likely benign (ii) are graphically summarized in Figure 1. Higher values of

expand the range of Prior_P under which a Bayesian interpretation is viable. For example, if

, this re-interpretation is compatible with Prior_Ps of 0.10–0.32. At each of the (Prior_P,

) combinations specified above, broad consistency within the combining rules was evident. Five of the six likely pathogenic rules had Post_Ps of exactly 0.90 and multiple pairs of pathogenic rules had identical Post_Ps that were >0.99 with

and Prior_P = 0.10. We noted that there were two problematic ACMG/AMP combining rules: pathogenic (ii) and likely pathogenic (i) (boldface entries in Table 1). We could identify no combination of Prior_P and O PVSt that would make all 18 rules internally consistent and conclude that the ACMG/AMP framework has a degree of internal inconsistency.
Sequence variants will sometimes present with a mix of evidence for and against pathogenicity. One weakness of the ACMG/AMP combining criteria was that VUS rule (ii) given in the ACMG/AMP guidelines Table 5 was “the criteria for benign and pathogenic are contradictory,”3 without defining the relative strengths of “contradictory” evidence. Although the ACMG/AMP guidelines noted that “expert judgment must be applied when evaluating the full body of evidence to account for differences in the strength of variant evidence,” more guidance on how to address conflicting evidence would reduce variation in the application of expert judgment. Indeed, with the exponent X set to 2.0, equations (1) and (2) can be combined into a single equation (5):

which allows such combinations of evidence types to be combined and calculated. We evaluated four plausible situations where several moderate or strong pathogenic criteria coexist with a supporting benign criterion (e.g., in silico). For example, the combination of one very strong pathogenic criterion (PVS1, null variant), plus two moderate pathogenic criteria (PM2, absent from controls and PM6, assumed de novo) is designated as pathogenic (combination ib from Table 5 in Richards et al.3). If one were to add one supporting benign criterion to these (e.g., adding a benign evidence BP5 “variant found in a case with an alternate molecular basis for disease” to pathogenic rule ib), this would yield a Post_P of 0.997, which remains in the pathogenic range. In contrast, the combination of two strong pathogenic criteria (ACMG/AMP pathogenic combination (ii)), plus one strong benign (adding BS1 “allele frequency is greater than expected for disorder”), yields a posterior probability of 0.675, which is VUS. Indeed, a variety of combinations led to either pathogenic, likely pathogenic, or VUS; these and additional examples are explored in Table 2.
Discussion
While there was no a priori reason for a consistent Bayesian interpretation to emerge from the ACMG/AMP guidelines, it did. Interestingly, the ACMG/AMP committee did not consider Bayes rule when it was formulating its guidelines (H. Rehm and E. Lyon, personal communication). Our analysis showed that the ACMG/AMP guidelines3 delineated a heuristic system for variant classification that is compatible with a formal, quantitative, naive Bayesian classifier. This is an important observation because it provides a mathematical foundation to what could be considered to be (or dismissed as) simply a pragmatic description of existing clinical laboratory practice. Our most important conclusion is that the ACMG/AMP framework is Bayesian in character and fundamentally sound.
We set out to understand the repeating structure within the ACMG/AMP combining criteria and to learn why it was compatible with scaling the relative strength of the ordered evidence categories to the power of 2.0. On close inspection, we noted that multiple pairs of the ACMG/AMP pathogenic combining criteria were related to each other through the rubric of “one criterion from a given strength of evidence category can be replaced with two criteria from the next weaker category.” Indeed, each pair of combining criteria that have the same OP and Post_P feature either an instance of this rubric or else its higher-order version “one criterion from a given strength of evidence category can be replaced with four criteria from the two-steps-weaker category.” Because in a naïve Bayesian calculation the OPs from all of the observational evidence are multiplied together to get an overall OP, this rubric is equivalent to the quantitative assertion that “OP attributed to a given strength of evidence category is equal to the square of OP attributed to the next weaker category.” Hence, the repeated use of this structural rubric is deeply compatible with the exponential scaling to the power 2 that we found through a grid search and used in our subsequent analyses of the Prior_P and

.
Having established that the ACMG/AMP guidelines were Bayesian in character and deriving an equation to formalize that rubric, we next set out to determine if the combining criteria were internally consistent. While our analysis supported most of the “rules for combining criteria” from the ACMG/AMP recommendations,3 two inconsistencies were observed, using a Prior_P of 0.10 and

