Abstract
Purpose
To describe examples of missed pathogenic variants on whole-exome sequencing (WES) and the importance of deep phenotyping for further diagnostic testing.
Methods
Guided by phenotypic information, three children with negative WES underwent targeted single-gene testing.
Results
Individual 1 had a clinical diagnosis consistent with infantile systemic hyalinosis, although WES and a next-generation sequencing (NGS)-based ANTXR2 test were negative. Sanger sequencing of ANTXR2 revealed a homozygous single base pair insertion, previously missed by the WES variant caller software. Individual 2 had neurodevelopmental regression and cerebellar atrophy, with no diagnosis on WES. New clinical findings prompted Sanger sequencing and copy number testing of PLA2G6. A novel homozygous deletion of the noncoding exon 1 (not included in the WES capture kit) was detected, with extension into the promoter, confirming the clinical suspicion of infantile neuroaxonal dystrophy. Individual 3 had progressive ataxia, spasticity, and magnetic resonance image changes of vanishing white matter leukoencephalopathy. An NGS leukodystrophy gene panel and WES showed a heterozygous pathogenic variant in EIF2B5; no deletions/duplications were detected. Sanger sequencing of EIF2B5 showed a frameshift indel, probably missed owing to failure of alignment.
Conclusion
These cases illustrate potential pitfalls of WES/NGS testing and the importance of phenotype-guided molecular testing in yielding diagnoses.
Similar content being viewed by others
Introduction
Whole-exome sequencing (WES) has revolutionized clinical genetics by providing a comprehensive and agnostic method for patient evaluation.1 Diagnostic rates vary from 25 to 50% and WES has allowed new disease-gene identification and insights into the phenotypic and genetic heterogeneity of Mendelian disorders.2, 3, 4 WES has quickly become part of the standard repertoire of genetic testing, with a prevailing sense that a negative result indicates that disorders in the differential diagnoses have been effectively excluded. We describe three individuals in whom WES and targeted next-generation sequencing (NGS)-based testing were nondiagnostic. Phenotype reassessment and use of additional data, such as the single-nucleotide polymorphism (SNP) microarray data, helped determine the next steps in the diagnostic process. Targeted single-gene Sanger sequencing and deletion/duplication analyses identified pathogenic variants for the clinically suspected genetic disorder in all three individuals. We provide insights into the reasons for negative WES results, and, in an era when genomic technology tends to drive the diagnostic process, we highlight the importance of revisiting clinical information for additional targeted testing.
Patients, methods, and results
Individuals 1 and 2 were evaluated at the Duke clinical site of the National Institutes of Health Undiagnosed Diseases Network (UDN) (https://undiagnosed.hms.harvard.edu) and individual 3 at the Duke Genome Sequencing Clinic.
Individual 1
An 18-month-old Mexican female with progressive joint contractures and related morbidity was evaluated due to extensive skin plaques and subcutaneous and gingival nodules; a biopsy demonstrated dermal accumulation of amorphous hyaline material. Review of a duodenal biopsy showed dilated lymphatics and mucosal edema, consistent with clinical symptoms of protein-losing enteropathy. She had intact cognitive skills, ruling out alternative diagnoses such as Farber and Winchester syndromes. The parents reported a common ancestor in Mexico.
This extended phenotype was consistent with infantile systemic hyalinosis (OMIM 228600), an autosomal recessive disorder due to loss-of-function variants in ANTXR2, leading to widespread progressive accumulation of hyaline material and childhood death.5
Pertinent previous genetic testing: A SNP microarray identified 64.2 Mb regions of homozygosity (ROH), including the ANTXR2 locus. An initial NGS-based sequencing of the exons and flanking splice junctions of ANTXR2, followed by a proband-only WES through a commercial laboratory, were negative.
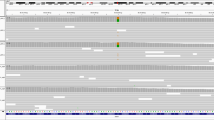
Results of post-WES genetic testing in the Undiagnosed Diseases Network: Review of the SNP microarray did not identify deletions in the regions of ROH. Review of the NGS-based ANTXR2 sequencing and WES data confirmed that 98.5% of the coding regions of ANTXR2 gene were covered at >10 ×, except exon 1 (85.6%). Sanger sequencing revealed a homozygous pathogenic variant in exon 13 (c.1073dupC). Parental studies confirmed trans configuration. Upon discussion with the commercial laboratory that had performed the WES and NGS-based testing, it appeared that their variant-calling software had not reported the variant. Its location adjacent to a homopolymeric repeat region and a common SNP could have contributed to low mapping quality, leading to failure of variant calling (Figure 1). Subsequently, we obtained the BAM files, and manual inspection of the data by the Undiagnosed Diseases Network bioinformatician confirmed the presence of the variant.
