Abstract
Purpose
CLAPO syndrome is a rare vascular disorder characterized by capillary malformation of the lower lip, lymphatic malformation predominant on the face and neck, asymmetry, and partial/generalized overgrowth. Here we tested the hypothesis that, although the genetic cause is not known, the tissue distribution of the clinical manifestations in CLAPO seems to follow a pattern of somatic mosaicism.
Methods
We clinically evaluated a cohort of 13 patients with CLAPO and screened 20 DNA blood/tissue samples from 9 patients using high-throughput, deep sequencing.
Results
We identified five activating mutations in the PIK3CA gene in affected tissues from 6 of the 9 patients studied; one of the variants (NM_006218.2:c.248T>C; p.Phe83Ser) has not been previously described in developmental disorders.
Conclusion
We describe for the first time the presence of somatic activating PIK3CA mutations in patients with CLAPO. We also report an update of the phenotype and natural history of the syndrome.
Similar content being viewed by others
Introduction
Vascular anomalies represent a broad spectrum of disorders produced by abnormal embryological development of blood vessels. The manifestations of these disorders may be prenatal, congenital, or postnatal. In general, they manifest with a gradual increase in size or extent, proportionally greater than the growth of the patient.1 Vascular anomalies frequently occur as part of recognizable, pleiotropic developmental syndromes,2 which include overgrowth syndromes.3
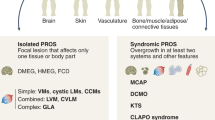
CLAPO syndrome (OMIM 613089)—capillary malformation of the lower lip, lymphatic malformation of the face and neck, asymmetry and partial/generalized overgrowth—is a rare vascular disorder of unknown etiology first described in 2008 in six unrelated patients.4 Only two additional patients have been described.5,6 The tissue distribution of the clinical manifestations in patients with CLAPO is similar to other vascular and overgrowth syndromes caused by somatic mutations such as Proteus syndrome caused by activating mutations in AKT1,7 Sturge–Weber syndrome caused by activating mutations in GNAQ,8 or PROS (PIK3CA related overgrowth spectrum) caused by activating mutations in PIK3CA.9
Somatic mosaicism is defined as the presence of more than one clone of cells with different genotypes, all of which are derived from a single cell. Mosaicism can contribute to variable clinical expressivity of a trait due to tissue-specific involvement. Consequently, the phenotypic spectrum of all these vascular and/or segmental overgrowth syndromes is heterogeneous and complex. Somatic mosaicism is a theoretical possibility in CLAPO, and even though it is a clinically well-distinguished syndrome, it shares several clinical manifestations with previously mentioned syndromes as well as with other entities also caused by mutations in genes within the PI3K-Akt-mTOR pathway. Here we tested the hypothesis that CLAPO could be caused by somatic mutations activating this pathway, and we evaluated the natural history and phenotype of CLAPO and its phenotypic overlap with other PI3K-Akt-mTOR entities.
Materials and methods
Patients and samples
The study was performed at the Institute of Medical and Molecular Genetics and the Vascular Anomalies Center at La Paz Hospital, Madrid (Spain). We retrospectively reviewed the clinical characteristics and follow-up of 13 patients diagnosed with CLAPO between February 2007 and March 2017, including the 6 patients reported in the original description of the syndrome.4 Clinical findings are summarized in Table 1.
Blood samples were collected from all patients, saliva samples from four patients and affected tissue samples from nine patients (Table 1). Affected tissue was defined as any type of vascular malformation obtained as part of the routine treatment, and in the case of cutaneous biopsies, these were obtained in regions with vascular malformation and/or true overgrowth (differentiating it from an asymmetry as a consequence of a vascular malformation). DNA extraction was performed by standard procedures. All studies in this project were approved by the Ethics Committee of the Hospital Universitario La Paz (PI-1919), with informed consent including consent to publish patient photos.