. First, likely pathogenic rule (i) (one very strong plus one moderate evidence of pathogenicity) was equivalent in strength to pathogenic rules (iiia, iiib, and iiic)—all of these yielded a Post_P of 0.994. Second, and of greater concern, pathogenic rule (ii) (minimally, two strong criteria supporting pathogenicity) was weaker than the other pathogenic rules, yielding a Post_P of 0.975. This was intermediate in strength between the five internally consistent likely pathogenic rules (ii–vi, which yield a Post_P of 0.900) and the next tier of pathogenic rules (iiia, iiib, and iiic). Indeed, likely pathogenic rules (ii and iv) supplemented with one additional moderate criterion in favor of pathogenicity yielded Post_Ps of 0.975; nonetheless, these combinations would be likely pathogenic under the ACMG/AMP guidelines. We could identify no combination of Prior_P and O PVSt that would allow all 18 of the specified evidence combinations (Table 5 in Richards et al.3) to be internally consistent. The two internal inconsistencies we identified could lead to over- or underestimating variant pathogenicity probability, leading to variant misclassification. Laboratories may choose to exercise their expert judgment by requiring that pathogenic rule ii (two strong criteria), which our analysis suggests yields a Post_P of 0.975, may need to be buttressed by the addition of two supporting or one moderate criteria that support pathogenicity to raise it above the threshold of 0.990. Alternatively, the addition of a single supporting criterion of pathogenicity would raise it from 0.975 to 0.988, which can arguably be considered sufficiently close to the 0.99 threshold to warrant a designation as pathogenic.
The clinical consequences of the potential errors that might result from these two inconsistencies are not necessarily symmetric or equivalent and adjustment or revision of these criteria should be considered. As noted above, the asymmetry of potential errors in classification leads us to have greater concern that combining rule pathogenic (ii) overestimates pathogenicity (relative to a somewhat lesser concern that likely pathogenic rule (i) underestimates pathogenicity). In general, we hold the view that incorrectly downgrading a variant from pathogenic to likely pathogenic is less likely to cause a serious clinical error than incorrectly overestimating the pathogenicity of a likely pathogenic variant to be pathogenic. That being said, it is important to remember that the ACMG/AMP criteria were guidelines, not practice standards, and they included the caveat that “expert judgment must be applied when evaluating the full body of evidence.” We encourage laboratories to take our analyses into consideration as a part of their expert judgment when they evaluate variants that fall into one of the two categories that we have identified as being inconsistent. Looking forward, this and other analyses should be taken into account as a part of the deliberative processes of the ClinGen consortium Genomic Variant Working Group and Sequence Variant Working Group as well as future revisions of the ACMG/AMP guidelines.
The second major implication of our work is that there are a number of scenarios where there is a mix of evidence, some supporting and some weighing against pathogenicity. The ACMG/AMP VUS criterion (ii) was defined as “the criteria for benign and pathogenic are contradictory.” This has been widely discussed and it is likely that most laboratories interpret this to apply to situations where the evidence for and against pathogenicity are relatively balanced in their strength. It does not seem reasonable that a single supporting criterion for benign (e.g., BP4, in silico or BP6, reputable source) would sufficiently contravene very strong evidence for pathogenicity, resulting in a determination of VUS. Our approach demonstrates that this is not the case and that supporting evidence against pathogenicity, in combination with strong evidence for pathogenicity, can lead to posterior probabilities in the range of likely pathogenic or even pathogenic. We have provided a single, unified equation that can yield an estimate of pathogenicity for any potential combination of criteria. We have included as a Supplementary File a simple spreadsheet calculator that uses this equation such that inputting any set of criteria leads to a calculated Post_P (Supplementary Table S1). As in the discussion above, expert judgment is always necessary and we do not intend this calculator to be a substitute for that.
The transition from the ACMG/AMP categorical heuristic to a formal, quantitative Bayesian framework provides a number of potential opportunities to refine and evolve these criteria. For example, we accepted the implicit assumption of the ACMG/AMP guidelines that each categorical type of evidence of the ACMG/AMP framework at the same evidence strength level had the same mathematical support. That is to say, e.g., PS1 (same amino acid change), PS2 (de novo), PS3 (functional data), and PS4 (case versus control) all had identical OP. This was a reasonable simplifying assumption for the ACMG/AMP heuristic, but it may be incorrect. Should it be determined that one of these criteria had somewhat more or less strength of evidence than the others, it would be trivial to adapt equation (5) to include any number of terms in the exponent section of the equation, adjusting the denominator of that term to reflect a more precise weighting of a particular piece of evidence. This is not practical in the current ACMG/AMP categorical heuristic because the number of combining criteria rules (Richards et al.3 Table 5) would rise exponentially with respect to the number of distinct evidence weights and the resulting combining criteria would be unwieldy.
A second example of an opportunity to improve the system is based on the recognition that some data types included in the ACMG/AMP guidelines are continuous variables (e.g., segregation), rather than categorical yes/no attributes of variants. Indeed, an inherently unsatisfying attribute of categorical systems is that they tend to misrepresent reality near the category threshold(s). A quantitative system allows continuous evidence types to be integrated into the system as continuous, rather than categorical, criteria. A straightforward extension of our work would be to replace the six exponential terms in equation (5) with a term for each evidence type, such that it expressed