Internal and external read alignment at the ANTXR2 variant site in individual 1. Integrated Genome Variant browser showing condensed read alignment for (a) internally realigned data and (b) the BAM file provided by the clinical laboratory. Reads are shown stacked together, with colors indicating a mismatch to the reference sequence. Almost all reads show the pathogenic insertion (c.1073dupC, 22:g.80905986dupG) as a purple mark. The insertion call may be considered low quality because of the adjacent homopolymer repeat and nearby variant. The nearby single-nucleotide polymorphism (c.1069G>C, 22:g.80905990C>G) can be seen by the majority of reads containing the orange G nucleotide change. This variant can also contribute to lower mapping quality and can affect variant quality at the insertion.
Individual 2
A 3.5-year-old girl of Pakistani origin exhibited developmental regression at 16 months of age, cerebellar atrophy, and a negative trio WES (proband and parents). She had lost the ability to cruise, crawl, sit, speak, and eat by mouth. The parents were first cousins, and the proband had two first cousins once removed who died at age two years after neurodevelopmental regression.
The patient had optic atrophy, profound generalized hypotonia, minimal spontaneous movements, tongue fasciculations, and diminished Achilles reflexes, in contrast to previously observed hypertonia with generalized hyperreflexia at age 2.5 years. Review of brain magnetic resonance images obtained at 2 and 2.5 years of age revealed stable white matter volume loss of the vermis and cerebellar hemispheres, a normal pons and no iron accumulation (Supplementary Materials and Methods, Supplementary Figure S1 online). The new clinical finding of peripheral nerve involvement led us to consider infantile neuroaxonal dystrophy (OMIM 256600), a disorder of neurodevelopmental regression in childhood and early death, caused by biallelic variants in PLA2G6.6, 7
Pertinent previous genetic testing: Metabolic laboratory tests and an ataxia gene panel (42 genes) were negative. Trio WES through a commercial laboratory detected a homozygous missense variant of unknown significance in RPGRIP1L, but the clinical course and brain magnetic resonance image findings were not consistent with Joubert syndrome.
Results of post-WES genetic testing in Undiagnosed Diseases Network: A review of the SNP microarray identified several ROH in > 4.6% of the genome; 12 genes within these regions, including PLA2G6, were associated with cerebellar atrophy and developmental regression. Manual inspection of the WES BAM files found no functionally significant single nucleotide or copy number variants in these 12 genes. Sanger sequencing and multiplex ligation-dependent probe amplification (MLPA) for deletions/duplications was performed for the five genes within the ROH with the greatest phenotypic overlap: PLA2G6, AC02, BCS1L, NDUFA11, and ADSL. Sanger sequencing was normal for all, but exon 1 of PLA2G6 failed to sequence. MLPA showed a novel homozygous deletion of the noncoding exon 1 in PLA2G6 (Figure 2a and Supplementary Figure S2). Follow-up MLPA analysis of the parents confirmed that they were carriers of the deletion.
PLA2G6 deletion mapping and mRNA expression. (a) Multiplex ligation-dependent probe amplification (MLPA) analysis of PLA2G6. Normalized MLPA data showing the homozygous deletion of the noncoding exon 1 of the PLA2G6 gene leading to total absence of amplification and hence a ratio of zero for the probe covering this region. (b) Quantitative real-time polymerase chain reaction (qRT-PCR) analysis of PLA2G6 messenger RNA expression in blood. Results are expressed as means ± SD; messenger RNA levels were quantitated with real-time PCR and normalized to the level of GAPDH. The figure represents real-time PCR quantification of PLA2G6 gene expression in the patient compared to controls. All results were done in triplicates. P value was calculated by Student’s t-test.
The breakpoint junction of the PLA2G6 deletion was amplified by long-range polymerase chain reaction (PCR; see Supplementary Methods). The deletion included 2431-bp in the 5’UTR region of PLA2G6 and revealed a 7-bp insertion at the breakpoint junction (c.-545_-46 + 1931delinsCGATCTC) (Supplementary Figure S3). Fine mapping analysis8 demonstrated that the deletion encompassed a portion of the promoter region of the gene. Quantitative real-time PCR showed that messenger RNA expression of PLA2G6 was significantly lower in the patient’s blood compared with unaffected controls (p < 0.01) (Figure 2b). Review of the WES data revealed that the capture kit did not include the noncoding exon 1.