High-throughput sequencing
High-throughput sequencing studies were performed using three distinct custom panels previously available and validated for use in our laboratory. The subsequent analyses of the sequencing data were based on the use of virtual panels (i.e., tiers) with an average expected reading depth of 500 ×. The first panel (Tier1: Mosaic panel) included 20 genes known to cause developmental syndromes in the form of somatic mosaicism. The second panel (Tier2: overgrowth, lymphatic and vascular syndromes (OGLYVAS) panel) was composed of 301 genes associated with lymphatic/vascular malformations and overgrowths of vascular origin, including PI3K-Akt-mTOR pathway genes, as well as 20 genes of the Mosaic panel. The third panel (Tier3: expanded mosaic candidate genes (EMG)) included 1,375 genes, including genes in the OGLYVAS panel. The complete lists of genes can be found in Supplementary Table 1 online.
The three panels were designed, captured, and analyzed with the same tools, protocols, and instruments. The custom panels were designed by using NimbleDesign software (Roche NimbleGen, WI, USA): hg19 NCBI Build 37.1/GRCh37.p13, targeting >98% of all exons (RefSeq) for these genes. For each sample, paired-end (2 × 150 bp reads) libraries were created according to the standard protocols KAPA HTP Library Preparation Kit for Illumina platforms (Roche Sequencing, CA, USA), SeqCap EZ Library SR (Roche NimbleGen, WI, USA) and NEXTflex-96 Pre Capture Combo Kit (Bioo Scientific, TX, USA) for indexing. The captured DNA samples were sequenced on a NextSeq 500 instrument (Illumina, CA, USA) using a HIGH v2 300 cycles cartridge, according to the standard operating protocol.
Bioinformatics analysis pipeline
Data generated by the NextSeq 500 Desktop Sequencer was analyzed using an in-house bioinformatics pipeline for somatic mosaicism detection. BCL files containing base calls were converted into paired FASTQ files using bcl2fastq-v2.15.0.4 software from Illumina (https://github.com/brwnj/bcl2fastq) and preprocessing using Trimmomatic10 for trimming and cropping FASTQ data as well as removing adapters. Subsequently, balanced reads were mapped to hg19/GRCh37 human genome by using Bowtie2 aligner,11 and polymerase chain reaction (PCR) duplicate reads were removed using Picard MarkDuplicates (http://broadinstitute.github.io/picard/). A subsequent local realignment and recalibration of reads was done to correct misalignments due to the presence of indels by using Genome Analysis Toolkit RealignerTargetCreator and IndelRealigner, and BaseRecalibrator methods, respectively. Once this preprocessing was achieved, the flowchart depicted in Supplementary Figure 1 was followed.
The specific approach for mosaic detection included the extraction of the base-pair information for each genomic position from the BAM files using SAMtools mpileup v1.3 (ref.12), facilitating the subsequent single-nucleotide polymorphism/indel calling. The base quality (q value) and mapping quality scores (Q value) were lowered to 0, to characterize mutated alleles amplified in very low proportion and to avoid losing variants in non-unique genomic sequences, since many genes in the capture design have pseudogenes or share high percent identity with other genomic sites.
Variant calling was performed using bcftools v1.3. The low constraints allowed the emergence of many alternate alleles, even though many could be false positives. The strategy of the analysis was to keep all multiallelic sites in the VCF file for later consideration. Some attributes were defined to filter out likely sequencing artifacts or variants with high frequency in samples sequenced in the same run, and others to keep mosaic variants to analyze further on. This was applied in all the samples individually, regardless of the tissue type, generating a corresponding VCF file (see Supplementary Figure 1a).
Subsequently, a germ-line versus somatic variant comparison was undertaken by analyzing the tissue and the blood VCF files from the same patient/sample (see Supplementary Figure 1b). The contrasting of matched tissue and blood samples was crucial to distinguish somatic from germ-line variants, bringing out low allelic fraction from background noise caused by the high-throughput sequencing error rate. The final files encoded global information about alignments. Manual filtering was applied to determine the candidate pathogenic variants. The resulting VCF files were manually visualized with the Integrative Genomics Viewer software to verify mutations and correct annotation.