, or odds of that evidence being observed if the variant were pathogenic. Examples of such approaches have been developed for several evidence types in both breast2,5,6,7 and colorectal8,9,10 cancer genetic analyses. Our Bayesian re-interpretation of ACMG/AMP could provide robust mathematical guidelines, e.g., how much segregation corresponds to each category of evidence in favor of pathogenicity. This could be extended to data types not yet quantitatively integrated, e.g., computational predictors and functional assays. In this way, the field of variant classification could transition from a primarily subjective endeavor to a primarily quantitative and objective endeavor.
A third opportunity is that by translating the ACMG/AMP system into a quantitative framework, we can begin to objectively evaluate if the various forms of evidence indeed have the relative weights that the system posits. As noted above, it is straightforward to ask the question of whether the four strong pathogenic criteria are equally strong. Our approach allows the various levels to be compared. Based on our assumptions, O PVSt has an odds of pathogenicity of 350, O PSt of 18.7 (i.e.,

), O PM 4.3 (i.e.,

), O PSu 2.08 (i.e.,

), O BSt 0.053 (i.e.,

), and O BSu 0.48 (i.e.,

). These various forms of evidence can then be tested against experimental data, such as functional assays or population constraint, that either support or argue against pathogenicity to determine if their relative weighting is valid. As well, it would be useful to determine whether real Prior_Ps fall within the range over which the classifiers are valid. With O Vst = 350 and other constraints of the ACMG/AMP structure, this range was 0.10 to 0.32. It is worth noting that the valid range of Prior_Ps is directly dependent on the strength of OP (Figure 1).
Re-interpretation of the ACMG/AMP guidelines in a quantitative Bayesian framework shows that the existing classification system is fundamentally sound, albeit with minor weaknesses. The analysis also identifies important developmental opportunities. Concerns about the relative weakness of the pathogenic rule (ii), and evidence that the stronger pathogenic rules could accommodate one or two supporting criteria for benign, yet still result in a likely pathogenic classification, underline the need for clinical judgment during variant classification using the existing system. Looking forward, transformation of the ACMG/AMP system into a quantitative Bayesian calculator—coupled to refined and more accurately quantitated evidence—could integrate additional data types, increase overall flexibility, and provide a pathway toward automation of the classification process.
References
Plon SE, Eccles DM, Easton D et al. Sequence variant classification and reporting: recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum Mutat 2008;29:1282–1291.
Goldgar DE, Easton DF, Deffenbaugh AM et al. Integrated evaluation of DNA sequence variants of unknown clinical significance: application to BRCA1 and BRCA2. Am J Hum Genet 2004;75:535–544.
Richards S, Aziz N, Bale S et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015;17:405–424.
Abkevich V, Zharkikh A, Deffenbaugh AM et al. Analysis of missense variation in human BRCA1 in the context of interspecific sequence variation. J Med Genet 2004;41:492–507.
Easton DF, Deffenbaugh AM, Pruss D et al. A systematic genetic assessment of 1,433 sequence variants of unknown clinical significance in the BRCA1 and BRCA2 breast cancer-predisposition genes. Am J Hum Genet 2007;81:873–883.
Thompson D, Easton DF, Goldgar DE. A full-likelihood method for the evaluation of causality of sequence variants from family data. Am J Hum Genet 2003;73:652–655.
Tavtigian SV, Byrnes GB, Goldgar DE et al. Classification of rare missense substitutions, using risk surfaces, with genetic- and molecular-epidemiology applications. Hum Mutat 2008;29:1342–1354.