Individual 3
An 8-year-old Caucasian female was evaluated for symptoms of ataxia, seizures, and white matter disease. At age 3 years, she developed frequent falls and progressive decline in fine motor skills, speech, and short-term memory. Generalized seizures started at 7.5 years. Exam revealed dysarthria, lower extremity spasticity, hyperreflexia, clonus, and a wide-based gait. Brain magnetic resonance image showed diffuse symmetric non-enhancing signal abnormalities involving both cerebral hemispheres with volume loss (Supplementary Figure S4). The features were consistent with leukoencephalopathy with vanishing white matter (VWM) (OMIM 603896), a disorder that presents with neurological regression, ataxia, spasticity, epilepsy, and progressively vanishing white matter in brain magnetic resonance images.9 Variants in five genes (EIF2B1–B5) can cause the disorder, most frequently biallelic variants in EIF2B5.10
Pertinent Previous Genetic Testing: SNP microarray analysis detected a paternally inherited 416-Kb deletion at 10p12.31, interpreted as a benign variant. An NGS panel of 62 genes for VWM leukodystrophy showed a heterozygous c.338 G > A, p. Arg113His pathogenic variant in the EIF2B5 gene. Subsequent deletion/duplication testing for the EIF2B5 gene via exon-targeted array comparative genome hybridization was normal. Trio WES re-identified the heterozygous p. Arg113His variant. Manual inspection of the reads for the EIF2B5 gene did not reveal any additional variants, with coverage of > 10 × for 100% of this gene.
Results of Post-WES Genetic Testing: Due to the continued clinical suspicion of VWM leukodystrophy, and the detection of one pathogenic variant in the gene, Sanger sequencing of EIF2B5 was pursued. A heterozygous insertion, c.1694delAins45; p.Lys565Ilefs*38 was detected. Although not previously reported in patients with VWM, the insertion had been detected once by the commercial laboratory, in an affected individual. Subsequent parental testing confirmed trans configuration for each variant. Difficulty in alignment of indels larger than 20–50 bp was likely the reason for missing this variant by WES, since retrospective manual inspection of the BAM files failed to detect it.
Discussion
WES is increasingly used as the premier and first-line test for rare and undiagnosed Mendelian disorders.1, 2, 4, 11, 12, 13, 14 The vast majority (>97%) of variants detected by Sanger sequencing can also be detected by WES and with increased detection of mosaicism, WES is a practical tool for comprehensive molecular evaluation.15 When WES is negative, reanalysis of the data can provide resolution in 10–30% of cases16 and this is likely to increase with improvements in technology and new disease associations. However, there is little clarity on the diagnostic options if WES and re-analysis remain negative. Although whole-genome sequencing may be an option, it is not currently widely available clinically.17 Therefore, in instances when WES is negative, clinicians may conclude that no diagnostic options remain for these patients.
WES is a complex high-throughput method, and data loss is possible at each step. Pathogenic variants in known disease-causing genes may be missed because of decreased coverage, locus-specific features such as GC-rich regions, homopolymeric repeats, sequencing biases, and indels that are >20–50 nucleotides.18 The most common reason for variants being missed is a lack of sufficient sequence coverage depth.19 Laboratories performing WES and NGS panels may use alternative methods to capture these.20 Clinicians do reconsider the fit of a phenotype when interpreting variants of uncertain significance on WES, but when WES is negative, adequate coverage of selected genes or exons may lead to a belief that the WES was truly comprehensive. However, in all three of our individuals, coverage was adequate (10 × at 98–100% of the bases), yet the pathogenic variants were missed. Thus, a negative WES result should be interpreted in the clinical context of the individual patient, to determine further testing.
In individual 1, the well-characterized phenotype was consistent with only one diagnosis (infantile systemic hyalinosis), and the ROH on the SNP array included ANTXR2, but molecular confirmation remained elusive despite repeated sequencing and adequate coverage. The single-nucleotide insertion detected by Sanger sequencing is located adjacent to a complex repetitive region and a SNP, both of which could have decreased the mapping quality of the region around the insertion, and resulted in failure of the variant caller program to detect the insertion. Updated variant-calling software and/or manual inspection of the reads would have identified this variant. Manual inspection is of particular importance when a limited set of specific genes is under consideration, but it is not standard practice in commercial laboratories.