Validation of high-throughput deep sequencing variants (Sanger, pyrosequencing, and droplet digital polymerase chain reaction)
Sanger sequencing was used to confirm variants present in more than 15% of the reads in the deep sequencing data. Standard PCR and Sanger sequencing were performed using the 96-capillary ABI 3730xl ADN analyzer (Applied Biosystems, CA, USA). Long-range PCR and Sanger sequencing were also used to verify chromosomal location in variants found with high identity to another part of the genome. Pyrosequencing primers were designed using PyroMark software, and QIAGEN reagents and the PyroMark Q96 MD instrument (Qiagen, Hilden, Germany) were used according to manufacturer’s protocol to confirm mosaic variants in the 5% and 15% read fraction range. Droplet digital polymerase chain reaction (ddPCR), able to detect and quantify somatic mosaic variants at frequencies as low as 0.1%,13 was used to detect/validate and quantify variants found on deep sequencing studies in less than 5% of reads or variants above 5% of reads but whose confirmation by pyrosequencing was not conclusive. We also used ddPCR to screen for the three common mutations described in PROS (PIK3CA: NM_006218.2: c.1624G>A, p.Glu542Lys; c.1633G>A, p. Glu545Lys; and c.3140A>G, His1047Arg) in all negative samples by high-throughput sequencing.14 The QX200 Droplet Digital PCR System (Bio-Rad, CA, USA) was used according to the manufacturer´s protocol. Nucleic acid molecules were quantified using a two-color fluorescence detector (FAM channel for mutant allele and VIC channel for wild type) and six assays with well-defined and validated somatic mosaic variants were used as controls to validate the technique.
Results
Clinical features
Longitudinal data for 13 patients with CLAPO was gathered for a total of 70.5 person-years (ranging from 0.8 to 10.2). The mean age at diagnosis was 8.0 years (0.6 to 30.4) and the mean age at the end of this study was 10.4 years (2.2 to 40.5). None of the patients had a relevant family history. Except for one patient of Arab descent whose parents were born in Morocco (P6), all patients in the cohort were Caucasian and had parents from the Iberian Peninsula. The cohort included eight females and five males. Among the main features of CLAPO the only common clinical manifestation was the capillary malformation (CM) of the lower lip (13/13; 100%) (Figure 1); all other features were present at birth or became evident postnatally. The CM of the lower lip was always present in the midline, with a symmetrical distribution, ranging from 2 to 11 cm, with well-defined borders in several patients, and often affecting the portion of the skin under the lip or the intra-oral mucosa. We observed three distinct patterns of midline CM on the lower lip: (i) five patients (P2, P5, P6, P8, P9) had a narrow midline CM with reddish color, (ii) seven patients (P1, P3, P4, P7, P10, P11, P12) had a CM that involved the entire lower lip with a predominant brown/purple color, and (iii) one patient (P13) had a CM including the central inner area of the lower lip without external skin involvement under the vermilion. In our experience, untreated isolated capillary malformations (classically called port-wine stains) evolve by a progressive thickening and darkening of the malformation due to a progressive vessel dilation, which can, in turn, produce skin overgrowth. Nonetheless, we did not observe this type of progression in the complete cohort of 13 patients with CLAPO. However, as some of the patients are very young and have been followed up for a short period of time, thickening and darkening of the CM on the lower lip in these patients cannot be ruled out in the future. Remarkably, what we did observe, based on photographs and information provided by patients and relatives, is a decreased intensity in the CM of the lower lip, usually between months 6 and 18 of age, in four (P1–3, P6) of the six patients followed more than 9 years. While midline CMs of the face are frequently correlated with lesions of the brain in patients with megalencephaly-capillary malformation syndrome (MCAP),15 no psychomotor delay or intellectual deficit was apparent in CLAPO patients.
Capillary malformation of the lower lip, the hallmark of CLAPO syndrome, in the 13 patients reviewed. It is constantly a congenital feature, always present in the midline, symmetrical and well defined. There are three patterns: narrow midline capillary malformation (i.e., P6), entire lower lip affected (i.e. P7), and central inner area without external skin involvement (P13).