Thompson BA, Goldgar DE, Paterson C et al. A multifactorial likelihood model for MMR gene variant classification incorporating probabilities based on sequence bioinformatics and tumor characteristics: a report from the Colon Cancer Family Registry. Hum Mutat 2013;34:200–209.
Thompson BA, Greenblatt MS, Vallee MP et al. Calibration of multiple in silico tools for predicting pathogenicity of mismatch repair gene missense substitutions. Hum Mutat 2013;34:255–265.
Thompson BA, Spurdle AB, Plazzer JP et al. Application of a 5-tiered scheme for standardized classification of 2,360 unique mismatch repair gene variants in the InSiGHT locus-specific database. Nat Genet 2014;46:107–115.
Acknowledgments
S.V.T. and M.S.G. are supported in part by R01 CA164944 (mismatch repair variants). S.V.T. and K.M.B. are supported in part by P30 CA042014 (Cancer Center Support grant). S.V.T. is supported in part by R01 CA121245 (BRCA gene variants). M.S.G. is supported in part by U01 HG007437 (Clinical Genome Resource). S.M.H. is supported in part by U41 HG006834 (Clinical Genome Resource). S.A.P. is supported in part by U01 HG007436 (Clinical Genome Resource). L.G.B. is supported in part by ZIA HG200387 03 and ZIA HG200388 03 (Intramural Research Program of the National Human Genome Research Institute).
Author information
Authors and Affiliations
Consortia
Contributions
Antonis Antoniou, Cambridge University, Cambridge, UK; Jonathan S. Berg, University of North Carolina, Chapel Hill, NC; Leslie G. Biesecker, National Human Genome Research Institute, National Institutes of Health, Bethesda, MD; co-chair; Steven E. Brenner, University of California, Berkeley, Berkeley, CA; Fergus Couch, Mayo Clinic, Rochester, MN; Garry Cutting, Department of Human Genetics, Johns Hopkins University School of Medicine, Baltimore, MD; Marc S. Greenblatt, University of Vermont; Robert Larner, College of Medicine, Burlington, VT; Steven M. Harrison, Partners HealthCare Laboratory for Molecular Medicine and Harvard Medical School, Boston, MA; co-chair; Christopher D. Heinen, University of Connecticut Health, Farmington, CT; Matthew E. Hurles, Wellcome Trust Sanger Institute, Hinxton, UK; H. Peter Kang, Counsyl, San Francisco, CA; Rachel Karchin, Johns Hopkins University School of Medicine, Baltimore, MD; Robert L. Nussbaum, Invitae, San Francisco, CA; Sharon E. Plon, Baylor College of Medicine, Houston, TX; Heidi L. Rehm, Partners HealthCare Laboratory for Molecular Medicine and Harvard Medical School, Boston, MA; Sean V. Tavtigian, Department of Oncological Science and Huntsman Cancer Institute, University of Utah School of Medicine, Salt Lake City, UT.
Corresponding author
Ethics declarations
Conflict of Interest
R.L.N. receives salary and equity from Invitae, serves as chair of the Rare Disease Therapeutic Area Scientific Review Panel for Pfizer, and is on the Advisory Board of Genome Medical. L.G.B. is an uncompensated adviser for Illumina. The other authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Tavtigian, S.V., Greenblatt, M.S., Harrison, S.M. et al. Modeling the ACMG/AMP variant classification guidelines as a Bayesian classification framework. Genet Med 20, 1054–1060 (2018). https://doi.org/10.1038/gim.2017.210
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.210
Keywords
This article is cited by
-
Explicable prioritization of genetic variants by integration of rule-based and machine learning algorithms for diagnosis of rare Mendelian disorders
Human Genomics (2024)
-
Rapid genomic sequencing for genetic disease diagnosis and therapy in intensive care units: a review
npj Genomic Medicine (2024)
-
Genetic testing for fetal loss of heterozygosity using single nucleotide polymorphism array and whole-exome sequencing
Scientific Reports (2024)
-
Genotype–phenotype correlation study of structural abnormalities in a fetal brain caused by a novel KDM4B variant
Molecular Biology Reports (2024)
-
An AI-based approach driven by genotypes and phenotypes to uplift the diagnostic yield of genetic diseases
Human Genetics (2024)