Individual 2 had clinical features that changed over a year, leading us to strongly consider a diagnosis of infantile neuroaxonal dystrophy. In this instance, the WES did not capture the deletion because the noncoding exon 1 was not included in the capture kit, as is commonly the case with WES capture kits. Furthermore, structural variants of this size (2.3 kb) would not be detectable by WES. The clinical phenotype and the ROH containing the PLA2G6 gene led us to pursue Sanger sequencing and MLPA, which detected the deletion. It is possible that whole-genome sequencing might have detected this structural variant, but its limited availability and lack of validation of whole-genome sequencing structural variant callers make this an impractical option.
Individual 3 had features consistent with VWM leukoencephalopathy. Although indels <50 bp are below the resolution of exon-level deletion/duplication analyses, we would expect detection by WES, if alignment works well. WES missed the 46-bp indel in the EIF2B5 gene completely, since the detection rate of indels decreases with sizes >20–50 bp.
Clinical geneticists are aware of WES being unsuitable for trinucleotide repeat disorders, mitochondrial DNA variants, epigenetic disorders, and large structural variants. However, for Mendelian disorders in which single-nucleotide variants and small indels are possible, the prevalent thinking is that if coverage of the genes of interest by WES is adequate, the disorders have been effectively excluded. Indeed, an increased depth of coverage would not have detected the missed variants in all three of our cases. The cases presented here underscore the importance of further testing if the clinical phenotype is strongly indicative of a specific condition when WES is negative.
In conclusion, these three case examples illustrate the importance of a multipronged approach when WES is negative. These include (i) obtaining detailed clinical phenotyping to create an accurate differential diagnosis, (ii) reconsidering the family history and mode of inheritance, (iii) reassessing SNP microarray data to identify potential causal genes, (iv) manual inspection of the WES reads for genes that are of interest and obtaining information on capture kits and coverage, and (v) pursuing alternative sequencing methodologies such as Sanger sequencing and deletion/duplication testing to detect single-nucleotide variants and indels that might have been missed with WES. Although WES is comprehensive, its limitations must be considered when negative results are obtained, and alternative diagnostic approaches should be pursued if the phenotype is compelling.
References
Chong JX, Buckingham KJ, Jhangiani SN et al. The genetic basis of Mendelian phenotypes: discoveries, challenges, and opportunities. Am J Hum Genet 2015;97:199–215.
Need AC, Shashi V, Hitomi Y et al. Clinical application of exome sequencing in undiagnosed genetic conditions. J Med Genet 2012;49:353–361.
Yang Y, Muzny DM, Xia F et al. Molecular findings among patients referred for clinical whole-exome sequencing. JAMA. 2014;312:1870–1879.
Lee H, Deignan JL, Dorrani N et al. Clinical exome sequencing for genetic identification of rare Mendelian disorders. JAMA. 2014;312:1880–1887.
El-Kamah GY, Fong K, El-Ruby M et al. Spectrum of mutations in the ANTXR2 (CMG2) gene in infantile systemic hyalinosis and juvenile hyaline fibromatosis. Br J Dermatol 2010;163:213–215.
Kurian MA, Morgan NV, MacPherson L et al. Phenotypic spectrum of neurodegeneration associated with mutations in the PLA2G6 gene (PLAN). Neurology. 2008;70:1623–1629.
Crompton D, Rehal PK, MacPherson L et al. Multiplex ligation-dependent probe amplification (MLPA) analysis is an effective tool for the detection of novel intragenic PLA2G6 mutations: implications for molecular diagnosis. Mol Genet Metab 2010;100:207–212.
Larsson Forsell PK, Kennedy BP, Claesson HE . The human calcium-independent phospholipase A2 gene multiple enzymes with distinct properties from a single gene. Eur J Biochem 1999;262:575–585.
Bugiani M, Boor I, Powers JM, Scheper GC, van der Knaap MS . Leukoencephalopathy with vanishing white matter: a review. J Neuropathol Exp Neurol 2010;69:987–996.
van der Knaap MS, Leegwater PA, Konst AA et al. Mutations in each of the five subunits of translation initiation factor eIF2B can cause leukoencephalopathy with vanishing white matter. Ann Neurol. 2002;51:264–270.
Bowdin S, Gilbert A, Bedoukian E et al. Recommendations for the integration of genomics into clinical practice. Genet Med. 2016;18:1075–1084.