The second major clinical feature was the presence of lymphatic malformations (LMs), observed in 12 of 13 patients (92%). In 10 patients (P3–8, P10–13) the LM involved the lip, oral mucosa, neck, and tongue, seen in 5 of 10 with right sided predominance (P5, P6, P10–12). Another two patients (P1, P2) had unilateral LM on the lower limbs, one of which (P1) was associated with lymphedema. At birth, the tongue had a symmetrical midline combined capillary/lymphatic/venous malformation in 8 of 13 patients (P3–6, P8, P10, P12, P13) with CLAPO, which was mild, well-defined, and affected the tip of the tongue. In 5 of 8 patients (P3–5, P8, P13), the tongue lesions evolved over time, causing growth of the affected area and severe hemorrhagic events with episodes of acute inflammation, triggering and aggravating symptoms. On the other hand, P2 had a left thigh LM that was not noted until 15 years of age, indicating that an apparent absence of congenital LM does not exclude a later onset. Sporadic LMs are rare and arise more often with craniofacial venous malformations.16 In our cohort LMs of the neck, tongue, and/or limb were also observed in association with venous malformations in 11 patients (all but P2 and P7), which showed progressive increase in size and in no case involution.
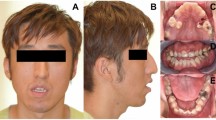
In this cohort the asymmetry in 8 of 13 patients (P4–6, P8, P10–13) actually reflected a direct consequence of the presence of a LM on the face/neck, although in three patients (P1–3) there was a true asymmetric overgrowth with the presence of bony hypertrophy. In addition to the facial asymmetry, P10 had asymmetry caused by left limb undergrowth. P1 had macrodactyly and overgrowth of the right side of the body (Figure 2), and P2 had a postnatal increase in the length of the left leg. P1 and P2 were initially referred with a diagnosis of Klippel–Trenaunay syndrome.
Recently, P9 suffered a large presacral ganglioneuroma that was successfully resected, and P10 had severe failure to thrive needing gastrostomy. Those findings can eventually expand the phenotype of CLAPO syndrome.
Molecular results
Our main hypothesis was that somatic mutations were the genetic cause of CLAPO. Accordingly, we screened 20 DNA blood/saliva/tissue samples from 9 patients (P1, P2, P4, P6–10, and P13) with CLAPO using custom high-throughput deep sequencing panels including genes associated with vascular malformations and overgrowth, and a custom bioinformatic pipeline for somatic mosaic detection with a tiered analysis approach. Average read depth for Tier1 was 535 ×, for Tier2 342 ×, and for Tier3 404 ×, after removing duplicates. Variants were excluded as disease candidates by their presence in >0.01 population frequency (1000 Genomes project (http://www.internationalgenome.org/), the National Heart, Lung, and Blood Institute Exome Variant Server (http://evs.gs.washington.edu/EVS), the Exome Aggregation Consortium (ExAC; http://exac.broadinstitute.org)), by pathogenicity predictors, and by using new custom parameters developed here for somatic mosaic detection (AVAF, SampleRun). After filtering, mean variants per sample for Tier1 was 5.2, for Tier2 31.3, and for Tier3 175.6.
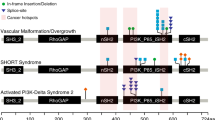
Our analysis identified PIK3CA variants in six of nine (66%) patients with CLAPO (P1, P2, P6, P9, P10, and P13). All mutations were somatic missense single-nucleotide variations with a range of mosaicism between 1.26% and 16%. We identified a total of five distinct mutations in PIK3CA (NM_006218.2), including one hotspot mutation (c.1624G>A;p.Glu542Lys); two recurrent, strong (gain of function) mutations (c.3140A>T;p.His1047Leu, c.1258T>C;p.Cys420Arg); one previously described mutation in patients with macrodactyly (c.344G>C;p.Arg115Pro); and one novel somatic PIK3CA mutation (c.248T>C; p.Phe83Ser) not previously described in developmental disorders (Figure 3).
All variants detected were confirmed using at least one orthogonal method based on the mosaicism percentage of the alternative allele from the next-generation candidate gene sequencing. None of the mutations detected in the patients were present (by Sanger sequencing) in the blood samples of the patient’s parents. Molecular data and variant information were collected in Table 2. In the affected tissue samples from patients P4, P7, and P8 no mutation was detected. Mean PIK3CA reading depth of 427 × (171–603) for P4, 399 × (151–573) for P7, and 311 × (60–583) for P8 was obtained.
Discussion
In the present report we describe for the first time the presence of somatic activating PIK3CA mutations in patients with CLAPO, and we report an update of the phenotype and natural history of the syndrome.
Somatic mosaic mutations in PIK3CA have been detected in six of nine patients with CLAPO. All mutations were detected in affected tissue, in percentages ranging from 3 to 16%, and in one of the patients (P9) the mutation was also mosaic at a level of 1.26% in a blood sample. All the mutations detected are missense variants, four of which have previously been reported as causing the PROS spectrum9,14 and detected in different types of cancer (COSMIC database; http://cancer.sanger.ac.uk/). We also identified a novel PIK3CA mutation (Phe83Ser), not previously described in development disorders, but then again reported in the COSMIC database in three cancer samples.
The family of syndromes caused by activating PIK3CA mutations includes well-known and distinct entities such as MCAP, CLOVES (congenital lipomatous overgrowth, vascular malformations, epidermal nevi, and skeletal/spinal abnormalities), isolated LM, isolated adipose fibrodysplasia, and hemimegalencephaly, among others.9 In addition to the distribution pattern, some of the syndromes included in PROS have overlapping clinical findings with CLAPO, mainly capillary and lymphatic malformations and segmental overgrowth. However, sharing a common mutated gene does not make their formerly differentiated classification obsolete since it has implications for clinical diagnosis, evolution, and follow-up.
Our findings suggest that somatic activating PIK3CA mutations can be responsible for the combination of capillary, lymphatic, and venous malformation as well as the overgrowth features characteristic of CLAPO syndrome. Given the overlapping phenotypic features between CLAPO and PROS (mainly with MCAP and CLOVES), and the presence of somatic activating PIK3CA mutations, we postulate that CLAPO syndrome might be included into the PROS spectrum.
The multiple or reticulate and diffuse CM present in different parts of the body appear to be similar in both CLAPO and MCAP. However, one of the main clinical manifestations in CLAPO is the CM of the lower lip, a feature that is not considered characteristic of MCAP, in which midline CMs are most frequently described in upper lip, glabella, and philtrum. Anatomically, the differential involvement of the lower lip or the upper lip is distinctive and is relevant as they have distinct embryological origins and clinical implications. The forehead, the dorsum apex of the nose, and the central part of the upper lip derive from the frontonasal prominence, whereas the chin, lower lip, and lower cheek regions derive from the mandibular prominences. These distinct embryological origins might also explain the co-occurrence of brain alterations and CM of the upper lip in MCAP and the absence of neurological involvement in CLAPO.
The second main feature in CLAPO, the LM, is also frequent in CLOVES and facial infiltrating lipomatosis. While 2 of the 13 patients described here had a LM of the legs, this type of malformation occurs in CLAPO more frequently in the cervicofacial region and tongue, where there is a clear nonrandom association with capillary or venous malformations. Venous malformations were not originally described as a main feature in CLAPO, but we now know that they are a rather frequent feature, and it has recently been described that this type of malformation can arise sporadically as a consequence of somatic activating mutations in PIK3CA,17 which has also been described for isolated LM.18 Since it is possible that both lymphatic and venous malformations are not apparent congenitally, the presence of CM in the lower lip should make us aware of a possible later appearance of these two types of vascular malformations. This specific and nonrandom combination of vascular malformations highlights the utility of the clinical diagnosis in patients with CLAPO.