Gilissen C, Hoischen A, Brunner HG, Veltman JA . Unlocking Mendelian disease using exome sequencing. Genome Biol. 2011;12:228.
Yang Y, Muzny DM, Reid JG et al. Clinical whole-exome sequencing for the diagnosis of Mendelian disorders. N Engl J Med 2013;369:1502–1511.
Need AC, Shashi V, Schoch K, Petrovski S, Goldstein DB . The importance of dynamic re-analysis in diagnostic whole exome sequencing. J Med Genet 2017;54:155–156.
Hamilton A, Tetreault M, Dyment DA et al. Concordance between whole-exome sequencing and clinical Sanger sequencing: implications for patient care. Mol Genet Genomic Med 2016;4:504–512.
Wenger AM, Guturu H, Bernstein JA, Bejerano G . Systematic reanalysis of clinical exome data yields additional diagnoses: implications for providers. Genet Med. 2017;19:209–214.
Meynert AM, Ansari M, FitzPatrick DR, Taylor MS . Variant detection sensitivity and biases in whole genome and exome sequencing. BMC Bioinformatics. 2014;15:247.
Ross MG, Russ C, Costello M et al. Characterizing and measuring bias in sequence data. Genome Biol. 2013;14:R51.
Lelieveld SH, Spielmann M, Mundlos S, Veltman JA, Gilissen C . Comparison of exome and genome sequencing technologies for the complete capture of protein-coding regions. Hum Mutat. 2015;36:815–822.
Tiwari A, Lemke J, Altmueller J et al. Identification of novel and recurrent disease-causing mutations in retinal dystrophies using whole exome sequencing (WES): benefits and limitations. PLoS One. 2016;11:e0158692.
Acknowledgements
Research reported in this article was supported by the National Institutes of Health (NIH) Common Fund, through the Office of Strategic Coordination/Office of the NIH Director under award 1U01HG007672-01 to V.S. and D.B.G. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
D.B.G. is a founder of and holds equity in Pairnomix and Praxis, serves as a consultant to AstraZeneca, and has research supported by Janssen, Gilead, Biogen, AstraZeneca, and UCB. The other authors declare no conflict of interest.
Additional information
Members of the Undiagnosed Diseases Network Baylor College of Medicine (Clinical): Mercedes E. Alejandro, Carlos A. Bacino, Ashok Balasubramanyam, Bret L. Bostwick, Lindsay C. Burrage, Shan Chen, Gary D. Clark, William J. Craigen, Shweta U. Dhar, Lisa T. Emrick, Brett H. Graham, Neil A. Hanchard, Mahim Jain, Seema R. Lalani, Brendan H. Lee, Richard A. Lewis, Mashid S. Azamian, Paolo M. Moretti, Sarah K. Nicholas, Jordan S. Orange, Jennifer E. Posey, Lorraine Potocki, Jill A. Rosenfeld, Susan L. Samson, Daryl A. Scott, Alyssa A. Tran, Tiphanie P. Vogel, Jing Zhang; Baylor College of Medicine (Model Organism Screening Center): Hugo J. Bellen, Michael F. Wangler, Shinya Yamamoto; Baylor College of Medicine (Sequencing): Christine M. Eng, Donna M. Muzny, Patricia A. Ward, Yaping Yang; Columbia University: David B. Goldstein, Nicholas Stong; Duke University: Yong-hui Jiang, Allyn McConkie-Rosell, Loren D.M. Pena, Kelly Schoch, Vandana Shashi, Rebecca C. Spillmann, Jennifer A. Sullivan, Nicole M. Walley; Harvard University (Clinical Site): Alan H. Beggs, Lauren C. Briere, Cynthia M. Cooper, Laurel A. Donnell-Fink, Elizabeth L. Krieg, Joel B. Krier, Sharyn A. Lincoln, Joseph Loscalzo, Richard L. Maas, Calum A. MacRae, J. Carl Pallais, Lance H. Rodan, Edwin K. Silverman, Joan M. Stoler, David A. Sweetser, Chris A. Walsh; Harvard University (UDN Coordinating Center): Cecilia Esteves, Ingrid A. Holm, Isaac S. Kohane, Paul Mazur, Alexa T. McCray, Matthew Might, Rachel B. Ramoni, Kimberly Splinter; HudsonAlpha: David P. Bick, Camille L. Birch, Braden E. Boone, Donna M. Brown, Daniel C. Dorset, Lori