Overgrowth is not always evident in CLAPO, as none of the patients had macrocephaly, the involvement was segmental and rarely generalized, and in the case of facial asymmetry it appears to be caused by a vascular component. As with all overgrowth syndromes, it is important to distinguish in CLAPO and PROS what is apparent asymmetry from true overgrowth caused by hyperplasia or hypertrophy. In addition, no patient with CLAPO has lipomatous overgrowth, which is a common feature in the PROS spectrum. In spite of the fact that partial/generalized overgrowth is part of the CLAPO acronym, its low frequency in this cohort of patients suggests that it should not be considered a major clinical criterion for the diagnosis of CLAPO, despite being a frequent (but not mandatory) feature in PROS.
Considering the clinical characteristics and the natural history described in our cohort of patients, CLAPO could be classified as an entity with intermediate severity in the PROS spectrum, sharing characteristics mainly with MCAP and CLOVES, and certainly with isolated forms of lymphatic and venous malformations.
One of the major challenges in somatic mosaicism is the detection of altered alleles in low mosaics in affected tissues. Correspondingly, the molecular diagnosis of CLAPO patients is not trivial. One limitation comes from the requirement to obtain affected tissue. Thus, the lack of success detecting PIK3CA mutations in three of nine of the molecularly studied patients with CLAPO does not implicate the absence of mutations in this gene, nor does it exclude the molecular diagnosis of CLAPO.
Another challenge is the bioinformatic method for the detection of somatic mosaicism. There are several computational approaches to detect somatic variants, but they cannot be applied to this problem since the starting data or the genomic landscape is different. Strelka19 for single-nucleotide variation discovery or Mutect20 for indel discovery are two of the most used variant callers for somatic variant detection. However, neither of them was developed to detect variants in mosaic disorders. Thanks to the computational approach applied here, we could rescue reads with low mapping quality score due to high percent identity with other genomic regions, and call variants that were present both in somatic and mosaic state.
In conclusion, this study documents the clinical features and natural history in a well-defined cohort of patients with CLAPO, and detected activating PIK3CA mutations in six of nine studied affected tissues. We detected PIK3CA activating variants previously described as causing the PROS spectrum, together with a clinical distribution pattern distinctive of somatic mosaicism, and a constellation of nonrandom clinical manifestations that, although in combination are specific to CLAPO, are also frequent in other syndromes within the PROS spectrum. We conclude that CLAPO belongs to the PROS family of somatic syndromes. In clinical terms, the differences between CLAPO and other PROS spectrum phenotypes demonstrate that the presence of a CM in the upper lip should raise suspicion of possible neurological involvement, whereas the CM in the lower lip may be associated with later occurrence of lymphatic/venous malformations. This could reflect their distinct embryological origin but does not imply that a patient with PROS and without upper lip CM should not be monitored for possible neurological compromise. In diagnostic terms, the approach in CLAPO requires the use of appropriate samples and bioinformatic algorithms, allowing the detection of somatic mosaic variants. Because CLAPO is caused by activating mutations in PIK3CA, affected patients may benefit from inhibitors of this pathway. Therapeutic trials of this disorder should be undertaken.
References
Del Pozo J, Gómez-Tellado M, López-Gutiérrez JC. Vascular malformations in childhood. Actas Dermosifiliogr 2012;103:661–678.
Duffy K. Genetics and syndromes associated with vascular malformations. Pediatr Clin North Am 2010;57:1111–1120.
Hagen SL, Hook KP. Overgrowth syndromes with vascular malformations. Semin Cutan Med Surg 2016;35:161–169.
López-Gutiérrez JC, Lapunzina P. Capillary malformation of the lower lip, lymphatic malformation of the face and neck, asymmetry and partial/generalized overgrowth (CLAPO): report of six cases of a new syndrome/association. Am J Med Genet A 2008;146A:2583–2588.
Krämer D, Cossio ML, Whittle C. CLAPO syndrome: case report. Int J Dermatol 2016;55:1149–1150.
Flores-Terry MÁ, Zamberk-Majlis P, Cortina-de la Calle MP, García-Arpa M. CLAPO syndrome. Actas Dermosifiliogr; e-pub ahead of print 15 May 2017.
Lindhurst MJ, Sapp JC, Teer JK et al. A mosaic activating mutation in AKT1 associated with the Proteus syndrome. N Engl J Med 2011;365:611–619.
Shirley MD, Tang H, Fallione CJ et al. Sturge-Weber syndrome and port-wine stains caused by somatic mutation in GNAQ. N Engl J Med 2013;368:1971–1979.
Keppler-Noreuil KM, Rios JJ, Parker VE et al. PIK3CA-related overgrowth spectrum (PROS): diagnostic and testing eligibility criteria, differential diagnosis, and evaluation. Am J Med Genet A 2015;167A:287–295.
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 2014;30:2114–2120.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods 2012;9:357–359.
Li H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 2011;27:2987–2993.
Hindson BJ, Ness KD, Masquelier DA et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal Chem 2011;83:8604–8610.
Mirzaa G, Timms AE, Conti V et al. PIK3CA-associated developmental disorders exhibit distinct classes of mutations with variable expression and tissue distribution. JCI Insight 2016;1:e87623.
Martínez-Glez V, Romanelli V, Mori MA et al. Macrocephaly-capillary malformation: analysis of 13 patients and review of the diagnostic criteria. Am J Med Genet A 2010;152A:3101–3106.
Ewing MJ, Zreik RT, Donner LR, Zehr KJ. Large lymphaticovenous malformation resection. Interact Cardiovasc Thorac Surg 2013;17:205–206.
Limaye N, Kangas J, Mendola A et al. Somatic activating PIK3CA mutations cause venous malformation. Am J Hum Genet 2015;97:914–921.
Osborn AJ, Dickie P, Neilson DE et al. Activating PIK3CA alleles and lymphangiogenic phenotype of lymphatic endothelial cells isolated from lymphatic malformations. Hum Mol Genet 2015;24:926–938.
Saunders CT, Wong WS, Swamy S, Becq J, Murray LJ, Cheetham RK. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 2012;28:1811–1817.
Cibulskis K, Lawrence MS, Carter SL et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol 2013;31:213–219.
Yeung KS, Ip JJ, Chow CP et al. Somatic PIK3CA mutations in seven patients with PIK3CA-related overgrowth spectrum. Am J Med Genet A 2017;173:978–984.
Kuentz P, St-Onge J, Duffourd Y et al. Moleculardiagnosis of PIK3CA-related overgrowth spectrum (PROS) in 162 patients and recommendations for genetic testing. Genet Med 2017;19:989–997.
Hucthagowder V, Shenoy A, Corliss M et al. Utility of clinical high-depth next generation sequencing for somatic variant detection in the PIK3CA-related overgrowth spectrum. Clin Genet 2017;91:79–85.
Chang F, Liu L, Fang E et al. Molecular diagnosis of mosaic overgrowth syndromes using a custom-designed next-generation sequencing panel. J Mol Diagn 2017;19:613–624.
Vahidnezhad H, Youssefian L, Baghdadi T et al. Phenotypic heterogeneity in PIK3CA-related overgrowth spectrum (PROS). Br J Dermatol 2016;175:810–814.
Tripolszki K, Knox R, Parker V et al. Somatic mosaicism of the PIK3CA gene identified in a Hungarian girl with macrodactyly and syndactyly. Eur J Med Genet 2016;59:223–226.
Youssefian L, Vahidnezhad H, Baghdadi T et al. Fibroadipose hyperplasia versus Proteus syndrome: segmental overgrowth with a mosaic mutation in the PIK3CA gene. J Invest Dermatol 2015;135:1450–1453.
Luks VL, Kamitaki N, Vivero MP et al. Lymphatic and other vascular malformative/overgrowth disorders are caused by somatic mutations in PIK3CA. J Pediatr 2015;166:1048–1054.