H. Handley, Howard J. Jacob, Angela L. Jones, Jozef Lazar, Shawn E. Levy, J. Scott Newberry, Molly C. Schroeder, Kimberly A. Strong (deceased), Elizabeth A. Worthey; National Institutes of Health: Jyoti G. Dayal, David J. Eckstein, Sarah E. Gould, Ellen M. Howerton, Donna M. Krasnewich, Laura A. Mamounas, Teri A. Manolio, John J. Mulvihill, Tiina K. Urv, Anastasia L. Wise; National Institute of Neurological Disorders and Stroke (NINDS): Ariane G. Soldatos; Oregon Health & Science University Metabolomics: Matthew Brush, Jean-Philippe F. Gourdine, Melissa Haendel, David M. Koeller; Pacific Northwest National Laboratory Metabolomics: Jennifer E. Kyle, Thomas O. Metz, Katrina M. Waters, Bobbie-Jo M. Webb-Robertson; Stanford University: Euan A. Ashley, Jonathan A. Bernstein, Annika M. Dries, Paul G. Fisher, Jennefer N. Kohler, Daryl M. Waggott, Matthew T. Wheeler, Patricia A. Zornio; University of California–Los Angeles: Patrick Allard, Hayk Barseghyan, Esteban C. Dell’Angelica, Ani Dillon, Katrina M. Dipple, Naghmeh Dorrani, Emilie D. Douine, Ascia Eskin, Brent L. Fogel, Matthew R. Herzog, Hane Lee, Allen Lipson, Sandra K. Loo, Julian A. Martínez-Agosto, Stan F. Nelson, Christina G.S. Palmer, Jeanette C. Papp, Neil H. Parker, Janet S. Sinsheimer, Eric Vilain, Allison Zheng; Undiagnosed Diseases Program (UDP): Christopher J. Adams, Elizabeth A. Burke, Katherine R. Chao, Mariska Davids, David D. Draper, Tyra Estwick, Trevor S. Frisby, Kate Frost, Valerie Gartner, Rena A. Godfrey, Mitchell Goheen, Gretchen A. Golas, Mary G. Gordon, Catherine A. Groden, Mary E. Hackbarth, Isabel Hardee, Jean M. Johnston, Alanna E. Koehler, Lea Latham, Yvonne L. Latour, C. Christopher Lau, Denise J. Levy, Adam P. Liebendorfer, Ellen F. Macnamara, Valerie V. Maduro, Thomas C. Markello, Alexandra J. McCarty, Jennifer L. Murphy, Michele E. Nehrebecky, Donna Novacic, Barbara N. Pusey, Sarah Sadozai, Katherine E. Schaffer, Prashant Sharma, Sara P. Thomas, Nathanial J. Tolman, Camilo Toro, Zaheer M. Valivullah, Colleen E. Wahl, Mike Warburton, Alec A. Weech, Guoyun Yu; UDP, Children’s National Medical Center: Andrea L. Gropman; UDP, National Human Genome Research Institute: David R. Adams, William A. Gahl, May Christine V. Malicdan, Cynthia J. Tifft, Lynne A. Wolfe; UDP, NINDS: Paul R. Lee; University of Oregon The Model Organisms Screening Center (MOSC): John H. Postlethwait, Monte Westerfield; Vanderbilt University: Anna Bican, Joy D. Cogan, Rizwan Hamid, John H. Newman, John A. Phillips III, Amy K. Robertson.
Supplementary material is linked to the online version of the paper at
Supplementary information
Rights and permissions
About this article
Cite this article
Pena, L., Jiang, YH., Schoch, K. et al. Looking beyond the exome: a phenotype-first approach to molecular diagnostic resolution in rare and undiagnosed diseases. Genet Med 20, 464–469 (2018). https://doi.org/10.1038/gim.2017.128
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.128
Keywords
This article is cited by
-
Beyond the exome: utility of long-read whole genome sequencing in exome-negative autosomal recessive diseases
Genome Medicine (2023)
-
The diagnostic yield, candidate genes, and pitfalls for a genetic study of intellectual disability in 118 middle eastern families
Scientific Reports (2022)
-
Fundoscopy-directed genetic testing to re-evaluate negative whole exome sequencing results
Orphanet Journal of Rare Diseases (2020)
-
Regulatory genome variants in human susceptibility to infection
Human Genetics (2020)
-
Genotype–phenotype analysis of 523 patients by genetics evaluation and clinical exome sequencing
Pediatric Research (2020)