Loconte DC, Grossi V, Bozzao C et al. Molecular and functional characterization of three different postzygotic mutations in PIK3CA-related overgrowth spectrum (PROS) patients: effects on PI3K/AKT/mTOR signaling and sensitivity to PIK3 inhibitors. PloS One 2015;10:e0123092.
Boscolo E, Coma S, Luks VL et al. AKT hyper-phosphorylation associated with PI3K mutations in lymphatic endothelial cells from a patient with lymphatic malformation. Angiogenesis 2015;18:151–162.
Maclellan RA, Luks VL, Vivero MP et al. PIK3CA activating mutations in facial infiltrating lipomatosis. Plast Reconstr Surg 2014;133:12e–19e.
Keppler-Noreuil KM, Sapp JC, Lindhurst MJ et al. Clinical delineation and natural history of the PIK3CA-related overgrowth spectrum. Am J Med Genet A 2014;164A:1713–1733.
Emrick LT, Murphy L, Shamshirsaz AA et al. Prenatal diagnosis of CLOVES syndrome confirmed by detection of a mosaic PIK3CA mutation in cultured amniocytes. Am J Med Genet A 2014;164A:2633–2637.
Bornstein E, Bacino CA, Maliszewski K et al. Isolated fetal macrodactyly: phenotypic and genetic disparities in mosaic overgrowth syndrome. J Ultrasound Med 2014;33:1305–1307.
Rios JJ, Paria N, Burns DK et al. Somatic gain-of-function mutations in PIK3CA in patients with macrodactyly. Hum Mol Genet 2013;22:444–451.
Lindhurst MJ, Parker VE, Payne F et al. Mosaic overgrowth with fibroadipose hyperplasia is caused by somatic activating mutations in PIK3CA. Nat Genet 2012;44:928–933.
Kurek KC, Luks VL, Ayturk UM et al. Somatic mosaic activating mutations in PIK3CA cause CLOVES syndrome. Am J Hum Genet 2012;90:1108–1115.
Leiter SM, Parker VE, Welters A et al. Hypoinsulinaemic, hypoketotic hypoglycaemia due to mosaic genetic activation of PI3-kinase. Eur J Endocrinol 2017;177:175–186.
Acknowledgments
This research was supported by the project “Genetics of vascular and lymphatic malformations” financed with funds donated by Asociación Ultrafondo and Villareal FC, cofinanced by project IP-17 from the funding call “Todos Somos Raros” (Telemaraton TVE promoted by Fundación Isabel Gemio, Federación ASEM, and Federación Española de Enfermedades Raras), cofinanced by the Instituto de Salud Carlos III, FEDER FUNDS FIS PI15/01481, and IIS-Fundación Jiménez Díaz UAM Genome Medicine Chair. We are grateful to the patients and their families. We want to honor the memory of Angel Llopis (affected by a non-CLAPO-related vascular anomaly) and thank his father, Fernando. We also thank Leslie Biesecker of the National Human Genome Research Institute, National Institutes of Health, for his invaluable contribution to critically revising this work.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Rodriguez-Laguna, L., Ibañez, K., Gordo, G. et al. CLAPO syndrome: identification of somatic activating PIK3CA mutations and delineation of the natural history and phenotype. Genet Med 20, 882–889 (2018). https://doi.org/10.1038/gim.2017.200
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.200
Keywords
This article is cited by
-
Somatic mutation spectrum of a Chinese cohort of pediatrics with vascular malformations
Orphanet Journal of Rare Diseases (2023)
-
French national diagnosis and care protocol (PNDS, protocole national de diagnostic et de soins): cystic lymphatic malformations
Orphanet Journal of Rare Diseases (2023)
-
Genetik, Diagnostik und Klinik des primären Lymphödems
Die Dermatologie (2023)
-
PIK3CA-related overgrowth with an uncommon phenotype: case report
Italian Journal of Pediatrics (2022)
-
When, where and which PIK3CA mutations are pathogenic in congenital disorders
Nature Cardiovascular Research (2022)